Abstract
Temozolomide (TMZ) is an alkylating agent used to treat glioblastoma. This tumor type synthesizes the antioxidant glutathione through system X −c , which is inhibited by sulfasalazine (SAS). We exposed A172 and T98G human glioblastoma cells to a presumably clinically relevant concentration of TMZ (25 µM) and/or 0.5 mM SAS for 1, 3, or 5 days and assessed cell viability. For both cell lines, TMZ alone did not alter viability at any time point, while the coadministration of TMZ and SAS significantly reduced cell viability after 5 days. The drug combination exerted a synergistic effect on A172 cells after 3 and 5 days. Therefore, this particular lineage was subjected to complementary analyses on the genetic (transcriptome) and functional (glutathione and proliferating cell nuclear antigen (PCNA) protein) levels. Cellular pathways containing differentially expressed genes related to the cell cycle were modified by TMZ alone. On the other hand, SAS regulated pathways associated with glutathione metabolism and synthesis, irrespective of TMZ. Moreover, SAS, but not TMZ, depleted the total glutathione level. Compared with the vehicle-treated cells, the level of PCNA protein was lower in cells treated with TMZ alone or in combination with SAS. In conclusion, our data showed that the association of TMZ and SAS is cytotoxic to T98G and A172 cells, thus providing useful insights for improving TMZ clinical efficacy through testing this novel drug combination. Moreover, the present study not only reports original information on differential gene expression in glioblastoma cells exposed to TMZ and/or SAS but also describes an antiproliferative effect of TMZ, which has not yet been observed in A172 cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glioblastoma is a high-grade, diffuse astrocytoma with a poor prognosis [1]. Temozolomide (TMZ), an alkylating agent, is the first-line treatment for glioblastoma patients. Adjuvant and concomitant TMZ administration with postoperative radiotherapy has been shown to lead to enhanced median and 5-year survival rates relative to those resulting from postoperative radiotherapy alone [2, 3]. However, acquired or intrinsic cellular resistance demonstrated by tumor cells may hamper TMZ efficacy [4, 5].
Many experimental approaches have been tested to combat glioblastoma cells. Aside from DNA alkylation by TMZ, the induction of oxidative stress is another strategy that has been investigated. Glutathione is part of an important intracellular antioxidant system that neutralizes reactive oxygen species through reactions involving such enzymes as glutathione peroxidase, reductase, and transferase [6]. Glutathione biosynthesis in astrocytes involves system X −c , which is a cytoplasmic membrane protein complex that imports cystine and releases glutamate. Cystine is then converted to cysteine, which is used for glutathione production [7]. This biosynthetic pathway of glutathione has also been described in primary cultures and in cultures of different glioblastoma cell lines [8]. Although present in normal glia, system X −c is overexpressed by and displays specific functions in gliomas [9]. In this context, sulfasalazine (SAS), which is currently used to treat inflammatory bowel disease, was found to inhibit system X −c in gliomas. In fact, SAS has been shown to decrease cell growth and induce apoptosis in primary cultures obtained from glioblastoma patients, as well as in other cell lines [7–11].
To the best of our knowledge, there are no data on the experimental in vitro exposure of glioma cells to TMZ associated with SAS. Therefore, we investigated whether combining TMZ, an alkylating agent, with SAS, an inhibitor of glutathione synthesis, would result in a synergistic inhibitory effect on A172 and T98G human glioblastoma cells in comparison with effects from the isolated administration of each drug. Our aim was to evaluate an approach for improving TMZ clinical efficacy by testing this hypothesis using cell lines commonly employed as in vitro glioblastoma models [4, 12].
Materials and methods
Cell lines and treatments
A172 and T98G human glioblastoma cells (American Type Culture Collection, Manassas, VA, USA) were cultured in Dulbecco’s modified Eagle’s medium containing 2 g/L glucose (DMEM, Vitrocell, cat# D0460, Campinas, SP, Brazil) supplemented with 10 % fetal bovine serum and 1 % penicillin/streptomycin (Vitrocell, cat# P0403) at 37 °C in a humidified atmosphere with 5 % CO2. One day after the cells were plated (10,000 cells/cm2), the culture medium was substituted with an equal volume of solution containing TMZ (25 µM; Sigma, cat# T2577, St Louis, MO, USA) and/or SAS (0.5 mM; Sigma, cat# S0883), in which the cells were cultured for 1, 3, or 5 days. In other to avoid pH changes due to cellular growth, the solution of each group of cells treated for 5 days was replaced by a new identical solution with an equal volume at the third day. Stock solutions of 50 mM TMZ [dissolved in dimethyl sulfoxide (DMSO; Sigma, cat# D2650)] or 20 mM SAS (solubilized in 0.2 M NaOH and then neutralized to pH 7.4 by titration with 0.2 M HCl) had been previously prepared. The cells were exposed to a final concentration of 0.1 % DMSO. The SAS concentration (0.5 mM) was chosen according to previous studies on glioma cells [10, 11] and pilot experiments that had been performed in our laboratory. Although SAS has been used in clinical practice for treating inflammatory illnesses, such as Crohn’s disease, no data on the corresponding in vitro concentrations were found. Regarding TMZ, 25 µM corresponds to a clinically relevant dose. In fact, based on pharmacokinetic studies performed by other researchers, 25 µM corresponds to the plasmatic concentration of an oral dose of 75 mg/m2, which is concomitantly used with radiation therapy to treat patients newly diagnosed with glioblastoma [13, 14].
Cell viability
After the cells were incubated in a 24-well plate, the medium was substituted with a MTT (Sigma, cat# M5655) solution (1 mg/mL in DMEM, without phenol (Vitrocell, cat# D0462); 250 µL/well) for 1.5 h (37 °C; 5 % CO2) [15]. Then, SDS acid solution was added (10 % SDS, 0.01 M HCl; 250 µL/well; 24 h) to dissolve the resulting formazan salts. The absorbance of the samples was read at 570 nm subtracting the value measured at 650 nm (PowerWave XS 2, BioTek Instruments, Winooski, VT, USA).
