Abstract
Treatment during early tumor development has greater success because tissue growth remains largely confined to its original locus. At later stages, malignant cells migrate from their original location, invade surrounding normal areas, and can disseminate widely throughout the body. Remodeling of the extracellular matrix (ECM) serves as a key facilitator of this dissemination. Proteolytic enzymes including plasmin and matrix metalloproteinases (MMPs) play an integral role in degrading the surrounding ECM proteins and clearing a path for tumor cell migration. Specific MMPs are highly expressed late during malignant tumor invasion. It is not understood whether early changes in MMPs influence apoptotic and necrotic cell death, processes known to govern the early stages of carcinogenesis. Similarly, the interaction between MDM2 and p53 is tightly controlled by a complex array of post-translational modifications, which in turn dictates the stability and activity of both p53 and MDM2. The present studies examine the hypothesis that model hepatotoxin dimethylnitrosamine (DMN), which is also a model carcinogen, will induce the MMP family of proteins after administration in hepatotoxic doses. Doses of 25, 50, and 100 mg/kg DMN were administered i.p. to male C3H mice. Changes in parameters associated with apoptotic and necrotic cell death, DNA damage, cell proliferation, and extracellular proteinases were examined in liver at 24 h. Serum ALT activity, oxidative stress [malondialdehyde], and caspase-activated DNAse mediated DNA laddering increased in a dose-dependent manner, as did the level of MDM2 protein. MMP-9, -10 and -12 (gelatinase-B, stromelysin-2, macrophage elastase), and p53 protein levels increased following 25 mg/kg DMN, but were successively decreased after higher DMN doses. The results of this study demonstrate changes in MDM2 and MMPs during DMN-induced acute liver injury and provide a plausible linkage between DMN-induced oxidative stress-mediated genomic injury and its likely involvement in setting the stage for initiating subsequent metastatic disease at later circumstances.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dimethylnitrosamine (DMN) is a genotoxic carcinogen and the simplest representative of the dialkylnitrosamines. It has been used for decades as a model hepatotoxin to understand bioactivation-dependent liver injury [1–4] and to explore mechanisms of hepatocarcinogenesis in rodents [4, 5]. DMN is carcinogenic in all animal species. Its organ and species specificities are attributed to differences in metabolism and bioactivation [5–9]. The primary target organs for DMN-induced carcinogenesis are liver, kidney, and lung [4, 10–12].
DMN toxicity is attributed to its ability to alkylate numerous cellular components including DNA, RNA, protein, and phospholipids in multiple target organs. The nitrosamine is metabolically activated when it undergoes α-hydroxylation. The unstable α-hydroxy metabolite spontaneously decomposes to yield reactive electophiles that are methylating agents, including methyl cation. Other pathways to reactive metabolites may include oxidation of α-hydroxydimethylnitrosamine to the corresponding aldehyde, N-nitroso-N-methylformamide [13]. Historically, these biological reactive intermediates were thought to cause genomic alkylation, DNA single-strand breaks, DNA double-strand breaks, and other carcinogenic alterations. Until the late 1980s, most DMN-induced hepatotoxic changes were interpreted as necrotic. In 1989, Pritchard and Butler [14] demonstrated that DMN can also induce apoptosis. Following this observation, several mechanistic studies disclosed the ability of DMN to induce DNA damage that was directly related to various forms of cell death [2–4, 15–18] having differing implications for hepatocarcinogenesis [4, 12].
Along with the model hepatotoxicant acetaminophen [15, 19–23], DMN is a classical hepatotoxin that interferes with processes in every intracellular compartment, including mitochondria [24, 25] nuclei [4, 15, 26–29], the plasma membrane [28, 30, 31], and cytoplasm [32, 33]. The alkylating ability of DMN has been considered to be its most injurious property relating to liver cell death [2, 26].
A limited number of studies focus on the early molecular and cellular events bridging DMN toxicity and carcinogenicity in the liver. However, how DMN impacts extracellular matrix (ECM) architecture and inherent cell cycle regulatory elements that have been tied to neoplastic propagation and metastatic tissue invasion remains unexplored to date. Various effects resulting from DMN bioactivation could contribute to these processes, including oxidative stress in most intracellular compartments, extensive genome alteration, partial or full arrest of the DNA repair machinery, and an imbalance in the pro-oxidant and antioxidant status of the cell. Certain effects alone, or several in concert, would be expected to alter cell adhesion molecules, their receptor sites on basement membranes, and communication across cells and ECM.
Tissues consist of sheets of epithelial cells that are carefully arranged on a basement membrane by different adhesion receptors. Properly functioning receptors prevent cells from migrating to other destinations. When a cell becomes transformed and tumorigenic, it loses most adhesive properties. As damage progresses, cells migrate from their origin and invade neighboring and underlying tissues [34, 35]. Cancerous cells invade blood vessels to find their way into the bloodstream, where they gain access to all parts of the body and are able to take shelter in preferred organs. Each type of cancer has its own propagation and metastasis characteristics. DMN is a potent inducer of liver metastasis in rodent models [4].
Cell death due to cancer often becomes apparent in secondary metastatic sites and not from the site where the cancer originated. During this process of malignancy, transformed cells detach from the primary site, migrate, and cross structural barriers, including basement membranes and surrounding extracellular membranes. A key step in metastasis, tumor-induced angiogenesis, relies upon degradation of the stromal collagenous ECM. The role of matrix metalloproteinases (MMPs) in tumor invasion stems from high expression of specific MMPs in metastatic tumors [36, 37]. Evidence demonstrating the activity of distinct MMPs in vivo in metastatic tumorigenesis is now considerable. ECM remodeling by MMP-driven pericellular tissue proteolysis patterns differs for single-cell and collective-cell migration and invasion [38]. The present studies were undertaken to investigate how DMN induces liver cell death and whether this agent has the capacity to upregulate members of the MMP family that have the capacity to alter long-term signaling pathways involved in tumorigenesis. Such extracellular changes may also play a role in acute cell death induced by DMN.
