Abstract
Mammalian hibernators undergo significant physiological and biochemical changes when confronted with cold temperatures. Metabolic depression and translational repression are two examples of the various processes impacted during a torpor bout. MicroRNAs (miRNAs), non-coding transcripts that bind to mRNAs, are known regulators of mRNA translation and a growing number of these molecules have been found to be differentially expressed during hibernation. We hypothesized that a group of six miRNAs, with targets involved in various metabolic cascades, is modulated in selected tissues of the hibernating thirteen-lined ground squirrel Ictidomys tridecemlineatus. Expression levels of these miRNAs were assessed in the liver, heart, and skeletal muscle ground squirrel tissues using qRT-PCR. miR-29a, miR-152, miR-195, miR-223, and miR-486 were shown to be up-regulated in the hibernating liver, while miR-378 was shown to be down-regulated in hibernating skeletal muscle tissue samples. Interestingly, fatty acid synthase (FAS), an enzyme involved in fatty acid biosynthesis and a miR-195 target, was shown to be down-regulated in hibernating squirrel liver. This data add to the growing signature of differentially expressed miRNAs during hibernation and puts the light on the potential regulation of fatty acid homeostasis by a miRNA in torpid animals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hibernation is a strategy that involves tightly regulated metabolic suppression and is instigated by several mammals to survive severe environmental conditions. Mammalian hibernators undergo sustained torpor periods at low body temperature (T b ~4 °C) and can experience a metabolic rate reduction of more than 20-fold, when compared to the euthermic state [1–3]. Other physiological changes observed in animals undergoing hibernation notably include reduced blood flow, breathing rate, and heart rate [4, 5]. In hibernating ground squirrels, this hypometabolic state translates to energy savings of almost 90 % that would otherwise be consumed if the animals were to remain active during the cold season [1]. Such changes are correlated with considerable regulation of ATP-consuming processes including transcription and translation [6, 7]. Large-scale approaches performed on hibernating animals have indeed highlighted significant modulation of global transcript and protein levels during torpor [8, 9]. Selected studies on translation factors such as eIF4E and eEF2 have further emphasized the importance of translational control in hibernation [10, 11].
MicroRNAs (miRNAs) are non-coding RNAs that regulate translation of selected transcript targets in animals, plants, and protozoa [12]. miRNAs are initially transcribed as primary miRNAs (pri-miRNAs) by the RNA polymerase II and subsequently cleaved by the enzyme Drosha to yield precursor miRNAs (pre-miRNAs). Pre-miRNAs are exported to the cytoplasm and subsequently cleaved to generate mature miRNAs. miRNAs associate with several proteins to form the miRNA-induced silencing complex (miRISC). This complex interacts with target mRNAs to repress translation or to promote transcript degradation [12]. Recent predictions suggested the likelihood that miRNAs can regulate 60 % of all protein-coding genes [13]. Thus, it is not surprising that these short RNAs, usually 19–25 nucleotides in length, influence a myriad of metabolic cascades and physiological processes. This includes miRNAs that can influence insulin signaling and glucose metabolism as well as miRNAs that can regulate the expression of target transcripts involved in lipid homeostasis [14, 15].
The present study was undertaken to assess the expression of six miRNAs in a small mammalian hibernator. The characterized miRNAs have been linked, in non-hibernating models, with processes relevant to hibernation including glucose metabolism (miR-29a and miR-223), lipid metabolism, (miR-195) and muscle atrophy (miR-378). miR-152 and miR-486, miRNAs previously shown to be differentially regulated in another hibernating ground squirrel, were also selected for this study. We report the expressions of these miRNAs in three tissues of the euthermic and hibernating thirteen-lined ground squirrels, Ictidomys tridecemlineatus, and notably highlight the differential expression of miR-195, a miRNA with potential implications in fatty acid homeostasis.