Transcriptome sequencing
Total RNA extraction and purification were performed with TRIzol (Thermo Fisher Scientific, cat# 15596018, Waltham, MA, USA). RNA (400 ng) was converted to a cDNA library using a TruSeq Stranded LT mRNA kit (Illumina, cat# 122-2103-RS, San Diego, CA, USA); the cDNA libraries were quantified by qPCR using primers specific for Illumina universal adapters. One unique identifier sequence was added for sample separation after each library was sequenced, thus allowing all samples to be sequenced during the same run and minimizing technical variations. Libraries were sequenced using a HiSeq 2500 platform (Illumina) in high-output mode, producing sequences of 2 × 100 nucleotides for each sequenced molecule. Sequence alignment was performed with TopHat2 (http://ccb.jhu.edu/software/tophat/index.shtml) to the Homo sapiens UCSC hg19 assembly. The average sequence alignment rate was 80 %.
Total glutathione levels
Total glutathione (GSH plus GSSG) was assessed through an enzymatic recycling assay (Cayman Chemical, cat# 703002, Ann Arbor, MI, USA). Cell lysate was prepared from each sample (60 cm2 dish) according to the manufacturer’s instructions, and total protein concentration was measured by using the Bradford colorimetric method (Sigma, cat# B6916) [16]. Kinetic absorbance was obtained at 405 nm (PowerWave XS 2).
Western blotting
Cells were lysed on ice using a sonicator (MISONIX, Sonicator® 3000, Farmingdale, NY, USA). For each sample, total protein (10 µg) was separated by electrophoresis on a 12 % SDS-polyacrylamide gel and electroblotted onto a 0.45 µm nitrocellulose membrane (Bio-Rad Laboratories, cat# 162-0115, Hercules, CA, USA) [17, 18]. The membranes were stained with Ponceau S solution (Sigma, cat# P7170), incubated with primary antibody against proliferating cell nuclear antigen (PCNA) (1:1000; BD Biosciences, cat# 610665, Franklin Lakes, NJ, USA) and with peroxidase-conjugated secondary antibody (1:10,000; BD Biosciences, cat# 554002, Franklin Lakes, NJ, USA). Immunoreactive bands were detected using a SuperSignal® West Pico chemiluminescence kit (Thermo Fisher Scientific, cat# 34080) and then quantified (ImageJ software, version 1.49). The optical density value of all bands stained with Ponceau S was used as internal control [19–21].
Statistical analyses
For the levels of cell viability, total glutathione, and PCNA, the data are presented as the mean value ± standard error of the mean. For multiple comparisons, one-way ANOVA was used, followed by the Bonferroni test (GraphPad Prism 5, version 5.00). The significance level was defined as p < 0.05.
HTSeqCount and DESeq 2 packages (http://www-huber.embl.de/users/anders/HTSeq/doc/overview.html and http://www.bioconductor.org/packages/release/bioc/html/DESeq2.html) were used for the transcriptome analyses. HTSeqCount estimates gene expression by counting sequences aligned to genome elements, such as exons and genes. DESeq 2 employs a negative-binomial distribution for data normalization and corrects for the effects of outliers and the contribution of genes with low expression to the analysis of variance. In addition, DESeq 2 uses differential expression statistical Wald test and corrects for multiple tests employing the Benjamini and Hochberg procedure. A list of differentially expressed genes was generated, and the enrichment of pathways for the set of differentially expressed genes was calculated using Metacore® (Thomas Reuters). The significance level was defined as p < 0.05 (after correction for multiple tests, i.e., the adjusted p value).
Results
SAS intensified the cytotoxic effect of TMZ on both T98G and A172 lineages
As regards viability of both T98G and A172 lineages at all time points, no statistical difference was detected after treatment with TMZ 25 µM compared to cells cultured in supplemented medium containing TMZ vehicle (DMSO 0.1 %). Moreover, the viability of cells exposed to 0.1 % DMSO was not different from that of cells only exposed to the supplemented DMEM (Figs. 1a, 2a). The observations of cellular density paralleled those of cell viability from day 1 to 5 (Figs. 1b, 2b).
Viability of T98G human glioblastoma cells after 1, 3, or 5 days of treatment with temozolomide (TMZ) and/or sulfasalazine (SAS). a Values of cell viability are expressed as a percentage of MTT reduction compared to that of cells treated with supplemented medium (DMEM group: 100 %); data are presented as the mean ± standard error of the mean of four independent experiments performed in triplicate. Data were analyzed using one-way ANOVA with the Bonferroni post hoc test; p < 0.05 was considered statistically significant. Letters indicate significantly different groups; a versus DMEM; b versus 0.1 % DMSO. b Cellular density demonstrating reduced viability after cotreatment with SAS and TMZ for 5 days. Representative images captured after 5 days. Scale bar = 100 µm (applies for all pictures)
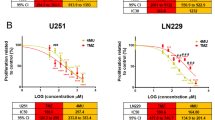
Viability of A172 human glioblastoma cells after 1, 3, or 5 days of treatment with temozolomide (TMZ) and/or sulfasalazine (SAS). a Values of cell viability are expressed as a percentage of MTT reduction compared to that of cells treated with supplemented medium (DMEM group: 100 %); data are presented as the mean ± standard error of the mean of four independent experiments performed in triplicate. Data were analyzed using one-way ANOVA with the Bonferroni post hoc test; p < 0.05 was considered statistically significant. Letters indicate significantly different groups; a versus DMEM; b versus 0.1 % DMSO; c versus 25 µM TMZ; d versus 0.5 mM SAS. b Cellular density reflecting reduced viability during the assessed period; lower confluence levels were observed after cotreatment with SAS and TMZ for 5 days. Representative images captured after 5 days. Scale bar = 100 µm (applies for all pictures)
Regarding A172 cells, treatment with SAS did not significantly alter cell viability after 1 day. However, the combination of TMZ and SAS resulted in lower cell viability than DMEM alone and containing 0.1 % DMSO. After 3 and 5 days, SAS alone and in combination with TMZ resulted in significantly lower viability than DMEM alone and containing 0.1 % DMSO. Moreover, after 3 and 5 days, the viability of the cells cotreated with TMZ and SAS was lower than that resulting from TMZ administered alone. Particularly, after 3 days of treatment, the viability of cells cotreated with TMZ and SAS was statistically lower than that of cells treated with SAS alone (Fig. 2a).
