Abstract
Genetic adaptation is one of the key features of Escherichia coli (E. coli) that ensure its survival in different hostile environments. E. coli seems to initiate biofilm development in response to specific environmental cues. A number of properties inherent within bacterial biofilms indicate that their gene expression is different from that of planktonic bacteria. Two of the possible important genes are rpoS and bolA. The rpoS gene has been known as the alternative sigma (σ) factor, which controls the expression of a large number of genes, which are involved in responses to a varied number of stresses, as well as transition to stationary phase from exponential form of growth. Morphogene bolA response to stress environment leads to round morphology of E. coli cells, but little is known about its involvement in biofilms and its development or maintenance. The purpose of this study was to understand and analyse the responses of rpoS and bolA gene to sudden change in the environment. In this study, E. coli K-12 MG1655, rpoS, and bolA mutant strains were used and gene expression was studied. Results show that both genes contribute to the ability to respond and adapt in response to various types of stresses. RpoS response to various stress environments was somehow constant in both the planktonic and biofilm phases, whereas bolA responded well under various stress conditions, in both planktonic and biofilm mode, up to 5–6-fold change in the expression was noticed in the case of pH variation and hydrogen peroxide stress (H2O2) as compared with rpoS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacteria form biofilms as an adaptive mechanism in challenging environments. These can exist wherever surface contact is available to bacteria in naturally occurring fluids [1]. Biofilms are pervasive and problematic because they are more resistant to antibiotics, hydrodynamic shear forces, UV light, and chemical biocides; increased rates of genetic exchange, altered biodegradability, and increased secondary metabolite production than their planktonic counterparts [2, 3]. It is difficult to understand mechanisms of biofilm formation, as biofilms are heterogeneous in the environment and industrial settings and are composed of complex microbial communities [4].
It has been estimated that 65% of the infections are biofilm-associated [5, 6]. Reduced susceptibility of the biofilm bacteria to antimicrobial agents is a vital problem in the treatment of chronic infections [5, 6]. Single-species biofilm might exist in a variety of infections and on the surfaces of indwelling medical implants. The mechanism of biofilm formation can be better understood at the molecular level by studying single-species biofilm under controlled conditions.
Recently, research into the genetic control of biofilm formation has gained importance. Various intrinsic properties within bacterial biofilms indicate that their gene expression is different to their planktonic counterparts and numerous genes have been proposed to be important in biofilm formation. Vast arrays of genes are implicated in biofilm formation [7, 8]. Two of the possibly important genes are rpoS (RNA polymerase sigma factor) and morphogene bolA. RpoS is a sigma subunit of RNA polymerase in E. coli that is induced and can replace vegetative sigma factor rpoD to some extent, under several stress conditions. Consequently, transcription of numerous σS-dependent genes is activated [1].
Morphogene bolA was first described to be involved in adaptation to the stationary growth phase [9]. However, its function is still not fully understood. Its expression might be induced by different forms of stresses that result in the high-level expression of bolA mRNA and the formation of biofilms. It also has a major effect on the bacterial envelope and, therefore, may be implicated in cellular protection under adverse growth conditions [10]. Even though the significance of the rpoS gene in biofilm development has been suggested, the role of rpoS and bolA gene in the formation of biofilm and its expression under different types of stresses has not been investigated.
Stress may be defined as any detrimental factor that adversely affects the growth or survival of microorganisms. Outcomes of stresses applied to microorganisms vary. Sublethal levels of stress reduce or stop the growth of the microorganism and do not result in viability loss [11]. In the case of moderate stress environments, the outcome leads to loss in cell viability and stops the growth of microorganism. Acute or extreme stress is lethal to cells and causes the death of the mainstream of the population. The increase in resistance of an organism to one stress, after application of a different and unrelated sublethal stress, is known as cross-protection [12]. Stress responses are extremely imperative to microorganisms as their habitats are subject to continuous change [11].
In response to changes in their environment, bacteria have the ability to regulate the expression of genes that control their growth and physiology quickly [13]. Because bacterial gene expression is strongly regulated at the transcriptional level [14] and prokaryotic RNAs have short half-lives [15], transcriptional profiling has been widely used in characterization of bacterial responses to various environmental conditions [14, 16]. Reverse transcription followed by quantitative real-time PCR (qRT–PCR) is a sensitive tool to quantitatively analyze RNA levels transcribed from a relatively large number of genetic regions. In addition, it can quantify low-abundance RNAs and, with slight modification, can be applied to measure all categories of RNAs [17]. Moreover, direct measurement of RNA levels from a set of responsive genes that either get induced or repressed under a specific environmental condition can reveal information about bacterial responses and be critical to understanding conditions in microenvironments around bacteria at the time of expression profiling.
