Abstract
The aim of the study was to understand the role of homologous recombination repair (HRR) pathway genes in development of chemotolerance in breast cancer (BC). For this purpose, chemotolerant BC cells were developed in MCF-7 and MDA MB 231 cell lines after treatment with two anthracycline anti-tumor antibiotics doxorubicin and nogalamycin at different concentrations for 48 h with differential cell viability. The drugs were more effective in MCF-7 (IC50: 0.214–0.242 µM) than in MDA MB 231 (IC50: 0.346–0.37 µM) as shown by cell viability assay. The drugs could reduce the protein expression of PCNA in the cell lines. Increased mRNA/protein expression of the HRR (BRCA1, BRCA2, FANCC, FANCD2, and BRIT1) genes was seen in the cell lines in the presence of the drugs at different concentrations (lower IC50, IC50, and higher IC50) irrespective of the cell viability (68–41%). Quantitative methylation assay showed an increased percentage of hypomethylation of the promoters of these genes after drug treatment in the cell lines. Similarly, chemotolerant neoadjuvant chemotherapy (NACT) treated primary BC samples showed significantly higher frequency of hypomethylation of the genes than the pretherapeutic BC samples. The drugs in different concentrations could reduce m-RNA and protein expression of DNMT1 (DNA methyltransferase 1) in the cell lines. Similar phenomenon was also evident in the NACT samples than in the pretherapeutic BC samples. Thus, our data indicate that reduced DNMT1 expression along with promoter hypomethylation and increased expression of the HRR genes might have importance in chemotolerance in BC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast carcinoma (BC) is frequent cancer among women worldwide [1] which accounts for 25% of cancer burden in Eastern Indian population [2]. Chemotherapy is a cornerstone treatment in patients with early and advanced breast cancer apart from surgery and radiotherapy. The recent trend in therapy of BC is the administration of neoadjuvant chemotherapy (NACT) to shrink the locally advanced tumors for better prognosis of the disease [3]. However, characteristics of the chemotolerant BC cells are not well understood.
Chemosensitivity of the tumors is associated with competency of DNA damage response (DDR) activation [4,5,6,7]. Among different DDR pathways, homologous recombination repair (HRR) pathway is important due to the presence of several BC susceptible genes [8]. In pretherapeutic BC samples, frequent (41–47%) alteration (deletion/methylation) of BRIT1 and its reduced mRNA/protein expression were reported [9, 10]. Our previous studies showed variable frequencies of deletion (50–70%) and methylation (30–60%) in BRCA1/BRCA2 genes in pretherapeutic BC [11, 12]. In addition, 40–65% deletion and methylation in FANCC and FANCD2 genes as well as 20–40% of their reduced protein expression were observed in BC [12,13,14]. Similarly, high frequency of alterations (deletion/methylation) of some HRR genes has been seen in BC of NACT-treated patients in spite of low proliferation index and stem cell prevalence [12]. It seems that epigenetics may play an important role in modulating cancer cells to acquire drug tolerance [15, 16].
In NACT treatment of BC patients, doxorubicin is mainly used along with other chemotherapeutic drugs [17]. Doxorubicin is a common anthracycline anti-tumor antibiotic with DNA intercalation property and inhibits nucleic acid synthesis as well as topoisomerase II [18]. Like doxorubicin, nogalamycin is another anthracycline anti-tumor antibiotic with severe toxicity and also inhibits nucleic acid biosynthesis [18]. It has been reported that doxorubicin (1 µM) could inhibit the expression of BRCA1/BRCA2 in different BC cell lines [19]. In addition, it could induce apoptosis in a BC cell line MCF-7 [20]. On the other hand, increased expression of the HRR protein like BRIT1 and foci formation at the DNA damaged site was found in MCF-7 after exposure of ionization radiation [10]. Thus, characterization of doxorubicin/nogalamycin-tolerant BC cells in vitro may mimic the NACT-tolerant BC cells in vivo.
For this reason, we have analyzed the association of the key regulatory genes of the HRR pathway with doxorubicin/nogalamycin-tolerant BC cell lines MCF-7 and MDA MB 231 at first, followed by validation in NACT-treated primary BC samples. In this study, the quantitative expression (RNA/Protein) and methylation profiles of the HRR genes were analyzed in the BC cell lines with increasing concentration of the drugs followed by correlation with the NACT samples. Then, the methylation patterns of the genes were correlated with DNMT1 expression in BC cell lines and samples. Our study showed that increased expression of the HRR genes due to their promoter hypomethylation and reduced expression of the DNMT1 were associated with the drug tolerance of BC cells.
Materials and methods
Cell culture
MCF-7 and MDA MB 231 cell lines were obtained from NCCS, Pune, India. MCF-7 is a luminal A type (ER+, PR+, HER2−) BC cell, whereas MDA MB 231 is of triple-negative (ER−, PR−, HER2−) type. MCF-7 and MDA MB 231 were grown, respectively, in RPMI 1640 (GIBCO, USA) and Leibovitz’s L-15 (Himedia, India) Medium, supplemented with 10% FBS and 1% penicillin/streptomycin at 37 °C, 5% CO2 according to the supplied protocol.
Treatment of chemotherapeutic drugs on BC Cell lines
The effect of anthracycline anti-tumor antibiotics doxorubicin and nogalamycin on MCF-7 and MDA MB 231 cell lines was analyzed by dye Sulforhodamine B (SRB) to do the drug kinetics and dose selection according to the standard procedure [21]. Doxorubicin was purchased from Sigma-Aldrich, USA and nogalamycin was a kind gift from Upjohn Company, USA. The drugs were dissolved in water to prepare a stock concentration and diluted with respective cell culture media as par the dose. Approximately, 5000 BC cells were plated in 96-well plate and allowed to attach to the bottom of the plate overnight. After 24 h of plating, cells were treated with increasing doses of doxorubicin and nogalamycin. Cells were incubated at 37 °C for 48 h and fixed with 50% chilled TCA for 45 min at 4 °C and dried at 37 °C overnight. Then, absorbance at 510 nm (A510) was taken in the presence of 0.4% SRB. The 50% inhibitory concentration (IC50) was calculated as described by Khafif et al. [22] from the mean OD values obtained in triplicate independent experiments. Similarly for further experiments, the cells were harvested after 48 h of the drug treatment and DNA/RNA/protein were isolated.
m-RNA expression analysis in BC cells
Total RNA was isolated from BC cell lines (MCF-7 and MDA MB 231) using TRIzOL reagent (Invitrogen, USA) according to manufacturer’s protocol. Purity of RNA was checked by spectrophotometry from A260/280 ratio and concentration was determined from A260 value.