The effect (i.e., the percent reduction in cell viability compared to that of the DMSO group) of cotreating A172 cells with TMZ and SAS was found to be synergistic. In fact, after 3 and 5 days of treatment, the effect of the drug combination was higher than the arithmetic sum of the effects observed after the isolated administration of TMZ or SAS (Day 3: 25 µM TMZ = 9.0 %, 0.5 mM SAS = 33.2 %, 25 µM TMZ + 0.5 mM SAS = 64.0 %; Day 5: 25 µM TMZ = 7.6 %, 0.5 mM SAS = 52.7 %, 25 µM TMZ + 0.5 mM SAS = 76.3 %). On the other hand, after 1 day of treatment, although TMZ administered with SAS also reduced cell viability, the effect was not synergistic (Day 1: 25 µM TMZ = 19.2 %, 0.5 mM SAS = 24.0 %, 25 µM TMZ + 0.5 mM SAS = 41.4 %).
Regarding T98G cells, the coadministration of 25 µM TMZ with 0.5 mM SAS reduced cell viability; however, the effect was not synergistic at any time point. Specifically, compared with the 0.1 % DMSO treatment, there was a reduction in the viability of cells cotreated with 25 µM TMZ and 0.5 mM SAS after 1 and 5 days. The effect of the drug combination was lower than the arithmetic sum of the effects observed after the isolated administration of TMZ or SAS (Day 1: 25 µM TMZ = 6.5 %, 0.5 mM SAS = 11.8 %, 25 µM TMZ + 0.5 mM SAS = 14.6 %; Day 5: 25 µM TMZ = 13.9 %, 0.5 mM SAS = 26.4 %, 25 µM TMZ + 0.5 mM SAS = 39.8 %). No difference in viability was found among the groups after day 3 (Fig. 1).
In conclusion, decreases in A172 and T98G cell viability were not detected after treatment with the presumably clinically relevant dose of 25 µM TMZ. Conversely, reduced viability was noted after cotreatment with 25 µM TMZ and 0.5 mM SAS. These results indicate that the combination of these two drugs in vitro exerts a positive therapeutic effect. As we observed a more intense and synergistic effect in A172 cells, we proceeded with gene expression and functional analyses specifically in this cell line. Particularly, because the most intense viability reduction was detected after 5 days of treatment, we investigated alterations in mRNA expression (transcriptome) and total glutathione levels during the process of cellular demise (i.e., after 3 days of treatment). Proliferation (PCNA protein expression) was assessed in the remaining cells after the last time point (i.e., 5 days).
Transcriptome analyses
As mentioned above, we observed a more intense and synergistic effect of TMZ and SAS in A172 cells and decided to proceed with whole-transcriptome analysis via RNA sequencing specifically with this cell line. Indeed, this technique, which can be defined as the evaluation of all gene transcripts produced in a given cell line, enables the simultaneous quantification of the expression of thousands of genes [22]. Such a large-scale analysis of gene expression not only allowed us to infer the enriched biological pathways associated with these genes but also provided insights into the molecular mechanisms involved in cellular responses to the treatments. These mechanisms may be assessed in future studies to develop targeted glioblastoma therapies.
Based on the MTT data, cells treated with 25 µM TMZ alone or in conjunction with 0.5 mM SAS (both solutions containing 0.1 % DMSO) and cells exposed to medium with 0.1 % DMSO were selected for transcriptome analyses. Sequencing provided a total of 162,248,078 paired-end 100 bp reads (~80 % >Q30; ~10 Mi reads/sample). A list of significantly differentially expressed genes was generated using the DESeq 2 pipeline (Online Resource 1). Comparisons between the 0.1 % DMSO and TMZ groups indicated 1018 differentially expressed genes. Groups receiving SAS without or with TMZ showed 575 and 2368 differentially expressed genes (vs 0.1 % DMSO), respectively. Among these three sets of differentially expressed genes, 139 genes were common to all groups, while 233, 173, and 1334 were exclusive to the TMZ, SAS, and coadministered TMZ and SAS groups, respectively (Fig. 3).
a Venn diagram showing the number of differentially expressed genes in each experimental group compared with cells that received supplemented media containing 0.1 % DMSO. Note that TMZ combined with SAS had the highest number of exclusively differentially regulated genes (1334), followed by TMZ alone (233), and SAS alone (173). b Table displaying the number of up- or downregulated genes of the total number of differentially expressed genes. c Gene expression data were processed using the PCA dimensionality reduction method. The results are graphically shown and demonstrate sample segregation. Each symbol indicates a treatment (n = 4 for each group)
Considering that the action mechanisms of TMZ and SAS are DNA alkylation and system X −c inhibition, respectively, we calculated pathway enrichment for the set of differentially expressed genes. An enriched pathway is defined as a group of functionally related genes that present a number of differentially expressed components greater than what would be expected by chance. In such an analysis, a smaller p value indicates a larger degree of enrichment. For example, “Cell cycle_The metaphase checkpoint” in the TMZ group has a −log(adjusted p value) of 14.36 (p value = 4.37 × 10−15) and 50 % differentially expressed genes (18/36 genes). On the other hand, “Cell cycle_Sister chromatid cohesion” in the TMZ group has a −log(adjusted p value) of 1.6 (p value = 2.54 × 10−2) and approximately 30 % differentially expressed genes (4/15 genes) (Fig. 4; Online Resource 1).
a–d Graphs showing enriched pathways containing the differentially expressed genes represented in the venn diagram in Fig. 3, after treatment with TMZ and/or SAS for 3 days. The X-axis presents values corresponding to −log(adjusted p values). Enriched pathways associated with values higher than −log(0.05) = 1.30 were considered statistically significant. Further enriched pathways and details on individual genes are reported in Online Resource 1
For analyzing the transcriptome data, we grouped differentially expressed genes as up- or downregulated. Thus, for downregulated genes, TMZ alone significantly enriched 17 pathways, which were mostly involved in cell cycle regulation, such as “Cell cycle_The metaphase checkpoint” (Fig. 4; Online Resource 1). TMZ coadministered with SAS significantly enriched 20 pathways, whose majority of genes and pathways coincided with those modulated by TMZ alone (Fig. 4; Online Resource 1). SAS alone did not enrich any pathways (Fig. 4; Online Resource 1).