Materials and methods
Animals, housing, and treatments
Two-month-old male C3H mice (Harlan Sprague–Dawley, Inc., Indianapolis, IN) were given access to rodent chow (Purina, St. Louis, MO) and tap water ad libitum. Animals acclimated in an environment of controlled temperature (22–25 °C), humidity and light/dark cycle in the Long Island University animal care facility for 3 weeks prior to study. Pilot experiments were conducted to standardize DMN exposure and minimize mortality. Use of less than 8-week-old male C3H resulted in excessive animal death. Animals of age 8–9 weeks were chosen for this investigation. DMN in doses of 25, 50, and 100 mg/kg was administered intraperitoneally to male mice. Animals (n = 10) were killed by decapitation after 24 h. Blood was collected for serum chemistry analysis that monitored liver and kidney function. All animal procedures received prior approval by the Institutional Laboratory Animal Care and Use Committee and met or exceeded all local, state, and federal standards.
Liver gross pathology and histopathology
Liver was photographed using a digital camera. Portions from the largest lobe were immediately sectioned, kept on a paraffin rectangle, and wetted with 10 % phosphate-buffered formalin solution. The remaining tissues were immediately frozen in liquid nitrogen and stored at −70 °C for future analysis. Liver sections were processed, sectioned, and stained by American Histolabs (Gaithersburg, MD). Sections stained with H&E were examined for normal, apoptotic, and necrotic cells according to previously established guidelines [4, 16] at 400× and 1,000× magnification and imaged with video microscopy for abnormalities (Carl-Zeiss bright field microscope fitted with a Minolta Digital Rebel SLR camera). Each section was examined three times in its entirety. Livers from three different animals and a minimum of three sections per liver were evaluated.
Chemicals, reagents, and serum chemistry
Reagents were of molecular biology grade and chemicals were of analytical grade or higher and used without purification (Sigma Chemical Co., St. Louis, MO, USA). Sigma or Bioxy Tech kits were used for analytical studies. Hepatotoxicity was determined by measuring the serum activity of alanine aminotransferase (ALT, EC. 2.6.1.2) and evaluating liver histopathology. ALT activity was determined at room temperature using a Raichem kit (ALT Procedure No. 59-UV) based on the method of Wroblewski and Ladue [39]. MMP-9, -10, and -2, and MDM2, PCNA, and p53 antibodies were obtained from Santa Cruz Biotech, Santa Cruz, CA.
Quantitative assessment of genomic injury
DNA damage was assessed quantitatively by density-dependent high-speed centrifugation based on previously published methods [23, 40]. To quantitate liver DNA fragmentation using spectrophotometry, a portion of the liver frozen in liquid N2 was homogenized in chilled lysis buffer [10 mM Tris-HCl, 20 mM EDTA, 0.5 % Triton X-100, pH 8.0]. Homogenates were centrifuged at 27,000g for 20 min to separate intact chromatin in the pellet from fragmented DNA in the supernatant [23]. Pellets were resuspended in 0.5 N perchloric acid and supernatants were treated with concentrated perchloric acid to reach a final concentration of 0.5 N. All the samples were heated at 90 °C for 15 min, and centrifuged at 3000g for 10 min to coprecipitate protein along with other debris. Portions of the resulting supernatants were then treated with diphenylamine (DPA; 1.5 g plus 1 ml concentrated H2SO4 plus 100 ml glacial acetic acid plus 50 mM CH3CHO) for 16–20 h at room temperature [23]. Absorbance was measured at λ600 nm with a Beckman DU 640 spectrophotometer. DNA fragmentation in control samples is expressed as percentage total DNA appearing in the supernatant fraction. Treatment effects were reported as % of control fragmentation.
Qualitative assessment of genomic injury
Caspase-Activated DNAse-dependent genomic injury (laddering) was assessed using agarose gel electrophoresis [40, 41]. To detect the presence of DMN-induced oligonucleosome-length DNA ladders, frozen liver (~200 mg) was homogenized in pre-chilled ~5 ml Sarkosyl lysis buffer (0.1 M Tris-HCl/10 mM EDTA containing 0.5 % Sarkosyl, pH 7.8), kept for 15 min at 4 °C, and treated with buffer-saturated phenol (pH 8.0), with a phenol–chloroform–isoamyl alcohol mixture (25:24:1-V/V/V), and with chloroform-isoamyl alcohol mix (49:1, V/V) to remove protein, fatty materials, and tissue debris, respectively. After the final extraction, the DNA solution was mixed with 3.5 M sodium acetate (pH 4.0) followed by pre-chilled 200-proof absolute ethanol to precipitate DNA. DNA samples were washed with 66 % ethanol, dissolved in TE buffer (0.1 M Tris-HCl/20 mM EDTA, pH 8.0), and the purity verified by the ratio of absorbance at 260/280 nm. Dissolved DNA samples (10 μg/lane) were loaded onto ethidium bromide (0.4 μg/ml) containing agarose gels. The degree of activation of CAD was determined by constant voltage mode electrophoresis (BioRad) at 60v on 1.3 % agarose gels using HindIII digested λ-DNA as molecular weight standard. Electrophoretograms were illuminated with UV light (300 nm) on a UV-transilluminator and photographed using instant Polaroid film #667 [41].