Real-time quantification of the candidate TSGs was performed using a power SYBR-green assay (Applied Biosystems, USA) with β2-microglobulin as control. Each sample was loaded in triplicate and run at 40 cycles on an ABI prism 7500 machine (Applied Biosystem, USA). After each run, melting curves were generated to confirm amplification of specific transcripts. To determine the relative level of gene expression, the comparative threshold cycle (ddCT) method was employed after normalization of respective gene of interest expression level against that of B2M, (dCT) [23,24,25,26]. The relative gene expression of the genes in terms of fold expression change (2-ddC T) was calculated from the ddCT values obtained using the formula:
The primer sequences are listed in Supplementary Table S1a.
Protein expression analysis in BC cells
Protein expression was analyzed by western blot analysis and immunocytochemistry in MCF-7 and MDA MB 231 cell lines. For protein extraction and western blotting, MCF-7 and MDA MB 231 cells were lysed by sonication with RIPA buffer [21, 23, 24]. Proteins (80 µg) were separated by 10% SDS–PAGE and then transferred to polyvinylidene difluoride membrane (PVDF: Millipore, MA). To reduce non-specific binding, membranes were incubated with 3% non-fat dry milk for 1–2 h at room temperature. Membranes were then incubated overnight at 4 °C with desired primary antibodies (at dilution of 1:500–1:1000 in 1% non-fat dry milk) followed by corresponding HRP-conjugated secondary antibodies (1:2000–1:10,000 in 1% non-fat dry milk). The target protein bands were then visualized using luminal reagent and autoradiographed on X-ray film (Kodak, Rochester, NY). All the immunoblotting experiments were performed in triplicates. The band intensities were quantified using densitometric scanner (BioRad GS-800, Hercules, CA). Peak densities of the proteins of interest were normalized using peak density of loading control α-tubulin.
For immunocytochemistry, monolayer of cells were grown on sterile cover slips and fixed with methanol at − 20 °C. Then the cells were reacted with the same dilution of primary antibody of these proteins after permeabilization with 0.5% Triton X-100 and blocking with 5% BSA. After washing, the cover slips were incubated with fluorescein isothiocyanate (FITC)-conjugated corresponding secondary antibody at 1:500 dilutions and mounted with glycerol after thorough washing. Imaging of the cover slip was performed in florescence microscope (Leica DM4000 B, Germany) [21].
The primary antibodies used for BRCA1 (IMG-80317) was of Imagenex, India. Other antibodies like BRCA2 (H-299, sc-28235), FANCC (C-14, sc-18110), FANCD2 (H-300, sc-28194), MCPH1/BRIT1 (N-15, sc-48882), DNMT1 (H-300, sc-20701), PCNA (F-2, sc-25280), appropriate HRP-conjugated secondary antibodies, FITC-conjugated secondary antibodies, and DAPI of M/s Santa Cruz Biotechnology, Santa Cruz, CA, USA were used.
Demography of the clinical specimens
Freshly operated 12 pretherapeutic and 10 NACT-treated primary BC samples were collected from the hospital section of Chittaranjan National Cancer Institute (CNCI), Kolkata. NACT-treated samples were operated after 2–6 months of NACT treatment. All tumors were staged according to the International Union Against Cancer (UICC) tumor-node-metastasis (TNM) classification. Molecular subtyping of the samples based on estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER2) status was done by immunohistochemistry using specific antibodies sc-787, sc-7208, and sc-7301, respectively, of M/s Santa Cruz Biotechnology, Santa Cruz, CA, USA [12]. Scoring was done as per the recommended American Society of Clinical Oncology (ASCO) guidelines. Detailed clinicopathological information like stage/grade/nodal status/age of onset/molecular subtypes is listed in Supplementary Table S1b.
Microdissection and DNA extraction
The contaminant normal cells in the BC lesions were removed by microdissection from cryosections (5 µm) using surgical knives under a dissecting microscope (Leica MZ16, Germany) [27]. The representative sections from various regions of the samples were stained with hematoxylin and eosin for pathological diagnosis as well as for marking the tumor-rich regions. Microdissected samples containing > 60% tumor cells were taken for DNA isolation. The cells from the BC cell lines were collected after trypsinization and used for DNA isolation. High molecular weight DNA from the cells of BC cell lines and microdissected BC tumors were isolated by phenol–chloroform extraction [27, 28]. The quality and quantity of DNA were checked by measuring O.D at 260–280 nm [28].
Promoter methylation analysis
Promoter methylation status of the HRR genes was analyzed by PCR-based methylation-sensitive restriction analysis (MSRA) using HpaII (Promega, WI, USA), Msp1 (Sibenzyme, Russia), and Hha1 (Sibenzyme, Russia) enzymes [12, 29]. The β-3A adaptin gene (K1) and RARβ2 (K2) were used as digestion and integrity controls, respectively [12, 30]. The PCR was run for 35 cycles. The primer sequences are listed in Supplementary Table S1c.
