Abstract
As a m6A methylation modifier, METTL3 is functionally involved in various biological processes. Nevertheless, the role of METTL3 in osteogenesis is not determined up to date. In the current study, METTL3 is identified as a crucial regulator in the progression of osteogenic differentiation. Loss of METTL3 significantly augments calcium deposition and enhances alkaline phosphatase activity of mesenchymal stem cells, uncovering an inhibitory role of METTL3 in osteogenesis. More importantly, the underlying molecular basis by which METTL3 regulates osteogenesis is illustrated. We find that METTL3 positively regulates expression of MYD88, a critical upstream regulator of NF-κB signaling, by facilitating m6A methylation modification to MYD88-RNA, subsequently inducing the activation of NF-κB which is widely regarded as a repressor of osteogenesis and therefore suppressing osteogenic progression. Moreover, the METTL3-mediated m6A methylation is found to be dynamically reversed by the demethylase ALKBH5. In summary, this study highlights the functional importance of METTL3 in osteogenic differentiation and METTL3 may serve as a promising molecular target in regenerative medicine, as well as in the field of bone tissue engineering.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As it is well known, the osteogenic differentiation of mesenchymal stem cells (MSCs) is a fundamental process for repair and regeneration of bone defects, as well as embryonic development. Due to the strong differentiation ability into multiple cell types, MSCs are widely accepted as a promising candidate for regenerative medicine. Numerous studies have shown that the stem cell-based therapy is an ideal choice for these bone tissue-related disorders [1,2,3,4,5]. Therefore, investigating the underlying molecular basis of osteogenesis is a central point for promoting the development of stem cell-based clinical treatment of these dysfunctional bone defects.
A number of signaling pathways are implicated in mediating osteogenesis, especially NF-κB which is a critical regulator consisting homo- or heterodimers of RelA (p65), RelB, or c-Rel. Increasing evidence showed that the activation of NF-κB signaling has an inhibitory effect on the progression of osteogenic differentiation [6,7,8], emphasizing the functional importance of NF-κB in osteogenic progression. Thus, targeting NF-κB pathway may be an effective strategy to restore those bone tissue dysfunctions.
Recently, RNA modifications, especially N6-methyladenosine (m6A) RNA methylation, have been reported to play critical roles in multiple biological processes such as stem cell renewal, development, tissue homeostasis, and disease progression [9,10,11,12]. To date, m6A is widely regarded as one of the most widespread modifications of mRNAs in higher eukaryotes, contributing to the regulation of gene expressions through mediating mRNA export, stability, translation, and other mRNA processing steps [13,14,15,16]. m6A methylation, often catalyzed by the methyltransferase METTL3 and its partner METTL14 and WTAP [17,18,19,20,21], can be dynamically reversed by the demethylase ALKBH5 or FTO [22,23,24]. Thus, the methyltransferases usually cooperate with the demethylases to regulate the m6A status of specific mRNA, maintaining a balanced expression level of some key regulators.
In this project, we identified METTL3 as a crucial regulator of osteogenic differentiation. Attenuation of METTL3 significantly enhanced both calcium deposition and alkaline phosphatase activity of MSCs, demonstrating an inhibitory role of METTL3 in the regulation of osteogenesis. More interestingly, we find that METTL3 is involved in modulating the activity of NF-κB which is an important inhibitor for osteogenic differentiation. Therefore, these findings illustrate the regulatory function of METTL3 in the progression of osteogenesis via NF-κB signaling.
Materials and methods
Isolation and culture of MenSCs
The menstrual bloods were first isolated from female donors and then subjected to standard Ficoll procedures. Subsequently, the detailed protocol for MenSCs isolation and culture was performed as previously described [25].
Antibodies and reagents
Anti-IκBα (#10268-1-AP), anti-METTL3 (15073-1-AP), anti-MYD88 (#23230-1-AP), anti-ALKBH5 (#16837-1-AP), and anti-p65 (#10745-1-AP) antibodies were purchased from Proteintech Group Inc. Anti-m6A (#202003) antibody was purchased from Synaptic Systems. Anti-β-actin (#A5441) was purchased from Sigma. The chemical reagents Bay 11-7082 (#B5556), Alizarin Red S (#A5533), BCIP/NBT liquid substrate (#B1911), and osteogenic differentiation medium (#SCM121) were all purchased from Sigma.