Western blot analyses
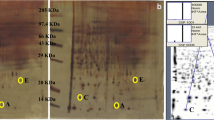
Expression of p53, MMP-9, MMP-10, MMP-12, MDM2, and PCNA proteins was determined using Western blotting, with all steps at room temperature. Liver samples were homogenized in protein extraction buffer (Tris-HCl, NP-40, NaCl, EDTA, NaN3, and PMSF at pH 7.5) with freshly added protease inhibitors (DTT, leupeptin and aprotinin) and stood at 4 °C for 30 min before centrifugation at 27,000g for 20 min. Protein concentration was determined using Pierce Protein Reagent Assay BCA Kit (Thermo Scientific, Rockford, IL). Lysate protein (150 μg) was subjected to 15 % SDS-PAGE at room temperature [42]. Proteins were transferred onto nitrocellulose membrane (ECL, Amersham) at 176v for 1 h using Towbin transfer buffer [43] and a transblot apparatus (Bio-Rad Inc, Hercules, CA). Membranes were blocked with 5 % Carnation® nonfat dry milk in PBS-T (PBS solution, 0.1 % Tween 20) overnight at 4 °C. After one 10-min and two 5-min washes with PBS-T, membranes were incubated overnight with specific primary antibodies diluted in TBS-T for p53, MMPs, MDM2, and PCNA proteins, respectively. After one 10-min and two 5-min washes with PBS-T, membranes were incubated with diluted HRP-conjugated species-specific secondary antibody for 2 h. After one 10-min and four 5-min washes with PBS-T, membranes were developed using ECL Western blotting detection reagents (Amersham). All antibodies were from Santa Cruz Biotech (Santa Cruz, CA). Final images were captured using X-ray film (Kodak).
Hepatic oxidative stress (TBA reactive substances)
Liver samples (~150 mg) frozen and preserved in liquid nitrogen were homogenized in a fixed volume of pre-chilled buffer (10 mM PBS pH 7.4 containing 4 mM BHT), and 250 μl of homogenates was assayed [4, 23]. Samples were treated with 500 μl 10 % TCA, 50 μl 2 % BHT, and 1 ml freshly prepared 0.67 % (Thiobarbituric acid) boiled in a pre-heated water bath (95 °C) for 15–20 min, and brought to room temperature by keeping for 15–20 min on ice. Samples were gently vortexed and centrifuged at 3,500 rpm for 15 min. Debris-free supernatant samples were used for measuring absorbance at 532 nm. Tissue malondialdehyde (MDA) concentrations were determined using a standard curve prepared with known concentrations of MDA (nM range).
Statistical analyses
Results are presented as mean ± SEM unless otherwise indicated. Data were analyzed for significant differences using analysis of variance (ANOVA) followed by Scheff’s F-test (Stat View 512TM+ for Macintosh, Version 1.04, Abacus Concepts Inc., Berkeley, CA) or evaluated by linear regression analysis and correlation. Differences were attributed to treatment rather than chance variance when p ≤ 0.05.
Results
Single doses of DMN caused deaths in male C3H mice at 24 h (Fig. 1). The number of deaths rose directly with dose, reaching 50 % at the highest dose of DMN. The intent was to induce optimal hepatocellular injury in the absence of significant mortality. DMN produced considerable liver injury as reflected in elevation of serum ALT activities (Fig. 2). ALT activity increased smoothly with the amount of administered DMN in a dose-dependent manner. Whereas all doses of DMN damaged the liver, the degree of liver injury following the 100 mg/kg dose of DMN was considerable. The increase in serum ALT activity after this dose exceeded the control value (13 ± 2 U/l) by more than 500-fold (7,027 ± 156 U/l). The amount of ALT leakage is directly dependent on the degree of plasma membrane injury to liver cells.
DMN-induced hepatotoxicity. Single-dose treatment with DMN induced necrosis, as assessed from elevated activity of serum alanine aminotransferase (U/l) and confirmed by histopathology, in the liver of mice in treatment groups at 24 h. Hepatotoxicity was dose-dependent. Results are mean ± SEM with n = 4–10 mice per group. *p ≤ 0.05 versus control
DMN exposure resulted in oxidative stress-mediated lipid peroxidation in liver for all treatment groups, as quantified by the resulting accumulation of malondialdehyde (MDA) (Fig. 3). The 25-mg/kg dose of DMN caused a 2.7-fold increase in MDA. Doses of 50 and 100 mg/kg produced 3.9- and 5.6-fold increases in MDA accumulation, respectively. The increase in tissue MDA concentration closely followed the dose-related effect seen for liver damage via the increase in serum ALT activity.
DMN-induced oxidative stress. DMN treatment increased the level of oxidative stress in the liver of mice in treatment groups at 24 h. The effect was dose-dependent. Oxidative stress was inferred from tissue concentration of malondialdehyde (MDA). Results are mean ± SEM with n = 4–10 mice per group. *p ≤ 0.05 versus control
Quantitative changes in the status of genomic DNA during exposure to hepatotoxic doses of DMN are shown in Fig. 4. Chromatin damage was assessed through the extent of DNA fragmentation. This was based on comparing the accumulation of fragmented or damaged DNA in the supernatant fraction to intact undamaged DNA that is readily pelleted by centrifugation. The quantitative assay reliably provides the extent of fragmentation. DMN exposure induced substantial DNA fragmentation at 24 h. DMN administered at 25 mg/kg produced a threefold (330 %) increase in fragmented DNA compared with control. The 50 mg/kg dose increased fragmentation to 510 % and the 100 mg/kg dose 660 % of control. This induction of DNA fragmentation mirrored the patterns seen for liver damage (serum ALT activity) and oxidative stress (tissue MDA) induced by DMN treatment (Figs. 2, 3).
DMN-induced DNA fragmentation. DMN treatment increased DNA fragmentation in the liver of mice in treatment groups at 24 h. This effect was dose-dependent. Fragmentation was calculated based on the percentage of DNA in the soluble fraction versus total DNA as measured by spectrophotometric analysis. Results are mean ± SEM with n = 4–10 mice per group. *p ≤ 0.05 versus control
Orderly cleavage of DNA to form a ladder of fragments resulting from nucleosomal periodicity of chromatin is primarily linked to CAD’s (endonucleolytic) ability to attack DNA during apoptosis. DNA that appears as a smear on electrophoresis is attributed to more random attack on DNA preceding cell death by necrosis. The presence of both these characteristic patterns on a gel signifies cell death by apoptosis and by necrosis. Data from qualitative experiments examining the effects of DMN on DNA are shown in Fig. 5. DMN treatment produced a distinct ladder pattern of DNA fragments when evaluated by agarose gel electrophoresis. These multiple repetitive bands on electrophoretograms were diagnostic of Ca2+–Mg2+-mediated endonuclease activity (or CAD activity; Lanes 3–5). The presence of this ladder pattern of DNA fragmentation was a clear-cut indication of the involvement of cell death by apoptosis. The highest dose of DMN resulted in extensive loss of large molecular weight DNA. The DNA damage patterns mirrored the dose–response seen with the quantitative measure of genomic injury.