Quantification of promoter methylation was done by real-time PCR using power SYBR-green assay (Applied Biosystems, USA) [31]. Each sample was loaded in triplicate and run at 40 cycles on an ABI prism 7500 machine (Applied Biosystem, USA). The percentage of hypomethylation was calculated by taking the ratio of CT values of the digested DNA sample with its respective undigested DNA, presuming the undigested DNA as 100% methylated. The percentage of hypomethylation is a comparative term which was plotted as the differences in the methylation percentages among the samples. The higher the percentage of hypomethylation, the lower the frequency of methylation in the sample and vice versa. The percentage of hypomethylation of particular gene was calculated using the following formula:
\(\left\{ {{\text{CT}}\left( {{\text{digested}}} \right)/{\text{CT}}\left( {{\text{undigested}}} \right) \times 100} \right\} - 100.\)
Immunohistochemical analysis in BC samples
The protein expression of DNMT1 was analyzed in pretherapeutic and NACT-treated BC samples by immunohistochemistry. To this end, appropriate HRP-conjugated secondary antibodies and Diaminobenzidine (sc-24982) (Santa Cruz Biotechnology, CA, USA) were used for color development and hematoxylin was used as counterstain. The detailed procedure and scoring was described by Perrone et al. [32]. The staining patterns of the proteins in primary tumors were compared to the staining patterns of the ducts of normal samples.
Statistical analysis
T test was performed to compare the m-RNA/protein expression and percentages of hypomethylation in BC cells and percentages of hypomethylation and DNMT1 expression between pretherapeutic and NACT-treated samples. P < 0.05 was considered to be statistically significant.
Results
Effect of drugs on viability of BC cell lines
The viability of cells of the MCF-7 and MDA MB 231 cell lines was done at different concentrations of doxorubicin and nogalamycin (Fig. 1a, b). It was evident that IC50 concentrations of the drugs were comparatively lower in MCF-7 (doxorubicin: 0.214 µM and nogalamycin: 0.242 µM) than in MDA MB 231 (doxorubicin: 0.346 µM and nogalamycin: 0.370 µM).
a–d Effect of anthracycline drugs on the cell viability and proliferation of BC cell lines. a, b Cell viability assay after treatment with doxorubicin and nogalamycin. a MCF-7: b MDA MB 231. The x-axis shows the doses of drugs in µM. The line denotes 50% cell viability. c, d Representative Immunocytochemical patterns of PCNA proteins in untreated control cells and cells treated with IC50 dose of doxorubicin/nogalamycin. c MCF-7: d MDA MB 231
Expression of PCNA in the BC cell lines at IC50 concentration of drugs
It was evident that at IC50 concentration of the drugs the nuclear expression of PCNA was found to be lower compared to the control (Fig. 1c, d). It indicates that the drugs reduce the proliferative potential of the BC cells.
Analysis of expression of the HRR pathway genes in the presence of drugs in BC cell lines
m-RNA expression
The effect of the drugs (doxorubicin/nogalamycin) on m-RNA expression of the HRR genes was analyzed by treating three different doses (lower to IC50 dose, IC50 dose, and higher to IC50 dose) of the drugs having cell viability between 68 and 41% in MCF-7 and MDA MB 231 cell lines. It was evident that the drugs could differentially increase the expression of HRR genes in the cell lines. In MCF-7, the expression of the HRR genes were increased (2.065–17.595-fold) at lower concentration of IC50 (0.1 µM) with about 53% cell viability followed by significantly reduced or comparable expression (0.207–11.073-fold) at IC50 and higher IC50 (0.4 µM) concentrations (41–45% cell viability) (P = 0.001–0.044) (Figs. 2a, b, Supplementary Fig. S1a, b). Similarly, in MDA MB 231 cells the expression of the genes was high (0.138–6.699-fold) at lower concentration of IC50 (0.2 µM) with about 60–68% cell viability (Fig. 2c, d, Supplementary Fig. S1a, b). The expression became comparable or significantly increased/decreased (0.361–6.776-fold) at IC50 and higher IC50 (0.6 µM) concentrations (46–48% cell viability) (P = 0.0009–0.015) (Fig. 2c, d).
a–d Effect of anthracycline drugs on the m-RNA expression of HRR genes in BC cell lines. a, b Histogram of m-RNA expression fold changes in MCF-7 cell line after treatment with three different doses of doxorubicin/nogalamycin. a Doxorubicin: b Nogalamycin. c, d Histogram m-RNA expression fold changes in MDA MB 231 cell line after treatment with three different doses of doxorubicin/nogalamycin. c Doxorubicin: d Nogalamycin. *Denotes significant difference. P < 0.05 was considered to be statistically significant
Western blot analysis
In western blot analysis, the expression of HRR proteins in the presence of the drugs showed concordance with m-RNA expression, and in most cases a significant increase in expression was observed after drug treatment compared to the control (P = 0.0002–0.01) (Fig. 3a, b). In MCF-7, the protein expressions were increased in lower IC50 concentration (1.236–5.132 times) followed by comparable or increase/decrease expression (1.019–5.423 times) in IC50 and higher IC50 concentration of the drugs (Fig. 3b). In MDA MB 231 cells, the expression of these proteins was mostly highest (1.063–4.484 times) in lower IC50 concentration of the drugs with comparable or slightly low expression in IC50 (1.168–2.859 times) and higher IC50 concentration (1.054–2.695 times) except FANCC protein after treatment with nogalamycin, where the highest increase in expression was shown by IC50 concentration (Fig. 3b).
a, b Effect of anthracycline drugs on the protein expression of HRR genes in BC cell lines. a Western blot analysis of HRR proteins in MCF-7 and MDA MB 231 cell line. b Histogram of mean peak densities of the western blot analysis depicting the protein expressions of HRR proteins in MCF-7 and MDA MB 231 cells normalized with the loading control α-tubulin. * Denotes significant difference. P < 0.05 was considered to be statistically significant. Lane-wise description of the samples: (1) Untreated control BC cells, (2) lower IC50 DX-treated cells, (3) IC50 DX-treated cells, (4) higher IC50 DX-treated cells, (5) lower IC50 NG-treated cells, (6) IC50 NG-treated cells, (7) higher IC50 NG-treated cells. DX doxorubicin, NG nogalamycin
Immunocytochemical expression
Similar to the western blot analysis, immunocytochemical expressions of the HRR genes were high at the lower IC50 concentration of the drugs which gradually become reduced or remained comparable in the IC50 and the higher IC50 concentration (Fig. 4a–e). In MCF-7 cells, the expressions of HRR proteins were predominant in nuclear membrane with diffused nuclear and cytoplasmic expression (Fig. 4a–e). Among the HRR proteins, predominant nuclear expression of FANCD2 was observed (Fig. 4d). Interestingly, high cytoplasmic and nuclear expression of the HRR proteins was seen at the lower IC50 concentration of the drugs (Fig. 4a–e). However, the expression of the proteins was reduced at IC50 and at the higher IC50 concentrations compared to the lower IC50 concentration of the drugs with predominant localization at the nuclear membrane (Fig. 4a–e). Similar patterns of expression were also found In MDA MB 231 cells (data not shown).
