Abstract
Islet 1 (ISL1), a marker of second heart field progenitors, plays a crucial role in cardiomyocyte differentiation and proliferation. However, little is known about transcriptional regulating mechanisms on Isl1 gene expression. Recent studies have demonstrated that Wnt/β-catenin signaling regulates Isl1 expression during heart development. However, the detailed mechanisms still remain unclear. In the present study performed during differentiation of P19CL6 into cardiomyocytes, we explored the underlying regulating mechanisms on Wnt/β-catenin-mediated Isl1 expression after we first confirmed that Wnt/β-catenin signaling promoted cardiomyocyte differentiation partly through Isl1 activation. We found a novel TCF/LEF1 binding site that was located 2300 bp upstream of the Isl1 ATG. Furthermore, Wnt/β-catenin signaling upregulated histone H3K9 acetylation on TCF/LEF1 binding sites on the Isl1 promoter, resulting in upregulation of Isl1 expression. This Wnt-mediated H3K9 acetylation on the Isl1 promoter was modulated by the acetyltransferase CREB-binding protein (CBP), instead of p300, through interaction with β-catenin. Collectively, these results suggest that in early stages of cardiomyocyte differentiation Wnt/β-catenin signaling promotes Isl1 expression via two ways: a novel TCF/LEF1 binding site and H3K9 acetylation conducted by CBP on the Isl1 promoter. To our knowledge, this is the first study reporting Wnt/β-catenin-regulated H3K9 acetylation on promoters of its target genes. And this study gives new insights into transcriptional regulating mechanisms of Wnt-mediated Isl1 expression during cardiomyocyte differentiation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Islet 1 (ISL1), a LIM-homeodomain transcription factor, plays a key role in cardiac development. Fate mapping experiments with Isl1 Cre indicated that ISL1 marks second heart field (SHF) progenitors, which contribute to the formation of outflow tract, right ventricle, and much of the atria [1]. Mice homozygous null for Isl1 die at approximately E10.5, failing to develop heart structures derived from the SHF [1]. Isl1 mRNA is expressed in progenitors, but is downregulated as cells differentiate and enter the heart, although expression is maintained in a few subpopulations [2]. In human fetal hearts, ISL1+ cells are found within SHF derivatives, consistent with data in murine embryos [3]. Due to its spatial and temporal expression profile, ISL1 is regarded as a marker of cardiac progenitor cells (CPC). In the myocardium of postnatal rats, mice, and humans, ISL1+ cardiac progenitors have been identified, with potential to give rise to cardiomyocytes [4]. In both mouse and human, fetal ISL1+ progenitors are capable of differentiating to diverse cardiovascular lineages, including cardiomyocytes, smooth muscle cells, and endothelial cells [3, 5]. Recent studies show that ISL1+ multipotent cardiovascular progenitors can be generated from mouse iPS cells and spontaneously differentiate into cardiovascular lineages in vivo without teratoma [6]. These findings may pave the way for new strategies to repair damaged myocardium after infarction.
In addition, ISL1 lies atop a conservative transcriptional network governing SHF development and is involved in setting in motion a cascade of downstream transcriptional events. Targets downstream of ISL1, including Mef2c, Nkx2.5, and Fgf10, ultimately determine cardiac development [7–10]. Some partners which interact with ISL1, including CIP and Ajuba, have been identified, and repress transcriptional activity of Isl1 [11, 12]. However, mechanisms responsible for dynamic regulation of Isl1 expression are still unclear. Loss of FGF8 signaling disrupts Isl1 expression, indicating a positive role in Isl1 expression regulation [13]. Also, Isl1 is a direct target of Forkhead transcription factors in SHF-derived mesoderm [14]. During cardiomyocyte differentiation of the pluripotent mouse embryonic carcinoma cell line, P19CL6, the POU homeodomain protein OCT1 plays a crucial role in promoting Isl1 expression [15]. Recently, studies have identified inhibitors of Isl1 expression. BMP2 and BMP4 signaling downregulates Isl1 expression via miRNA-17-92 cluster [16]. In addition, polycomb repressive complex2 (PRC2) represses Isl1 expression by EZH2-mediated H3K27 trimethylation of the Isl1 promoter [17].
Wnt/β-catenin signaling has biphasic effects on cardiac differentiation, depending on the stage of differentiation [18]. Wnt/β-catenin signaling is required for early stages of cardiac differentiation but is decreased during terminal differentiation. In contrast, Wnt/Planar Cell Polarity (PCP) signaling is activated during differentiation. Research on the role of Wnt/β-catenin signaling during cardiac development has been carried out in different models. In chick, forced expression of either Wnt3a or Wnt8c represses heart formation from anterior mesoderm and promotes development of primitive erythrocytes from the precardiac region. Reciprocally, Wnt inhibitors Crescent and DKK1 induce formation of heart muscle or heart-specific gene expression in posterior lateral plate mesoderm [19]. In P19CL6 cells, activation of Wnt/β-catenin signaling promotes cardiomyocyte differentiation [20]. In ES cells, Wnt signaling is essential for mesodermal development [21]. In mice, ablation of β-catenin severely disrupts cardiogenesis, with alterations in the proliferation and apoptosis of cardiac progenitors [22]. Similarly, stabilization of β-catenin in cardiac progenitors results in expansion of SHF progenitors and inhibition of differentiation [23–25]. Although loss of β-catenin results in decreased Isl1 expression in SHF progenitors [22, 25], abnormally prolonged exposure to stabilized β-catenin can also cause Isl1 downregulation in SHF progenitors [26], perhaps in a feedback loop.
Despite the foregoing which demonstrates the role for Wnt/β-catenin signaling in regulating Isl1 expression, questions still remain as to mechanisms underlying this regulation. To address this issue, we utilized pluripotent P19CL6 cells, a well-validated model for dissection of pathways determining cardiomyocyte differentiation [27]. Our data demonstrated that Wnt/β-catenin promoted cardiomyocyte differentiation partly through Isl1 activation. Wnt/β-catenin signaling promotes Isl1 expression via two ways: a novel TCF/LEF1 binding site located 2300 bp upstream of the ATG on the Isl1 promoter and acetylation of histone H3K9 on TCF/LEF1 binding sites on the Isl1 promoter by the acetyltransferase CREB-binding protein (CBP).