Materials and methods
Animals
Animal experiments on thirteen-lined ground squirrels, I. tridecemlineatus, were performed as previously described [16]. Squirrels weighing 150–300 g were captured by a licensed trapper (TLS Research, Bloomingdale, IL) and transported to the NIH facility (Bethesda, MD). Animals were kept in shoebox cages maintained at 21 °C and fed ad libitum until they accumulated sufficient lipid stores to enter torpor. A sensor chip was injected subcutaneously in anesthetized animals. To induce hibernation, squirrels were moved to a dark cold room maintained at 4–6 °C. Body temperature (Tb) and respiration rate were closely followed to identify the stage of torpor-arousal cycle. The animals constituting the hibernating group were sacrificed following a torpor period of at least 5 days (Tb between 5 and 8 °C), while the control animals had demonstrated stable Tb readings (between 34 and 37 °C) for at least 3 days. All squirrels were sacrificed by decapitation. Tissues were subsequently shipped to Université de Moncton in New Brunswick on dry ice and stored at −80 °C until use.
Total RNA isolation
Total RNA was isolated from the liver, heart, and skeletal muscle tissues of ground squirrels using TRIzol reagent (Invitrogen) according to manufacturer’s instructions. Briefly, 100 mg of tissue was homogenized in 1 ml of TRIzol using a Polytron homogenizer followed by the addition of 0.2 ml chloroform. Samples were subsequently centrifuged at 12,000 rpm for 15 min at 4 °C. Following upper layer transfer to a different tube, 0.5 ml of isopropanol was added and the mixture was left at room temperature for 10 min to precipitate the RNA. Samples were centrifuged at 12,000 rpm for 10 min at 4 °C. The resulting RNA pellet was washed with 75 % ethanol and centrifuged at 7,500×g for 5 min at 4 °C. Pellets were dried and resuspended in 100 μl of DEPC-treated water. The final products were analyzed using a NanoVue Plus Spectrophotometer to determine RNA concentration as well as RNA purity via the absorbance ratio at 260/280 nm. All samples were stored at −80 °C until use.
cDNA synthesis
Ictidomys tridecemlineatus miR-specific stem-loop primers were designed based on a consensus alignment sequence of known miR-29a, miR-107, miR-152, miR-195, miR-223, miR-378, and miR-486 sequences obtained from Homo sapiens, Mus musculus, and Rattus norvegicus miRNA species. Sequences for stem-loop primers used are listed in Table 1. miRNAs were subsequently amplified using a stem-loop based RT-PCR protocol [17] to confirm that the designed primers could specifically amplify the miRNAs of interest. First-strand synthesis was performed as described previously [18]. Briefly, 1 μg of total RNA was combined with 5 μl of 250 nM of miRNAs-specific stem-loop primer. This mixture was first incubated for 5 min at 95 °C, subsequently placed at 60 °C for 5 min and finally cooled on ice for 1 min. The remaining reagents—4 μl of 5× first-strand buffer, 2 μl of 0.1 M dithiothreitol, 1 μl of 10 mM dNTPs, 2 μl DEPC-treated water, and 1 μl M-MLV reverse transcriptase (Life Technologies)—were added to the reaction mixture. Samples were incubated at 16 °C for 30 min, 42 °C for 30 min, and 85 °C for 5 min. Serial dilutions of the cDNA were prepared (10−1–10−4) in water and were used for PCR amplification of the different miRNAs.
qRT-PCR amplification of miRNAs
The qRT-PCR reaction was performed by mixing 5 μl of cDNA template (10−2) to 5.5 μl of DEPC-treated water, 1 μl of 25 μM miRNA-specific forward primer, 1 μl of 25 μM universal reverse primer, and 12.5 μl of EconoTaq PLUS GREEN 2X Master Mix (Lucigen). Sequences for forward and universal primers used are listed in Table 1. The amplification protocol consisted of an initial denaturing step at 95 °C for 10 min, followed by 35 cycles at 95 °C for 15 s and 60 °C for 60 s. PCR products were separated on 2 % agarose gels and bands were visualized on a UV box. Products were sequenced by the Université Laval sequencing platform (Quebec City, QC), and sequences were confirmed as encoding miR-29a, miR-107, miR-152, miR-195, miR-223, miR-378, and miR-486 by sequence comparison using BLAST. Initial qRT-PCR runs were performed to assess the amplification efficiencies for each primer set. Briefly, each miRNA was amplified in tenfold dilutions (10−1–10−4) of cDNA samples using the same qRT-PCR conditions as described above. Reactions showed efficiencies between 90 and 97 %, and miR-107 was selected as a housekeeping miRNA. This miRNA displayed considerable stability between the euthermic and hibernating ground squirrel tissue samples as assessed by the Excel 2003 add-in NormFinder version 0.953 [19].