a–e Effect of anthracycline drugs on the protein expression and localization of HRR genes in BC cell lines. a–e Representative Immunocytochemical patterns of HRR proteins in control MCF-7 cells and MCF-7 cells with treatment of three different doses of doxorubicin/nogalamycin. a BRCA1: b BRCA2: c FANCC: d FANCD2: e BRIT1. Magnifications × 40, Scale bar = 50 µm
Analysis of promoter methylation of the HRR pathway genes in the presence of drugs in BC cell lines
To understand the molecular mechanism behind the increased expression of HRR pathway genes in the presence of the drugs (doxorubicin/nogalamycin), the promoter methylation status of the genes was analyzed. No qualitative differences in methylation patterns of the genes were seen due to treatment of the drugs compared to the respective controls (Fig. S2a-e). Interestingly, in quantitative methylation analysis of the genes, mostly significant increased frequencies of hypomethylation were observed in BC cell lines after drug treatment (P < 0.0001) (Fig. 5a–d). In MCF-7, increased frequencies of hypomethylation were observed at the lower IC50 (0.1–0.2 µM) concentrations of the drugs irrespective of the restriction enzymes used (Fig. 5a–d). However, with increased drug concentration (IC50 and higher IC50), the frequencies of hypomethylation of the genes were comparable or higher than control (Fig. 5a–d). In case of MDA MB 231, the frequencies of hypomethylation of the genes were comparable or higher at the lower IC50 concentration of the drugs than the control (Fig. 5e–f). Similar to MCF-7, comparable or higher frequencies of hypomethylation of the genes were seen in IC50 and higher IC50 concentrations than the control (Fig. 5e–f).
a–h Quantitative methylation analysis of HRR genes in BC cell lines after treatment with anthracycline drugs. a–h Representative histogram of qRT-PCR analysis showing increased percentage of hypomethylation of HRR genes in MCF-7 and MDA MB 231 cells after treatment with three different doses of doxorubicin and nogalamycin while digested with HpaII and HhaI enzymes. a Doxorubicin-treated HpaII digested MCF-7 cells: b doxorubicin-treated HhaI digested MCF-7 cells: c nogalamycin-treated HpaII digested MCF-7 cells: d nogalamycin-treated HhaI digested MCF-7 cells: e doxorubicin-treated HpaII digested MDA MB 231 cells: f doxorubicin-treated HhaI digested MDA MB 231 cells: g nogalamycin-treated HpaII digested MDA MB 231 cells: h nogalamycin-treated HhaI digested MDA MB 231 cells. DX doxorubicin, NG nogalamycin. *Denotes significant difference. P < 0.05 was considered to be statistically significant
Quantification of hypomethylation profile of the HRR genes in primary BC samples
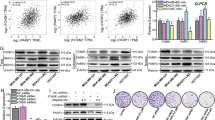
For validation of the hypomethylation profile of the HRR genes in the drug-tolerant BC cell lines, quantitative hypomethylation profiles of the genes were analyzed in pretherapeutic (n = 12) and NACT-treated (n = 10) BC samples, though no qualitative difference in methylation profile has been seen in the two groups of sample as seen in our previous study Dasgupta et al. [12]. Like the drug-tolerant BC cell lines, hypomethylation profiles of the HRR genes were significantly higher (P value = 0.003–0.048) in the NACT-treated BC samples than in pretherapeutic BC samples (Fig. 6a–e). This was concordant with the methylation status of the genes (Supplementary Table S2).
a–e Quantitative methylation analysis of HRR genes in pretherapeutic and NACT-treated BC samples. a–e Representative histogram of qRT-PCR analysis showing percentage of hypomethylation of HRR genes in pretherapeutic and NACT-treated BC samples while digested with HpaII enzyme. a BRCA1: b BRCA2: c FANCC: d FANCD2: e BRIT1. The line denotes the median percentage of hypomethylation
Expression of DNMT1 in the BC cells in the presence of the drugs
To understand the mechanism of increased hypomethylation of the HRR genes in the BC cell lines in the presence of the drugs, the m-RNA and protein expressions of DNMT1 (DNA methyltransferase 1) were analyzed (Fig. 7), due to its (DNMT1) importance in maintenance of tissue-specific methylation of cytosine residues [33]. It was evident that m-RNA expression of DNMT1 was considerably reduced at the lower IC50 concentration of the drugs in the cell lines than the controls (Fig. 7a, Supplementary Fig. S3). Similar pattern of expression of DNMT1 was observed at IC50/ higher IC50 concentrations of the drugs (Fig. 7a, Supplementary Fig. S3). At higher IC50 concentration of nogalamycin, the expression was significantly increased in MDA MB 231 cells than lower IC50 and IC50 concentration (P = 0.0002, 0.0003) (Fig. 7).