Results
The involvement of Isl1 in Wnt-promoted cardiomyocyte differentiation of P19CL6 cells
To investigate mechanisms of Wnt-mediated Isl1 expression during cardiomyocyte differentiation, we utilized an in-vitro cardiomyocyte differentiation model using induction of P19CL6 cells by DMSO. The cardiomyocyte differentiation of the cells was demonstrated by the expression of several cardiac-specific genes (Fig. 1a—Electronic supplementary material) and the contraction after DMSO induction for 12 days (Movie 1—Electronic supplementary material). Our results, consistent with previous reports [15, 28] indicated successful cardiomyocyte differentiation of P19CL6 cells. Also, our results revealed that the expression profile of Wnt/β-catenin signaling molecules and transcriptional activation of Wnt/β-catenin target genes were similar to the Isl1 expression pattern which peaked at day 4, but attenuated at day 8 after DMSO induction (Fig. 1b–f—Electronic supplementary material). Besides, Wnt/β-catenin signaling upregulated Isl1 expression in early stages of P19CL6 cardiomyocyte differentiation (Fig. 2—Electronic supplementary material).
To further clarify whether Wnt/β-catenin signaling regulated cardiac differentiation via Isl1, we maintained Wnt/β-catenin signaling activity at a high level by culturing P19CL6 cells with Wnt3a from day 0 to day 4. As a result, expression of three cardiac-specific marker genes (gata4, mef2c and troponin T) was significantly elevated at day 6 and day 14 (Fig. 1a). However, this elevation was attenuated when P19CL6 cells were transfected with Isl1 siRNA from day 0 to day 6 (Fig. 1c, d). The percentage of beating colonies was also counted at day 14. Wnt3a treatment from day 0 to day 4 promoted formation of beating cardiomyocytes. However, no beating clones were observed when P19CL6 cells were exposed to Isl1 siRNA from day 0 to day 6 even when treated with Wnt3a prior to day 4 (Fig. 1b). Therefore, these results indicate that in early stages of cardiomyocyte differentiation, Wnt/β-catenin signaling upregulates Isl1 expression. Additionally, knockdown of Isl1 attenuates Wnt/β-catenin-promoted cardiac differentiation.
The involvement of ISL1 in Wnt-promoted cardiac differentiation of P19CL6 cells. The cardiac differentiation of P19CL6 cells was induced by DMSO. a Real-time RT-PCR was conducted at day 6 and day 14 to evaluate the expression of Isl1, gata4, mef2c, and Troponin T in P19CL6 cells treated with Wnt3a (100 ng/ml) at day 0 and day 2. P19CL6 cells without Wnt3a treatment at day 6 or day 14 were used as controls. Data represent three independent experiments. Each bar represents mean ± SD (**p < 0.01, vs the control). b Quantification of beating colonies was conducted at day 14 of culture. P19CL6 cells were treated by transient transfection with Isl1 siRNA or non-silencer RNA (NS) at both day 0 and day 3. Wnt3a was added (100 ng/ml) at day 0 and day 2. Data represent three independent experiments. Each bar represents mean ± SD (*p < 0.05, **p < 0.01, vs the cells at day 14). c, d Real-time RT-PCR and Western blot were performed at day 6 (c) and day 14 (d) to detect the expression of Isl1, gata4, mef2c, and Troponin T in P19CL6 cells. Cells were treated as mentioned above (b). Data represent three independent experiments, each in triplicate. P19CL6 cells at day 6 or day 14 were used as the control. Each bar represents mean ± SD (*p < 0.05, **p < 0.01, vs the control). GAPDH was used as an internal control for Western blot analysis
A novel β-catenin/LEF1 binding site on the Isl1 promoter
As demonstrated above, Wnt/β-catenin signaling increased Isl1 expression in early stages of differentiation. To detect novel binding sites of TCF/LEF1 on the Isl1 promoter, bioinformatics analysis was performed. MatInspector software revealed a conserved TCF/LEF1 binding site 2300 bp upstream of the ATG translation start site on the Isl1 promoter (Fig. 2a). The sequence covering the TCF/LEF1 binding site was used as a probe (the Isl1 probe) for subsequent electrophoretic mobility shift assays (EMSA). Results showed that a specific complex was formed with the Isl1 probe. Complex specificity was confirmed by incubating the nuclear extract with a 100-fold molar excess of unlabeled Isl1 probe prior to addition of the labeled probe. A control Isl1 probe carrying a mutation of the TCF/LEF1 binding site failed to compete for complex formation. Although the addition of unrelated antibody did not affect formation of the protein-DNA complex, formation of the complex was significantly attenuated in the presence of anti-LEF1 antibody, attesting to the specificity of the protein-DNA complex (Fig. 2b).
