Abstract
Adult skeletal muscle fibers can be categorized into slow-oxidative and fast-glycolytic subtypes based on specialized metabolic and contractile properties. The Forkhead box O1 (FoxO1) transcription factor governs muscle growth, metabolism, and cell differentiation, and has been shown to be involved in regulating muscle fiber type specification. However, to date, the mechanism behind FoxO1-mediated fiber type diversity is still unclear. In this article, FoxO1 being expressed preferentially in fast twitch fiber enriched muscles is reported. Moreover, the autors also detected that FoxO1 expression decreased in both fast and slow muscles from mice undergoing endurance exercise which induced a fast-to-slow fiber type transition. Using C2C12 myoblast, constitutively active FoxO1 mutant altered the proportion of muscle fiber type composition toward a fast-glycolytic phenotype and attenuated calcineurin phosphatase activity. In addition, a transcriptionally inactive FoxO1 by resveratrol triggered the expression of genes related to slow-oxidative muscle but not sufficient to induce a complete slow fiber transformation. Taken together, these results suggest that FoxO1 up-regulates fast fiber-type formation and down-regulates muscle oxidative capacity at least in part through inhibition of the calcineurin pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Skeletal muscle is composed of heterogeneous specialized muscle fibers that differ in their biochemical, physiological, and metabolic properties [1]. On the basis of specific myosin heavy chain (MyHC) isoform expression, adult skeletal muscle fibers are generally categorized as MyHC I (slow oxidative), MyHC IIa (fast oxidative), MyHC IIx/d (fast glycolytic), and MyHC IIb (fast glycolytic) [2, 3]. Fiber composition in adult skeletal muscle is regulated in response to changes in physical activity, environment, or pathological conditions [1, 4]. For example, endurance exercise training increases the percentage of slow fiber transforming the myofibers to an increased oxidative metabolism [5–7].
The O subfamily of Forkhead/winged helix transcription factors (FoxO) has been shown to regulate various cell functions including metabolism [8, 9], cell cycle [10, 11], and muscle atrophy [12, 13]. FoxO1 transcriptional activity is regulated by a variety of signaling mechanisms including phosphatidylinositol-3 kinase (PI3K)-mediated activation of AKT, which directly phosphorylates FoxO, leading to nuclear export and inactivation [14, 15]. In addition, FoxO1 can modulate cell differentiation and stimulate myotube fusion of primary mouse myoblasts [16, 17]. However, the physiological role of FoxO1 in skeletal muscle is still unclear. Meanwhile, FoxO1 skeletal muscle transgenic mice showed a marked decrease in skeletal muscle mass, and the expression of slow fiber genes, but not that of fast fiber genes decreased [18]. Reciprocally, conditional ablation of FoxO1 expression in the soleus muscle leads to reduced slow fiber and increased fast fiber formation [19]. It seems to be controversial how FoxO1 plays a critical role in muscle fiber-type composition postnatally. Therefore, the effects of FoxO1 on the muscle fiber-type specification still need to be clarified.
At present, the intracellular pathways that transduce physiological stimuli into molecular signals that alter fiber-type gene expression are largely unknown. Several reports suggest that calcineurin is responsible for the formation of slow-twitch fibers [20, 21]. Furthermore, it has been reported that FoxO1 protein decreased calcineurin phosphatase activity in the cardiomyocytes [22, 23]. Moreover, using gain- and loss-of-function in C2C12 myoblasts demonstrated that calcineurin inhibited FoxO factors and prevented myotube atrophy [24]. Given that calcineurin is important for the formation of slow fibers and the interaction between FoxO1 and calcineurin, FoxO1 might be involved in calcineurin pathway regulating muscle fiber transition.