a–d Expression analysis of DNMT1 gene in BC cell lines. a Histogram of m-RNA expression fold changes of DNMT1 gene in MCF-7 and MDA MB 231 cell lines after treatment with three different doses of doxorubicin/nogalamycin. b Western blot analysis of DNMT1 protein in MCF-7 and MDA MB 231 cell line. c Histogram of mean peak densities of the western blot analysis depicting the protein expressions of DNMT1 protein in MCF-7 and MDA MB 231 cells normalized with the loading control α-tubulin. Lane-wise description of the samples: (1) Untreated control BC cells, (2) lower IC50 DX-treated cells, (3) IC50 DX-treated cells, (4) higher IC50 DX-treated cells, (5) lower IC50 NG-treated cells, (6) IC50 NG-treated cells, (7) higher IC50 NG-treated cells. DX doxorubicin, NG nogalamycin. *Denotes significant difference. P < 0.05 was considered to be statistically significant. (d) Representative immunocytochemical patterns of DNMT1 proteins in control MCF-7 cells and MCF-7 cells with treatment of three different doses of doxorubicin/nogalamycin. Magnifications × 40, Scale bar = 50 µm
The western blot analysis of DNMT1 protein showed concordance with the m-RNA expression patterns, and a significant decrease in expression was observed after drug treatment compared to the control (P = 0.0009–0.003) (Fig. 7b). In both the cell lines, the expression of DNMT1 protein was reduced to 0.166–0.79 times from the respective controls (Fig. 7c). In immunocytochemical analysis, reduced nuclear expression of DNMT1 was seen in the cell lines at the different concentrations of the drugs (Fig. 7d).
Expression of DNMT1 in pretherapeutic and NACT-treated BC samples
The protein expression of DNMT1 was analyzed in 35 (19 pretherapeutic and 16 NACT-treated) BC samples by immunohistochemistry to validate our in vitro data in vivo (Fig. 8a–h). In normal breast ducts, mostly nuclear expression of DNMT1 was present in 9–13% myoepithelial and luminal epithelium cells (Fig. 8b). In pretherapeutic samples, nuclear expression of DNMT1 was mostly moderate to high and found in 21–92% (mean 58%) BC cells in the samples (Fig. 8c–e). However, in NACT-treated BC, the expression was low to moderate, i.e., 5–51% (mean 25%) (Fig. 8f–h). The nuclear expression of DNMT1 was significantly higher (P = 0.009) in pretherapeutic samples than in NACT-treated samples.
a–h Protein expression of DNMT1 gene by immunohistochemistry in pretherapeutic and NACT-treated BC samples. a, b Representative Immunohistochemical pattern of DNMT1 protein in BC: a negative stain (#812T), b expression in luminal and myoepithelial cells of adjacent normal (#5000N). c–e Expression of DNMT1 in pretherapeutic samples: c representative moderate expression (#1904T), d representative high expression (#3503T), e histogram of percentage of expression positive cells in the samples. The line denotes the average percentage of expression. f–h Expression of DNMT1 in NACT-treated samples: f representative moderate expression (#6632T), g representative low expression (#1656T), h histogram of percentage of expression positive cells in the samples. The line denotes the average percentage of expression. Scale bar = 50 µm, magnification × 40
Discussion
The present study was undertaken to understand the importance of HRR pathway in development of chemotolerance in BC. For this reason, firstly expression (mRNA/protein) and methylation profile of some of the key regulatory genes of HRR pathway, i.e., BRCA1, BRCA2, FANCC, FANCD2, and BRIT1 were analyzed in two BC cell lines MCF-7 and MDA MB 231 in the presence of anthracycline group of drugs doxorubicin and nogalamycin.
The MCF-7 cells showed more sensitivity to the drugs due to low IC50 value compared to MDA MB 231. This might be due to the higher differentiation status of MCF-7, i.e., luminal A type, whereas to a triple-negative type with lower differentiation profile for MDA MB 231 [34]. However, at the IC50 concentration of the drugs, reduced expression of PCNA was seen in both the cell lines indicating inhibition of cellular proliferation by the drugs. Similarly, reduced PCNA expression has also been seen in NACT-treated BC samples [12, 35].
The expression (m-RNA/protein) of the HRR genes was low in the BC cell lines. Interestingly, expression (m-RNA/protein) of the HRR genes was increased at the lower IC50 concentrations of the drugs in MCF-7 (0.1 µM) and MDA MB 231 (0.2 µM) cells with about 53 and 60–68% cell viability, respectively. The protein expression of these genes was predominantly seen in the nucleus. This indicates that the sub-lethal doses of the drugs might induce genotoxic stress leading to the increased expression of the HRR genes irrespective of the differentiation status of BC cell lines. Similar phenomenon was also observed in MCF-7 after sub-lethal doses of gamma radiation with increased BRIT1 expression and nuclear foci formation [10]. Anticancer agents like isoflavones (phytoestrogens) increase the expression of HRR genes in mouse mammary glands and primary BC [36], and downregulation of HRR makes the glioma cells more sensitive to methylating agents like temozolomide [37]. This indicates that the increased expression of the HRR genes with nuclear localization seems to be needed for efficient DNA repair of the survived tumor cells. However, at IC50 and the higher IC50 concentration of the drugs, differential pattern of expression of the genes was seen in the cell lines. In MCF-7, majority of the genes showed low expression at IC50/higher concentrations compared to that of the lower IC50 concentration. In MDA MB 231, majority of the genes showed comparable/high expression at IC50/higher concentrations compared to the lower IC50 concentration. This indicates that the differences in expression profile of the HRR genes in the two cell lines at higher concentration of the drugs might be due to differences in differentiation status [38]. Importantly, the protein expression of the genes at the higher concentration of the drugs was concordant with m-RNA expression but localized gradually in the nuclear periphery. The mechanism behind the translocation of the proteins is not known. This might be due to some post translational modifications or blockage in nuclear transport of these proteins. It was evident that these drugs could inhibit the nuclear transport of Usn RNAs [39].
To find the reason behind the increased expression of HRR proteins in the chemotolerant BC cells, their promoter methylation status (qualitative/quantitative) of the genes was analyzed. In qualitative methylation analysis, no differences in methylation pattern of the genes were seen between untreated and drug-treated cells [12]. However, in quantitative methylation analysis, promoter hypomethylation of these genes was prevalent in different concentrations of the drugs and found to be concordant with mRNA/protein expression of the genes. Similar phenomenon of higher promoter hypomethylation of the genes was also evident in NACT-treated BC samples than in pretherapeutic samples. Similar phenomenon of hypomethylation was also evident in BRCA1/BRCA2 genes in MCF-7 and MDA MB 231 cells after treatment with Equol, a metabolite of anticancer phytoestrogens daidzein [40]. Likewise, global hypomethylation was reported in rat mammary glands after gamma radiation [41]. On the other hand, anthracycline-based therapies showed better response in BC cells with BRCA1 methylation [42].
