Abstract
Oleanolic acid (OA) is a natural compound from plants with anti-tumor activities. However, the mechanism of the inhibitory effect of OA on cell cycle progression has not been completely explored. We employed several lung carcinoma cell lines to investigate the cell cycle-related molecular pathway affected by OA. The data revealed that OA suppressed the proliferation of lung cancer cells in both dose- and time-dependent manners, along with an increase in miR-122 abundance. The suppression of miR-122 abolished the effect of OA on lung cancer cells. CCNG1 and MEF2D, two putative miR-122 targets, were found to be downregulated by OA treatment. Restoring their expression counteracted the effect of OA on lung carcinoma cells. OA was further shown to induce the expression of miR-122-regulating transcriptional factors in lung cancer cells. Collectively, OA induced cell cycle arrest in lung cancer cells through miR-122/Cyclin G1/MEF2D pathway. This finding may contribute to the understanding of the molecular mechanism of OA’s anti-tumor activity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oleanolic acid (OA), a natural pentacyclic triterpene, is widely distributed in a variety of herbs and vegetables. It has been reported that OA possesses many bioactivities, including hepatoprotection, anti-inflammation effect, and anti-tumor activity [1]. The anti-tumor activity of OA has been verified in many types of cancers, such as glioma [2], breast carcinoma [3], osteosarcoma [4], hepatocellular carcinoma [5], and lung carcinoma [6]. The induction of cell cycle arrest is among major mechanisms of OA’s anti-tumor activity. However, the molecular mechanism by which OA induces cell cycle arrest is still unclear. Previous investigations revealed that the activation of ERK/p53 pathway contributes to cell cycle arrest induced by OA in HepG2 cells [5]. However, other p53-independent mechanisms should exist, because p53 mutation and deletion is quite common in cancer cell lines and the tumor specimen from patients.
miRNAs have been well documented to be associated with cancer initiation and progression [7]. Therefore, they are also believed to be effective therapeutic targets [7]. miR-122 has been found to be an important tumor suppressor in some types of cancer. This miRNA can induce apoptosis [8], trigger cell cycle arrest [9], inhibit metastasis [10], and enhance drug resistance of cancer cells [11]. To inhibit cell cycle progression of cancer cells, miR-122 recognizes and binds miRNA response element (MRE), a sequence complementary to the “seed” of miR-122, which is located within 3’ untranslated region (3’ UTR) of its target mRNA, such as Cyclin G1 [9] and MEF2D [12]. Subsequently, miR-122 degrades these existing mRNA molecules and reduces the expression of their proteins. Eventually, cancer cells are subjected to G0/G1 or G2/M arrest and lose their high proliferation rate.
In this study, we employed several lung carcinoma cell lines and primary lung cancer cells to investigate if miR-122 and its downstream target genes are associated with the anti-proliferation effect of OA on cancer cells.
Materials and methods
Cell cultures and chemicals
Human lung cancer cell lines, A549, NCI-H460, and NCI-H1299, were purchased from American Type Culture Collection (Manassas, VA), as well as normal lung fibroblast cell line, MRC-5. The cells were cultured with DMEM containing 10 % fetal bovine serum under a 37 °C 5 % CO2-containing humidified condition. OA was purchased from Sigma Aldrich (St. Louis, MO).
Lung carcinoma specimen
Lung carcinoma tissues were obtained from the patients with their written informed consent following the procedures approved by Ethical Review Board in Shandong Cancer Hospital and Institute (Jinan, China). These patients were subjected to surgery at Department of Thoracic Surgery, Shandong Cancer Hospital and Institute (Jinan, China). The obtained tumors were cut into pieces. Then, trypsin was used to separate cells. The cells were filtered with steel net and cultured in DMEM containing 15 % FBS. The established primary lung cancer cell lines were defined as primary LC 1 and primary LC 2.