Alizarin Red S and ALP assays
For Alizarin Red assay, MenSCs were first fixed with 70% ethanol, followed by 1% Alizarin Red solution staining for 1 min. The detailed protocol was performed as previously described [25]. For ALP assay, a BCIP/NBT liquid substrate composed of 0.1 M 2-amino-2-methyl-1-propanol, 1 mM MgCl2, and 8 mM p-nitrophenyl phosphate disodium was incubated with cell homogenates at 37 °C for 30 min, and then was stopped by the addition of 0.1 N NaOH. Finally, the absorbance was measured at 405 nm.
Constructs and lentiviral infection
The shRNA targeting METTL3 was inserted into a pLV-H1 lentiviral vector. The sequence for shMETTL3 is 5′- CTGCAAGTATGTTCACTATGA -3′. The siRNAs for ALKBH5 and FTO were all purchased from Thermo Fisher. The overexpression vector of METTL3 was amplified by RT-PCR, and then cloned into a lentiviral vector as previously reported [25]. For lentivirus infection, the detailed protocol was performed as previously described [25].
Quantitative RT-PCR
RNAs were extracted from MenSCs by using Trizol reagent, followed by reverse transcription, according to the manufacturers’ instructions. Real-time quantitative PCR was carried out as previously reported [25]. The primer sequences for qRT-PCR are: F. 5′-GGACGAGGCAAGAGTTTCAC-3′, R. 5′-GAGGCGGTCAGAG AACAAAC-3′ (RUNX2); F. 5′-CACAGCTCTTCTGACTGTCTG-3′, R. 5′-CTGGTGAAATGCC TGCATGGAT-3′ (SP7); F. 5′-CATGAGAAGTATGACAACAGCCT-3′, R. 5′-AGTCCTTCCACGATACC AAAGT-3′ (GAPDH); F. 5′-CAAGCTGCACTTCAGACGAA-3′, R. 5′-GCTTGGCGTGTG GTCTTT-3′ (METTL3).
RNA immunoprecipitation (RIP) assay
Approximately 107 cells were scraped off from dishes using a cell scraper. After washing for three times with PBS, the cell pellets were resuspended with 1 ml Buffer A (10 mM Hepes pH 7.5, 1.5 mM MgCl2, 10 mM KCL, 1.0 mM DTT, 0.5 mM PMSF, 1 × protease inhibitor cocktail) and centrifuged at 1000×g for 5 min. the swollen cells were resuspended and then lysed with 0.8 ml RIP buffer (30 mM Hepes pH 7.5, 1.5 mM MgCl2, 0.3 M NaCl, 20% glycerol, 0.5% NP40, 1.0 mM DTT, 0.5 mM PMSF). The debris were pelleted by centrifugation at 13,000 RPM for 10 min, and the supernatants were collected and incubated with indicated antibodies for 4 h at 4 °C with gentle rotation. Add 30 μl protein A beads and rotate at 4 °C for additional 1 h for immunoprecipitation. The immunoprecipitates were washed three times with the RIP buffer and additional two times with a high salt buffer (50 mM Tris–HCl pH 7.5, 300 mM NaCl, 1 mM EDTA, 1% Triton X-100, 0.1% SDS, 0.5 mM PMSF). The co-immunoprecipitated RNAs were extracted and analyzed by qRT-PCR. The primer sequence of MYD88 used in RIP assays were F. 5′-GAGACACAAGCGGACCCC-3′, R. 5′-CTGTTCCAGTTGC CGGATCA-3′.
Statistical analysis
All results in this project are derived from at least three independent experiments and shown as mean ± SD. All statistical analyses were performed using Prism5 (GraphPad). Student’s t test was used for comparisons between two groups, and one-way ANOVA followed by Tukey post hoc test was used to compare more than three groups. p Value < 0.05 was considered statistically significant.
Results
METTL3 expression is decreased after osteogenic induction
To study the potential role of METTL3 in osteogenic differentiation, mesenchymal stem cells were isolated from menstrual bloods of the female donors, and then subjected to osteogenic induction, using a commercial osteogenic medium. We observed the dynamic changes of METTL3 expression after osteogenic induction for 7 and 14 days. Interestingly, expression of METTL3 was significantly downregulated when the menstrual blood-derived mesenchymal stem cells (MenSCs) were exposed to the osteogenic medium treatment (Fig. 1a), indicating a possible role of METTL3 in the regulation of osteogenesis. Meanwhile, we examined expressions of representative osteogenic marker genes, and found that RUNX2 and SP7 expressions were dramatically elevated after osteogenic induction (Fig. 1b).