Western blotting
The frozen ground squirrel liver, heart, or skeletal muscle tissue samples (~100 mg) were homogenized in 1 mL of buffer containing 100 mM MOPS, 25 mM HEPES, 25 mM β-glycerophosphate, 5 mM EDTA, 1 mM EGTA, and 250 μM Na3VO4, adjusted at pH 7.4, with 1 mM phenylmethylsulfonyl fluoride added immediately before homogenization. Homogenates were centrifuged at 10,000×g for 10 min at 4 °C, and supernatants were collected. Protein concentrations in lysates were determined using the Coomassie blue dye-binding method and the BioRad reagent (BioRad, Hercules, CA). SDS–polyacrylamide gel electrophoresis and blotting to polyvinylidene difluoride (PVDF) membranes were carried out as previously reported [20] with 8 % gels, 25 µg of protein lysates loaded per well, and electrophoresis at 110 V for 90 min. Proteins were subsequently transferred onto PVDF membranes for 70 min at 300 mA using a transfer buffer consisting of 25 mM Tris (pH 8.5), 192 mM glycine, and 10 % v/v methanol at 4 °C. Following transfer, membranes were blocked for 1 h in TBST (50 mM Tris–HCl pH 6.8, 150 mM NaCl, 0.05 % v/v Tween 20) with 5 % w/v powdered skim milk. This solution was removed and membranes were incubated overnight at 4 °C with primary antibodies solution. Antibodies specific for mammalian fatty acid synthase (FAS) were purchased from Abcam (#ab96863) and used at a 1:500 v:v dilution in TBST. The following day, membranes were incubated with HRP-linked anti-rabbit IgG secondary antibody (1:2,000 v:v dilution) in TBST for 1 h. Bands were visualized using the FluorChem imaging system and the FluorChem software (Alpha Innotech).
Quantification and statistics
For qRT-PCR results, quantification of the target miRNA products was performed using the Qbase + software version 2.5.1 (Biogazelle). Target miRNA levels were normalized against miR-107 amplified from the same cDNA sample. miRNA quantification was performed using the method described by Pfaffl, which accounts for differences in PCR efficiencies between the target and reference miRNAs [21]. For immunoblotting results, band intensity of protein of interest was normalized against band intensity of the housekeeping protein GAPDH in the same lane. Mean normalized band densities ± SEM were then calculated for aroused versus torpid samples, and significant differences between the groups were tested using the Student’s t test.
Results
Amplification and sequence of miRNAs in I. tridecemlineatus
Using RT-PCR and primers derived from consensus sequences of miRNAs obtained from other mammalian species, target miRNAs were amplified in ground squirrel tissues. The products were confirmed as encoding miR-29a, miR-107, miR-152, miR-195, miR-223, miR-378, and miR-486, and the sequences were submitted to GenBank with accession numbers KJ169551, KF856940, KF856942, KJ169552, KJ169553, KF856943, and KF856944, respectively. Figure 1 shows the nucleotide sequences of target miRNAs aligned with the corresponding miRNAs from three other mammalian sources sequenced to date. miRNAs are strongly conserved in mammals [22], and this was confirmed for the miRNAs amplified in this study.