Quantitative PCR
Total RNA were isolated from cells after the indicated treatment at certain time points with the help of Trizol solution (Sigma Aldrich), followed by reverse transcription reaction using All-in-One™ First-Strand cDNA Synthesis Kit (AORT-0020, GeneCopoeia) based on the manufacturer’s instructions. Quantitative PCR (qPCR) was performed using All-in-One™ miRNA qRT-PCR Detection Kit (AOMD-Q020, GeneCopoeia) on CFX96™ Real-Time PCR detection system with analytical software (Bio-Rad). The primers and probes specific for miR-122 and endogenous reference, U6, were available at the websites of GeneCopoeia.
miR-122 silencing
miR-122 antagomir was purchased from Genepharma (Shanghai, China). Cells were transfected with miR-122 antagomir with Lipofectamine™ 2,000 (Invitrogen), according to the instructions provided by manufacturers. After 48 h, subsequent experiments were performed for certain purposes.
Cell proliferation assay
5 × 103 A549, NCI-H460 and NCI-H1299 and two primary lung carcinoma cells were planted on 96-well plates. Six hr later, cells were treated with OA at different concentrations. At the indicated time points, 10 µl 3-(4,5-dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide (MTT) (5 mg l−1) was added to the media in the tested well and incubated for another 4 h. MTT solution was then discarded, and 150 µl DMSO was used to resolve the subsidence. A model 550 microplate reader (Bio-Rad) was used to detect the 570 nm absorptive value in each well. The proliferation rate was calculated by the following formula: Proliferation rates (folds) = absorbance value of tested wells/absorbance value of control wells.
Western blotting
Western blotting was used to detect the expression level of proteins in cells in our study. Proteins were isolated from cells with M-PER® mammalian protein extraction reagent (Thermo Scientific). BCA assays were used to determine the concentration of proteins in samples. Then polyacrylamide gel electrophoresis was performed according to routine procedures, followed by transferring onto 0.45 μm nitrocellulose membranes. The membrane was incubated with specific primary antibodies for 16 h at 4 °C. Appropriate secondary antibodies were then used to incubate the membranes. Finally, the blots were visualized using Western Blotting Substrate (Pierce). The used primary antibodies (anti-CCNG1, anti-MEF2D, anti-HNF1α, anti-HNF3β, anti-HNF4α, anti-HNF6, and anti-β-tubulin) were all purchased from Cell Signaling Technology (MA, USA).
CCNG1 and MEF2D overexpression vectors
pcDNA3.1-CCNG1 and pcDNA3.1-MEF2D were both purchased from Highene Biotechnology Company (Qingdao, China), as well as pcDNA3.1-GFP as control. 2 ng of the above plasmid was transfected into cells with Lipofectamine™ 2000 (Invitrogen) according to the manufacturer’s instructions. 48 h later, the cells were subjected to the subsequent experiments.
Animal experiments
The procedures of animal experiment were approved by Committee on the Use and Care on Animals of Shandong Cancer Hospital and Institute (Jinan, China). Lung carcinoma xenograft was established by injecting 5 × 106 primary lung cancer cells into the flanks of 4- to 5- week-old male BALB/c nude mice. 18 mice were divided into three groups (n = 6). Low dose (40 mg kg−1) or high dose (120 mg kg−1) of OA was orally administrated every day for 4 weeks. Tumor diameters were then periodically measured with calipers. Tumor volume (mm3) = maximal length (mm) × perpendicular width (mm)2/2.
Statistical analysis
We employed two-tailed Student’s test for statistical analysis in our study. * P < 0.05; ** P < 0.01.
Results
OA reduced the proliferation rates of lung cancer cells in time- and dose-dependent fashions
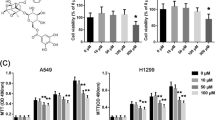
First of all, we detected if OA can affect the proliferation of lung carcinoma cell lines, A549, NCI-H460, and NCI-H1299, as well as two primary lung cancer cells. The results indicated that, 48 h after treatment, OA significantly reduced the proliferation of the tested cells by different extents (A549, 10.7–39 %; NCI-H460, 1.3–41.8 %; NCI-H1299, 2–50.4 %; primary lung cancer 1, 4.5–64.3 %; primary lung cancer 2, 1.8–46.9 %) in a dose-dependent manner (Fig. 1a). Also, OA suppressed their proliferation in a time-dependent manner at the concentration of 30 μg ml−1 (Fig. 1b). In addition, normal lung fibroblast cell lines, MRC-5, had no significant reduction in its proliferation rate when OA was added to the media (Fig. 1a, b).