The direct recruitment of β-catenin/LEF1 on the Isl1 promoter. a The putative TCF/LEF1 binding site at 2300 bp upstream of Isl1 ATG was identified using professional MatInspector database and indicated by a box. Alignment of Isl1 5′UTR was performed with vector NTI10.0 software among human, mouse, and rat species, and the conserved sequences between all the species are shaded light gray. b EMSA was performed with HeLa cell nuclear extracts (transfected with pcDNA3.1 [28] or pCG-LEF1) for the β-catenin/LEF1 proteins binding ability to the 32P-labeled oligonucleotides containing the putative LEF1 binding site on the Isl1 promoter. Lane 1 free probe, Lane 2 HeLa nuclear extracts transfected with pcDNA3.1 construct, Lane 3 HeLa nuclear extracts transfected with LEF1 expression construct, Lane 4 100× wild-type (wt) unlabeled oligonucleotide competition, Lane 5 100× mutant (mt) unlabeled oligonucleotide competition, Lane 6 anti-LEF1 antibody, Lane 7 unrelated antibody: goat normal IgG. c Luciferase assay was performed to evaluate the transcriptional activity of the Isl1 promoter. 293T cells were transiently transfected with Isl1 promoter-luciferase reporter constructs in combination with either pcDNA3.1/β-gal, LEF1, or β-catenin expression constructs. pGL(−2772/−65), wild-type Isl1 promoter-luciferase construct; Mut LEF1 pGL(−2772/−65), Isl1 promoter-luciferase construct with mutated LEF1 consensus sites. Data are presented as fold activity over basal promoter activity (relative activity). Each bar represents mean ± SD from three independent experiments, each in triplicate (**p < 0.01, vs the control). d ChIP assay was performed with antibodies against β-catenin at day 0 or day 4. The immunoprecipitated DNA fragments were amplified by real-time PCR for the Isl1 promoter region from −2940 to −2630 (−2900 bp left panel) or −2341 to −2203 (−2300 bp right panel), respectively. Input represents 10 % of the total input chromatin
To further characterize the functional significance of LEF1 binding to the Isl1 promoter, luciferase assays were performed with a −2772/−65 bp Isl1-promoter-luciferase reporter and expression vectors for LEF1 or β-catenin. Results demonstrated activation of the Isl1-promoter by recruitment of LEF1 and activated β-catenin. This activation was disrupted by a mutation of the LEF1 binding site located 2300 bp upstream of Isl1 ATG (Fig. 2c). To further evaluate recruitment of β-catenin to the Isl1 promoter, including a previously reported TCF/LEF1 binding site located 2900 bp upstream of the ATG translation start site on the Isl1 promoter [22], ChIP assays were conducted in P19CL6 cells at day 0 and day 4. Our data showed that recruitment of β-catenin onto two TCF/LEF1 binding sites on the Isl1 promoter was augmented after DMSO induction for 4 days (Fig. 2d). In summary, our data reveal that during cardiomyocyte differentiation of P19CL6 cells, Wnt-promoted Isl1 expression is partially dependent on a novel TCF/LEF1 binding site located 2300 bp upstream of Isl1 ATG.
Wnt-promoted H3K9 acetylation on TCF/LEF1 binding sites on the Isl1 promoter during cardiomyocyte differentiation of P19Cl6 cells
Increasing evidence shows that histone modifications take part in Wnt-responsive gene expression [29]. To investigate whether histone modification is involved in Wnt-promoted Isl1 expression, a series of ChIP assays were conducted. Our data showed that mono- and tri-methylation of histone H3K4 and trimethylation of H3K9 in the two Isl1 promoter regions containing TCF/LEF1 binding sites were not changed at day4 of DMSO induction. Acetylation of H3 also remained unchanged following treatment of P19CL6 with Wnt3a or LiCl (data not shown). However, acetylation of H3K9 on the Isl1 promoter was significantly increased at day 4 (Fig. 3a), relative to day 0. Stimulation with Wnt3a also resulted in increased H3K9 acetylation on the Isl1 promoter at day 0 (Fig. 3b). Consistently, the application of DKK1 and Frizzled-4/Fc showed inhibitory effect on H3K9 acetylation on the Isl1 promoter (Fig. 3c, d). These results suggest that Wnt/β-catenin signaling modulates H3K9 acetylation on the Isl1 promoter from day 0 to day 4 during P19CL6 cardiomyocyte differentiation.
Wnt-promoted H3K9 acetylation on the Isl1 promoter during cardiomyocyte differentiation of P19Cl6 cells. a ChIP assay was conducted in P19CL6 cells at day 0 and day 4, with an antibody against acetylated H3K9. b–d ChIP assay was performed to detect H3K9 acetylation in response to treatment with Wnt3a (100 ng/ml, 24 h) at day 0 (b), and DKK1 (100 ng/ml) (c) or Frizzled-4/Fc (100 ng/ml) (d) through day 0–4 during P19CL6 cells differentiation into cardiomyocytes. Immunoprecipitated DNA fragments were amplified by real-time PCR for two Isl1 promoter regions containing TCF/LEF1 binding sites (−2900 bp: from −2940 to −2630; −2300 bp: from −2341 to −2203). The data are normalized on the basis of the corresponding input. The data for the 2900 bp binding site from P19CL6 cells uninduced or induced at day 4 without any treatment were used as a control. Each bar represents mean ± SD from three independent experiments, each in triplicate (*p < 0.05, **p < 0.01, vs the control)
Roles of CBP in Wnt-promoted acetylation of H3K9 on TCF/LEF1 binding sites on the Isl1 promoter
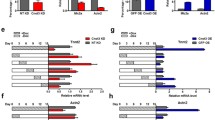
Histone acetylation is regulated by histone acetyltransferases (HATs) and histone deacetylases (HDACs). To find out which HATs or HDACs modulate Wnt-promoted acetylation of H3K9 on the Isl1 promoter, a series of ChIP assays were performed for the two aforementioned TCF/LEF1 binding sites on the Isl1 promoter. Our ChIP assays indicated that HDAC1 and PCAF had no significant effect on Wnt-induced H3K9 acetylation in the Isl1 promoter regions (data not shown). Although both CBP and p300 co-immunoprecipitated with β-catenin at day 4 (Fig. 4a), no significant enrichment for p300 was observed in the two Isl1 promoter regions at the same time, compared to nonspecific binding in the IgG IP samples. By contrast, CBP significantly recruited onto two TCF/LEF1 binding sites on the Isl1 promoter at day 4 (Fig. 3—Electronic supplementary material). Besides, accumulation of CBP was increased at day 4, compared with that at day 0 (Fig. 4b). Coincidently, Wnt3a stimulation increased accumulation of CBP on the Isl1 promoter at day 0 (Fig. 4c), consistent with attenuated recruitment of CBP when P19Cl6 cells were treated with DKK1 and Frizzled-4/Fc prior to day 4 (Fig. 4d, e). These results suggested that Wnt/β-catenin signaling promoted accumulation of CBP, on the Isl1 promoter. Also, overexpression of cbp increased Isl1 expression (Fig. 4f). To further investigate whether CBP is involved in Wnt-promoted H3K9 acetylation on the Isl1 promoter, we overexpressed cbp from day 0 to day 4. ChIP assays demonstrated that CBP attenuated negative effects of Dkk1 and Frizzled on H3K9 acetylation on the Isl1 promoter (Fig. 4g, h). Therefore, our results document that Wnt-mediated acetylation of H3K9 on the Isl1 promoter is conducted by CBP which interacts with endogenous β-catenin.