For upregulated genes, SAS alone significantly enriched 7 pathways, which were mainly related to the antioxidant defense system and amino acid and glutathione metabolism, e.g., “Glutathione metabolism/Human version” (Fig. 4; Online Resource 1). TMZ and SAS coadministration significantly enriched 5 pathways, including amino acid metabolism and the antioxidant defense system. Despite not enriching the glutathione metabolism pathway, TMZ and SAS cotreatment significantly upregulated individual genes in this pathway. TMZ alone did not enrich any pathways (Fig. 4; Online Resource 1).
SAS decreased total glutathione levels
After 3 days, total glutathione levels were not significantly different among the DMEM, 0.1 % DMSO, and TMZ groups. Conversely, SAS alone and in combination with TMZ significantly decreased the total glutathione levels to 7.3 and 5.4 %, respectively, compared with DMEM (Fig. 5).
Total glutathione content in A172 human glioblastoma cells after 3 days of treatment with temozolomide (TMZ) and/or sulfasalazine (SAS). The graph represents the mean ± standard error of the mean of three independent experiments. Data were analyzed using one-way ANOVA with the Bonferroni post hoc test; p < 0.05 was considered statistically significant. Letters indicate p < 0.001; a versus DMEM; b versus 0.1 % DMSO; c versus 25 µM TMZ
TMZ reduced PCNA expression
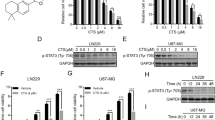
After 5 days, the proliferation of the remaining cells was assessed by Western blotting for PCNA protein. No significant differences were found between cells receiving DMEM only or with 0.1 % DMSO. However, PCNA expression was significantly lower in the TMZ group than in the 0.1 % DMSO group (53.5 vs 128.8 %, respectively). Additionally, TMZ coadministered with SAS significantly reduced PCNA expression compared with that resulting from treatment with 0.1 % DMSO (60 vs 128.8 %, respectively). The combination of drugs also decreased PCNA levels in comparison with that of cells treated only with SAS (60 vs 135.8 %, respectively). PCNA levels were not significantly altered by SAS alone (Fig. 6).
Western blotting was performed to detect proliferating cell nuclear antigen (PCNA) expression in A172 human glioblastoma cells after 5 days of treatment with temozolomide (TMZ) and/or sulfasalazine (SAS). a The graph represents PCNA expression after exposure to TMZ and/or SAS (mean ± standard error of the mean of six independent experiments). DMEM: cells receiving supplemented medium (considered 100 %). b Representative PCNA bands of each group are displayed on the membrane. c The same membrane was previously stained with Ponceau S to show the total protein content for each group. The ratio between the optical density (OD) of the PCNA band and that corresponding to all protein bands (whole lane) was calculated. Data are shown as OD values relative to that of the DMEM group, which was considered 100 %. MW: molecular weight. Data were analyzed using one-way ANOVA with the Bonferroni post hoc test; p < 0.05 was considered statistically significant. Symbols indicate the following: + versus 0.1 % DMSO; & versus 0.5 mM SAS. One or two symbols indicate p < 0.05 or p < 0.01, respectively
Discussion
In the current study, we found that SAS intensifies the effect of TMZ on reducing the viability of A172 and T98G human glioblastoma cells. This effect was detected by testing a presumably clinically relevant concentration of TMZ (25 µM) [13]; however, this concentration did not reduce cell viability when administered alone.
Notably, our findings on treating T98G cells with 25 µM TMZ are in accordance with previously reported data. Indeed, Kanzawa et al. [12] tested a higher concentration of TMZ (100 µM) for three days and found no change in the number of viable T98G cells. Similarly, Huang et al. [23] studied the effect of 100 µM TMZ on the same lineage after one and three days. The authors found that cell viability decreased by only 2 and 13 %, respectively. Moreover, the IC50 values for TMZ were determined to be >1.000 µM or 441.6 µM after counting T98G cells exposed to the drug for three days [4, 24]. Such high IC50 values corroborate our observation that 25 µM TMZ does not significantly affect T98G cell viability. In addition, when T98G cells were treated with SAS alone, we did not observe changes in cell viability after 1, 3, or 5 days. On the other hand, cotreatment with SAS and TMZ resulted in reduced cell viability after five days of treatment. To the best of our knowledge, there have been no previous reports on the effect of coadministered TMZ and SAS on T98G cells.
Regarding A172 cells, He et al. [25] described that TMZ concentrations of 0.4, 4, or 40 µM did not influence cell viability after treatment for two days, which is in agreement with our findings from the same glioblastoma cell lineage. After treatment with SAS alone, we observed reduced cell viability after 3 and 5 days; the effect on viability compared to that of 0.1 % DMSO was particularly intense after 5 days, reaching a reduction of 52.7 %. Likewise, Sleire et al. [26] detected an intense cell viability reduction (of approximately 90 %) after 8 days of incubation with 0.5 mM SAS. In our experiments, SAS coadministered with TMZ resulted in a synergistic cytotoxic effect after 3 and 5 days compared with the effect of each drug alone. These are novel findings as there have been no previous reports of a synergistic effect of SAS and TMZ on A172 cells.
Furthermore, for the first time, we report gene expression and function changes in A172 human glioblastoma cells demonstrating a synergistic cytotoxic effect after the coadministration of SAS and TMZ. In this context, SAS is known to inhibit system X −c , an antiporter that intakes cystine and releases glutamate, in glioma cells. Intracellular cystine is converted to cysteine and used for glutathione synthesis [7, 8, 27]. Our results show that decreased A172 cell viability is associated with reduced total glutathione levels, thus favoring the inhibition of system X −c . Although reduced glutathione levels via SAS has been previously described [8, 26], there are no reports on reduced glutathione levels after the exposure of A172 cells to TMZ in combination with SAS. In this sense, our data demonstrating that glutathione depletion also occurred after the cotreatment are original and in agreement with observations from other investigations of SAS alone.