METTL3 expression is decreased after osteogenic induction. a METTL3 expression in MenSCs was examined by qRT-PCR assay after osteogenic medium treatment for 14 and 21 days. b Expressions of representative osteogenic markers RUNX2 and SP7 were measured using qRT-PCR analysis after osteogenic medium treatment for 14 and 21 days. All data are from biological triplicates and are shown as mean ± SD. n = 3. *p < 0.05,**p < 0.01, ***p < 0.001 versus Ctrl
METTL3 represses osteogenesis of MenSCs
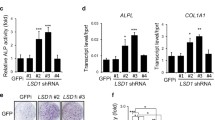
To determine the regulatory function of METTL3 in osteogenesis. We first depleted METTL3, using a specific shRNA, and then performed Alizarin Red S staining analysis. As a consequence, attenuation of METTL3 remarkably enhanced calcium deposition of the MenSCs (Fig. 2a). To further confirm this data, the activity of alkaline phosphatase (ALP) of MenSCs with or without METTL3 knockdown was measured by a BCIP/NBT liquid substrate. Consistently, ALP activity was found to be augmented when METTL3 was silenced (Fig. 2b). To further validate the inhibitory effect of METTL3 on osteogenesis, we next assayed expressions of the osteogenic markers. qRT-PCR analysis revealed that knockdown of METTL3 significantly upregulated the representative osteogenic marker genes RUNX2 and SP7 (Fig. 2c). Moreover, enforced overexpression of METTL3 inhibited both the calcium deposition and expressions of those osteogenic markers (Fig. 2d, e). Collectively, those observations suggest that METTL3 serve as a critical suppressor for osteogenic differentiation of mesenchymal stem cells.
METTL3 negatively regulates osteogenesis. a The effect of METTL3 on calcium deposition was determined by Alizarin Red S staining. MenSCs with or without METTL3 knockdown were subjected to Alizarin Red S staining analysis. b The effect of METTL3 on alkaline phosphatase activity, determined by ALP assay. METTL3 in MenSCs, with or without osteogenic induction for 14 days, was depleted and then lysed for measuring ALP activity. c qRT-PCR analysis in MenSCs showing the effect of METTL3 on expressions of osteogenic markers RUNX2 and SP7. d The effect of enforced METTL3 overexpression on calcium deposition in MenSCs with osteogenic induction for 21 days, determined by Alizarin Red S staining. e METTL3 overexpression was performed in MenSCs to examine the influence of METTL3 on RUNX2 and SP7 expressions in the presence of osteogenic induction for 14 days, using qRT-PCR assay. f, g The knockdown and overexpression efficiency of METTL3 were measured in MenSCs by Western blot analysis. All data are from biological triplicates and are shown as mean ± SD. n = 3. *p < 0.05, **p < 0.01, ***p < 0.001. (Color figure online)
METTL3 negatively regulates osteogenesis through NF-κB signaling
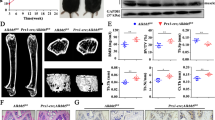
Previous studies reported that NF-κB is a critical regulator of osteogenesis. The activation of NF-κB signaling can inhibit osteogenic differentiation. Therefore, we ask whether METTL3 regulates osteogenesis, at least in part, through NF-κB signaling. To prove this concept, we first examined the effect of METTL3 on IκBα degradation and the phosphorylation status of p65 at S536 site, a crucial modification which is responsible for NF-κB activity [26]. Western blotting analysis showed that METTL3 knockdown not only blocked proteasome-mediated IκBα degradation but also restrained the phosphorylation level of p65 at S536 site (Fig. 3a). To confirm the regulatory role of METTL3 for the activation of NF-κB signaling, we conducted fractionation experiment to observe the effect of METTL3 on NF-κB nuclear translocation. Consequently, highly expressed METTL3 remarkably facilitated p65 accumulation in nuclear fraction (Fig. 3b), suggesting a positive role of METTL3 for the activation of NF-κB. Next, we assayed the functional importance of NF-κB signaling in METTL3-mediated osteogenesis. Consistent with the prior report, ALP activity assay showed that METTL3 overexpression had an inhibitory effect on the osteogenesis of MenSCs (Fig. 3c). However, this inhibitory effect disappeared after treatment with NF-κB-specific inhibitor Bay 11-7082 (Fig. 3c), demonstrating the crucial role of NF-κB signaling in METTL3-mediated osteogenesis of MenSCs. Furthermore, this finding was further validated by the expression changes of these osteogenic marker genes using a qRT-PCR assay (Fig. 3d, e). Overall, these observations suggest that METTL3 suppresses osteogenic differentiation, at least in part, through enhancing the activity of NF-κB signaling.