To gain insight into the potential role of FoxO1 in control of differentiation of muscle fiber type, the authors subjected mice to endurance swimming training which could promote the transformation of fast- to slow-twitch fiber. It was found that in controlled conditions, FoxO1 expression was much higher in fast than in slow muscle fibers, and it was significantly depressed in both slow-oxidative and fast-glycolytic muscles by long-lasting training. In this study, the authors demonstrated that in C2C12, myoblast overexpression of FoxO1 induced the transformation of slow to fast fiber type and markedly decreased calcineurin phosphatase activity. Accordingly, inhibition of FoxO1 by resveratrol (RSV) increased the expression of genes related to slow-oxidative muscles, but without a significant transition in muscle fiber types. Together, these results suggest that FoxO1 protein regulates slow to fast myofiber switching and attenuates muscle oxidative capacity at least in part through inhibition of the calcineurin pathway.
Materials and methods
Animals and endurance exercise program
Twelve male C57BL/6J mice, at 4 week old, were obtained from the Fourth Military Medical University (FMMU, China). They were reared at 25°C under artificial lighting for 12 h from 7 AM to 7 PM daily and had free access to water and food. After 2 week acclimation to the new environment, they were randomly assigned to a swimming training group (trained, n = 6) and a sedentary group (untrained, n = 6). They were then exercised by a modification of the procedure described previously [25]. In brief, the mice in the trained group were accustomed to swimming training for a period of 4 week, 6 days/week, during which time exercise was for 10 min initially and then was extended by 10 min daily until the animals swam continuously for 1 h. Water temperature was controlled every 10 min to be maintained at 35–36°C. Mice swam as a group in a bath measuring 51 × 71 cm2 (diameter, height). Group swimming was used because it promotes more vigorous exercise than when mice are allowed to swim alone [26].
Muscle dissection
About 24 h after the previous session, mice were euthanized by decapitation and were exsanguinated. Samples of slow-oxidative and fast-glycolytic muscles were obtained by dissection of soleus and gastrocnemius muscles, which were quickly excised from both hindlimbs of the animals. The specimens were immediately cooled in liquid nitrogen and stored at −80°C for subsequent analyses.
Plasmids, antibodies and reagents
Plasmids for pcDNA3-FLAG-tagged wild-type FoxO1 (FoxO1-WT) and Akt phosphorylation-resistant mutant of FoxO1 (FoxO1-A3, where the three Akt phosphorylation sites Thr24, Ser256, and Ser319 were converted to alanine) were gifted by Huang et al. [27]. The following antibodies were purchased: anti-FoxO1, anti-phospho-FoxO1 (Ser256), anti-MyoD, anti-β-actin, and anti-Flag (Santa Cruz Biotechnology, USA); anti-sarcomeric MyHC (clone MF-20, DSHB); anti-Myogenin (Millipore, USA); anti-MyHC slow (clone NOQ7.5.4D, Sigma); and anti-MyHC fast (clone MY-32, Sigma). Inhibitor reagent RSV was purchased from Sigma.
Cell culture and induction of differentiation
Mouse C2C12 myoblasts were grown on gelatin-coated dishes in Dulbecco’s modified Eagle’s medium (Gibco) supplemented with 4.5 g/l d-glucose, 1.5 g/l sodium bicarbonate, 1 mM sodium pyruvate, 10% fetal bovine serum (Hyclone Laboratories), and antibiotics (penicillin: 100 IU ml−1, streptomycin: 100 μg ml−1) in 5% CO2 at 37°C. When the cells achieved 70% confluence, they were differentiated in 2% horse serum (Gibco) for 3 days. Under these conditions, cells were found to be healthy, viable, and not undergoing necrosis or apoptosis, as observed using phase contrast microscope (Fig. 3a–c).