Materials and methods
Bacterial strains and growth conditions
E. coli K-12 MG1655 wild type (WT) and mutant strains (Δ) have been used in this study and were kindly provided by National Institute of Genetics, Japan. The WT strain was E. coli K-12 MG1655 and the mutants were E. coli K-12 MG1655 rpoS mutant (ΔrpoS) and E. coli K-12 MG1655 bolA mutant (ΔbolA). Cells were grown in LB (Luria–Bertani) medium. Samples were taken at OD600 = 1.0 and was considered as exponential growth phase, whereas OD600 = 2.2 was considered to be stationary growth phase.
Inoculum preparation
A bacterial suspension was prepared by gently removing bacteria from the solid medium using a sterile nichrome loop to inoculate the bacteria into a 500 ml flask containing 200 ml of sterile nutrient medium. This bacterial suspension was incubated at 37°C with agitation at 120 rpm for 18 h to have bacteria in the exponential phase of growth.
Stress response experiment
Heat shock, cold shock, pH stress, and H2O2 stress
A volume of 0.1 ml of E. coli K-12 MG1655 culture (WT, ∆bolA, and ΔrpoS) was withdrawn at 2 min intervals and plated out directly to determine the viable cell numbers. Percentage survival was defined as the percentage change in the CFU counts per ml obtained after incubation onto LB medium for 15 min following a sudden shift from optimal growth conditions, i.e., heat shock temperatures (42 and 46°C), cold shock temperatures (5 and 20°C), pH stress levels (pH 5, 6, 8, and 9), and different concentrations of H2O2 (3, 4, and 5 mM). This was done to check the rapid change in expression level of rpoS and bolA genes.
Glycogen assay
To confirm the rpoS mutant status, both E. coli WT and ΔrpoS strains were streaked on LB agar plates and incubated overnight at 37°C. After incubation, plates were left at 4°C for 24 h before they were flooded with concentrated iodine solution. Glycogen-deficient ΔrpoS gave a negative-staining reaction (white colonies), whereas the WT glycogen-excess strains generated a positive-staining reaction (dark brown colonies) [18].
Catalase activity
Cultures were also tested qualitatively for catalase activity by applying 6% (wt/vol) H2O2 directly onto colonies on Luria agar plates. Vigorous bubbling indicated WT rpoS activity and positive reaction to hydrogen peroxide.
Biofilm formation assay: crystal violet staining
A biofilm formation assay was performed using a microtitre plate. A volume of 20 μl aliquots of an overnight culture with OD600 of 1.0 were inoculated into 200 μl medium in a PVC microtitre plate. After 72 h incubation, the medium was removed from wells, which were then washed five times with sterile distilled water, and unattached cells were removed. Plates were air-dried for 45 min and each well with attached cells were stained with 1% crystal violet (CV) solution in water for 45 min. After staining, plates were washed with sterile distilled water five times. At this point, biofilms were visible as purple rings formed on the side of each well. The quantitative analysis of biofilm production was performed by adding 200 μl of 95% ethanol to destain the wells. About 100 μl from each well was transferred to a new microtiter plate, and the level (OD) of the crystal violet present in the destaining solution was measured at 595 nm.
Experimental replication
Data from all experiments, including control treatments for both the planktonic and biofilm phase, represent the averages of three or more independent experiments.
Isolation of RNA
RNA was routinely isolated using the RNeasy® ProtectTM Bacteria Mini Kit (Qiagen Ltd., UK), which comprises two steps: (i) immediate stabilization of bacterial RNA and (ii) subsequent isolation and purification of total RNA.
Analysis of RNA integrity
The integrity of total RNA samples was determined by using denaturing (formaldehyde) agarose gel electrophoresis. RNA samples, used for RT-PCR analysis, were routinely checked using this method for the presence of two clear sharp bands of 16S and 23S E. coli ribosomal RNA, which are indicative of intact RNA.
cDNA synthesis for real-time two-step RT-PCR
Messenger RNA was reverse transcribed into cDNA using the QuantiTect® Reverse Transcription kit (Qiagen Ltd., UK). RNA was converted to cDNA with 15 min incubation at 42°C and 3 min inactivation at 95°C. The cDNA was subjected to real time PCR using ABI 7500 (Applied Biosystems). Reactions were performed in a 12.5 μl reaction volume.