MiR-29a, miR-152, miR-195, miR-223, miR-378, and miR-486 complete sequences in I. tridecemlineatus. MiR-29a (a), miR-152 (b), miR-195 (c), miR-223 (d), miR-378 (e), and miR-486 (f) sequences amplified from I. tridecemlineatus compared to sequences of H. sapiens, M. musculus, R. norvegicus, and/or E. caballus. Genbank accession numbers are AJ421751.1, AJ459720.1, NR_031836.1, and KJ169551 for miR-29a; NR_029687.1, AJ459764.1, NR_031894.1, and KF856942 for miR-152; NR_029712.1, AJ560737.1, NR_031912.1, and KJ169552 for miR-195; NR_029637.1, NR_029801.1, NR_031936.1, and KJ169553 for miR-223; NR_029870.1, NR_029879.1, NR_032139.1, and KF856943 for miR-378; and NR_030161.1, NR_030254.1, NR_033059.1, and KF856944 for miR-486, respectively. Dashes indicate identical nucleotides between sequences; spacer dots indicate missing nucleotides in one or more sequence
miRNA expression in hibernating tissues of I. tridecemlineatus
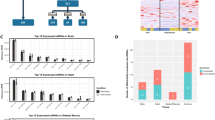
miRNA levels of miR-29a, miR-152, miR-195, miR-223, miR-378, and miR-486 were assessed via qRT-PCR in three tissues of the euthermic and hibernating I. tridecemlineatus (Fig. 2). miR-29a, miR-152, miR-195, miR-223, and miR-486 content was significantly higher (P < 0.05) in the liver of hibernating I. tridecemlineatus; values were 1.33 ± 0.06-, 1.79 ± 0.26-, 1.68 ± 0.19-, 4.15 ± 0.58-, and 1.62 ± 0.17-fold higher than in tissues from the euthermic animals, respectively. In skeletal muscle tissue of hibernating I. tridecemlineatus, miR-378 levels decreased significantly to 66.2 ± 5.7 % of the euthermic value (P < 0.05), while the other miRNAs quantified remained unchanged between torpid and the euthermic samples. None of the miRNAs tested in this study were differentially expressed in the hibernating heart.
Effects of hibernation on the levels of six miRNA species in the liver, skeletal muscle, and heart of I. tridecemlineatus. Histogram shows the ratios of normalized miRNA levels, versus the housekeeping miR-107, in tissues from hibernating versus the euthermic animals. Data are mean ± SEM for n = 5–6 independent trials on tissue from different animals. Expression ratios and standard errors were calculated using the relative expression software tool (Qbase + software version 2.5.1 from Biogazelle). *Significantly different from control group (P < 0.05)
FAS protein expression in the hibernating liver
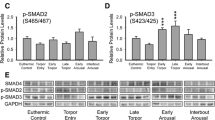
FAS protein levels were measured by immunoblotting in three ground squirrel tissues. Western blot analysis of FAS in ground squirrel liver showed that the content of the 273-kDa FAS protein decreased significantly in this tissue during hibernation to values that are 55.9 ± 20.4 % of the ones observed in the euthermic samples (Fig. 3; P < 0.05).
Discussion
miRNAs can regulate translation of several target transcripts and concurrently impact a plethora of cellular processes including metabolism, cell cycle regulation, and apoptosis [12, 23, 24]. In recent years, a growing body of evidence has highlighted the potential importance of this family of nucleotides in modulating responses to different stresses including anoxia and cold temperatures [25]. In this study, we hypothesized that the inhibitory control of transcript translation by miRNAs could be a significant mechanism to impact gene and protein expression of targets involved in key metabolic cascades in a small mammalian hibernator. The differential expression of miR-29a, miR-195, miR-223, miR-378, and miR-486 in the hibernating liver and/or skeletal muscle ground squirrel tissues reported in the current work supports this statement.
Increased miR-29a levels are observed in hibernating ground squirrel liver tissues. This miRNA has been shown to target numerous molecules involved in glucose homeostasis in the non-hibernating models. miR-29a overexpressions in primary hepatocytes and mouse livers were associated with a down-regulation in protein levels of PGC-1α and G6Pase, characterized targets of this miRNA, leading to reduced hepatic glucose production [26]. Previous studies have reported elevated PGC-1α protein levels in the hibernating liver and skeletal muscle tissues of M. lucifugus as well as in the hibernating heart and skeletal muscle tissues of I. tridecemlineatus suggesting that other miRNAs could be involved in regulating this target in hibernators [27, 28]. Measurements of miR-27 and miR-696 levels, two miRNAs that have been proposed as potential PGC-1α regulators, would yield interesting information on this topic [29, 30]. In addition, miR-29a overexpression was also correlated with insulin resistance in mouse adipocytes [31]. Interestingly, a recent publication has highlighted the fact that several pathways differentially modulated in hibernating animals are also frequently found to be deregulated during insulin resistance in human type 2 diabetes, and thus making hibernators prospective models to study this condition [32]. miR-29a might yet be another commonality between these two models.