Our transcriptome analyses revealed the enrichment of several pathways, which are detailed in the Online Resource. Regarding the TMZ treatment alone, pathways related to the cell cycle and DNA repair were enriched compared with 0.1 % DMSO alone. During the mitotic spindle checkpoint of the cell cycle, abnormal attachment between microtubules and chromosomes leads to anaphase delay or inhibition. Many proteins regulate this checkpoint, including those encoded by genes of the budding uninhibited by benzimidazole (BUB) and mitotic arrest deficient (MAD) families [i.e., BUB1, BUB3, MAD1, MAD2, and MAD3 (BUBR1 in humans)] [28, 29]. Morales et al. [29] observed BUB1 and BUBR1 upregulation in glioblastoma cells from commercial cell lines and tumor samples compared with nonneoplastic white matter. Moreover, these authors reported that BUB1 and BUBR1 inhibition decreased the proliferation and increased the radiosensitization of pediatric glioblastoma cells (SF188). We verified that TMZ reduced BUB1 and BUBR1 expression levels alone and after coadministration with SAS (“Cell cycle: The metaphase checkpoint” pathway, sheets 4 and 5 in Online Resource 1). Although TMZ alone did not significantly reduce cell viability after 3 or 5 days, TMZ coadministered with SAS led to a significant decrease in viability. Therefore, the reduced viability of A172 cells might have at least partially resulted from decreased BUB1 and BUBR1 expression. Taken together, these findings support focusing on BUB1 and BUBR1 as targets for therapeutic approaches to glioblastoma.
TMZ alone or in conjunction with SAS enriched the “Role of Anaphase Promoting Complex (APC) in cell cycle regulation” pathway with downregulated genes. The APC, which is activated by the cell division cycle 20 (Cdc20) protein, regulates cell cycle progression by the ubiquitination of proteins. Cdc20, which is necessary for chromosome separation, plays an oncogenic role in carcinomas [30] and is upregulated in glioblastoma [31]. Moreover, the pharmacological inhibition of APC/Cdc20 causes mitotic arrest in metaphase in several cancer cell lines [32]. We observed that TMZ alone or with SAS reduced CDC20 gene expression (sheets 4 and 5 in Online Resource 1). Considering that Cdc20 activates the APC, CDC20 downregulation after 3 days of both treatments might have induced mitotic arrest in A172 glioblastoma cells.
TMZ alone or coadministered with SAS also enriched the “Cell cycle: Spindle assembly and chromosome separation” and “Cell cycle: Sister chromatid cohesion” pathways. During the cell cycle, the cohesin protein complex holds newly replicated sister chromatids together. The endopeptidase separase cleaves a specific cohesin subunit and allows accurate chromosomal separation [28, 33]. Considering this context, Mukherjee et al. [34] studied glioblastoma samples from adults and verified the occurrence of separase overexpression, which was negatively correlated with overall survival. Our results showed reduced separase gene (ESPL1) expression after the administration of TMZ alone or with SAS (sheets 4 and 5 in Online Resource 1). This finding supports not only an antiproliferative action of TMZ but also a potentially beneficial effect of the current tested combination, as separase overexpression may correlate with reduced overall survival [34].
Because our transcriptome data highlighted the enrichment of pathways related to cell cycle progression, we searched for TMZ and/or SAS actions on cell proliferation. We evaluated PCNA expression after 5 days and observed decreased protein levels in cells receiving either TMZ or TMZ combined with SAS in comparison with the vehicle (0.1 % DMSO). Moreover, a similar decrease was observed in cells receiving TMZ coadministered with SAS compared with SAS alone. However, the transcriptome analysis showed no significant PCNA gene expression alteration after 3 days of any treatment (sheets 1, 2, and 3 in Online Resource 1). Although the TMZ treatment alone did not result in cell viability significantly different from that resulting from treatment with vehicle for 5 days, our data on PCNA protein levels suggest that the cells remaining after the TMZ treatment exhibited less proliferative activity. Indeed, TMZ arrests the cell cycle in the G2/M phase and reduces the proliferation of U251 and U87MG glioblastoma cells [35, 36]. Furthermore, a reduction in the number of PCNA-immunopositive cells was detected in the brain of rodents grafted with C6 or U87MG glioma cells and treated with TMZ [37, 38]. Together with those previous data, our PCNA protein expression results corroborate the antiproliferative effect of TMZ in A172 cells, which has not been previously reported.
Regarding the effect of SAS on gene expression, we observed the enrichment of glutathione metabolism and oxidative stress pathways. SAS inhibits system X −c , a protein complex constituted by light (xCT) and heavy (4F2) chains [7, 8, 27]. In our study, SAS alone or with TMZ increased the expression of the xCT gene (SLC7A11), which is present in the “Oxidative stress: Role of Sirtuin1 and PGC1-alpha in activation of antioxidant defense system” pathway (sheets 6 and 7 in Online Resource 1). The same pathway exhibited alterations in the expression of genes encoding different subunits of glutamate-cysteine ligase (GCL), an enzyme that is essential for glutathione synthesis. The subunits of GCL are known as regulatory (GCL reg) and catalytic (GCL cat) components [39, 40]. SAS-treated cells showed increased expression of the GCL regulatory subunit gene (GCLM). Cotreatment with SAS and TMZ induced higher expression levels of both GCL catalytic (GCLC) and regulatory subunit genes (sheets 6 and 7 in Online Resource 1).
Oxidized glutathione (GSSG) is converted to its reduced form (GSH) by glutathione reductase (GSHR) [39, 41, 42]. We found an increased expression of the GSHR gene (GSR) after treatment with SAS only or in conjunction with TMZ, but not after treatment with TMZ alone. Additionally, SAS upregulated some genes of the “Glutathione metabolism/Human version” pathway, such as glutathione S-transferases (GSTM2, GSTM3, GSTK1, and MGST) (sheet 6 in Online Resource 1). These enzymes depend on glutathione and participate in the detoxification of products generated by oxidative stress [43].