Primer designing
Specific primers for rpoS, bolA, and 16S rRNA (house-keeping gene) were designed using Primer 3 software (http://www.genome.wi.mit.edu/cgi-bin/primer/primer3_www.cgi) (Table 1). Primers were ordered from Invitrogen™ life technologies, UK. On receipts, all primers were rehydrated in nuclease-free water and dispensed into 10 μM aliquots of working stock solution before storage at −20°C.
Optimization of the PCR
Optimal PCR conditions were determined using Veriti Thermal Cycler (Applied Biosystems). The optimum concentrations of magnesium chloride and primers for both sets of rpoS and bolA primers were found to be 1.5 mM and 0.3 μM, respectively. These concentrations were subsequently used in all real-time RT-PCR experiments to maintain reaction stringency. The optimum annealing temperature for the amplification of rpoS and bolA was determined to be 60°C.
Real-time quantitative RT-PCR
QuantiTectTM SYBR® Green I PCR (Qiagen) assays were run on ABI 7500 Real-Time PCR machine for quantitative analysis of rpoS and bolA mRNA. Initial assays were carried out according to the reaction conditions recommended by the manufacturer in conjunction with the optimum parameters determined using standard PCR.
Melting curve analysis
The identity of PCR products was confirmed by melting curve analysis, which was performed after the amplification stage of every experiment.
Analysis of gene expression using 2−ΔΔCT method (relative quantification)
The polymerase chain reaction is an exponential process whereby the specifically amplified product ideally doubles each cycle. As such, the measured Ct (cycle threshold) value is a logarithmic value that needs to be converted into a linear relative quantity [19]. The average Ct was calculated for both the target genes and 16S rRNA, and the ΔCt (delta threshold) was determined as (the mean of the triplicate Ct values for the target gene) minus (the mean of the triplicate Ct values for 16S rRNA). The ΔΔCT (delta delta threshold) represented the difference between samples. The expression levels of the gene of interest were normalized by dividing it by the relative expression level for the housekeeping gene for the same sample. The fold-change in gene expression was calculated by dividing the normalized expression level for the experimental sample by the normalized number for the control sample.
Results
Growth curve was plotted to check the differences in the growth rate of +rpoS/−bolA, +bolA/−rpoS, and WT. It was found that E. coli with ΔrpoS and ΔbolA gene can grow at the same rate as WT does in planktonic cells (Fig. 1).
Planktonic growth curve of wild type (WT), rpoS mutant (filled square, rpoS), and bolA mutant (filled diamond, bolA) strains in LB media. Optical density was measured at A 600. OD600 = 1.0 (exponential growth phase) and OD600 = 2.2 (stationary growth phase). The data used are an average of three individual experiments
The analysis of integrity of RNA was routinely checked using formaldehyde agarose gel electrophoresis (Fig. 2). The product sizes for rpoS, bolA, and 16S rRNA were 273, 216, and 201 bp, respectively (Fig. 3). Throughout in this study, ribosomal gene 16S rRNA was used as a reference gene.
Preparation of DNA standards and a standard curve for quantification using real-time PCR
Relative quantification was employed for determination of the relative level of expression of the genes of interest and the housekeeping gene for all experimental samples. Absolute quantification was also performed to generate the Ct values for relative quantification. The advantage of absolute quantification is the quality of results, which provide information on actual levels of a given mRNA, in this case rpoS and bolA mRNA. Furthermore, the results can be compared as independent results, and are not linked to parameters specific to the experiment. The calibration curve was obtained during the runs performed with the DNA standards, and the original screenshot of a standard curve generated during the experiment was taken as an example (Fig. 4). The PCR amplification efficiency can be determined from the slope of the calibration curve. A slope equal to −3.3 indicates 100% efficiency. It should be noted that absolute quantities of each template are calculated based on individual calibration curves generated during individual PCR runs. The optimal baseline and threshold setting for each experiment was set to manual Ct (i.e., threshold 0.02). Ct values were generated for preparation of standard curve for each standard using seven independent experiments.