It seems that the hypomethylation of the genes in the presence of the drugs might be due to the inhibition/reduced expression of the DNA methylating enzymes. We have seen reduced DNMT1 m-RNA and protein expression in MCF-7 and MDA MB 231 cell lines after drug treatment. Doxorubicin inhibits DNMT1 in colon carcinoma cell line HCT116 [43]. It was evident that epigenetic modifications of chromatin could modulate the drug response of the tumor cells, as derivatives of valproic acid, an inhibitors of the class I histone deacetylases (HDAC-I), induce the sensitivity of lung cancer cells to cisplatin [44].
Conclusion
Thus, our study indicates that the chemotolerance of the BC cells in the presence of the anthracycline anti-tumor antibiotics doxorubicin and nogalamycin might be due to the increased expressions of HRR genes through promoter hypomethylation and reduced DNMT1 expression. However, detailed analysis in the epigenetic modifications of chromatin in the presence of the drugs is warranted to understand the molecular mechanism of drug tolerance of this tumor.
Abbreviations
- BC:
-
Breast carcinoma
- ER:
-
Estrogen receptor
- PR:
-
Progesterone receptor
- HER2:
-
Human epidermal growth factor receptor 2
- NACT:
-
Neoadjuvant chemotherapy
- HRR:
-
Homologous recombination repair
- UICC:
-
International Union Against Cancer
- TNM:
-
Tumor size, lymph node, metastasis
- MSRA:
-
Methylation-sensitive restriction analysis
- qRT-PCR:
-
Real-time PCR quantification
- DNMT1:
-
DNA methyltransferase 1
- PCNA:
-
Proliferating cell nuclear antigen
- HRP:
-
Horse-radish-peroxidase
- FITC:
-
Fluorescein isothiocyanate
References
Ochayon L, Tunin R, Yoselis A, Kadmon I (2014) Symptoms of hormonal therapy and social support: is there a connection? Comparison of symptom severity, symptom interference and social support among breast cancer patients receiving and not receiving adjuvant hormonal treatment. Eur J Oncol Nurs 14:192–196
Sinha S, Chunder N, Mukherjee N, Alam N, Roy A, Roychoudhury S, Panda CK (2008) Frequent deletion and methylation in SH3GL2 and CDKN2A loci are associated with early- and late-onset breast carcinoma. Ann Surg Oncol 15:1070–1080
Gampenrieder SP, Rinnerthaler G, Greil R (2013) Neoadjuvant chemotherapy and targeted therapy in breast cancer: past, present, and future. J Oncol 2013:732047
Asakawa H, Koizumi H, Koike A, Takahashi M, Wu W, Iwase H, Fukuda M, Ohta T (2010) Prediction of breast cancer sensitivity to neoadjuvant chemotherapy based on status of DNA damage repair proteins. Breast Cancer Res 12:R17
Kriege M, Seynaeve C, Meijers-Heijboer H, Collee JM, Menke-Pluymers MB, Bartels CC, Tilanus-Linthorst MM, Blom J, Huijskens E, Jager A, van den Ouweland A, van Geel B, Hooning MJ, Brekelmans CT, Klijn JG (2009) Sensitivity to first-line chemotherapy for metastatic breast cancer in BRCA1 and BRCA2 mutation carriers. J Clin Oncol 27:3764–3771
Kennedy RD, D’Andrea AD (2006) DNA repair pathways in clinical practice: lessons from pediatric cancer susceptibility syndromes. J Clin Oncol 24:3799–3808
Mulligan JM, Hill LA, Deharo S, Irwin G, Boyle D, Keating KE, Raji OY, McDyer FA, O’Brien E, Bylesjo M, Quinn JE, Lindor NM, Mullan PB, James CR, Walker SM, Kerr P, James J, Davison TS, Proutski V, Salto-Tellez M, Johnston PG, Couch FJ, Paul Harkin D (2014) Identification and validation of an anthracycline/cyclophosphamide-based chemotherapy response assay in breast cancer. J Natl Cancer Inst 106:djt335
Rassool FV, Tomkinson AE (2010) Targeting abnormal DNA double strand break repair in cancer. Cell Mol Life Sci 67:3699–3710
Bhattacharya N, Mukherjee N, Singh RK, Sinha S, Alam N, Roy A, Roychoudhury S, Panda CK (2013) Frequent alterations of MCPH1 and ATM are associated with primary breast carcinoma: clinical and prognostic implications. Ann Surg Oncol 3:S424–S432
Rai R, Dai H, Multani AS, Li K, Chin K, Gray J, Lahad JP, Liang J, Mills GB, Meric-Bernstam F, Lin SY (2006) BRIT1 regulates early DNA damage response, chromosomal integrity, and cancer. Cancer Cell 10:145–157
Chunder N, Mandal S, Roy A, Roychoudhury S, Panda CK (2004) Differential association of BRCA1 and BRCA2 genes with some breast cancer-associated genes in early and late onset breast tumors. Ann Surg Oncol 11:1045–1055
Dasgupta H, Mukherjee N, Islam S, Bhattacharya R, Alam N, Roy A, Roychoudhury S, Biswas J, Panda CK (2017) Frequent alterations of HRR pathway in primary and chemotolerant breast carcinomas: clinical importance. Future Oncol 13:159–174
Sinha S, Singh RK, Alam N, Roy A, Roychoudhury S, Panda CK (2008) Alterations in candidate genes PHF2, FANCC, PTCH1 and XPA at chromosomal 9q22.3 region: pathological significance in early- and late-onset breast carcinoma. Mol Cancer 7:84
van der Groep P, Hoelzel M, Buerger H, Joenje H, de Winter JP, van Diest PJ (2008) Loss of expression of FANCD2 protein in sporadic and hereditary breast cancer. Breast Cancer Res Treat 107:41–47