Although TMZ coadministered with SAS did not enrich the “Glutathione metabolism/Human version” pathway, we verified the individual upregulation of the GSR, GSTM2, GCLM, and GCLC genes (sheet 7 in Online Resource 1). In addition, we detected significantly decreased glutathione after 3 days of treatment with either SAS alone or combined with TMZ. Therefore, as SAS alone or with TMZ induced an increased expression of genes encoding xCT, glutathione reductase, glutathione S-transferases, or glutamate-cysteine ligase, it is possible that compensatory intracellular mechanisms can be activated to restore normal glutathione levels. Altogether, the alterations we observed in gene expression and glutathione level could have been involved in the process of cell viability reduction we observed after 3 and 5 days of treating cells with SAS with or without TMZ.
Mandal et al. [27] showed that under glutathione depletion, the thioredoxin/thioredoxin reductase 1 system reduces the cystine imported by system X −c , thus serving as a substitute for glutathione. We found that thioredoxin reductase 1 gene (TXNRD1) was induced in groups with total glutathione depletion (SAS with or without TMZ). Therefore, this induction could be an additional compensatory response to deal with oxidative stress.
Although our study showed reduced A172 and T98G cell viability after exposure to TMZ with SAS, a clinical trial phase 1/2 recommended caution with the use of SAS as a therapy for glioblastoma [44]. Specifically, Robe et al. [44] recruited patients who had previously undergone surgery, standard radiation therapy, and a course of chemotherapy with an alkylating agent. Moreover, the patients had not been taking the cytotoxic medications for at least 4 weeks prior to SAS treatment. One tumor was stabilized after two months of treatment with SAS; however, glioblastoma progression was not affected in the other patients. Additionally, all individuals (n = 10) exhibited grade 1–3 adverse effects. Four patients developed grade 4 toxicity, and two subsequently developed grade 5 (i.e., lethal) toxicity. In spite of such negative results, it is important to note that this study included only patients with progressive or recurrent high-grade glioma (i.e., those who were severely ill or had serious neurological impairment). Thus, the small number of patients and their deteriorated clinical conditions may have hampered the detection of tumor progression reduction by SAS. More recently, another clinical trial was conducted with newly diagnosed glioblastoma patients (n = 12) who were given TMZ and SAS with radiation therapy after surgery [45]. This study detected no changes in overall survival, progression-free survival, or seizure-free survival compared with patients who received radiation and TMZ. Grade 3 or 4 adverse effects occurred during the treatment in nine patients. Nevertheless, the data suggested that the administration of an adequate dosage of SAS might improve seizure control. To the best of our knowledge, these are the only clinical trials that have been reported to date. As both studied a limited number of individuals, further investigations are needed to more accurately assess the toxicity and possible therapeutic role of glutathione-depleting agents, such as SAS, in high-grade glioma patients.
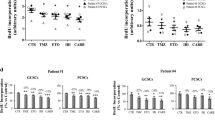
In fact, the action of erastin, a potent system X −c inhibitor, on GBM-N15 primary human glioblastoma cells was recently described [46]. Specifically, GBM-N15 cells were subjected to treatment with increasing concentrations of TMZ (from 50 to 600 µM) combined with 0.3 mM erastin for 48 h. Irrespective of TMZ concentration, the cell viability reduction induced by TMZ coadministered with erastin was higher than that resulting from treatment with either TMZ or erastin alone. Thus, Chen et al. [46] have laid a foundation for clinical trials of another system X −c inhibitor for treating glioblastoma patients.
In summary, our data show that both A172 and T98G human glioblastoma cells exhibited reduced viability after five days of treatment with a combination of TMZ and SAS. Furthermore, we present original data on A172 glioblastoma cell exposure to TMZ and/or SAS; we verified the occurrence of not only a synergistic cytotoxic effect but also gene expression alteration and cellular pathway enrichment resulting from these drugs. Indeed, as TMZ modulated the cell cycle and SAS regulated antioxidant mechanisms, their association altered both cell cycle and antioxidant pathways. The combination of these mechanisms could explain the more intense cytotoxic effect observed after coadministering TMZ and SAS compared with that resulting from each drug alone. Therefore, our study provides insights into potential therapeutic targets for treating glioblastoma, along with corresponding molecular data. These targets could be considered in future investigations for the improvement of the clinical efficacy of TMZ against glioblastoma.
References
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, Scheithauer BW, Kleihues P (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114:97–109. doi:10.1007/s00401-007-0243-4
Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, Taphoorn MJ, Belanger K, Brandes AA, Marosi C, Bogdahn U, Curschmann J, Janzer RC, Ludwin SK, Gorlia T, Allgeier A, Lacombe D, Cairncross JG, Eisenhauer E, Mirimanoff RO (2005) European organisation for research and treatment of cancer brain tumor and radiotherapy groups radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352:987–996. doi:10.1056/NEJMoa043330