The melting temperature of the specific product amplified from the initial 16S rRNA, rpoS, and bolA mRNA template had a predicted melting temperature of 83, 84, and 80°C (Fig. 5). From the melting curve plot, it could be deduced that no primer dimers or secondary products were formed because only one peak was seen, which corresponds to the desired product. The products of all real-time PCR experiments presented in this report were confirmed using melting curve analysis and by agarose gel electrophoresis analysis.
The graph illustrates data from a typical real-time RT-PCR experiment with melting curve analysis. Two-step RT-PCR was carried out according to the optimised protocol. It illustrates the calculated plot of fluorescence against temperature. Using this plot, the melting temperature of the amplification product can be determined, which in this case is 83, 84, and 80°C for 16S rRNA, rpoS, and bolA. The data collected also include no template control
Analysis of rpoS and bolA gene expression using the relative quantification method under heat, cold, pH, and oxidative stress
A noticeable difference in gene expression of rpoS and bolA gene under various stress-induced environments in both the planktonic and biofilm phases was seen. In this study, the data are presented as the fold change in target gene expression in various stress-induced environments normalized to the internal control gene (16S rRNA) and relative to the normal control. The N-fold differential expression in the target gene of a stress-induced samples compared with the normal sample counterpart was expressed as 2−ΔΔCT in this study. The rpoS and bolA gene expression level was seen higher in biofilms than the exponential planktonic cells. Expression analysis of mRNA of rpoS and bolA genes under various stress environments was performed using relative quantification method. Results showed the N-fold change in the expression of both rpoS (Fig. 6) and bolA (Fig. 7) genes under heat shock temperatures (42 and 46°C), cold shock temperatures (5 and 20°C), pH stress levels (pH 5, 6, 8, and 9), and different concentrations of H2O2 (3, 4, and 5 mM).
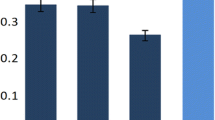
Bar graph represents the expression of rpoS gene under various stress conditions in planktonic and biofilm phase. The cultures were grown overnight in LB at 37°C and percent survival was calculated. The values shown are the means of three independent experiments and the error bars indicate the range. Increased mRNA expression was defined as N-fold > 1.0, “normal” expression (control) was an N-fold = 1, and decreased mRNA expression was N-fold < 1.0
Bar graph represents the expression of bolA gene under various stress conditions in planktonic and biofilm phase. The cultures were grown overnight in LB at 37°C and percent survival was calculated. The values shown are the means of three independent experiments and the error bars indicate the range. Increased mRNA expression was defined as N-fold > 1.0, “normal” expression was an N-fold = 1 (control), and decreased mRNA expression was N-fold < 1.0
Discussion
Earlier studies on rpoS and bolA genes have investigated long-term stress conditions and biofilm formation under several forms of stress, including nutrient starvation at stationary phase, where the increased level of expression has been seen. This study assessed whether rpoS and bolA gene could express under suddenly changing stress conditions, i.e., 15 min intervals from optimal condition to the various stress-induced conditions (i.e., heat, cold, pH fluctuation, and oxidative stress) in both planktonic and biofilm phase. Morphogene bolA is known to express in the stationary phase. Its expression in the biofilm phase at exponential level of growth and the possible role of bolA gene under sudden change in environment was therefore investigated.
E. coli frequently encounters various types of stresses in natural and man-made environments. In this study, real-time RT-PCR was performed to investigate the expression profiles of rpoS and bolA genes in response to similar stresses. The stress-induced conditions used in this study were chosen to represent some scenarios that this bacterium may encounter during natural shifts. These results indicate that the bolA and rpoS respond to different conditions quite distinctly, and have distinct expression patterns under various stress conditions.
RpoS is a conserved stress regulator that plays a significant role in survival under stress conditions in E. coli. The rpoS mutation had a pronounced effect on gene expression in stationary phase, and more than 1,000 genes were differentially expressed. Even in exponential phase when rpoS is expressed at low levels, mutation in rpoS affects the expression of a large set of genes [20]. On the other hand, bolA expression is also confined to stationary phase. Its involvement in biofilm formation and expression under stationary phase is two different events, which are related to stress. So the purpose here was to analyse the expression of rpoS and its dependent gene bolA under biofilm mode of growth, as a sudden response to stress.