The roles of CBP in Wnt-promoted acetylation of H3K9 on the Isl1 promoter. a CoIP assay was performed with cell lysates from P19CL6 cells at day 4. Immunocomplex was precipitated with anti-CBP or p300 antibodies and then detected by anti-β-catenin antibody. 10 % of total cell lysates were used as an input. Normal IgG was used as a control antibody. b–e ChIP assay was carried out to detect CBP recruitment on two TCF/LEF1 binding sites on the Isl1 promoter in P19CL6 cells at day 0 and day 4 (b), or treated with Wnt3a (100 ng/ml, 24 h) at day 0 (c), or exposure to DKK1 (100 ng/ml) (d) and Frizzled-4/Fc (100 ng/ml) (e) through day 0–4. The data for the 2900 bp binding site from P19CL6 cells uninduced or induced at day 4 without any treatment were used as controls (*p < 0.05, **p < 0.01, vs the control). f Real-time RT-PCR and Western blot were performed to evaluate the expression of Isl1 in P19CL6 cells transfected with CBP plasmids (*p < 0.05, vs the cells transfected with pcDNA3.1/β-gal). GAPDH was used as an internal control. g–h ChIP assay was conducted to analyze the H3K9 acetylation on two TCF/LEF1 binding sites on the Isl1 promoter after treated with the inhibitors of canonical Wnt/β-catenin signaling. P19CL6 cells overexpressing cbp were incubated with DKK1 (100 ng/ml) (g) or Frizzled-4/Fc (100 ng/ml) (h) through day 0–4. The cells were harvested at day 4. The DNA extractions were amplified using the primers that covered two TCF/LEF1 binding sites on the Isl1 promoter (−2900 bp: from −2940 to −2630; −2300 bp: from −2341 to −2203) by real-time PCR. The data are normalized on the basis of the corresponding input control. The data for the 2900 bp binding site from P19CL6 cells induced at day 4 without any treatment were used as controls. Each bar represents mean ± SD from three independent experiments, each in triplicate (*p < 0.05, **p < 0.01, vs the cells transfected with pcDNA3.1/β-gal)
Discussion
ISL1, as a marker of SHF progenitors, plays a crucial role in cardiomyocyte differentiation and proliferation [1]. ISL1+ CPCs give rise to cardiomyocytes, smooth muscle, and endothelial cell lineages in both mice and humans [3, 5]. Isl1 mutant mice die due to lethal cardiac defects. Studies of human populations demonstrate that genetic variants and specific haplotypes in the ISL1 gene are associated with congenital heart disease [30, 31], partly via effecting activation of Mef2c [32]. Aside from roles in cardiac development, increasing evidence shows that ISL1+ cells give rise to adult resident cardiac stem cells. Studies of adult mice hearts find that ISL1+ progenitors persist into adulthood in mice and rats [33, 34]. Recent studies have been focused on the role of Isl1 after myocardial infarction. However, the results seem contradictory. In the spleen and left ventricle, Isl1 is re-expressed as a response to myocardial infarction, suggesting a potential role in cardiac repair [35]. Other studies show that ISL1+ cells are scarcely detected in the infarct zone after myocardial infarction, raising questions as to whether they contribute to cardiac repair [34]. To sum up, ISL1 is crucial in cardiac development, and may be a potential target for myocardial infarction treatment. However, many questions remain unanswered. To understand mechanisms that regulate Isl1 expression is to understand mechanisms that drive heart development as well as to understand potential strategies for heart regeneration.
Wnt/β-catenin signaling exerts developmental stage-specific effects on cardiac development by promoting cardiogenesis but inhibiting terminal cardiomyocyte differentiation [36]. Several transcription factors are regulated by Wnt/β-catenin signaling during cardiac development, including BRY, NKX2.5, and GATA6 [37–39]. Aside from selective TCF/LEF1 occupancy, chromatin around TCF-bound states in Wnt-inducible promoters shows active histone marks, including DNA hypomethylation, H3 hyperacetylation, and H3K4 trimethylation [29]. Several epigenetic modifiers recruited by TCF/LEF1 have been identified to be involved in Wnt-regulated gene expressions. For example, when Wnt/β-catenin signaling is poised for activation, HDAC1 dissociates from LEF1 to facilitate binding of β-catenin to LEF1, indicating that repression by LEF1 requires HDAC activity [40]. In addition, recruitment of β-catenin with different acetyltransferases, including p300, CBP, and PCAF has been reported [41, 42].
In this study, we investigated mechanisms involved in Wnt-mediated Isl1 expression during P19CL6 cardiomyocyte differentiation. We first confirmed that Wnt/β-catenin molecules and Isl1 exhibited similar expression pattern and Wnt-upregulated Isl1 expression in early stages of cardiomyocyte differentiation. In addition, we found that expression of three cardiac-specific markers, gata4, mef2c, and Troponin T, which mark early, middle, and late stages of cardiac differentiation, respectively, increased significantly following incubation of P19CL6 cells with Wnt3a. However, knocking down Isl1 inhibited expression of all three markers to varying degrees, even when Wnt3a was added to P19CL6 cells prior to day 4. Addition of Wnt3a significantly increased the percentage of beating clones. However, in Isl1-deficient P19CL6 cells, no beating cardiomyocytes were observed. These results suggested that activation of Wnt/β-catenin signaling in early stages promoted the generation of cardiomyocytes partly through Isl1.