Transfection
C2C12 skeletal muscle cells were transfected at 70–80% confluence with 8 μg of DNA and 20 μl of Lipofectamine 2000 (Invitrogen, USA), according to the manufacturer’s protocol. In breif, transfection reagent was incubated with plasmid DNA constructs in Opti-MEM (Invitrogen) at room temperature for 20 min. Then, this transfection mixture was applied to the proliferating cells and incubated for 5 h at 37°C. For stable transfections, following incubation, the transfection medium was removed and replaced with growth medium containing 15% FBS and 400 μg/ml G418 (Gibco) over several days until 7 days after all the cells in the control untransfected flask had been eliminated by G418. All subsequent experiments on stably transfected cells were conducted in the absence of G418. At this stage, cells were subjected to differentiation medium consisting of DMEM plus 2% horse serum for 3 days to induce differentiation into myotubes. After differentiation, multiple clones were isolated and subjected to preliminary analysis, and a single clone that was representative of the pool was chosen on the basis of FoxO1 expression.
RNA extraction and quantitative real time RT-PCR
Muscle tissue or C2C12 myotube lysates were homogenized and extracted for total RNA using TRIzol reagent (TaKaRa) by standard techniques. The integrity of RNA was checked on 2% agarose gels, and total RNA concentration was estimated by a spectrophotometer (Thermo). Two micrograms of total RNA was reverse-transcribed to synthesize cDNA by using the PrimeScript RT-reagent Kit for RT-PCR (TaKaRa) after treatment with DNAse I (TaKaRa) to remove contaminating genomic DNA. Real time PCR amplification reactions were carried out by Bio-Rad iQ5 by SYBR Premix Ex Taq™ II (TaKaRa) chemistry detection under amplification conditions. Quantification of the mRNA data was done by the comparative threshold cycle (ΔΔCT) method, with the modification that the relative efficiency of each primer was included in the calculation. The specificity of the PCR amplification was always verified with melting curve analysis. Table 1 provides details of primers of the genes studied.
Western analysis
Protein was collected from mice skeletal muscles and C2C12 myotubes. Protein lysate from each sample was denatured at 95°C for 5 min. Total protein for each sample was diluted in a loading dye consisting of Tris, 10% SDS, 2-mercaptoethanol and bromophenol blue and then separated on 7.5% SDS-PAGE gel at 75 V for 6 h. The gels were then transferred onto PVDF membranes (Millipore) at 30 V for 16 h at 4°C. Blots were incubated in blocking solution consisting of 4% non-fat dry milk and 0.05% Tween-20 in PBS for 2 h. The primary monoclonal antibodies were diluted in blocking solution and incubated with the blots for 2 h at room temperature. Blots were washed three times in PBS/Tween for 5 min each. Blots were then incubated with secondary antibodies (Santa Cruz Biotechnology) for 1 h at room temperature. Blots were washed as before and detection with Bio-Rad GS-800 densitometer and analyzed using Quantity One software. All Western analyses were repeated as two independent experiments.
Immunofluorescence
The immunostaining of muscle fibers in culture was performed by using mAbs against total MyHC. On the third day of culture, myotubes were fixed and processed for immunohistochemistry. Cells were rinsed with Dulbecco’s PBS (DPBS), fixed in 4% paraformaldehyde for 60–90 min, washed in DPBS, incubated in DPBS supplemented with 1% BSA and 0.05% permeabilization solution, and blocked for 30 min with 10% goat serum and 1% BSA in DPBS. Cultures were then incubated overnight with primary antibodies against sarcomeric MyHC (clone MF-20, DSHB), followed by the secondary antibody Cy3 anti-mouse IgG1. The secondary antibody was diluted 1:100 in DPBS containing 10% goat serum, and incubation was performed for 1 h at room temperature.
Calcineurin phosphatase activity assay
Calcineurin activity was measured spectrophotometrically using the Calcineurin Cellular Assay Kit Plus (BIOMOL). In brief, cells were lysed in lysis buffer provided by the kit. Phosphatase activity was measured by detection of free phosphate released from the calcineurin-specific RII phosphopeptide, and normalized to protein content. Assays were performed in quadruplicate in three independent experiments.