Studies in non-hibernating animal models have highlighted the involvement of miR-195 in both the pathogenesis of type II diabetes as well as its ability to target a key player in fatty acid synthesis, the enzyme fatty acid synthase (FAS) [33, 34]. These findings prompted us to assess miR-195 levels in the current study. Overexpression of miR-195 was observed in the hibernating liver tissues of I. tridecemlineatus as well as reduced FAS protein levels in the same tissue suggesting a role for miR-195 in regulating a key node of the fatty acid synthesis pathway. A recent proteomics-based approach performed in hibernating I. tridecemlineatus has reported a reduction in enzyme levels notably involved in the lipid biosynthetic pathways further emphasizing the importance of this cascade in the hibernation phenotype [35]. The current findings add to this knowledge and provide yet another means by which hibernators can regulate fatty acid homeostasis during the torpor bouts. It will be interesting to further investigate additional miRNAs that can impact, directly or indirectly, the expression of enzymes involved in fatty acid synthesis including FAS and acetyl-CoA carboxylase (ACC). A list of miRNAs that can target key players of the fatty acid synthesis pathway is presented in Table 2. These notably include miR-424 and miR-451, which have been reported to impact FAS expression in osteosarcomas and ACC expression in smooth muscle cells, respectively [36, 37].
Increased miR-223 levels in the hibernating liver tissue samples were also reported in this study. A potential impact of this differential expression might be the translational regulation of FOXO1, a transcription factor with diverse functions. miR-223 can regulate FOXO1 expression in colorectal cancer cells [38]. FOXO transcription factors can modulate the transcription of genes leading to diverse cellular effects including oxidative stress and glucose metabolism [39]. Interestingly, differential transcript and protein levels of selected FOXO proteins were associated with stress resistance in the anoxia-tolerant turtle Trachemys scripta elegans [40]. Another relevant target that miR-223 could potentially regulate is the hepatic scavenger receptor class B type I (SR-BI), a protein involved in high-density lipoprotein cholesterol (HDL-C) uptake. A recent study demonstrated that miR-223 could interact with the 3′ UTR of SR-BI and prevent its translation, thus suggesting a potential implication of this miR in cholesterol metabolism [41]. Additional work will be required to further characterize miR-223 function in the torpid mammals, yet the strong overexpression observed in this study suggests that it should not be overlooked.
Interestingly, hibernating mammals are perceived as excellent natural models to assess the mechanisms underlying skeletal muscle plasticity and metabolism during prolonged periods of inactivity [42]. Differential expressions of MEF2 and MyoD, transcription factors with functions in skeletal muscle myogenesis, were reported in hibernating I. tridecemlineatus and provided a glimpse of the molecular switches involved in skeletal muscle dynamics at low temperatures [43]. A set of differentially expressed miRNAs likely involved in preserving pectoral muscle mass of the hibernating little brown bat Myotis lucifugus was recently reported [44]. The current study shows down-regulation of miR-378 in hibernating ground squirrel tissues and contributes to the growing list of miRNAs with a suspected implication in muscle integrity maintenance during the prolonged periods of inactivity as experienced by the torpid mammals. Interestingly, it was shown that MyoD could up-regulate miR-378 during myogenic differentiation in mouse myoblasts [45]. While a miR-378 transcript target with a role in skeletal muscle dynamics remains to be characterized to fully measure the significance of this measured change, differential expression of the MyoD-miR-378 axis in hibernating skeletal muscle tissues nevertheless reinforces its potential importance during hibernation.
The current work reported up-regulation of miR-152 and miR-486 levels in the hibernating liver tissues of I. tridecemlineatus. Differential expressions were also reported for these miRNAs in a study that looked at expressions of more than 200 miRNAs in liver tissue of another hibernating species, the Arctic ground squirrel Spermophilus parryii [46]. The transcripts targeted by these miRNAs as well as the significance of these variations during hibernation remain to be clarified. It is worth mentioning that work in non-hibernating model has linked miR-486 to a potential role in tumor progression and cell growth [47]. In addition, a recent study reported that miR-486 can directly target key components of insulin growth factor (IGF) signaling including insulin-like growth factor 1 (IGF1) and IGF1 receptor (IGF1R) in lung cancer cells [48]. Interestingly, IGF axis has been demonstrated to be significantly down-regulated in hibernating golden-mantled ground squirrels, Spermophilus lateralis [49]. Further work will be required to determine if these effects on IGF signaling are mediated by miR-486.