DMN-induced cleavage of DNA. DMN treatment induced endonuclease-mediated internucleosomal DNA cleavage in the liver of mice in treatment groups at 24 h. This electrophoretogram demonstrates rising degrees of DNA damage with increasing DMN dose. Lane 1, BSTEII-digested λDNA standard; Lane 2, vehicle-treated control; Lane 3, DMN 25 mg/kg; Lane 4, DMN 50 mg/kg; Lane 5, DMN 100 mg/kg
DMN treatment produced changes in the morphology in the liver for all treatment groups at 24 h (Fig. 6a–d). Gross damage was easily observed on the surface of the liver and reflected DMN-induced acute cell death and hemorrhagic necrosis of hepatocytes radiating toward the distal regions of individual lobes of the liver. This was established independently by histopathological analysis in Fig. 6e–h. Damage was clearly discernable to the naked eye in liver of mice treated with 25 mg/kg DMN (Fig. 6b). The extent of macroscopic damage increased successively with DMN doses of 50 mg/kg (Fig. 6c) and 100 mg/kg (Fig. 6d). This increase in gross pathological injury which was observable macroscopically roughly paralleled the dose-dependent increases observed in serum ALT activity (Fig. 2), a measure of hepatic necrosis, oxidative stress (Fig. 3), and genomic injury (Figs. 4, 5), a common initiator and contributor to cell death by apoptosis and necrosis.
DMN-induced gross and histopathological liver damage: DMN treatment produced readily observed changes in gross morphology in the liver of mice in all treatment groups at 24 h. Damage appearing on the surface of the liver reflected hemorrhagic acinar infarct and associated necrosis of hepatocytes radiating to the distal regions of liver lobes. Gross liver damage roughly paralleled the dose-dependent increases observed in serum ALT activity, a measure of hepatic necrosis, and oxidative stress. Hepatocellular cell death was confirmed by histopathological evaluation showing dose-dependent necrotic and apoptotic cell death induced by DMN. a, e—untreated control liver; b, f—DMN 25 mg/kg; nel c, g—DMN 50 mg/kg; d, h—DMN 100 mg/kg
Analysis of liver histopathology showed that DMN treatment produced cell death by apoptosis and necrosis 24 h after doses of 50 and 100 mg/kg administered intraperitoneally to C3H mice. No morphological abnormalities were found in liver from control mice 24 h after vehicle treatment (Fig. 6e). At 24 h after treatment with 100 mg/kg DMN (Fig. 6h), H&E-stained liver sections demonstrated extensive hemorrhagic necrosis with loss of basophilic staining in centrilobular regions and extended midzonally. Swollen hepatocyte remnants with degraded nuclei were depleted of glycogen and accumulated large lipid vacuoles. Located adjacent to these necrotic cells were diverse inflammatory cells and red blood cells. DMN-induced hepatocellular damage was present but was less severe after the 50 mg/kg dose (Fig. 6g), and was infrequent after the 25 mg/kg dose (Fig. 6f). Representative swollen hepatocyte with disintegrated cytoplasm, disintegrated nuclei, and nuclei with typical features of apoptosis are presented in Fig. 7a. Apoptotic nuclei with characteristic heterochromatin condensation (Fig. 7a, b) and cells with disintegrated nuclei and cytoplasm are shown in Fig. 7c. Glycogen-depleted and glycogen loaded-cells were frequently observed in the vicinity of injury.
DMN-induced hepatocellular apoptosis and necrosis. a shows various stages of chromatin condensation and the extensive presence of necrosis. b shows numerous apoptotic nuclei (condensed, fragmented and marginated chromatin—hallmarks of apoptotic death). c shows several hepatocytes in varying states, including (i) damaged cytoplasm (double arrow) but intact nuclei, (ii) normal overall morphology with a normal nucleus (single arrow), and (iii) normal cytoplasm with a disintegrated nucleus (arrowhead)
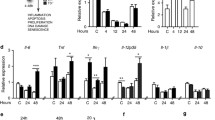
Western blot analysis was performed to assess changes in several MMP proteins during hepatotoxicity caused by the genotoxicant DMN and compared with protein levels in liver extracts from control mice (Fig. 8). Control liver samples showed modest expression of a range of MMP-9 (Fig. 8a), -10 (Fig. 8b), and -12 (Fig. 8c) protein levels. The 25 mg/kg dose of DMN produced an increase in each MMP at 24 h. Interestingly, both DMN doses, 50 and 100 mg/kg, decreased the protein level of each MMP in a dose-dependent manner, to the point that MMP proteins became nearly undetectable. A similar pattern was noted for changes in p53 (Fig. 9a). Total p53 increased following the 25 mg/kg dose of DMN, but declined from this peak with 50 and 100 mg/kg doses. At 100 mg/kg dose, the expression was lower than 50 mg/kg dose. A different pattern was observed for the negative regulator of p53, MDM2 (Fig. 9b). This protein increased steadily with increasing DMN doses to reach a maximum at 100 mg/kg DMN as compared with control. Hence, when p53 and MDM2 expressions are compared with increase in MDM2 expression, the p53 expression decreased at 50 and 100 mg/kg DMN doses—indicating the inhibitory role of MDM2 on p53. In contrast to above observation, the expression of PCNA did not increase at any dose as compared with control, but decreased with increase in DMN doses. Although surprising, the PCNA protein expression was lowest at 100 mg/kg dose.