Furthermore, we investigated mechanisms of Wnt-mediated Isl1 expression and found a novel TCF/LEF1-binding site located 2300 bp upstream of the Isl1 ATG. EMSA and luciferase assays showed that Wnt/β-catenin signaling positively regulated transcription of Isl1 by direct binding of β-catenin to the promoter. Before our study, another TCF/LEF1 binding site located 2900 bp upstream of the Isl1 ATG had been reported [22]. Our ChIP assays of these two TCF/LEF1 binding sites revealed β-catenin binding during DMSO-induced P19CL6 cardiomyocyte differentiation. In addition, our data showed that Wnt/β-catenin signaling promoted histone H3K9 acetylation on two TCF/LEF1 binding sites on the Isl1 promoter, but no significant changes in H3K4 de- or tri-methylation, H3K9 trimethylation or H3 acetylation.
Two possible mechanisms might be involved in Wnt-mediated H3K9 acetylation. Wnt/β-catenin signaling may activate HAT or facilitate HAT binding to target chromatin regions. Alternatively, Wnt/β-catenin signaling may inhibit HDAC activity or the binding of HDAC proteins to chromatin. At first, we were inclined toward the second possibility, because Wnt/β-catenin signaling promotes Nkx2-5 expression via HDAC1 [38]. However, we failed to observe any changes in accumulation of HDAC1 on the Isl1 promoter after P19CL6 cells were treated with Wnt3a, none did an acetyltransferase PCAF. Previous study reported interaction of β-catenin with two acetyltransferases coactivators, p300 and CBP, with diverse functions in murine embryonic stem cells [43]. In our study, we observed interaction between β-catenin and CBP and p300 at day 4. However, we failed to observe significant recruitment of p300 to the Isl1 promoter at the same time. In contrast, CBP was significantly recruited to the two TCF/LEF1 binding sites on the Isl1 promoter in P19CL6 induced at day 4. Furthermore, Wnt/β-catenin signaling regulated recruitment of CBP to the Isl1 promoter when P19CL6 cells treated with a Wnt activator or inhibitors. Overexpression of cbp promoted expression of Isl1, indicating the regulation of CBP to Isl1. These data suggested that Wnt/β-catenin signaling promoted the recruitment of CBP to the Isl1 promoter, as well as CBP-promoted H3K9 acetylation in the two TCF/LEF1 binding sites on the Isl1 promoter. Additionally, overexpression of cbp partially rescued H3K9 hypoacetylation on the Isl1 promoter caused by DKK1 or Frizzled-4/Fc, indicating that Wnt/β-catenin signaling promoted H3K9 acetylation on the Isl1 promoter partly via CBP. Based on a previous study showing that interaction between β-catenin and CBP induces ES cell proliferation and interaction between β-catenin and p300 initiates a differentiation program [43], our data indicated that ISL1 might play a role in CPCs proliferation, which is consistent with the previous study [1]. Interestingly, we found that at day 4, on the 2300 bp TCF/LEF1 binding site, in most situations, both H3K9 acetylation and CBP recruitment are more than that on the 2900 bp binding site, indicating the possible preference between these two binding sites.
In conclusion, we found that Wnt/β-catenin signaling regulates cardiomyocyte differentiation partially via Isl1. Wnt/β-catenin signaling regulates Isl1 expression through a novel TCF/LEF1 binding site and H3K9 acetylation on TCF/LEF1 binding sites on the Isl1 promoter modulated by CBP. This study first reports that Wnt/β-catenin-regulated H3K9 acetylation on promoters of its target genes and provides new insights into mechanisms of Wnt-mediated Isl1 expression during cardiomyocyte differentiation.
Experimental procedures
Cell culture, induction of differentiation and reagents
P19CL6 cells were cultured and induced to cardiomyocytes by DMSO as described previously [38]. After induction for 14 days, the spontaneously contracting colonies were counted, and the percentage of beating colonies was calculated. At least, three independent experiments were performed. As activators of Wnt pathway, recombinant mouse Wnt3a (100 ng/ml, 24 h) (catalog no. 1324-WN) and LiCl (10 mM, 12 h) were purchased from R&D Systems, Inc., (Minneapolis, MN, USA) and Sigma-Aldrich Corporation (St. Louis, MO, USA). Frizzled-4/Fc (100 ng/ml) (catalog no. 194-fz) and DKK1 (100 ng/ml) (catalog no. 5897-DK/CF), as inhibitors of Wnt pathway, were purchased from R&D Systems, Inc., (Minneapolis, MN, USA). The concentrations of these reagents were optimized in our pilot studies (data not shown) and previous reports [28, 38].
RT-PCR and real-time PCR
Total RNA extraction from P19CL6 cells and real-time RT-PCR were performed as previously described [15]. To quantify each experiment, the C t value was relatively quantified using Gapdh as an internal reference control. Transcript levels were normalized to the average expression levels of the controls or undifferentiated samples. Each value represented the average of at least three independent experiments, each in triplicate. The primers are listed in Table 1.
Western blot
Western blot was performed as previously described [15, 28]. Antibodies were used, including Islet1 (ab-109517, Abcam, Cambridge, MA, USA), Gata4 (sc-25310, Santa Cruz, CA, USA), GAPDH (ab-9484, Abcam, Hong Kong, China), β-catenin (sc-7199, Santa Cruze, CA, USA), Troponin T (ab-10214, Abcam, Cambridge, MA, USA), and horseradish peroxidase-conjugated secondary antibody from Santa Cruz.