Statistical analysis
All transfection experiments were carried out 3–5 times in triplicate. Main and interactive effects were analyzed by One-way ANOVA using SPSS13.0 software. When justified by One-way ANOVA, differences between individual group means were analyzed by Fisher’s PLSD test. Differences were considered statistically significant at P < 0.05.
Results
Endogenous FoxO1 and MyHCs genes expression in skeletal muscle
In mammalian skeletal muscle, myofibers are mainly classified into fast-glycolytic and slow-oxidative fibers based on specific MyHC isoform expression. As a first step to explore the potential relationship between FoxO1 and muscle fiber expression in muscle, the authors examined the expression pattern of FoxO1 and MyHCs in fast- and slow-twitch skeletal muscles by quantitative real time RT-PCR. As shown in Fig. 1, mouse slow-oxidative soleus consists of about 60% MyHC I slow fibers and 40% MyHC IIa fast oxidative fibers, while fast-glycolytic gastrocnemius is mainly composed of MyHC IIb and MyHC IIx fast fibers (>90%, together).
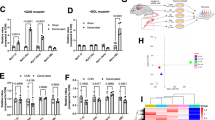
Myosin heavy chain (MyHC) isoforms mRNA distribution in soleus and gastrocnemius of untrained (gray bars) and mice swimming trained (black bars) for 4 weeks normalized against β-actin. a Quantitative real-time RT-PCR (qRT-PCR) showed soleus consists of MyHC I and MyHC IIa fibers. After chronic training, the percentage of MyHC I was increased but MyHC IIa was decreased in soleus. b Gastrocnemius is mainly composed of MyHC IIb and MyHC IIx fibers. Compared to untrained mice, MyHC IIa and IIx were up-regulated but MyHC IIb was down-regulated in gastrocnemius. Values are means ± SEM; n = 6 in each group. *P < 0.05
The real-time PCR also showed that FoxO1 expression was significantly related to muscle fiber type distribution and expressed preferentially in muscle enriched in fast fibers (P < 0.05) (Fig. 2a). Accordingly, the comparison between gastrocnemius and soleus in control mice showed that FoxO1 protein expression was 3–4 times greater in the fast than in the slow muscles (P < 0.05) (Fig. 2b, c).
FoxO1 loses activity after swimming training. a qRT-PCR showed FoxO1 mRNA expression pattern in soleus and gastrocnemius of untrained and swimming trained mice. FoxO1 is expressed preferentially in gastrocnemius and a significant reduction of FoxO1 mRNA contents both in two muscles compared to untrained mice. b–d Western blot analysis of FoxO1 protein expression in soleus and gastrocnemius of untrained (NC) and trained (SW). Western blot results showed FoxO1 protein expression was higher in gastrocnemius than in soleus muscles. An increase in FoxO1 phosphorylation at Ser256 compared to untrained mice. Values are means ± SEM; n = 6 in each group. *P < 0.05, **P < 0.01
Inactivation of FoxO1 after long-term swimming training
To further understand the relationship between FoxO1 and the specification of muscle fiber type, the authors subjected mice to swimming training [25, 28]. After an chronic training based on 4 weeks with daily swimming training, the autors found an increase in the percentage of MyHC I and a concomitant decrease in MyHC IIa in soleus while an up-regulation of MyHC IIa and IIx, but a down-regulation of MyHC IIb in gastrocnemius of trained mice compared to untrained mice (P < 0.05) (Fig. 1a, b). These data suggest that the endurance swimming training program was enough to switch the fiber from fast to slow-twitch type.
FoxO1 expression at the level of mRNA was markedly affected by training, since a significant reduction of FoxO1 mRNA was detected in both slow-oxidative and fast-glycolytic muscles of trained mice when compared to untrained mice (Fig. 2a). Surprisingly, Western blot did not detect any change in the amount of total FoxO1 protein in both soleus and gastrocnemius muscles of the trained and the untrained mice (Fig. 2b, c). However, an increase in FoxO1 phosphorylation at Ser256 was observed in both slow and fast muscles after long-term endurance exercise (Fig. 2b, d). These results may reflect that the loss of FoxO1 function could contribute to the transformation of fast to slow twitch fibers in vivo.