DMN-induced changes in matrix metalloproteinase protein expression. DMN treatment induced changes in the expression of MMPs in the liver of mice in treatment groups at 24 h. Expression of MMPs was increased following 25 mg/kg DMN, but appeared to be decreased after higher DMN doses. a—MMP-9, b—MMP-10, c—MMP-12
Discussion
Drug and chemical hepatotoxins that undergo intracellular conversion to reactive metabolites initiate damage at various sites within the cell leading to cell death by necrosis and apoptosis [44]. Comparatively little attention has focused on whether these drugs and chemicals also produce damage outside the cell which is relevant to acute cell death and to later untoward events such as cancer and metastatic disease. In this study, DMN produced cell death in the liver by both necrosis and apoptosis. Previous studies have shown that and DMN causes Ca2+ dysregulation, DNA fragmentation, and apoptotic and necrotic cell death, where the importance of DNA damage to necrosis is illustrated by its increase with treatments that inhibit DNA repair and its decrease with treatment to inhibit Ca2+-endonuclease activity [2, 14, 45].
The present studies extend these observations showing that treatment with 25 mg/kg DMN produced increases in PCNA and p53 proteins but that higher doses caused these proteins to decrease. The p53 inhibitor, MDM2, increased in a dose-dependent manner with DMN treatment, in parallel with increasing DNA damage. Damage leading to loss of hepatocytes by cell death initiates repair and recovery pathways in proportion to the magnitude of damage and cell loss, and with a delay period that can be dictated by the degree of initial damage and by the extent of ongoing damage. Because MMPs are proteins that play a central role in recovery and repair after organ insult by drugs and chemicals, and because they are also positioned to participate in longer term adverse effects such as morphogenesis and metastatic cancer, they constitute a family of proteins that merit closer evaluation in DMN hepatotoxicity.
The activation of p53 is regulated by environmental milieu, nature of stress, and cell types. In response to severe or sustained stress signals, p53 drives irreversible apoptosis or senescence programs. However, under the acute conditions, when cell repair is possible, p53 engages a temporary program of cell-cycle arrest and allows cells to repair damaged DNA, thereby limiting the propagation of oncogenic mutations. In contrast, inappropriate p53 activity can be detrimental. However, several E3 ubiquitin ligases’ including MDM2 are present upstream, which negatively regulates p53 protein levels keeping its activity low when not required. MDM2 not only regulates p53 transcriptional activity and stability, but also inhibits its transactivation function by blocking the requirement of essential transcriptional machinery components.
MMPs are involved in fundamental biological processes—morphogenesis, tissue remodeling and repair, angiogenesis, proliferation, and host defenses. They are central to diseases like cirrhosis and arthritis, and are the most prominent family of proteases associated with tumorigenesis [36, 46]. Results of the present study offer a picture of the intracellular effects of DMN associated with acute liver injury, including oxidative stress and DNA damage. The effects of DMN on matrix metalloproteinase proteins may give insights into the potential for extracellular actions to contribute to acute events like cell death as well as later events like metastasis. A representative panel of MMPs was selected to evaluate the effects of DMN in the current studies, including MMP-9 or gelatinase-B which degrades type IV and V collagens and gelatin and participates in cell invasion and metastasis [37], MMP-10 or stromelysin-2 which degrades proteoglycans and fibronectin but not collagen and plays a role in hypertension-associated end-stage renal disease [47], and MMP-12 or macrophage elastase which degrades soluble and insoluble elastin and contributes to aneurysm and emphysema [48].
Matrix metalloproteinases are central modulators of homeostasis in the liver. They contribute to the occurrence of metastatic liver cancers [49], resolution of hepatic cirrhosis [50], and hepatic tissue repair after other insults [51]. Substantial increases in MMP-9, -10, and -12 proteins were observed in liver 24 h after treatment with 25 mg/kg DMN. These effects suggest that DMN has the capacity to alter the amounts of these proteins in the ECM where they are involved in proteolysis and are associated with tissue repair, metastasis, and tissue remodeling. DMN is not alone in this regard among chemical hepatotoxicants and carcinogens. MMP-9 and MMP-10 increase by as early as 3 h during acute liver injury caused by CCl4 and peak around 24 h during high levels of inflammatory reaction [51]. MMP-2 and MMP-9 increases in the liver in response to acetaminophen [52], and in the heart, after doxorubicin exposure [53]. Induction of these MMP proteins occurs during injury to other organs including the lung, skin, brain, and peripheral nerves [54–57].
It was interesting to note here that the increases in MMP proteins which were observed at the low acute dose of DMN (25 mg/kg), and which are consistent with those changes associated with cell proliferation and metastasis, were not seen following the higher doses of 50–100 mg/kg DMN. Rather, the levels of MMP-9, -10, and -12 proteins were substantially decreased by DMN. There are several plausible explanations for this observation, although there are no data that directly address the mechanism(s) underlying this finding. Perhaps the most likely basis for these results is that the extensive toxicity induced by high doses of DMN was accompanied by a broad inhibition of protein synthesis, including the synthesis of new MMP-9, -10, and -12 proteins. Inhibition of protein synthesis is a common general effect under conditions of high toxicity. It is also possible that the high levels of oxidative stress and protein damage in liver cells were accompanied by an increased rate of protein degradation by the proteosomal system and an associated reduction in the rate of MMP proenzyme protein export.
Damage to hepatocytes by intoxication with drugs or alcohol, chronic hepatitis, and other factors elicits inflammatory reactions in the liver. Metabolites of toxic agents, infiltrating inflammatory cells, and membrane components of the damaged hepatocytes activate Kupffer cells. Once activated, Kupffer cells release soluble agents including cytokines and reactive oxygen species which activate kinases [58] and induce transcription of the MMPs that assist in morphological modification of the ECM. MAP kinase signal transduction appears to play a role of MMP-9 and MMP-13 expression in other cells [59].