OA reduced the proliferation rates of lung cancer cells in time- and dose-dependent fashions. A549, NCI-H460, NCI-1299, primary LC 1, primary LC 2, and MRC-5 were treated with OA at 15, 30, or 60 μg ml−1 for 48 h (a), or at 30 μg ml−1 for 24, 48, or 72 h (b). The proliferation rates of these cells were tested by MTT assays and shown as mean ± SD of three independent experiments
OA elevated the expression level of miR-122 in lung carcinoma cells
To explore the molecular mechanism of OA’s anti-proliferation effect, we detected the expression levels of miR-122 in lung cancer cell under this treatment. The data revealed that OA was able to induce the expression of miR-122, 48 h after OA treatment (6.02–9.86 folds for 60 μg/ml of OA) (Fig. 2a). Time course analysis also indicated that OA elevated the abundance of miR-122 in a time-dependent manner (Fig. 2b).
OA elevated the expression level of miR-122 in lung carcinoma cells. A549, NCI-H460, NCI-1299, primary LC 1, and primary LC 2 were treated with OA at 15, 30, or 60 μg ml−1 for 48 h (a), or at 30 μg ml−1 for 24, 48, or 72 h (b). The expression level of miR-122 was detected by qPCR assay and shown as mean ± SD of three independent experiments. U6 was selected as endogenous reference
The decline in miR-122 level abolished the effect of OA on lung cancer cells
To confirm the role of miR-122 overexpression in the anti-proliferation effect of OA on lung cancer cells, we used antagomir to suppress the expression of miR-122 in cells treated with OA (Fig. 3a). Antagomir abolished the inhibitory effect of OA on the proliferation of lung cancer cells, evidenced by the increased proliferation rates of the cells co-treated with OA and antagomir, compared with that treated with OA alone (Fig. 3b).
Reducing miR-122 levels abolished the effect of OA on lung cancer cells. (a) A549 and primary LC 1 were treated with OA (30 μg ml−1) or/and miR-122 antagomir. 48 h later, qPCR assay was performed to examine the abundance of miR-122 in these cells. The results were shown as mean ± SD of three independent experiments. U6 was selected as endogenous reference. *P < 0.05; **P < 0.01. (b) Under the above treatments, proliferation rates of A549 and primary LC 1 cells were also evaluated by MTT assays and shown as mean ± SD of three independent experiments. *P < 0.05
OA suppressed the expressions of CCNG1 and MEF2D in lung cancer cells
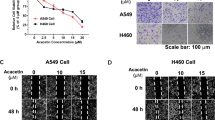
To furthermore investigate the mechanism by OA affected cell proliferation by miR-122, we evaluated the expression of its target genes that have been identified to be related to cell cycle progression. CCNG1, a promoter of G0/G1 transition that has been verified as a miR-122 target, was found to be downregulated in lung cancer cells treated with OA in both dose- and time-dependent ways (Fig. 4a, b). The expression of another miR-122 target, MEF2D, was also found to be silenced in the tested cells under the treatment of OA (Fig. 4a, b).
OA suppressed the expressions of CCNG1 and MEF2D. A549 and primary LC 1 were treated with OA at 15, 30, or 60 μg ml−1 for 48 h (a), or at 30 μg ml−1 for 24, 48, or 72 h (b). The expression level of CCNG1 and MEF2D was detected by immunoblot assays. The densities of these blots were determined with ImageJ software
The restoration of CCNG1 and MEF2D reversed the effect of OA on lung carcinoma cells
To confirm if CCNG1 and MEF2D mediated the inhibitory effect of OA on A549 and primary lung cancer cells, we transfected these cell with plasmids that overexpressed CCNG1 and MEF2D (pcDNA3-CCNG1 and pcDNA-MEF2D) (Fig. 5a). The results revealed that the overexpression of these two proteins was able to protect lung cancer cells from the action of OA, evidenced by increased proliferation of lung cancer cells under co-treatment of OA and these plasmids (Fig. 5b).