Sharma SV, Lee DY, Li B, Quinlan MP, Takahashi F, Maheswaran S, McDermott U, Azizian N, Zou L, Fischbach MA, Wong KK, Brandstetter K, Wittner B, Ramaswamy S, Classon M, Settleman J (2010) A chromatin-mediated drug-tolerant state in cancer cell subpopulations. Cell 141:69–80
Glasspool RM, Teodoridis JM, Brown R (2006) Epigenetics as a mechanism driving polygenic clinical drug resistance. Br J Cancer 94:1087–1092
Trudeau M, Charbonneau F, Gelmon K, Laing K, Latreille J, Mackey J, McLeod D, Pritchard K, Provencher L, Verma S (2005) Selection of adjuvant chemotherapy for treatment of node-positive breast cancer. Lancet Oncol 6:886–898
Siitonen V, Selvaraj B, Niiranen L, Lindqvist Y, Schneider G, Metsä-Ketelä M (2016) Divergent non-heme iron enzymes in the nogalamycin biosynthetic pathway. Proc Natl Acad Sci 113:5251–5256
Andres JL, Fan S, Turkel GJ, Wang JA, Twu NF, Yuan RQ, Lamszus K, Goldberg ID, Rosen EM (1998) Regulation of BRCA1 and BRCA2 expression in human breast cancer cells by DNA-damaging agents. Oncogene 16:2229–2241
Chen JS, Konopleva M, Andreeff M, Multani AS, Pathak S, Mehta K (2004) Drug-resistant breast carcinoma (MCF-7) cells are paradoxically sensitive to apoptosis. J Cell Physio 200:223–234
Sur S, Pal D, Banerjee K, Mandal S, Das A, Roy A, Panda CK (2015) Amarogentin regulates self renewal pathways to restrict liver carcinogenesis in experimental mouse model. Mol Carcinog 55:1138–1149
Khafif A, Schantz SP, Chou TC, Edelstein D, Sacks PC (1998) Quantitation of chemopreventive synergism between (-)-epigallocatechin-3-gallate and curcumin in normal, premalignant and malignant human oral epithelial cells. Carcinogenesis 19:419–424
Veroni C, Marnetto F, Granieri L, Bertolotto A, Ballerini C, Repice AM, Schirru L, Coghe G, Cocco E, Anastasiadou E, Puopolo M, Aloisi F (2015) Immune and Epstein-Barr virus gene expression in cerebrospinal fluid and peripheral blood mononuclear cells from patients with relapsing-remitting multiple sclerosis. J Neuroinflammation 14:12:132
Di Napoli A, Al-Jadiri MF, Talerico C, Duranti E, Pilozzi E, Trivedi P, Anastasiadou E, Alsaadawi AR, Al-Darraji AF, Al-Hadad SA, Testi AM, Uccini S, Ruco L (2013) Epstein-Barr virus (EBV) positive classical Hodgkin lymphoma of Iraqi children: an immunophenotypic and molecular characterization of Hodgkin/Reed-Sternberg cells. Pediatr Blood Cancer 60:2068–2072
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108
Dasgupta S, Mukherjee N, Roy S, Roy A, Sengupta A, Roychowdhury S, Panda CK (2002) Mapping of the candidate tumor suppressor genes’ loci on human chromosome 3 in head and neck squamous cell carcinoma of an Indian patient population. Oral Oncol 38:6–15
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor
Loginov VI, Maliukova AV, SereginIu A, Khodyrev DS, Kazubskaia TP, Ermilova VD, Gar’kavtseva RF, Kiselev LL, Zabarovskii ER, Braga EA (2004) Methylation of the promoter region of the RASSF1A gene, a candidate tumor suppressor, in primary epithelial tumors. MolBiol (Mosk) 38:654–667
Ivanova T, Petrenko A, Gritsko T, Vinokourova S, Eshilev E, Kobzeva V, Kisseljov F, Kisseljova N (2002) Methylation and silencing of the retinoic acid receptor-beta 2 gene in cervical cancer. BMC Cancer 2:4
Thomassin H, Kress C, Grange T (2004) MethylQuant: a sensitive method for quantifying methylation of specific cytosines within the genome. Nucleic Acids Res 32:e168
Perrone F, Suardi S, Pastore E, Casieri P, Orsenigo M, Caramuta S, Dagrada G, Losa M, Licitra L, Bossi P, Staurengo S, Oggionni M, Locati L, Cantu G, Squadrelli M, Carbone A, Pierotti MA, Pilotti S (2006) Molecular and cytogenetic subgroups of oropharyngeal squamous cell carcinoma. Clin Cancer Res 12:6643–6651
Klein CJ, Botuyan MV, Wu Y, Ward CJ, Nicholson GA, Hammans S, Hojo K, Yamanishi H, Karpf AR, Wallace DC, Simon M, Lander C, Boardman LA, Cunningham JM, Smith GE, Litchy WJ, Boes B, Atkinson EJ, Middha S, Dyck B, Parisi PJ, Mer JE, Smith G, Dyck DI PJ (2011) Mutations in DNMT1 cause hereditary sensory neuropathy with dementia and hearing loss. Nat Genet 43:595–600
Perou CM (2010) Molecular stratification of triple-negative breast cancers. Oncologist 16:61–70
Alfaro Y, Delgado G, Carabez A, Anguiano B, Aceves C (2013) Iodine and doxorubicin, a good combination for mammary cancer treatment: antineoplastic adjuvancy, chemoresistance inhibition, and cardioprotection. Mol Cancer 12:45
Vissac-Sabatier C, Bignon YJ, Bernard-Gallon DJ (2003) Effects of the phytoestrogens genistein and daidzein on BRCA2 tumor suppressor gene expression in breast cell lines. Nutr Cancer 45:247–255
Chai KM, Wang CY, Liaw HJ, Fang KM, Yang CS, Tzeng SF (2014) Downregulation of BRCA1-BRCA2-containing complex subunit 3 sensitizes glioma cells to temozolomide. Oncotarget 5:10901–10915
Liedtke S, Biebernick S, Radke TF, Stapelkamp D, Coenen C, Zaehres H, Fritz G, Kogler G (2015) DNA damage response in neonatal and adult stromal cells compared with induced pluripotent stem cells. Stem Cells Transl Med 4:576–589