Stupp R, Hegi ME, Mason WP, van den Bent MJ, Taphoorn MJ, Janzer RC, Ludwin SK, Allgeier A, Fisher B, Belanger K, Hau P, Brandes AA, Gijtenbeek J, Marosi C, Vecht CJ, Mokhtari K, Wesseling P, Villa S, Eisenhauer E, Gorlia T, Weller M, Lacombe D, Cairncross JG, Mirimanoff RO (2009) Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol 10:459–466. doi:10.1016/S1470-2045(09)70025-7
Yoshino A, Ogino A, Yachi K, Ohta T, Fukushima T, Watanabe T, Katayama Y, Okamoto Y, Naruse N, Sano E, Tsumoto K (2010) Gene expression profiling predicts response to temozolomide in malignant gliomas. Int J Oncol 36:1367–1377. doi:10.3892/ijo_00000621
Sun S, Wong TS, Zhang XQ, Pu JK, Lee NP, Day PJ, Ng GK, Lui WM, Leung GK (2012) Protein alterations associated with temozolomide resistance in subclones of human glioblastoma cell lines. J Neurooncol 107:89–100. doi:10.1007/s11060-011-0729-8
Rahman I, Kode A, Biswas SK (2006) Assay for quantitative determination of glutathione and glutathione disulfide levels using enzymatic recycling method. Nat Protoc 1:3159–3165. doi:10.1038/nprot.2006.378
Sontheimer H (2008) A role for glutamate in growth and invasion of primary brain tumors. J Neurochem 105:287–295. doi:10.1111/j.1471-4159.2008.05301.x
Chung WJ, Lyons SA, Nelson GM, Hamza H, Gladson CL, Gillespie GY, Sontheimer H (2005) Inhibition of cystine uptake disrupts the growth of primary brain tumors. J Neurosci 25:7101–7110. doi:10.1523/JNEUROSCI.5258-04.2005
Lyons SA, Chung WJ, Weaver AK, Ogunrinu T, Sontheimer H (2007) Autocrine glutamate signaling promotes glioma cell invasion. Cancer Res 67:9463–9471. doi:10.1158/0008-5472.CAN-07-2034
Robe PA, Bentires-Alj M, Bonif M, Rogister B, Deprez M, Haddada H, Khac MT, Jolois O, Erkmen K, Merville MP, Black PM, Bours V (2004) In vitro and in vivo activity of the nuclear factor-kappaB inhibitor sulfasalazine in human glioblastomas. Clin Cancer Res 10:5595–5603. doi:10.1158/1078-0432.CCR-03-0392
Chung WJ, Sontheimer H (2009) Sulfasalazine inhibits the growth of primary brain tumors independent of nuclear factor-kappaB. J Neurochem 110:182–193. doi:10.1111/j.1471-4159.2009.06129.x
Kanzawa T, Bedwell J, Kondo Y, Kondo S, Germano IM (2003) Inhibition of DNA repair for sensitizing resistant glioma cells to temozolomide. J Neurosurg 99:1047–1052. doi:10.3171/jns.2003.99.6.1047
Ostermann S, Csajka C, Buclin T, Leyvraz S, Lejeune F, Decosterd LA, Stupp R (2004) Plasma and cerebrospinal fluid population pharmacokinetics of temozolomide in malignant glioma patients. Clin Cancer Res 10:3728–3736. doi:10.1158/1078-0432.CCR-03-0807
Barazzuol L, Jena R, Burnet NG, Jeynes JC, Merchant MJ, Kirkby KJ, Kirkby NF (2012) In vitro evaluation of combined temozolomide and radiotherapy using X rays and high-linear energy transfer radiation for glioblastoma. Radiat Res 177:651–662. doi:10.1667/RR2803.1
Liu Y, Peterson DA, Kimura H, Schubert D (1997) Mechanism of cellular 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) reduction. J Neurochem 69:581–593. doi:10.1046/j.1471-4159.1997.69020581.x
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. doi:10.1016/0003-2697(76)90527-3
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685. doi:10.1038/227680a0
Towbin H, Staehelin T, Gordon J (1979) Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci USA 76:4350–4354. doi:10.1073/pnas.76.9.4350
Vieira AS, Rezende AC, Grigoletto J, Rogério F, Velloso LA, Skaper SD, Negro A, Langone F (2009) Ciliary neurotrophic factor infused intracerebroventricularly shows reduced catabolic effects when linked to the TAT protein transduction domain. J Neurochem 110:1557–1566. doi:10.1111/j.1471-4159.2009.06259.x
Ignarro RS, Vieira AS, Sartori CR, Langone F, Rogério F, Parada CA (2013) JAK2 inhibition is neuroprotective and reduces astrogliosis after quinolinic acid striatal lesion in adult mice. J Chem Neuroanat 48–49:14–22. doi:10.1016/j.jchemneu.2013.02.005
Romero-Calvo I, Ocón B, Martínez-Moya P, Suárez MD, Zarzuelo A, Martínez-Augustin O, de Medina FS (2010) Reversible Ponceau staining as a loading control alternative to actin in western blots. Anal Biochem 401:318–320. doi:10.1016/j.ab.2010.02.036
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10:57–63. doi:10.1038/nrg2484
Huang H, Lin H, Zhang X, Li J (2012) Resveratrol reverses temozolomide resistance by downregulation of MGMT in T98G glioblastoma cells by the NF-κB-dependent pathway. Oncol Rep 27:2050–2056. doi:10.3892/or.2012.1715
Kanzawa T, Germano IM, Komata T, Ito H, Kondo Y, Kondo S (2004) Role of autophagy in temozolomide-induced cytotoxicity for malignant glioma cells. Cell Death Differ 11:448–457. doi:10.1038/sj.cdd.4401359
He W, Liu R, Yang SH, Yuan F (2015) Chemotherapeutic effect of tamoxifen on temozolomide-resistant gliomas. Anti Cancer Drugs 26:293–300. doi:10.1097/CAD.0000000000000197
Sleire L, Skeie BS, Netland IA, Førde HE, Dodoo E, Selheim F, Leiss L, Heggdal JI, Pedersen PH, Wang J, Enger PØ (2015) Drug repurposing: sulfasalazine sensitizes gliomas to gamma knife radiosurgery by blocking cystine uptake through system Xc−, leading to glutathione depletion. Oncogene 34:5951–5959. doi:10.1038/onc.2015.60
Mandal PK, Seiler A, Perisic T, Kölle P, Banjac Canak A, Förster H, Weiss N, Kremmer E, Lieberman MW, Bannai S, Kuhlencordt P, Sato H, Bornkamm GW, Conrad M (2010) System x(c)- and thioredoxin reductase 1 cooperatively rescue glutathione deficiency. J Biol Chem 285:22244–22253. doi:10.1074/jbc.M110.121327
Musacchio A, Hardwick KG (2002) The spindle checkpoint: structural insights into dynamic signalling. Nat Rev Mol Cell Biol 3:731–741. doi:10.1038/nrm929
Morales AG, Pezuk JA, Brassesco MS, de Oliveira JC, de Paula Queiroz RG, Machado HR, Carlotti CG, Neder L, de Oliveira HF, Scrideli CA, Tone LG (2013) BUB1 and BUBR1 inhibition decreases proliferation and colony formation, and enhances radiation sensitivity in pediatric glioblastoma cells. Childs Nerv Syst 29:2241–2248. doi:10.1007/s00381-013-2175-8