Restoration of CCNG1 and MEF2D reversed the effect of OA on lung carcinoma cells. (a) A549 and primary LC 1 were treated with OA (30 μg ml−1) as well as pcDNA-GFP (−, 2 ng) or pcDNA-CCNG1 plus pcDNA-MEF2D (2 ng together). Forty-eight hr later, immunoblot assay was used to assess the abundance of CCNG1 and MEF2D proteins in these cells. The densities of these blots were determined with ImageJ software. (b) Under the above treatments, proliferation rates of A549 and primary LC 1 cells were also evaluated by MTT assays and shown as mean ± SD of three independent experiments. *P < 0.05
OA elevated the expression of miR-122-regulating transcriptional factors
To further identify the mechanisms by which OA induced miR-122 overexpression, we detected the expression of hepatocyte nuclear factor (HNF)1α, HNF3β, HNF4α, and HNF6, the known miR-122-regulating transcriptional factors, in A549 lung carcinoma cells treated with OA of different doses. Immunoblot assay revealed that OA was able to increase the expression of these transcriptional factors in both dose- and time-dependent manners (Fig. 6a, b), implying that OA may exert its stimulatory effect on miR-122 expression through inducing these transcriptional factors.
OA induced the expressions of HNF1α, HNF3β, HNF4α, and HNF6. A549 cells were treated with OA at 15, 30, or 60 μg ml−1 for 48 h (a), or at 30 μg ml−1 for 24, 48, or 72 h (b). The expression levels of HNF1α, HNF3β, HNF4α, and HNF6 were detected by immunoblot assays. The densities of these blots were determined with ImageJ software
OA suppressed the growth of lung carcinoma xenografts in vivo
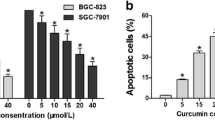
To investigate if OA exerts anti-tumor activity on lung carcinoma cells in vivo, we established a mouse model bearing tumor xenografts by subcutaneously injecting primary lung carcinoma cells. The determination of tumor volumes indicated that high dose of OA significantly inhibited the growth of lung cancer xenograft in mice (Fig. 7a). Although there was no statistical significance, low dose of OA also modestly reduced the growth of tumor xenograft (Fig. 7a). qPCR assays revealed that miR-122 expression was restored in tumor xenograft in mice by OA administration (Fig. 7b). CCNG1 and MEF2D were consistently found to underexpressed in these tumors (Fig. 7c).
OA suppressed the growth of lung carcinoma xenografts in vivo. (a) Low dose (40 mg kg−1) or high dose (120 mg kg−1) of OA was orally administrated every day for 4 weeks. Tumor diameters were periodically measured. The volumes were calculated based on the formula described in Materials and Methods and shown as mean ± SD. The expression level of miR-122 (b), as well as CCNG1 and MEF2D (c), was detected in tumor xenografts. The densities of these blots were determined with ImageJ software
Discussion
OA is a well-recognized natural compound with anti-tumor activity, which is derived from plants. However, the molecular mechanism by which OA suppresses the proliferation of cancer cells has not been completely explored yet. miRNA has been identified as therapeutic targets of many compounds for cancer treatment. For instances, rhamnetin and cirsiliol sensitize non-small cell lung cancer cells to radiotherapy in a miR-34-dependent manner [13]. Zhang et al. found that waltonitone can suppress the proliferation of A549 lung cancer cells by affecting 8 miRNAs associated with cell cycle progression [14]. To our knowledge, the implication of miRNA in the anti-tumor activity of OA has not been reported. Here, we verified that OA can increase the expression of tumor suppressor miRNA, miR-122, in lung cancer cells.
miR-122 attracts much attention from researchers, because of its significant inhibitory effect on cancer formation and progression. miR-122 can suppress almost all the aspects of cancer cells, including survival, proliferation, invasion, metastasis, drug resistance, metabolism, and angiogenesis [15]. Therefore, its restoration should be an effective strategy for cancer treatment [16]. However, there has been no confirmed natural compound that affects miR-122 level or function yet. Our data confirmed that miR-122 is an authentic therapeutic target for cancer treatment. Thus, more compounds are encouraged to be tested for their effect on miR-122.