METTL3 regulates the activity of NF-κB signaling. a MenSCs, with or without METTL3 knockdown, were subjected to Western blot analysis to observe the effect of METTL3 on IκBα degradation and the phosphorylation status of p65 at S536 site. b The effect of METTL3 on cellular location of NF-κB was examined by Western blot assay. Fractionation experiments were conducted in MenSCs after METTL3 depletion, follow by cellular localization analysis of p65 and RelB using Western blot assay. c The effect of NF-κB on alkaline phosphatase activity was examined by ALP assay. METTL3-overexpressed MenSCs in the presence or absence of Bay 11-7082 (0.5 μM, 7 days) treatment were subjected to ALP assay. d, e Enforced overexpression of METTL3 was performed to examine the effect of METTL3 on expressions of osteogenic marker genes RUNX2 and SP7. MenSCs were treated with or without Bay 11-7082 (0.5 μM, 7 days) upon osteogenic induction condition after METTL3 overexpression. All data are from biological triplicates and are shown as mean ± SD. n = 3. *p < 0.05, **p < 0.01
METTL3 regulates MYD88 expression in a m6A modification manner
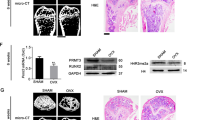
Given the critical role of METTL3 in the regulation of NF-κB signaling, we next sought to define the precise molecular basis by which METTL3 regulates the activity of NF-κB. We first examined the influence of METTL3 on the expression of MYD88, a widely accepted regulator of NF-κB. Western blot analysis evidenced that METTL3 knockdown significantly repressed MYD88 expression (Fig. 4a). Thus, we hypothesized that METTL3 is involved in MYD88-RNA m6A methylation. To test the hypothesis, we performed RNA immunoprecipitation (RIP) asssay, with an antibody against METTL3. The RIP result showed that METTL3 obviously interacts with MYD88-RNA (Fig. 4b). Moreover, anti-m6A RIP assays were employed to further confirm the effect of METTL3 on m6A modification of MYD88-RNA. Consequently, we found that attenuation of METTL3 dramatically abrogated the m6A status of MYD88 (Fig. 4c). Increasing evidences reveal that m6A modification is a dynamic reversable process, which can be reversed by specific demethylase, such as FTO and ALKBH5. Therefore, we next performed shRNA-mediated knockdown of FTO and ALKBH5, respectively, and then measured MYD88 expression. Interestingly, ALKBH5, but not FTO, knockdown significantly augmented the expression of MYD88 (Fig. 4d). RIP assays further confirmed the significant association of ALKBH5 with MYD88-RNA (Fig. 4e). In summary, those findings suggest that METTL3 positively regulates MYD88 expression through controlling m6A methylation status of MYD88-RNA, therefore leading to the activation of NF-κB signaling.
METTL3 regulates m6A methylation status of MYD88-RNA. a Western blot assays showing the effect of METTL3 on MYD88 expression. Knockdown of METTL3 was performed in MenSCs, and then subjected to Western blot analysis. b Anti-METTL3 RIP assays were performed in MenSCs to test the association of METTL3 with MYD88-RNA. c RIP assays, with an antibody against m6A, were conducted in MenSCs to determine the effect of METTL3 on m6A status of MYD88-RNA. d ALKBH5 and FTO knockdown were performed in MenSCs, respectively, and then subjected to Western blot assay to examine the effect of ALKBH5 and FTO on MYD88 expression. e RIP assays showing the association of ALKBH5 with MYD88-RNA. All data are from biological triplicates and are shown as mean ± SD. n = 3. *p < 0.05, **p < 0.01, ***p < 0.001
Discussion
The regulation of osteogenesis is a central step for stem cell-based therapy in bone tissue repair and regeneration. In this study, we first characterized METTL3 as a critical regulator for osteogenic differentiation. Inhibition of METTL3 can enhance calcium deposition, using Alizarin Red S staining analysis. Moreover, we further examined the effect of METTL3 on the activity of alkaline phosphatase and found that METTL3 knockdown significantly augmented alkaline phosphatase activity. These results demonstrate the critical inhibitory role of METTL3 in regulation of osteogenic differentiation.