Luciferase assay
The DNA fragment upstream the Isl1 start codon was amplified and cloned into pGL3-basic vector (Promega). Primers were 5′ primer 5′-AAGGTACCACATAGCAATCAACTAGGGGAGGAGGAGAAA-3′ and 3′ primer 5′-AACTCGAGAGAGGATGCTGGTGCTGTGGCTAGG-3′. The QuikChange sited-directed mutagenesis kit (Stratagene) was used to make a point mutation in the conserved LEF1-binding site in the Isl1 5′-promoter region according to the manufacturer’s protocol. The mutation primers were 5′ primer 5′-GAAGAGTGAGGGTCAGGTGTGGGTCTCAAGTC-3′ and 3′ primer 5′-GACTTGAGACCCACACCTGACCCTCACTCTTC-3′. Transfections were carried out in P19CL6 or HeLa cells using Lipofectamine™2000 (Invitrogen, Carlsbad, CA, USA). For luciferase reporter analysis, pcDNA3.1/β-gal was used as a control for measuring transfection efficiency. Data are presented as fold activity over Renilla luciferase activity (relative activity). The data represent three independent experiments, each in triplicate. Each bar represents mean ± SD from three samples.
Plasmids transfection and RNA interference
P19CL6 cells were transfected with plasmids or synthesized siRNA for gene overexpression or knockdown, respectively. The PCG-LEF1-HA construct was provided by Dr. Klaus Wolff (Department of Experimental Dermatology at the University of Vienna, Austria). Constitutively active β-catenin was obtained from Dr. Roel Nusse (Stanford University School of Medicine, USA). CBP plasmid was kindly provided by Dr. Kendall Nettles (Department of Cancer Biology, the Scripps Research Institute Florida, USA). Non-silencer siRNA (sense: UUCUCCGAACGUGUCACGUTT; antisense: ACGUGACACGUUCGGAGAAT) and Isl1 siRNA (sense: UCCUUCAUGAGCGCAUCUGTT; antisense: CAGAUGCGCUCAUGAAGGATT) were synthesized by Sigma. Transfection of the plasmids or siRNA was carried out by LipofectamineTM2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocols. After 48 h of transfection, cells transfected with the plasmids were harvested at indicated days of differentiation. For RNA interference, cells were transfected with the siRNA at day 0 and day 3, and harvested at indicated days. All the transfections were performed at least three times.
Eletrophoretic mobility shift assay (EMSA)
Nuclear extracts were prepared from HeLa cells. Oligonucleotides used in EMSA assay were labeled with [γ-32P] ATP as described previously [44]. The sequences of the sense strand of these oligonucleotides were as follows. Wild-type Isl1 probe: 5′-GAA GAG TGA GGG TCC TTT GTG GGT CTC AAG TCA C-3′; Mutant Isl1 probe: 5′-GAA GAG TGA GGG TCA GGT GTG GGT CTC AAG TCA C-3′. Antibody analyses were performed by pre-incubation of nuclear protein extract with specific LEF1 antibody (sc-8591, Santa Cruze, CA, USA).
Chromatin immunoprecipitation (ChIP) assays
For ChIP experiments, extracts were prepared from P19CL6 cells treated as indicated, and the lysates were immunoprecipitated with nonspecific IgG, β-catenin (sc-7199, Santa Cruze, CA, USA), acetylated-H3K9 (ab-108112, Abcam, Cambridge, MA), and CBP (sc-369, Santa Cruze, CA, USA) antibodies for 12 h at 4 °C. Immune complexes were incubated with Protein A/G-Sepharose CL-4B (Amersham Biosciences, Uppsala, Sweden) for 2 h at 4 °C. Protein-DNA cross-linking was reversed by overnight incubation at 65 °C. The precipitated DNA was amplified by real-time PCR for fragments of the Isl1 promoter. The following PCR primers were used: primers P-2940 (5′-GCG CCA GGA ACT GTG CTC CAA-3′) and P-2630 (5′-AGG GGC GAC CTC TTG TGT TCA ATG-3′); primers P-2341 (5′-GGT GAA TGC CTG TAT ATG TTT GGA-3′) and P-2203 (5′-AAA TAG GGG ATC TGA ACC AAT TCT-3′).
Co-immunoprecipitation (CoIP) assay
For CoIP experiments, the lysates derived from P19CL6 cells at day 4 were incubated with anti-CBP (sc-369, Santa Cruze, CA, USA) or p300 (RW-128, upstate Biotechnology, NY, USA) or normal IgG overnight at 4 °C, and then with 50 μl of Protein A/G-Sepharose CL-4B (Amersham Biosciences, Uppsala, Sweden) for 3 h at 4 °C. Immunoprecipitates were washed four times with wash buffer and subjected to SDS-PAGE electrophoresis, then analyzed by Western blot using anti-β-catenin (sc-7199, Santa Cruze, CA, USA) antibody.
Statistic analysis
The data were displayed as mean ± standard deviation (SD). Comparisons between groups were analyzed by Student’s t test or ANOVA. The significance was analyzed with SPSS10.0 software, and a p-value <0.05 was considered to be statistically significant.