Results from the present study demonstrate added features of DMN toxicity related to DNA and to damage produced after high doses. Only the lowest dose of DMN induced MMP-9, but compared to the control PCNA expression decreased in a dose dependent manner. At the low dose, increases were observed in p53 and MDM2 proteins that also reflect underlying changes in cell proliferation in the liver after DMN treatment. The tumor suppressor protein p53 is a potent anti-proliferative and pro-apoptotic factor. MDM2 contributes to the control of p53 by participating in an autoregulatory feedback loop in which each protein controls the cellular level of the other. Increases in MDM2 protein result in binding to and inactivation of p53 by directly blocking the p53 transactivation domain and targeting p53 protein for ubiquitin-dependent degradation by the proteasome. The upregulation of MDM2 protein that was seen following the low dose of DMN suggests that p53, although stabilized and also increased in protein level, was inactivated and unable to contribute to the apoptotic elimination of cells that suffered DNA damage and may have undergone initiation. Together with increased MMP proteins induced by 25 mg/kg DMN, these cellular changes in concert contribute to an environment that is more favorable to the induction cancer as well as metastatic disease.
The liver provides some essential body functions and is an internal organ in the body with what may be the most robust ability to regenerate. Although tissue regeneration kinetics under the influence of DMN was not a focus of this investigation, factors that negatively influence this exceptional property should not be ignored. Similarly, roles played by other types of cells (Kupffer cells, stellate cells, endothelial cells, Ito cells, etc.) during toxic reactions and cycling of MMP-TIMP should also be kept in focus while interpreting short-term reactions that can set the stage for long-term events. For example, cell-regenerating ability can become paralyzed in the seriously damaged liver when hepatic stellate cells produce a fibrillar collagen type I and III flood during liver injury [60–62]. MMPs play important, yet perplexing roles during regenerative processes. Considerable expression of MMP-2, MMP-3, MMP-9, MMP-10, MMP-13, and MMP-14 is observed after liver injury in both human and animal model studies, with peak expression coinciding with inflammatory cytokine upregulation [63–65]. The expression of MMP and TIMP can positively or negatively impact short-term and long-term tissue status. Greater understanding of the ability of extracellular processes such as proteolysis by MMPs to impact acute events including drug- and chemical-induced cell death by necrosis and apoptosis would provide valuable insights into these processes.
References
Magee P (1956) Toxic liver injury; the metabolism of dimethylnitrosamine. Biochem J 68:676–682
Ray SD, Sorge CL, Kamendulis LM, Corcoran GB (1992) Ca2+-activated DNA fragmentation and dimethylnitrosamine-induced hepatic necrosis: effects of Ca2+-endonuclease and poly(ADP-ribose) polymerase inhibitors in mice. J Pharmacol Exp Ther 263:387–394
Ray SD, Lam TS, Rotollo JA, Phadke S, Patel C, Dontabhaktuni A et al (2004) Oxidative stress is the master operator of drug and chemically-induced programmed and unprogrammed cell death: implications of natural antioxidants in vivo. BioFactors 21:223–232
Ray SD, Parikh H, Bagchi D (2005) Proanthocyanidin exposure to B6C3F1 mice significantly attenuates dimethylnitrosamine-induced liver tumor induction and mortality by differentially modulating programmed and unprogrammed cell deaths. Mutat Res 579:81–106
Mirvish SS, Kaufman L (2008) A study of nitrosamines and S-carboxyl derivatives of cysteine as lung carcinogens in adult SWR mice. Int J Cancer 6:69–73
Fong LY, Lin HJ, Lee CH (1979) Methylation of DNA in target and non-target organs of the rat with methylbenzylnitrosamine and dimethylnitrosamine. Int J Cancer 23:679–682
Engelse LD, Oomen LC, Valk V, Hart AA, Dux A, Emmelot P (1981) Studies on lung tumours. V. Susceptibility of mice to dimethylnitrosamine-induced tumour formation in relation to H-2 haplotype. Int J Cancer 28:199–208
Meier-Batschi A, Lutz WK, Schlatter C (1983) Methylation of liver DNA of rat and mouse by N-nitrosodimethylamine formed in vivo from dimethylamine and nitrite. Food Chem Toxicol 21:285–289
Iqbal ZM, Dhal K, Epstein SS (1981) Biosynthesis of dimethylnitrosamines in dimethylamine-treated mice after exposure to nitrogen dioxide. J Natl Cancer Inst 67:137–141
Tomatis L, Magee PN, Shubik P (1964) Induction of liver tumors in the Syrian golden hamster by feeding dimethylnitrosamine. J Nat Cancer Inst 33:341–345
Suzuki T, Itoh T, Hayashi M, Nishikawa Y, Ikezaki S, Furukawa F et al (1996) Organ variation in the mutagenicity of dimethylnitrosamine in Big Blue mice. Environ Mol Mutagen 28:348–353
Khodos I, Ray SD (2009) Exposure to a proanthocynidin mixture significantly reduces dimethylnitrosamine (DMN)-induced nephrocarcinogenesis in vivo (abstract). Toxicologist 108:S163
Guttenplan JB (1993) Effects of cytosol on mutagenesis induced by N-nitrosodimethylamine, N-nitrosomethylurea and alpha-acetoxy-N-nitrosodimethylamine in different strains of Salmonella: evidence for different ultimate mutagens from N-nitrosodimethylamine. Carcinogenesis 14:1013–1019
Pritchard DJ, Butler WH (1989) Apoptosis—the mechanism of cell death in dimethylnitrosamine-induced hepatotoxicity. J Pathol 158:253–260