Multiple signaling pathways are functionally involved in the progression of osteogenic differentiation, especially NF-κB signaling. Increasing evidences revealed that the activation of NF-κB can suppress osteogenesis [8]. Therefore, we tested the relationship between METTL3 and NF-κB. Intriguingly, we identified METTL3 as a crucial activator of NF-κB. Knockdown of METTL3 can inhibit the proteasome-mediated IκBα degradation and the phosphorylation status of p65 at S536 site, therefore restraining NF-κB nuclear translocation and leading to its transcriptional repression. Here, we demonstrated the significant role of METTL3 in the regulation of NF-κB signaling activity.
As a methyltransferase responsible for m6A modification, METTL3 is implicated in the post-transcriptional regulation of many key mediators [24, 27]. In the current study, we discovered that METTL3 specially interacts with MYD88-RNA and regulates its expression through a m6A methylation mechanism. Depletion of METTL3 significantly downregulates the expression of MYD88, a widely accepted upstream regulator of NF-κB. Furthermore, RIP assay, with an antibody specially against m6A, showed that the m6A methylation status is repressed after METTL3 depletion, indicating that METTL3 regulates the activity of NF-κB signaling via controlling m6A methylation-mediated MYD88 expression (Fig. 5). More interestingly, we observed that METTL3-mediated m6A methylation of MYD88-RNA can be reversed by the demethylase ALKBH5 (Fig. 5), which is consistent with the prior reports that m6A methylation modification is a dynamically reversible process [22, 24]. Taken together, this study highlights the functional importance of METTL3 in osteogenesis through regulating MYD88-mediated NF-κB activity, and we conclude that METTL3 may be an effective therapeutic target for bone tissue repair and regeneration.
Schematic representation of the molecular basis by which METTL3 regulates the activity of NF-κB signaling. As a repressor of osteogenic differentiation, METTL3 interacts with MYD88-RNA and positively regulates its expression through a m6A methylation modification manner. The upregulated MYD88 will then activate its downstream regulator NF-κB, a critical repressor of osteogenesis. The activated NF-κB signaling will ultimately suppress the osteogenic differentiation of MSCs
Data availability
All data generated or analyzed during this study are shown in this article.
Abbreviations
- MSCs:
-
Mesenchymal stem cells
- m6A:
-
N6-methyladenosine
- MenSCs:
-
Menstrual blood-derived mesenchymal stem cells
- ALP:
-
Alkaline phosphatase
References
Barba M, Di Taranto G, Lattanzi W (2017) Adipose-derived stem cell therapies for bone regeneration. Expert Opin Biol Ther 17:677–689. https://doi.org/10.1080/14712598.2017.1315403
Chen M, Xu Y, Zhang T, Ma Y, Liu J, Yuan B, Chen X, Zhou P, Zhao X, Pang F, Liang W (2019) Mesenchymal stem cell sheets: a new cell-based strategy for bone repair and regeneration. Biotechnol Lett 41:305–318. https://doi.org/10.1007/s10529-019-02649-7
Fujii Y, Kawase-Koga Y, Hojo H, Yano F, Sato M, Chung UI, Ohba S, Chikazu D (2018) Bone regeneration by human dental pulp stem cells using a helioxanthin derivative and cell-sheet technology. Stem Cell Res Ther 9:24. https://doi.org/10.1186/s13287-018-0783-7
Shah AR, Cornejo A, Guda T, Sahar DE, Stephenson SM, Chang S, Krishnegowda NK, Sharma R, Wang HT (2014) Differentiated adipose-derived stem cell cocultures for bone regeneration in polymer scaffolds in vivo. J Craniofac Surg 25:1504–1509. https://doi.org/10.1097/SCS.0000000000000755