References
Cai CL, Liang X, Shi Y, Chu PH, Pfaff SL, Chen J, Evans S (2003) Isl1 identifies a cardiac progenitor population that proliferates prior to differentiation and contributes a majority of cells to the heart. Dev Cell 5:877–889
Sun Y, Liang X, Najafi N, Cass M, Lin L, Cai CL, Chen J, Evans SM (2007) Islet 1 is expressed in distinct cardiovascular lineages, including pacemaker and coronary vascular cells. Dev Biol 304:286–296
Bu L, Jiang X, Martin-Puig S, Caron L, Zhu S, Shao Y, Roberts DJ, Huang PL, Domian IJ, Chien KR (2009) Human ISL1 heart progenitors generate diverse multipotent cardiovascular cell lineages. Nature 460:113–117
Laugwitz KL, Moretti A, Lam J, Gruber P, Chen Y, Woodard S, Lin LZ, Cai CL, Lu MM, Reth M, Platoshyn O, Yuan JX, Evans S, Chien KR (2005) Postnatal isl1+ cardioblasts enter fully differentiated cardiomyocyte lineages. Nature 433:647–653
Moretti A, Caron L, Nakano A, Lam JT, Bernshausen A, Chen Y, Qyang Y, Bu L, Sasaki M, Martin-Puig S, Sun Y, Evans SM, Laugwitz KL, Chien KR (2006) Multipotent embryonic isl1+ progenitor cells lead to cardiac, smooth muscle, and endothelial cell diversification. Cell 127:1151–1165
Moretti A, Bellin M, Jung CB, Thies TM, Takashima Y, Bernshausen A, Schiemann M, Fischer S, Moosmang S, Smith AG, Lam JT, Laugwitz KL (2010) Mouse and human induced pluripotent stem cells as a source for multipotent Isl1+ cardiovascular progenitors. FASEB J 24:700–711
Takeuchi JK, Mileikovskaia M, Koshiba-Takeuchi K, Heidt AB, Mori AD, Arruda EP, Gertsenstein M, Georges R, Davidson L, Mo R, Hui CC, Henkelman RM, Nemer M, Black BL, Nagy A, Bruneau BG (2005) Tbx20 dose-dependently regulates transcription factor networks required for mouse heart and motoneuron development. Development 132:2463–2474
Dodou E, Verzi MP, Anderson JP, Xu SM, Black BL (2004) Mef2c is a direct transcriptional target of ISL1 and GATA factors in the anterior heart field during mouse embryonic development. Development 131:3931–3942
Golzio C, Havis E, Daubas P, Nuel G, Babarit C, Munnich A, Vekemans M, Zaffran S, Lyonnet S, Etchevers HC (2012) ISL1 directly regulates FGF10 transcription during human cardiac outflow formation. PLoS One 7:e30677
Watanabe Y, Zaffran S, Kuroiwa A, Higuchi H, Ogura T, Harvey RP, Kelly RG, Buckingham M (2012) Fibroblast growth factor 10 gene regulation in the second heart field by Tbx1, Nk2-5, and Islet1 reveals a genetic switch for down-regulation in the myocardium. Proc Natl Acad Sci USA 109:18273–18280
Witzel HR, Jungblut B, Choe CP, Crump JG, Braun T, Dobreva G (2012) The LIM protein Ajuba restricts the second heart field progenitor pool by regulating Isl1 activity. Dev Cell 23:58–70
Huang ZP, Young Seok H, Zhou B, Chen J, Chen JF, Tao Y, Pu WT, Wang DZ (2012) CIP, a cardiac Isl1-interacting protein, represses cardiomyocyte hypertrophy. Circ Res 110:818–830
Park EJ, Ogden LA, Talbot A, Evans S, Cai CL, Black BL, Frank DU, Moon AM (2006) Required, tissue-specific roles for Fgf8 in outflow tract formation and remodeling. Development 133:2419–2433
Kang J, Nathan E, Xu SM, Tzahor E, Black BL (2009) Isl1 is a direct transcriptional target of Forkhead transcription factors in second-heart-field-derived mesoderm. Dev Biol 334:513–522
Liu Y, Li Y, Li T, Lu H, Jia Z, Wang W, Chen P, Ma K, Zhou C (2011) POU homeodomain protein OCT1 modulates islet 1 expression during cardiac differentiation of P19CL6 cells. Cell Mol Life Sci 68:1969–1982
Wang J, Greene SB, Bonilla-Claudio M, Tao Y, Zhang J, Bai Y, Huang Z, Black BL, Wang F, Martin JF (2010) Bmp signaling regulates myocardial differentiation from cardiac progenitors through a MicroRNA-mediated mechanism. Dev Cell 19:903–912
He A, Ma Q, Cao J, von Gise A, Zhou P, Xie H, Zhang B, Hsing M, Christodoulou DC, Cahan P, Daley GQ, Kong SW, Orkin SH, Seidman CE, Seidman JG, Pu WT (2012) Polycomb repressive complex 2 regulates normal development of the mouse heart. Circ Res 110:406–415
Ueno S, Weidinger G, Osugi T, Kohn AD, Golob JL, Pabon L, Reinecke H, Moon RT, Murry CE (2007) Biphasic role for Wnt/beta-catenin signaling in cardiac specification in zebrafish and embryonic stem cells. Proc Natl Acad Sci USA 104:9685–9690
Marvin MJ, Di Rocco G, Gardiner A, Bush SM, Lassar AB (2001) Inhibition of Wnt activity induces heart formation from posterior mesoderm. Genes Dev 15:316–327
Nakamura T, Sano M, Songyang Z, Schneider MD (2003) A Wnt- and beta-catenin-dependent pathway for mammalian cardiac myogenesis. Proc Natl Acad Sci USA 100:5834–5839
Lindsley RC, Gill JG, Kyba M, Murphy TL, Murphy KM (2006) Canonical Wnt signaling is required for development of embryonic stem cell-derived mesoderm. Development 133:3787–3796
Lin L, Cui L, Zhou W, Dufort D, Zhang X, Cai CL, Bu L, Yang L, Martin J, Kemler R, Rosenfeld MG, Chen J, Evans SM (2007) Beta-catenin directly regulates Islet1 expression in cardiovascular progenitors and is required for multiple aspects of cardiogenesis. Proc Natl Acad Sci USA 104:9313–9318
Ai D, Fu X, Wang J, Lu MF, Chen L, Baldini A, Klein WH, Martin JF (2007) Canonical Wnt signaling functions in second heart field to promote right ventricular growth. Proc Natl Acad Sci USA 104:9319–9324