Ray R, Chakraborty BK, Ray K, Mukherji S, Chowdhury JR, Panda CK (1996) Effect of anthracycline antitumor antibiotics (adriamycin and nogalamycin) and cycloheximide on the biosynthesis and processing of major UsnRNAs. Mol Cell Biochem 162l:75–82
Bosviel R, Durif J, Déchelotte P, Bignon YJ, Bernard-Gallon D (2012) Epigenetic modulation of BRCA1 and BRCA2 gene expression by equol in breast cancer cell lines. Br J Nutr 108:1187–1193
Takabatake M, Blyth BJ, Daino K, Imaoka T, Nishimura M, Fukushi M, Shimada Y (2016) DNA methylation patterns in rat mammary carcinomas induced by pre-and post-pubertal irradiation. PLoS ONE 11:e0164194
Ignatov T, Poehlmann A, Ignatov A, Schinlauer A, Costa SD, Roessner A, Kalinski T, Bischoff J (2013) BRCA1 promoter methylation is a marker of better response to anthracycline-based therapy in sporadic TNBC. Breast Cancer Res Treat 141:205–212
Yokochi T, Robertson KD (2004) Doxorubicin inhibits DNMT1, resulting in conditional apoptosis. Mol Pharmacol 66:1415–1420
Tesei A, Brigliadori G, Carloni S, Fabbri F, Ulivi P, Arienti C, Sparatore A, Del Soldato P, Pasini A, Amadori D, Silvestrini R, Zoli W (2012) Organosulfur derivatives of the HDAC inhibitor valproic acid sensitize human lung cancer cell lines to apoptosis and to cisplatin cytotoxicity. J Cell Physiol 227:3389–3396
Acknowledgements
We thank the director of Chittaranjan National Cancer Institute, Kolkata, India. We are also thankful to the Upjohn Company, USA for gifting nogalamycin. Financial support for this work was provided by UGC-NET Fellowship Grant F.2-3/2000 (SA-I) (Sr. No. 2061030813, Ref. No.: 20-06/2010(i)EU-IV dated 22.10.2010) to H. Dasgupta.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There are no conflicts of interest.
Informed consent
Informed consent from the patients and approval from the Research Ethics Committee of the institute were obtained for sample collection.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11010_2018_3442_MOESM1_ESM.tif
Supplementary Figure S1: (a, b) Post real-time representative agarose gel images of m-RNA expression patterns of HRR genes. (a) Untreated and nogalamycin-treated MCF-7 cDNA, 1. untreated/control, 2. 0.1µM NG treated, 3. 0.242µM (IC50) NG treated, 4. 0.4µM NG treated: (b) Untreated and doxorubicin-treated MDA MB 231 cDNA, 1. untreated/control, 2. 0.2µM DX treated, 3. 0.346µM (IC50) DX treated, 4. 0.6µM DX treated. NG = Nogalamycin, DX = Doxorubicin. (TIF 883 KB)
11010_2018_3442_MOESM2_ESM.tif
Supplementary Figure S2: (a, b) Representative promoter methylation patterns of HRR genes after treatment with doxorubicin and nogalamycin in MCF-7 cells. (a) K1: (b) K2 (c) BRCA2: (d) FANCC (e) FANCD2. Lane-wise description of DNA: 1. Undigested, control, 2. HpaII digested, control, 3. HhaI digested, control, 4. Undigested, 0.1µM DX treated, 5. HpaII digested, 0.1µM DX treated, 6. HhaI digested, 0.1µM DX treated, 7. Undigested, 0.214µM (IC50) DX treated, 8. HpaII digested, 0.214µM (IC50) DX treated, 9. HhaI digested, 0.214µM (IC50) DX treated, 10. Undigested, 0.4µM DX treated, 11. HpaII digested, 0.4µM DX treated, 12. HhaI digested, 0.4µM DX treated, 13. Undigested, control, 14. HpaII digested, control, 15. HhaI digested, control, 16. Undigested, 0.1µM NG treated, 17. HpaII digested, 0.1µM NG treated, 18. HhaI digested, 0.1µM NG treated, 19. Undigested, 0.242µM (IC50) NG treated, 20. HpaII digested, 0.242µM (IC50) NG treated, 21. HhaI digested, 0.242µM (IC50) NG treated, 22. Undigested, 0.4µM NG treated, 23. HpaII digested, 0.4µM NG treated, 24. HhaI digested, 0.4µM NG treated. DX=Doxorubicin, NG=Nogalamycin. (TIF 1970 KB)
11010_2018_3442_MOESM3_ESM.tif
Supplementary Figure S3: Post real-time representative agarose gel images of m-RNA expression patterns of DNMT1 cDNA in BC cell lines after doxorubicin/nogalamycin treatment. Lane-wise description of the samples: 1. untreated/control, 2. lower IC50 treated, 3. IC50 treated, 4. higher IC50 treated. (TIF 809 KB)
11010_2018_3442_MOESM4_ESM.tif
Supplementary Table S1: (a) Primers for m-RNA expression analysis of HRR genes (b) Clinicopathological features of pretherapeutic and NACT-treated BC patients. (c) Primers for promoter methylation analysis of HRR genes. (TIF 2391 KB)
11010_2018_3442_MOESM5_ESM.tif
Supplementary Table S2: Concordance of Qualitative methylation status of HRR genes previously determined in pretherapeutic and NACT-treated samples (Dasgupta et al. 2017) with the quantitative methylation status (dCT). + = Methylation positive, − = Methylation negative, ND = Not determined. (TIF 1893 KB)
Rights and permissions
About this article
Cite this article
Dasgupta, H., Islam, M.S., Alam, N. et al. Induction of HRR genes and inhibition of DNMT1 is associated with anthracycline anti-tumor antibiotic-tolerant breast carcinoma cells. Mol Cell Biochem 453, 163–178 (2019). https://doi.org/10.1007/s11010-018-3442-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-018-3442-5