In conclusion, results presented here evaluated the expression levels of six miRNAs with potential metabolic relevance in three tissues of a small mammalian hibernator. Five of them displayed differential expressions in hibernating tissue samples including miR-195 in liver, a miRNA that can target a key enzyme involved in fatty acid synthesis. Reduced protein levels of this enzyme were also detected in the same tissue. Overall, this work further reinforces the potential implications of miRNAs in torpid animals and paves the way for a more detailed analysis of the specific pathways and transcripts targeted by miRNAs during hibernation.
References
Wang LCH, Lee TF (1996) Torpor and hibernation in mammals: metabolic, physiological, and biochemical adaptations. In: Fregley MJ, Blatteis CM (eds) Handbook of physiology: environmental physiology. Oxford University Press, New York
Geiser F (2004) Metabolic rate and body temperature reduction during hibernation and daily torpor. Annu Rev Physiol 66:239–274
Storey KB, Storey JM (2004) Metabolic rate depression in animals: transcriptional and translational controls. Biol Rev Camb Philos Soc 79:207–233
Wickler SJ, Hoyt DF, van Breukelen F (1991) Disuse atrophy in the hibernating golden-mantled ground squirrel, Spermophilus lateralis. Am J Physiol 261:1214–1217
Frerichs KU, Kennedy C, Sokoloff L, Hallenbeck JM (1994) Local cerebral blood flow during hibernation, a model of natural tolerance to “cerebral ischemia”. J Cereb Blood Flow Metab 14:193–215
Frerichs KU, Smith CB, Brenner M, DeGracia DJ, Krause GS, Marrone L, Dever TE, Hallenbeck JM (1998) Suppression of protein synthesis in brain during hibernation involves inhibition of protein initiation and elongation. Proc Natl Acad Sci USA 95:14511–14516
Osborne PG, Gao B, Hashimoto M (2004) Determination in vivo of newly synthesized gene expression in hamsters during phases of the hibernation cycle. Jpn J Physiol 54:295–305
Shao C, Liu Y, Ruan H, Li Y, Wang H, Kohl F, Goropashnaya AV, Fedorov VB, Zeng R, Barnes BM, Yan J (2010) Shotgun proteomics analysis of hibernating arctic ground squirrels. Mol Cell Proteomics 9:313–326
Xu Y, Shao C, Fedorov VB, Goropashnaya AV, Barnes BM, Yan J (2013) Molecular signatures of mammalian hibernation: comparisons with alternative phenotypes. BMC Genomics 14:567
Chen Y, Matsushita M, Nairn AC, Damuni Z, Cai D, Frerichs KU, Hallenbeck JM (2001) Mechanisms for increased levels of phosphorylation of elongation factor-2 during hibernation in ground squirrels. Biochemistry 40:11565–11570
van Breukelen F, Sonenberg N, Martin SL (2004) Seasonal and state-dependent changes of eIF4E and 4E-BP1 during mammalian hibernation: implications for the control of translation during torpor. Am J Physiol Regul Integr Comp Physiol 287:349–353
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Friedman RC, Farh KK, Burge CB, Bartel DP (2009) Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19:92–105
Guay C, Roggli E, Nesca V, Jacovetti C, Regazzi R (2011) Diabetes mellitus, a microRNA-related disease? Transl Res 157:253–264
Sacco J, Adeli K (2012) MicroRNAs: emerging roles in lipid and lipoprotein metabolism. Curr Opin Lipidol 23:220–225
McMullen DC, Hallenbeck JM (2010) Regulation of Akt during torpor in the hibernating ground squirrel, Ictidomys tridecemlineatus. J Comp Physiol B 180:927–934