Kwon C, Arnold J, Hsiao EC, Taketo MM, Conklin BR, Srivastava D (2007) Canonical Wnt signaling is a positive regulator of mammalian cardiac progenitors. Proc Natl Acad Sci USA 104:10894–10899
Qyang Y, Martin-Puig S, Chiravuri M, Chen S, Xu H, Bu L, Jiang X, Lin L, Granger A, Moretti A, Caron L, Wu X, Clarke J, Taketo MM, Laugwitz KL, Moon RT, Gruber P, Evans SM, Ding S, Chien KR (2007) The renewal and differentiation of Isl1+ cardiovascular progenitors are controlled by a Wnt/beta-catenin pathway. Cell Stem Cell 1:165–179
Kwon C, Qian L, Cheng P, Nigam V, Arnold J, Srivastava D (2009) A regulatory pathway involving Notch1/beta-catenin/Isl1 determines cardiac progenitor cell fate. Nat Cell Biol 11:951–957
Habara-Ohkubo A (1996) Differentiation of beating cardiac muscle cells from a derivative of P19 embryonal carcinoma cells. Cell Struct Funct 21:101–110
Li B, Jia Z, Wang T, Wang W, Zhang C, Chen P, Ma K, Zhou C (2012) Interaction of Wnt/beta-catenin and notch signaling in the early stage of cardiac differentiation of P19CL6 cells. J Cell Biochem 113:629–639
Wohrle S, Wallmen B, Hecht A (2007) Differential control of Wnt target genes involves epigenetic mechanisms and selective promoter occupancy by T-cell factors. Mol Cell Biol 27:8164–8177
Cresci M, Vecoli C, Foffa I, Pulignani S, Ait-Ali L, Andreassi MG (2013) Lack of association of the 3′-UTR polymorphism (rs1017) in the ISL1 gene and risk of congenital heart disease in the white population. Pediatr Cardiol 34:938–941
Stevens KN, Hakonarson H, Kim CE, Doevendans PA, Koeleman BP, Mital S, Raue J, Glessner JT, Coles JG, Moreno V, Granger A, Gruber SB, Gruber PJ (2010) Common variation in ISL1 confers genetic susceptibility for human congenital heart disease. PLoS One 5:e10855
Friedrich FW, Dilanian G, Khattar P, Juhr D, Gueneau L, Charron P, Fressart V, Vilquin JT, Isnard R, Gouya L, Richard P, Hammoudi N, Komajda M, Bonne G, Eschenhagen T, Dubourg O, Villard E, Carrier L (2013) A novel genetic variant in the transcription factor Islet-1 exerts gain of function on myocyte enhancer factor 2C promoter activity. Eur J Heart Fail 15:267–276
Genead R, Danielsson C, Andersson AB, Corbascio M, Franco-Cereceda A, Sylven C, Grinnemo KH (2010) Islet-1 cells are cardiac progenitors present during the entire lifespan: from the embryonic stage to adulthood. Stem Cells Dev 19:1601–1615
Weinberger F, Mehrkens D, Friedrich FW, Stubbendorff M, Hua X, Muller JC, Schrepfer S, Evans SM, Carrier L, Eschenhagen T (2012) Localization of Islet-1-positive cells in the healthy and infarcted adult murine heart. Circ Res 110:1303–1310
Barzelay A, Hochhauser E, Entin-Meer M, Chepurko Y, Birk E, Afek A, Barshack I, Pinhas L, Rivo Y, Ben-Shoshan J, Maysel-Auslender S, Keren G, George J (2012) Islet-1 gene delivery improves myocardial performance after experimental infarction. Atherosclerosis 223:284–290
Gessert S, Kuhl M (2010) The multiple phases and faces of wnt signaling during cardiac differentiation and development. Circ Res 107:186–199
Vonica A, Gumbiner BM (2002) Zygotic Wnt activity is required for Brachyury expression in the early Xenopus laevis embryo. Dev Biol 250:112–127
Liu Z, Li T, Liu Y, Jia Z, Li Y, Zhang C, Chen P, Ma K, Affara N, Zhou C (2009) WNT signaling promotes Nkx2.5 expression and early cardiomyogenesis via downregulation of Hdac1. Biochim Biophys Acta 1793:300–311
Afouda BA, Martin J, Liu F, Ciau-Uitz A, Patient R, Hoppler S (2008) GATA transcription factors integrate Wnt signalling during heart development. Development 135:3185–3190
Billin AN, Thirlwell H, Ayer DE (2000) Beta-catenin-histone deacetylase interactions regulate the transition of LEF1 from a transcriptional repressor to an activator. Mol Cell Biol 20:6882–6890
Hecht A, Vleminckx K, Stemmler MP, van Roy F, Kemler R (2000) The p300/CBP acetyltransferases function as transcriptional coactivators of beta-catenin in vertebrates. EMBO J 19:1839–1850
Ge X, Jin Q, Zhang F, Yan T, Zhai Q (2009) PCAF acetylates {beta}-catenin and improves its stability. Mol Biol Cell 20:419–427
Miyabayashi T, Teo JL, Yamamoto M, McMillan M, Nguyen C, Kahn M (2007) Wnt/beta-catenin/CBP signaling maintains long-term murine embryonic stem cell pluripotency. Proc Natl Acad Sci USA 104:5668–5673
Guo T, Wang W, Zhang H, Liu Y, Chen P, Ma K, Zhou C (2011) ISL1 promotes pancreatic islet cell proliferation. PLoS One 6:e22387
Acknowledgments
We thank Prof. Yunzeng Zou, Fu Dan University, Shanghai, China for providing us the P19CL6 cell line. This work was supported by the National Natural Sciences Foundation of China (81370236, 30871253, 90919022, 81170713, 30400242), the Leading Academic Discipline Project of Beijing Education Bureau, and the 111 Project of Ministry of Education of China (B07001). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Conflict of interest
The authors declare no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 1 (MPG 1014 kb)
Rights and permissions
About this article
Cite this article
Lu, H., Li, Y., Wang, Y. et al. Wnt-promoted Isl1 expression through a novel TCF/LEF1 binding site and H3K9 acetylation in early stages of cardiomyocyte differentiation of P19CL6 cells. Mol Cell Biochem 391, 183–192 (2014). https://doi.org/10.1007/s11010-014-2001-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-014-2001-y