MEF2D, a member of myocyte enhancer factor 2 family transcript factors, has recently been shown to be overexpressed in HCC cells and promotes their proliferation by accelerating G2/M transition. Interestingly, MEF2D is also a miR-122 target [12]. This finding raises the possibility that MEF2D may be a therapeutic target for cancer treatment. Actually, 6-Hydroxydopamine can reduce the survival of normal neuron cell, PC-12, by downregulating MEF2D [17]. Our results further confirmed that OA-induced MEF2D reduction can also decelerate the proliferation of tumor cells, reinforcing the notion that MEF2D is an effective target for cancer treatment.
In addition, other proteins may also be involved in the role of miR-122 in the effect of OA on the proliferation of lung carcinoma cells. For instances, pyruvate kinase type M (PKM) may be associated with the effect of OA on lung cancer cells. PKM has been shown as a confirmed target of miR-122 in liver cancer cells [18]. Furthermore, OA has been verified to regulate PKM function and suppress the proliferation of cancer cells [3]. Therefore, we hypothesized that PKM may also mediate the anti-tumor activity of OA on lung cancer cells in a miR-122-dependent manner. Other regulators, such as autophagy-related proteins [19, 20], AMPK [21], and p38 MAPK [22], may also be associated with the miR-122-involved inhibitory effect of OA on lung cancer cells.
Taken together, we identified that OA exerts its anti-proliferation effect on lung cancer cells through miR-122/CCNG1/MEF2D pathway. Furthermore, targeting miR-122 and its downstream oncogenic targets may be effective therapeutic anticancer strategies.
References
Liu J (2005) Oleanolic acid and ursolic acid: research perspectives. J Ethnopharmacol 100:92–94. doi:10.1016/j.jep.2005.05.024
Guo G, Yao W, Zhang Q, Bo Y (2013) Oleanolic acid suppresses migration and invasion of malignant glioma cells by inactivating MAPK/ERK signaling pathway. PLoS One 8:e72079. doi:10.1371/journal.pone.0072079
Liu J, Wu N, Ma L, Liu M, Liu G, Zhang Y, Lin X (2014) Oleanolic acid suppresses aerobic glycolysis in cancer cells by switching pyruvate kinase type m isoforms. PLoS One 9:e91606. doi:10.1371/journal.pone.0091606
Zhou R, Zhang Z, Zhao L, Jia C, Xu S, Mai Q, Lu M, Huang M, Wang L, Wang X, Jin D, Bai X (2011) Inhibition of mTOR signaling by oleanolic acid contributes to its anti-tumor activity in osteosarcoma cells. J Orthop Res 29:846–852. doi:10.1002/jor.21311
Wang X, Bai H, Zhang X, Liu J, Cao P, Liao N, Zhang W, Wang Z, Hai C (2013) Inhibitory effect of oleanolic acid on hepatocellular carcinoma via ERK-p53-mediated cell cycle arrest and mitochondrial-dependent apoptosis. Carcinogenesis 34:1323–1330. doi:10.1093/carcin/bgt058
Way TD, Tsai SJ, Wang CM, Ho CT, Chou CH (2014) Chemical constituents of rhododendron formosanum show pronounced growth inhibitory effect on non-small-cell lung carcinoma cells. J Agric Food Chem 62:875–884. doi:10.1021/jf404243p
Shah MY, Calin GA (2014) MicroRNAs as therapeutic targets in human cancers. Wiley Interdiscip Rev RNA. doi:10.1002/wrna.1229
Ma L, Liu J, Shen J, Liu L, Wu J, Li W, Luo J, Chen Q, Qian C (2010) Expression of miR-122 mediated by adenoviral vector induces apoptosis and cell cycle arrest of cancer cells. Cancer Biol Ther 9:554–561