Biggar KK, Kornfeld SF, Storey KB (2011) Amplification and sequencing of mature microRNAs in uncharacterized animal models using stem-loop reverse transcription-polymerase chain reaction. Anal Biochem 416:231–233
Lyons PJ, Poitras JJ, Courteau LA, Storey KB, Pier Jr Morin (2013) Identification of differentially regulated microRNAs in cold-hardy insects. Cryo Letters 34:83–89
Andersen CL, Jensen JL, Ørntoft TF (2004) Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64:5245–5250
Pier Jr Morin, Ni Z, McMullen DC, Storey KB (2008) Expression of Nrf2 and its downstream gene targets in hibernating 13-lined ground squirrels, Spermophilus tridecemlineatus. Mol Cell Biochem 312:121–129
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29:e45
Berezikov E, Guryev V, van de Belt J, Wienholds E, Plasterk RH, Cuppen E (2005) Phylogenetic shadowing and computational identification of human microRNA genes. Cell 120:21–24
Subramanian S, Steer CJ (2010) MicroRNAs as gatekeepers of apoptosis. J Cell Physiol 223:289–298
Bueno MJ, Malumbres M (2011) MicroRNAs and the cell cycle. Biochim Biophys Acta 1812:592–601
Lyons PJ, Lang-Ouellette D, Pier Jr Morin (2013) CryomiRs: towards the identification of a cold-associated family of microRNAs. Comp Biochem Physiol D 8:358–364
Liang J, Liu C, Qiao A, Cui Y, Zhang H, Cui A, Zhang S, Yang Y, Xiao X, Chen Y, Fang F, Chang Y (2013) MicroRNA-29a–c decrease fasting blood glucose levels by negatively regulating hepatic gluconeogenesis. J Hepatol 58:535–542
Eddy SF, Storey KB (2003) Differential expression of Akt, PPARgamma, and PGC-1 during hibernation in bats. Biochem Cell Biol 81:269–274
Eddy SF, Pier Jr Morin, Storey KB (2005) Cloning and expression of PPAR-gamma and PGC-1alpha from the hibernating ground squirrel, Spermophilus tridecemlineatus. Mol Cell Biochem 269:175–182
Aoi W, Naito Y, Mizushima K, Takanami Y, Kawai Y, Ichikawa H, Yoshikawa T (2010) The microRNA miR-696 regulates PGC-1{alpha} in mouse skeletal muscle in response to physical activity. Am J Physiol Endocrinol Metab 298:799–806
Sun L, Trajkovski M (2014) MiR-27 orchestrates the transcriptional regulation of brown adipogenesis. Metabolism 63:272–282
He A, Zhu L, Gupta N, Chang Y, Fang F (2007) Overexpression of micro ribonucleic acid 29, highly up-regulated in diabetic rats, leads to insulin resistance in 3T3-L1 adipocytes. Mol Endocrinol 21:2785–2794
Wu CW, Biggar KK, Storey KB (2013) Biochemical adaptations of mammalian hibernation: exploring squirrels as a perspective model for naturally induced reversible insulin resistance. Braz J Med Biol Res 46:1–13
Herrera BM, Lockstone HE, Taylor JM, Ria M, Barrett A, Collins S, Kaisaki P, Argoud K, Fernandez C, Travers ME, Grew JP, Randall JC, Gloyn AL, Gauguier D, McCarthy MI, Lindgren CM (2010) Global microRNA expression profiles in insulin target tissues in a spontaneous rat model of type 2 diabetes. Diabetologia 53:1099–1109
Mao JH, Zhou RP, Peng AF, Liu ZL, Huang SH, Long XH, Shu Y (2012) microRNA-195 suppresses osteosarcoma cell invasion and migration in vitro by targeting FASN. Oncol Lett 4:1125–1129
Rose JC, Epperson LE, Carey HV, Martin SL (2011) Seasonal liver protein differences in a hibernator revealed by quantitative proteomics using whole animal isotopic labeling. Comp Biochem Physiol D 6:163–170
Long XH, Mao JH, Peng AF, Zhou Y, Huang SH, Liu ZL (2013) Tumor suppressive microRNA-424 inhibits osteosarcoma cell migration and invasion via targeting fatty acid synthase. Exp Ther Med 5:1048–1052