Wang L, Zhang J, Wan L, Zhou X, Wang Z, Wei W (2015) Targeting Cdc20 as a novel cancer therapeutic strategy. Pharmacol Ther 151:141–151. doi:10.1016/j.pharmthera.2015.04.002
Marucci G, Morandi L, Magrini E, Farnedi A, Franceschi E, Miglio R, Calò D, Pession A, Foschini MP, Eusebi V (2008) Gene expression profiling in glioblastoma and immunohistochemical evaluation of IGFBP-2 and CDC20. Virchows Arch 453:599–609. doi:10.1007/s00428-008-0685-7
Zeng X, Sigoillot F, Gaur S, Choi S, Pfaff KL, Oh DC, Hathaway N, Dimova N, Cuny GD, King RW (2010) Pharmacologic inhibition of the anaphase-promoting complex induces a spindle checkpoint-dependent mitotic arrest in the absence of spindle damage. Cancer Cell 18:382–395. doi:10.1016/j.ccr.2010.08.010
Ivanchuk SM, Rutka JT, Piepmeier JM, Parsa AT, Boulis N (2004) The cell cycle: accelerators, brakes, and checkpoints. Neurosurgery 54:692–700. doi:10.1227/01.NEU.0000109534.28063.5D
Mukherjee M, Byrd T, Brawley VS, Bielamowicz K, Li XN, Merchant F, Maitra S, Sumazin P, Fuller G, Kew Y, Sun D, Powell SZ, Ahmed N, Zhang N, Pati D (2014) Overexpression and constitutive nuclear localization of cohesin protease separase protein correlates with high incidence of relapse and reduced overall survival in glioblastoma multiforme. J Neurooncol 119:27–35. doi:10.1007/s11060-014-1458-6
Hirose Y, Berger MS, Pieper RO (2001) p53 effects both the duration of G2/M arrest and the fate of temozolomide-treated human glioblastoma cells. Cancer Res 61:1957–1963
Shen W, Hu J-A, Zheng J-S (2014) Mechanism of temozolomide-induced antitumour effects on glioma cells. J Int Med Res 42:164–172. doi:10.1177/0300060513501753
Jo MY, Kim YG, Kim Y, Lee SJ, Kim MH, Joo KM, Kim HH, Nam DH (2012) Combined therapy of temozolomide and ZD6474 (vandetanib) effectively reduces glioblastoma tumor volume through anti-angiogenic and anti-proliferative mechanisms. Mol Med Rep 6:88–92. doi:10.3892/mmr.2012.868
Son MJ, Kim JS, Kim MH, Song HS, Kim JT, Kim H, Shin T, Jeon HJ, Lee DS, Park SY, Kim YJ, Kim JH, Nam DH (2006) Combination treatment with temozolomide and thalidomide inhibits tumor growth and angiogenesis in an orthotopic glioma model. Int J Oncol 28:53–59. doi:10.3892/ijo.28.1.53
Backos DS, Franklin CC, Reigan P (2012) The role of glutathione in brain tumor drug resistance. Biochem Pharmacol 83:1005–1012. doi:10.1016/j.bcp.2011.11.016
Aquilano K, Baldelli S, Ciriolo MR (2014) Glutathione: new roles in redox signaling for an old antioxidant. Front Pharmacol 5:196. doi:10.3389/fphar.2014.00196
Balendiran GK, Dabur R, Fraser D (2004) The role of glutathione in cancer. Cell Biochem Funct 22:343–352. doi:10.1002/cbf.1149
Traverso N, Ricciarelli R, Nitti M, Marengo B, Furfaro AL, Pronzato MA, Marinari UM, Domenicotti C (2013) Role of glutathione in cancer progression and chemoresistance. Oxid Med Cell Longev 2013:972913
Hayes JD, Flanagan JU, Jowsey IR (2005) Glutathione transferases. Annu Rev Pharmacol Toxicol 45:972913. doi:10.1146/annurev.pharmtox.45.120403.095857
Robe PA, Martin DH, Nguyen-Khac MT, Artesi M, Deprez M, Albert A, Vanbelle S, Califice S, Bredel M, Bours V (2009) Early termination of ISRCTN45828668, a phase 1/2 prospective, randomized study of sulfasalazine for the treatment of progressing malignant gliomas in adults. BMC Cancer 9:372. doi:10.1186/1471-2407-9-372
Takeuchi S, Wada K, Nagatani K, Otani N, Osada H, Nawashiro H (2014) Sulfasalazine and temozolomide with radiation therapy for newly diagnosed glioblastoma. Neurol India 62:42–47. doi:10.4103/0028-3886.128280
Chen L, Li X, Liu L, Yu B, Xue Y, Liu Y (2015) Erastin sensitizes glioblastoma cells to temozolomide by restraining xCT and cystathionine-γ-lyase function. Oncol Rep 33:1465–1474. doi:10.3892/or.2015.3712
Funding
This work was supported by Grants from FAPESP (2011/50400-0; 2013/02618-1; 2013/07559-3) and FAEPEX/UNICAMP (379/13; 554/14; 621/14). RSI was a recipient of a scholarship from CAPES.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
We have read and have abided by the statement of ethical standards for manuscripts submitted to Molecular and Cellular Biochemistry. This paper is not concurrently under consideration for publication in any other journal.
Conflicts of interest
The authors declare that all authors are in agreement with the present submission and have no conflicts of interest.
Research involving human and animal participants
In addition, this research does not encompass any studies involving human participants or animals performed by any of the authors.
Ethical approval
This research does not encompass any studies involving human participants or animals performed by any of the authors.
Additional information
Raffaela Silvestre Ignarro and Gustavo Facchini contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11010_2016_2742_MOESM1_ESM.xls
Online Resource 1 Gene expression data in A172 human glioblastoma cells after 3 days of treatment with temozolomide (TMZ) and/or sulfasalazine (SAS). Columns in Tables 1–3 show the gene symbol, log2 fold change relative to 0.1 % DMSO-treated cells, p value and adjusted p value. Tables 4–7 contain the enriched pathways for the up- and downregulated genes for each group. Fold change for each gene in the enriched pathways is presented in parenthesis after the name of the gene (XLS 6870 kb)
Rights and permissions
About this article
Cite this article
Ignarro, R.S., Facchini, G., Vieira, A.S. et al. Sulfasalazine intensifies temozolomide cytotoxicity in human glioblastoma cells. Mol Cell Biochem 418, 167–178 (2016). https://doi.org/10.1007/s11010-016-2742-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-016-2742-x