Fornari F, Gramantieri L, Giovannini C, Veronese A, Ferracin M, Sabbioni S, Calin GA, Grazi GL, Croce CM, Tavolari S, Chieco P, Negrini M, Bolondi L (2009) MiR-122/cyclin G1 interaction modulates p53 activity and affects doxorubicin sensitivity of human hepatocarcinoma cells. Cancer Res 69:5761–5767. doi:10.1158/0008-5472.CAN-08-4797
Coulouarn C, Factor VM, Andersen JB, Durkin ME, Thorgeirsson SS (2009) Loss of miR-122 expression in liver cancer correlates with suppression of the hepatic phenotype and gain of metastatic properties. Oncogene 28:3526–3536. doi:10.1038/onc.2009.211
Xu Y, Xia F, Ma L, Shan J, Shen J, Yang Z, Liu J, Cui Y, Bian X, Bie P, Qian C (2011) MicroRNA-122 sensitizes HCC cancer cells to adriamycin and vincristine through modulating expression of MDR and inducing cell cycle arrest. Cancer Lett 310:160–169. doi:10.1016/j.canlet.2011.06.027
Ma L, Liu J, Liu L, Duan G, Wang Q, Xu Y, Xia F, Shan J, Shen J, Yang Z, Bie P, Cui Y, Bian XW, Prieto J, Avila MA, Qian C (2014) Overexpression of the transcription factor MEF2D in hepatocellular carcinoma sustains malignant character by suppressing G2-M transition genes. Cancer Res 74:1452–1462. doi:10.1158/0008-5472.CAN-13-2171
Kang J, Kim E, Kim W, Seong KM, Youn H, Kim JW, Kim J, Youn B (2013) Rhamnetin and cirsiliol induce radiosensitization and inhibition of epithelial-mesenchymal transition (EMT) by miR-34a-mediated suppression of Notch-1 expression in non-small cell lung cancer cell lines. J Biol Chem 288:27343–27357. doi:10.1074/jbc.M113.490482
Zhang Y, Zhang GB, Xu XM, Zhang M, Qu D, Niu HY, Bai X, Kan L, He P (2012) Suppression of growth of A549 lung cancer cells by waltonitone and its mechanisms of action. Oncol Rep 28:1029–1035. doi:10.3892/or.2012.1869
Hu J, Xu Y, Hao J, Wang S, Li C, Meng S (2012) MiR-122 in hepatic function and liver diseases. Protein Cell 3:364–371. doi:10.1007/s13238-012-2036-3
Thomas M, Deiters A (2013) MicroRNA miR-122 as a therapeutic target for oligonucleotides and small molecules. Curr Med Chem 20:3629–3640
Kim MK, Kim SC, Kang JI, Hyun JH, Boo HJ, Eun SY, Park DB, Yoo ES, Kang HK, Kang JH (2011) 6-Hydroxydopamine-induced PC12 cell death is mediated by MEF2D down-regulation. Neurochem Res 36:223–231. doi:10.1007/s11064-010-0309-x
Liu AM, Xu Z, Shek FH, Wong KF, Lee NP, Poon RT, Chen J, Luk JM (2014) miR-122 targets pyruvate kinase M2 and affects metabolism of hepatocellular carcinoma. PLoS One 9:e86872. doi:10.1371/journal.pone.0086872
Liu J, Zheng L, Ma L, Wang B, Zhao Y, Wu N, Liu G, Lin X (2014) Oleanolic acid inhibits proliferation and invasiveness of Kras-transformed cells via autophagy. J Nutr Biochem. doi:10.1016/j.jnutbio.2014.06.006
Liu J, Zheng L, Zhong J, Wu N, Liu G, Lin X (2014) Oleanolic acid induces protective autophagy in cancer cells through the JNK and mTOR pathways. Oncol Rep 32:567–572. doi:10.3892/or.2014.3239
Liu J, Zheng L, Wu N, Ma L, Zhong J, Liu G, Lin X (2014) Oleanolic acid induces metabolic adaptation in cancer cells by activating the AMP-activated protein kinase pathway. J Agric Food Chem 62:5528–5537. doi:10.1021/jf500622p
Liu J, Wu N, Ma LN, Zhong JT, Liu G, Zheng LH, Lin XK (2014) p38 MAPK signaling mediates mitochondrial apoptosis in cancer cells induced by oleanolic acid. Asian Pac J Cancer Prev 15:4519–4525
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhao, X., Liu, M. & Li, D. Oleanolic acid suppresses the proliferation of lung carcinoma cells by miR-122/Cyclin G1/MEF2D axis. Mol Cell Biochem 400, 1–7 (2015). https://doi.org/10.1007/s11010-014-2228-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-014-2228-7