Zigdon H, Levin L (2012) Stem cell therapy for bone regeneration: present and future strategies. Alpha Omegan 105:35–38
Lin TH, Gibon E, Loi F, Pajarinen J, Cordova LA, Nabeshima A, Lu L, Yao Z, Goodman SB (2017) Decreased osteogenesis in mesenchymal stem cells derived from the aged mouse is associated with enhanced NF-kappaB activity. J Orthop Res 35:281–288. https://doi.org/10.1002/jor.23270
Liu C, Zhang H, Tang X, Feng R, Yao G, Chen W, Li W, Liang J, Feng X, Sun L (2018) Mesenchymal stem cells promote the osteogenesis in collagen-induced arthritic mice through the inhibition of TNF-alpha. Stem Cells Int 2018:4069032. https://doi.org/10.1155/2018/4069032
Wang N, Zhou Z, Wu T, Liu W, Yin P, Pan C, Yu X (2016) TNF-alpha-induced NF-kappaB activation upregulates microRNA-150-3p and inhibits osteogenesis of mesenchymal stem cells by targeting beta-catenin. Open Biol. https://doi.org/10.1098/rsob.150258
Batista PJ, Molinie B, Wang J, Qu K, Zhang J, Li L, Bouley DM, Lujan E, Haddad B, Daneshvar K, Carter AC, Flynn RA, Zhou C, Lim KS, Dedon P, Wernig M, Mullen AC, Xing Y, Giallourakis CC, Chang HY (2014) m(6)A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15:707–719. https://doi.org/10.1016/j.stem.2014.09.019
Geula S, Moshitch-Moshkovitz S, Dominissini D, Mansour AA, Kol N, Salmon-Divon M, Hershkovitz V, Peer E, Mor N, Manor YS, Ben-Haim MS, Eyal E, Yunger S, Pinto Y, Jaitin DA, Viukov S, Rais Y, Krupalnik V, Chomsky E, Zerbib M, Maza I, Rechavi Y, Massarwa R, Hanna S, Amit I, Levanon EY, Amariglio N, Stern-Ginossar N, Novershtern N, Rechavi G, Hanna JH (2015) Stem cells. m6A mRNA methylation facilitates resolution of naive pluripotency toward differentiation. Science 347:1002–1006. https://doi.org/10.1126/science.1261417
Lin S, Choe J, Du P, Triboulet R, Gregory RI (2016) The m(6)A methyltransferase METTL3 promotes translation in human cancer cells. Mol Cell 62:335–345. https://doi.org/10.1016/j.molcel.2016.03.021
Zhang C, Chen Y, Sun B, Wang L, Yang Y, Ma D, Lv J, Heng J, Ding Y, Xue Y, Lu X, Xiao W, Yang YG, Liu F (2017) m(6)A modulates haematopoietic stem and progenitor cell specification. Nature 549:273–276. https://doi.org/10.1038/nature23883
Cheng M, Sheng L, Gao Q, Xiong Q, Zhang H, Wu M, Liang Y, Zhu F, Zhang Y, Zhang X, Yuan Q, Li Y (2019) The m(6)A methyltransferase METTL3 promotes bladder cancer progression via AFF4/NF-kappaB/MYC signaling network. Oncogene. https://doi.org/10.1038/s41388-019-0683-z
Maity A, Das B (2016) N6-methyladenosine modification in mRNA: machinery, function and implications for health and diseases. FEBS J 283:1607–1630. https://doi.org/10.1111/febs.13614
Vu LP, Pickering BF, Cheng Y, Zaccara S, Nguyen D, Minuesa G, Chou T, Chow A, Saletore Y, MacKay M, Schulman J, Famulare C, Patel M, Klimek VM, Garrett-Bakelman FE, Melnick A, Carroll M, Mason CE, Jaffrey SR, Kharas MG (2017) The N(6)-methyladenosine (m(6)A)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nat Med 23:1369–1376. https://doi.org/10.1038/nm.4416
Yue Y, Liu J, He C (2015) RNA N6-methyladenosine methylation in post-transcriptional gene expression regulation. Genes Dev 29:1343–1355. https://doi.org/10.1101/gad.262766.115
Lin Z, Hsu PJ, Xing X, Fang J, Lu Z, Zou Q, Zhang KJ, Zhang X, Zhou Y, Zhang T, Zhang Y, Song W, Jia G, Yang X, He C, Tong MH (2017) Mettl3-/Mettl14-mediated mRNA N(6)-methyladenosine modulates murine spermatogenesis. Cell Res 27:1216–1230. https://doi.org/10.1038/cr.2017.117
Liu J, Yue Y, Han D, Wang X, Fu Y, Zhang L, Jia G, Yu M, Lu Z, Deng X, Dai Q, Chen W, He C (2014) A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat Chem Biol 10:93–95. https://doi.org/10.1038/nchembio.1432