Expression of rpoS and bolA in various stress conditions
No activity of rpoS was found under oxidative stress, which suggests that cells in mature biofilms do not require expression of the rpoS gene under oxidative stress in either the planktonic or in biofilm phases (Fig. 6). RpoS might be able to respond in later stages/higher concentration (H2O2) to oxidative stress but not suddenly (in this study). An interesting result was seen in the case of bolA, which showed a 5–6-fold increase in expression under oxidative stress in the planktonic phase when compared with rpoS expression. Decreased expression of bolA in the biofilm phase is seen under oxidative stress when compared with the planktonic phase, which shows that cells can respond well in the planktonic phase in presence of bolA but not in biofilms, whereas rpoS cannot respond in either phase. The data indicate that gene expression within biofilm is different from that observed in standard planktonic growth cultures. Nearly, 1.6-fold increase in the expression of rpoS and 2.2-fold increase in the expression of bolA were seen after 15 min of heat stress, i.e., shift from 37 to 46°C, under the biofilm mode of growth. In the planktonic phase, a minor change was seen after the shift to 42 and 46°C from 37°C (Fig. 6). Sudden decrease in the expression of rpoS and bolA both under cold shock condition suggests that low temperature does not induce the expression of both genes, or it can be said that rpoS and bolA cannot respond suddenly to the cold shock condition, whereas on the other hand, variation in the pH change induces the expression of rpoS and bolA up to 3.5- and 5.5-fold increase under biofilm mode of growth, which in turn shows the necessity for both genes when the pH is changed. It also hypothesizes that cells in biofilms were in stress conditions and requires the expression of rpoS and bolA as a sudden response to environmental change.
Overall, results from this study suggest a new phenotype for the bolA and rpoS gene. In addition to its ability to produce round cell morphology, bolA is implicated in biofilm development [21]. The fact that bolA is expressed under unfavorable conditions (i.e., stress and stationary phase) suggests that biofilm formation is a mode of action by which the bacteria protect themselves against the environment. The expression of bolA is under the transcriptional control of σ S (encoded by rpoS). The presence or absence of σ S has an impact on biofilms [22]. In rpoS mutant strains, the biofilm cell density is reduced by 50%, and there are differences in biofilm structure [23]. Interestingly, deletion of bolA also reduces biofilm formation by E. coli K-12 MG1655. Considering the fact that the levels of bolA depend on σ S, we can still hypothesize that bolA may facilitate the biofilm development. As the expression level of bolA was higher than that of rpoS alone shows that the sudden change in environment could increase the expression of bolA. This might indicate that σ S may act through bolA to facilitate biofilm development.
The study showed that both rpoS and bolA genes can respond and express under sudden change in environment. Change in pH suggests the importance of rpoS and bolA and their response to the pH fluctuation is constructive, which may lead to increased bolA and rpoS mRNA levels resulting in biofilm formation and development. In general, the study demonstrated that temperature, pH, and hydrogen peroxide have a dramatic effect on gene expression, signifying that adaptation to various environmental change conditions requires a coordinated multifunctional response. This study concludes that rpoS gene and its coordinated expression with bolA gene possibly play a major role in biofilm development.