Ray SD, Kamendulis LM, Gurule MW, Yorkin RD, Corcoran GB (1993) Ca2+ antagonists inhibit DNA fragmentation and toxic cell death induced by acetaminophen. FASEB J 7:453–463
Ray SD, Jena N (2000) A hepatotoxic dose of acetaminophen modulates expression of Bcl-2, Bcl-xL, and Bcl-xS during apoptotic and necrotic death of mouse liver cells in vivo. Arch Toxicol 73:594–606
Kamendulis LM, Corcoran GB (1995) Dimethylnitrosamine-induced DNA damage and toxic cell death in cultured mouse hepatocytes. J Toxicol Environ Health 46:31–46
Kamendulis LM, Corcoran GB (1994) DNA as a critical target in toxic cell death: enhancement of dimethylnitrosamine cytotoxicity by DNA repair inhibitors. J Pharmacol Exp Ther 271:1695–1698
Mitchell JR, Jollow DJ, Potter WZ, Davis DC, Gillette JR, Brodie BB (1973) Acetaminophen-induced hepatic necrosis. I. Role of drug metabolism. J Pharmacol Exp Ther 187:185–194
Corcoran GB, Mitchell JR, Vaishnav YN, Horning EC (1980) Evidence that acetaminophen and N-hydroxyacetaminophen form a common arylating intermediate, N-acetyl-p-benzoquinoneimine. Mol Pharmacol 18:536–542
Ray SD, Mumaw VR, Raje RR, Fariss MW (1996) Protection of acetaminophen-induced hepatocellular apoptosis and necrosis by cholesteryl hemisuccinate pretreatment. J Pharmacol Exp Ther 279:1470–1483
Ray SD, Balasubramanian G, Bagchi D, Reddy CS (2001) Ca2+-calmodulin antagonist chlorpromazine and poly(ADP-ribose)polymerase modulators 4-aminobenzamide and nicotinamide influence hepatic expression of Bcl-xL and p53 and protect against acetaminophen-induced programmed and unprogrammed cell death in mice. Free Rad Biol Med 31:277–291
Ray SD, Patel N, Shah N, Nagori A, Naqvi A, Stohs SJ (2006) Pre-exposure to a novel nutritional mixture containing a series of phytochemicals prevents acetaminophen-induced programmed and unprogrammed cell deaths by enhancing Bcl-xL expression and minimizing oxidative stress in the liver. Mol Cell Biochem 293:119–136
Friedman MA, Watt KM, Higgins ES (1977) In vitro and in vivo effects of dimethylnitrosamine on mouse liver mitochondrial function. Proc Soc Exp Biol Med 154:530–533
Yamada K, Yamamiya I, Utsumi H (2006) In vivo detection of free radicals induced by diethylnitrosamine in rat liver tissue. Free Radic Biol Med 40:2040–2046
Corcoran GB, Ray SD (1992) The role of the nucleus and other compartments in toxic cell death produced by alkylating hepatotoxicants. Toxicol Appl Pharmacol 113:167–183
Horn TL, Bhattacharjee A, Schook LB, Rutherford MS (2000) Altered hepatic mRNA expression of apoptotic genes during dimethylnitrosamine exposure. Toxicol Sci 57:240–249
Hsu YC, Chiu YT, Lee CL, Lin YL, Huang YT (2004) Increases in fibrosis-related gene transcripts in livers of dimethylnitrosamine-intoxicated rats. J Biomed Sci 11:408–417
Cheng K, Ye Z, Guntaka RV, Mahato RI (2005) Biodistribution and hepatic uptake of triplex-forming oligonucleotides against type alpha1(I) collagen gene promoter in normal and fibrotic rats. Mol Pharm 2:206–217
Ismail S, Rathod J, Parmar M et al (2007) Modulation of expression of matrix metalloproteinases (MMPs -9, -10, & -12) during dimetylnitrosamine-induced liver cell death (abstract). Toxicologist 96:158
Rathod J, Ismail S, Parmar M et al (2007) Modulation of matrix metalloproteinases (MMPs) and MDM2 during acute dimethylnitrosamine (DMN)-induced nephrotoxicity in mice (Abstract). FASEB J 21(730):2
Lin HL, Parsels LA, Maybaum J, Hollenberg PF (1999) N-Nitrosodimethylamine-mediated cytotoxicity in a cell line expressing P4502E1: evidence for apoptotic cell death. Toxicol Appl Pharmacol 157:117–124
Jin YL, Enzan H, Kuroda N, Hayashi Y, Toi M, Miyazaki E et al (2006) Vascularization in tissue remodeling after rat hepatic necrosis induced by dimethylnitrosamine. Med Mol Morphol 33:33–43
Clark EA, Golub TR, Lander ES, Hynes RO (2000) Genomic analysis of metastasis reveals an essential role for RhoC. Nature 406:532–535
Taverna D, Ullman-Culleré M, Rayburn H, Bronson RT, Hynes RO (1998) A test of the role of α5 integrin/fibronectin interactions in tumorigenesis. Cancer Res 58:848–853
Kessenbrock K, Plaks V, Werb Z (2010) Matrix metalloproteinases: regulators of the tumor microenvironment. Cell 141:52–67
Xu D, McKee CM, Cao Y, Ding Y, Kessler BM, Muschel RJ (2010) Matrix metalloproteinase-9 regulates tumor cell invasion through cleavage of protease nexin-1. Cancer Res 70:6988–6998
Wolf K, Wu YI, Liu Y, Geiger J, Tam E, Overall C et al (2007) Multi-step pericellular proteolysis controls the transition from individual to collective cancer cell invasion. Nat Cell Biol 9:893–904
Wroblewski F, Ladue JS (1955) Serum glutamic oxalacetic transaminase activity as an index of liver cell injury: a preliminary report. Ann Intern Med 43:345–360
Patel N, Joseph C, Corcoran GB, Ray SD (2010) Silymarin modulates doxorubicin-induced oxidative stress, Bcl-xL and p53 expression while preventing apoptotic and necrotic cell death in the liver. Toxicol Appl Pharmacol 245:143–152
Ray SD, Patel D, Wong V, Bagchi D (2000) In vivo protection of DNA damage associated apoptotic and necrotic cell deaths during acetaminophen-induced nephrotoxicity, amiodarone-induced lung toxicity and Doxorubicin-induced cardiotoxicity by a novel IH636 grape seed proanthocyanidin extract. Res Commun Mol Pathol Pharmacol 107:137–166
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Towbin H, Staehelin T, Gordon J (1979) Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci USA 76:4350–4354