Yao QJ, Sang L, Lin M, Yin X, Dong W, Gong Y, Zhou BO (2018) Mettl3-Mettl14 methyltransferase complex regulates the quiescence of adult hematopoietic stem cells. Cell Res 28:952–954. https://doi.org/10.1038/s41422-018-0062-2
Kobayashi M, Ohsugi M, Sasako T, Awazawa M, Umehara T, Iwane A, Kobayashi N, Okazaki Y, Kubota N, Suzuki R, Waki H, Horiuchi K, Hamakubo T, Kodama T, Aoe S, Tobe K, Kadowaki T, Ueki K (2018) The RNA methyltransferase complex of WTAP, METTL3, and METTL14 regulates mitotic clonal expansion in adipogenesis. Mol Cell Biol. https://doi.org/10.1128/MCB.00116-18
Selberg S, Blokhina D, Aatonen M, Koivisto P, Siltanen A, Mervaala E, Kankuri E, Karelson M (2019) Discovery of small molecules that activate RNA methylation through cooperative binding to the METTL3-14-WTAP complex active site. Cell Rep 26(3762–3771):e5. https://doi.org/10.1016/j.celrep.2019.02.100
Piette ER, Moore JH (2018) Identification of epistatic interactions between the human RNA demethylases FTO and ALKBH5 with gene set enrichment analysis informed by differential methylation. BMC Proc 12:59. https://doi.org/10.1186/s12919-018-0122-0
Shen F, Huang W, Huang JT, Xiong J, Yang Y, Wu K, Jia GF, Chen J, Feng YQ, Yuan BF, Liu SM (2015) Decreased N(6)-methyladenosine in peripheral blood RNA from diabetic patients is associated with FTO expression rather than ALKBH5. J Clin Endocrinol Metab 100:E148–E154. https://doi.org/10.1210/jc.2014-1893
Song H, Feng X, Zhang H, Luo Y, Huang J, Lin M, Jin J, Ding X, Wu S, Huang H, Yu T, Zhang M, Hong H, Yao S, Zhao Y, Zhang Z (2019) METTL3 and ALKBH5 oppositely regulate m(6)A modification of TFEB mRNA, which dictates the fate of hypoxia/reoxygenation-treated cardiomyocytes. Autophagy. https://doi.org/10.1080/15548627.2019.1586246
Zhu X, Yu J, Du J, Zhong G, Qiao L, Lin J (2019) LncRNA HOXA-AS2 positively regulates osteogenesis of mesenchymal stem cells through inactivating NF-kappaB signalling. J Cell Mol Med 23:1325–1332. https://doi.org/10.1111/jcmm.14034
Zou Z, Huang B, Wu X, Zhang H, Qi J, Bradner J, Nair S, Chen LF (2014) Brd4 maintains constitutively active NF-kappaB in cancer cells by binding to acetylated RelA. Oncogene 33:2395–2404. https://doi.org/10.1038/onc.2013.179
Wang H, Hu X, Huang M, Liu J, Gu Y, Ma L, Zhou Q, Cao X (2019) Mettl3-mediated mRNA m(6)A methylation promotes dendritic cell activation. Nat Commun 10:1898. https://doi.org/10.1038/s41467-019-09903-6
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (31502045) and Xinxiang Medical University Foundation (300-505307).
Author information
Authors and Affiliations
Contributions
JL, MC, and XZ are responsible for designing the project. JY and LS performed most of the experiments. YL contributed to data analysis. XZ wrote the draft of this manuscript. All authors take part in discussions.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Ethical approval
The MenSCs used in this study were obtained with the informed consent of the donors. All experiments in this manuscript meet the “Declaration of Helsinki” and were approved by the Ethics Committee of Xinxiang Medical University.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yu, J., Shen, L., Liu, Y. et al. The m6A methyltransferase METTL3 cooperates with demethylase ALKBH5 to regulate osteogenic differentiation through NF-κB signaling. Mol Cell Biochem 463, 203–210 (2020). https://doi.org/10.1007/s11010-019-03641-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-019-03641-5