References
Adnan M, Morton G, Singh J, Hadi S (2010) Contribution of rpoS and bolA genes in biofilm formation in Escherichia coli K-12 MG1655. Mol Cell Biochem 342:207–213
Latasa C, Solano C, Penades JR, Lasa I (2006) Biofilm-associated proteins. C R Biol 329:849–857
Sauer K (2003) The genomics and proteomics of biofilm formation. Genome Biol 4:219
Harvey J, Keenan KP, Gilmour A (2007) Assessing biofilm formation by Listeria monocytogenes strains. Food Microbiol 24:380–392
Costerton JW, Stewart PS, Greenberg EP (1999) Bacterial biofilms: a common cause of persistent infections. Science 284:1318–1322
Mah TF, O’Toole GA (2001) Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol 9:34–39
Beloin C, Michaelis K, Lindner K, Landini P, Hacker J, Ghigo J, Dobrindt U (2006) The transcriptional antiterminator RfaH represses biofilm formation in Escherichia coli. J Bacteriol 188:1316–1331
Richmond CS, Glasner JD, Mau R, Jin H, Blattner FR, Blattner FR, Bult CJ, Cole ST, Fleischmann RD, Himmelreich R, Kunst F, Chuang SE, Shalon D, Schena M, Welford SM, Khan J, DeRisi JL, Chu S, Zhu H, Wodicka L, Holstege FC, Spellman PT, Gingeras TR, Behr MA, Iyer VR, Eisen MB, Neidhardt FC, Sambrook J, Andre P, Lundberg KS, Cline J, Barnes WM, Webster C, Miller JH, Stokes HW, Wilbur WJ, Heller RA, Gross CA, Lindquist S, Lemaux PG, Yamamori T, Herendeen SL, Grossman AD, Erickson JW, Wang QP, Watson N, Yamanaka K, Missiakas D, De Las Penas A, Katayama Y, Gottesman S, Kroh HE, Conlin CA, Jiang X, Tsui HC, Herman C, Hall BG, Gold L, Henry MD, Riley M (1999) Genome-wide expression profiling in Escherichia coli K-12. Nucleic Acids Res 27:3821–3835
Santos JM, Freire P, Vicente M, Arraiano C, lia M (1999) The stationary-phase morphogene bolA from Escherichia coli is induced by stress during early stages of growth. Mol Microbiol 32:789–798
Dong T, Schellhorn H (2009) Control of RpoS in global gene expression of Escherichia coli in minimal media. Mol Gen Genomics 281:19–33
Vorob’eva LI (2004) Stressors, stress reactions, and survival of bacteria: a review. Appl Biochem Microbiol 40:217–224
Rowe MT, Kirk R (1999) An investigation into the phenomenon of cross-protection in Escherichia coli O157:H7. Food Microbiol 16:157–164
Hoch JA (2000) Two-component and phosphorelay signal transduction. Curr Opin Microbiol 3:165–170
Rhodius V, Van DTK, Gross C, LaRossa RA (2002) Impact of genomic technologies on studies of bacterial gene expression. Annu Rev Microbiol 56:599–624
Conway T, Schoolnik KG (2003) Microarray expression profiling: capturing a genome-wide portrait of the transcriptome. Mol Microbiol 47:879–889
Eriksson S, Lucchini S, Thompson A, Rhen M, Hinton JCD (2003) Unravelling the biology of macrophage infection by gene expression profiling of intracellular Salmonella enterica. Mol Microbiol 47:103–118
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT, Barbisin M, Xu NL, Mahuvakar VR, Andersen MR, Lao KQ, Livak KJ, Guegler KJ (2005) Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res 33:e179
Hengge-Aronis R, Fischer D (1992) Identification and molecular analysis of glgS, a novel growth-phase-regulated and rpoS-dependent gene involved in glycogen synthesis in Escherichia coli. Mol Microbiol 6:1877–1886
Hu N, Qian L, Hu Y, Shou ZJ, Wang C, Giffen C, Wang HQ, Wang Y, Glodstein MA, Buck EM, Taylor RP (2006) Quantitative real-time RT-PCR validation of differential mRNA expression of SPARC, FADD, Fascin, COL7A1, CK4, TGM3, ECM1, PPL and EVPL in esophageal squamous cell carcinoma. BMC Cancer 6:33
Dong T, Schellhorn H (2009) Global effect of RpoS on gene expression in pathogenic Escherichia coli O157:H7 strain EDL933. BMC Genomics 10:349
Vieira HLA, Freire P, Arraiano CM (2004) Effect of Escherichia coli morphogene bolA on biofilms. Appl Environ Microbiol 70:5682–5684
Corona-Izquierdo FP, Membrillo-Hernandez J (2002) A mutation in rpoS enhances biofilm formation in Escherichia coli during exponential phase of growth. FEMS Microbiol Lett 211:105–110
Adams JL, McLean RJC (1999) Impact of rpoS deletion on Escherichia coli biofilms. Appl Environ Microbiol 65:4285–4287
Acknowledgments
We gratefully acknowledge the supports from the National Bio Resource Project (NIG, Japan): E. coli for providing bacterial strains for use during this project and the Society for Applied Microbiology (sfam) for a research grant to complete this project.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Adnan, M., Morton, G. & Hadi, S. Analysis of rpoS and bolA gene expression under various stress-induced environments in planktonic and biofilm phase using 2−ΔΔCT method. Mol Cell Biochem 357, 275–282 (2011). https://doi.org/10.1007/s11010-011-0898-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-011-0898-y