Liebler DC, Guengerich FP (2005) Elucidating mechanisms of drug-induced toxicity. Nat Rev Drug Discov 4:410–420
Hirata K, Ogata I, Ohta Y, Fujiwara K (1989) Hepatic sinusoidal cell destruction in the development of intravascular coagulation in acute liver failure of rats. J Pathol 158:157–165
Heissig B, Nishida C, Tashiro Y, Sato Y, Ishihara M, Ohki M et al (2010) Role of neutrophil-derived matrix metalloproteinase-9 in tissue regeneration. Histol Histopathol 25:765–770
Friese RS, Rao F, Khandrika S, Thomas B, Ziegler MG, Schmid-Schonbein GW (2009) Matrix metalloproteinases: discrete elevations in essential hypertension and hypertensive end-stage renal disease. Clin Exp Hypertens 31:521–533
Cataldo DD, Gueders MM, Rocks N, Sounni NE, Evrard B, Bartsch P et al (2003) Pathogenic role of matrix metalloproteinases and their inhibitors in asthma and chronic obstructive pulmonary disease and therapeutic relevance of matrix metalloproteinase inhibitors. Cell Mol Biol 49:875–884
Zeng ZS, Guillem JG (1995) Distinct pattern of matrix metalloproteinase 9 and tissue inhibitor of metalloproteinase 1 mRNA expression in human colorectal cancer and liver metastases. Br J Cancer 72:575–582
Harty MW, Muratore CS, Papa EF, Gart MS, Ramm GA, Gregory SH et al (2010) Neutrophil depletion blocks early collagen degradation in repairing cholestatic rat livers. Am J Pathol 176:1271–1281
Knittel T, Mehde M, Grundmann A, Saile B, Scharf JG, Ramadori G (2000) Expression of matrix metalloproteinases and their inhibitors during hepatic tissue repair in the rat. Histochem Cell Biol 113:443–453
Ito Y, Abril ER, Bethea NW, McCuskey RS (2005) Inhibition of matrix metalloproteinases minimizes hepatic microvascular injury in response to acetaminophen in mice. Toxicol Sci 83:190–196
Goetzenich A, Hatam N, Zernecke A, Weber C, Czarnotta T, Autschbach R et al (2009) Alteration of matrix metalloproteinases in selective left ventricular adriamycin-induced cardiomyopathy in the pig. J Heart Lung Transplant 28:1087–1093
Lafleur M, Underwood JL, Rappolee DA et al (1996) Basement membrane and repair of injury to peripheral nerve—defining a potential role for macrophages, matrix metalloproteinases, and tissue inhibitor of metalloproteinases-1. J Exp Med 184:2311–2326
Swiderski RE, Dencoff JE, Floerchinger CS, Shapiro SD, Hunninghake GW (1998) Differential expression of extracellular matrix remodeling genes in a murine model of bleomycin-induced pulmonary fibrosis. Am J Pathol 152:821–828
Pagenstecher A, Stalder AK, Kincaid CL, Shapiro SD, Campbell IL (1998) Differential expression of matrix metalloproteinase and tissue inhibitor of matrix metalloproteinase genes in the mouse central nervous system in normal and inflammatory states. Am J Pathol 152:729–741
Madlener M, Parks WC, Werner S (1998) Matrix metalloproteinases (MMPs) and their physiological inhibitors (TIMPs) are differentially expressed during excisional skin wound repair. Exp Cell Res 242:201–210
Thakur V, Pritchard MT, McMullen MR, Wang Q, Nagy LE (2006) Chronic ethanol feeding increases activation of NADPH oxidase by lipopolysaccharide in rat Kupffer cells: role of increased reactive oxygen in LPS-stimulated ERK1/2 activation and TNF-alpha production. J Leukoc Biol 79:1348–1356
Eberhardt W, Huwiler A, Beck KF, Walpen S, Pfeilschifter J (2000) Amplification of IL-1β-induced matrix metalloproteinase-9 expression by superoxide in rat glomerular mesangial cells is mediated by increased activities of NF-kappa β and activating protein-1 and involves activation of the mitogen-activated protein kinase pathways. J Immunol 165:5788–5797
Iimuro Y, Nishio T, Morimoto T, Nitta T et al (2003) Delivery of matrix metalloproteinase-1 attenuates established liver fibrosis in the rat. Gastroenterology 124:445–458
Kanemura H, Iimuro Y, Takeuchi M, Ueki T, Hirano T, Horiguchi K et al (2008) Hepatocyte growth factor gene transfer with naked plasmid DNA ameliorates dimethylnitrosamine-induced liver fibrosis in rats. Hepatol Res 38:930–939
Wallace K, Burt AD, Wright MC (2008) Liver fibrosis. Biochem J 411:1–18
Bellayr IH, Mu X, Li Y (2009) Biochemical insights into the role of matrix metalloproteinases in regeneration: challenges and recent developments. Future Med Chem 1:1095–1111
Ray SD, Corcoran GB (2009) General and applied toxicology. In: Ballantyne B, Marrs T, Syversion T (eds) Cell death and apoptosis, 3rd edn. Wiley, New York, pp 248–312
Bulku E, Stohs SJ, Cicero L, Brooks T, Halley H, Ray SD (2012) Curcumin exposure in vivo modulates multiple pro-apoptotic and anti-apoptotic signaling pathways to antagonize acetaminophen-induced toxicity. Curr Neurovas Res 9:58–71
Acknowledgments
This study was supported by the Division of Pharmaceutical Sciences, AMS College of Pharmacy and Health Sciences, NY. Assistance provided by Ms. Dottie Young during the preparation of this manuscript is gratefully acknowledged. Animals used in this study was a generous gift by the Dean Dr. David Taft. This work was presented at the Annual meeting of the Society of Toxicology, 2007.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Syed, I., Rathod, J., Parmar, M. et al. Matrix metalloproteinase-9, -10, and -12, MDM2 and p53 expression in mouse liver during dimethylnitrosamine-induced oxidative stress and genomic injury. Mol Cell Biochem 365, 351–361 (2012). https://doi.org/10.1007/s11010-012-1277-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-012-1277-z