Turczyńska KM, Bhattachariya A, Säll J, Göransson O, Swärd K, Hellstrand P, Albinsson S (2013) Stretch-sensitive down-regulation of the miR-144/451 cluster in vascular smooth muscle and its role in AMP-activated protein kinase signaling. PLoS One 8:e65135
Wu L, Li H, Jia CY, Cheng W, Yu M, Peng M, Zhu Y, Zhao Q, Dong YW, Shao K, Wu A, Wu XZ (2012) MicroRNA-223 regulates FOXO1 expression and cell proliferation. FEBS Lett 586:1038–1043
Greer EL, Brunet A (2005) FOXO transcription factors at the interface between longevity and tumor suppression. Oncogene 24:7410–7425
Krivoruchko A, Storey KB (2013) Anoxia-responsive regulation of the FoxO transcription factors in freshwater turtles, Trachemys scripta elegans. Biochim Biophys Acta 1830:4990–4998
Wang L, Jia XJ, Jiang HJ, Du Y, Yang F, Si SY, Hong B (2013) MicroRNAs 185, 96, and 223 repress selective high-density lipoprotein cholesterol uptake through posttranscriptional inhibition. Mol Cell Biol 33:1956–1964
Cotton CJ, Harlow HJ (2010) Avoidance of skeletal muscle atrophy in spontaneous and facultative hibernators. Physiol Biochem Zool 83:551–560
Tessier SN, Storey KB (2010) Expression of myocyte enhancer factor-2 and downstream genes in ground squirrel skeletal muscle during hibernation. Mol Cell Biochem 344:151–162
Kornfeld SF, Biggar KK, Storey KB (2012) Differential expression of mature microRNAs involved in muscle maintenance of hibernating little brown bats, Myotis lucifugus: a model of muscle atrophy resistance. Genomics Proteomics Bioinform 10:295–301
Gagan J, Dey BK, Layer R, Yan Z, Dutta A (2011) MicroRNA-378 targets the myogenic repressor MyoR during myoblast differentiation. J Biol Chem 286:19431–19438
Liu Y, Hu W, Wang H, Lu M, Shao C, Menzel C, Yan Z, Li Y, Zhao S, Khaitovich P, Liu M, Chen W, Barnes BM, Yan J (2010) Genomic analysis of miRNAs in an extreme mammalian hibernator, the Arctic ground squirrel. Physiol Genomics 42A:39–51
Qian S, Ding JY, Xie R, An JH, Ao XJ, Zhao ZG, Sun JG, Duan YZ, Chen ZT, Zhu B (2008) MicroRNA expression profile of bronchioalveolar stem cells from mouse lung. Biochem Biophys Res Commun 377:668–673
Peng Y, Dai Y, Hitchcock C, Yang X, Kassis ES, Liu L, Luo Z, Sun HL, Cui R, Wei H, Kim T, Lee TJ, Jeon YJ, Nuovo GJ, Volinia S, He Q, Yu J, Nana-Sinkam P, Croce CM (2013) Insulin growth factor signaling is regulated by microRNA-486, an underexpressed microRNA in lung cancer. Proc Natl Acad Sci USA 110:15043–15048
Schmidt KE, Kelley KM (2001) Down-regulation in the insulin-like growth factor (IGF) axis during hibernation in the golden-mantled ground squirrel, Spermophilus lateralis: IGF-I and the IGF-binding proteins (IGFBPs). J Exp Zool 289:66–73
Shin D, Shin JY, McManus MT, Ptácek LJ, Fu YH (2009) Dicer ablation in oligodendrocytes provokes neuronal impairment in mice. Ann Neurol 66:843–857
Acknowledgments
We thank Dr. J. M. Hallenbeck and Dr. D. C. McMullen (National Institute of Neurological Disorders and Stroke, Bethesda, MD, USA) for gracefully providing ground squirrel tissues. This work was supported by a Discovery Grant from the Natural Sciences and Engineering Research Council of Canada awarded to P.J.M.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lang-Ouellette, D., Morin, P.J. Differential expression of miRNAs with metabolic implications in hibernating thirteen-lined ground squirrels, Ictidomys tridecemlineatus . Mol Cell Biochem 394, 291–298 (2014). https://doi.org/10.1007/s11010-014-2105-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-014-2105-4