FoxO1 induces slow to fast-twitch fiber transition
The authors have demonstrated that FoxO1 is selectively highly expressed in fast muscle, and changes after fiber-type transition in vivo. Next, the authors used C2C12 mouse cell line to investigate the role of FoxO1 in muscle fiber determination in vitro. First, the authors stably transfected C2C12 mouse cell line with plasmids expressing FoxO1-WT and FoxO1-A3 (the constitutively active FoxO1) followed by the exposure of the cells to differentiation medium (Fig. 3e). Then the authors detected the time-course expression of early (MyoD), intermediate (myogenin), and late (MyHC) myogenic transcription factors during the differentiation of skeletal muscle C2C12 myoblasts (Fig. 3g). Because MyoD is the predominant myogenic factor in fast fibers while myogenin is the predominant factor in slow fibers [29], the authors found that the overexpression of FoxO1 mutant had a little reduction in MyoD expression but an approximate 40% decrease in myogenin protein on day 3 of differentiation (Fig. 3f, h).
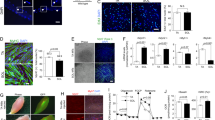
FoxO1 regulates C2C12 myoblast differentiation. Proliferating C2C12 cells were infected for 2 days in proliferation medium (PM) and followed by 3 day in differentiation medium (DM). a–d Differentiation and Morphometric analysis of C2C12 myoblast differentiation at 0 day (D0), 1 day (D1) and 3 day (D3). C2C12 cells were immunostained with anti-MyHC antibody. Bar 100 μm. e Cell lysates derived from control empty vector pcDNA3.1, FoxO1-WT, and FoxO1-A3 infected cells were subjected to Western analysis for FLAG and FoxO1 expression. f Western blot detected the time-course expression of myogenic markers (MyoD, myogenin, and MyHC) during the differentiation of C2C12 myoblasts transduced with empty pcDNA3.1 and FoxO1-A3 constructs. g Densitometric analyses showed MyoD and myogenin proteins significantly increased in the early differentiation time (D1) and then decreased both in the late differentiation stage (D3) in control cells. h MyoD expression had a little reduction and an almost 40% decrease in myogenin protein in FoxO1-expressing cells at 3 days differentiation (D3). Results are expressed as mean ± SEM from triplicate samples within the same experiment. *P < 0.05, **P < 0.01
The real-time PCR results obtained so far had described the relative abundance of each endogenous MyHC isoform and the changes of each MyHC gene isoform in FoxO1-A3 cells on day 3 of differentiation (D3). In control-infected C2C12 myotubes, MyHC IIx mRNA was the most highly expressed adult isoform, followed by MyHC IIb, MyHCI, and MyHC IIa (Fig. 4a). The authors demonstrated that C2C12 myotubes induced by FoxO1 was attributable to combine effects of the up-regulation of MyHC2x and the down-regulation of MyHC I, MyHC IIa and MyHC IIb (Fig. 4a).
FoxO1 induces the formation of fast-twitch fiber. Proliferating C2C12 cells were infected for 2 days in PM and followed by 3 day in DM (D3). a Real-time PCR was performed to quantify the relative levels of MyHC transcripts in C2C12 myotubes at day 3 of differentiation (D3). FoxO1 overexpression in C2C12 cells up-regulated MyHCIIx but down-regulated MyHC I, MyHC IIa, and MyHC IIb. Inhibition FoxO1 by resveratrol (RSV, 100 μM) did not switch the fiber from fast to slow twitch but could partial reverse FoxO1-induced muscle fiber transition. b Western blot detected fast MyHC, slow MyHC and total sarcomeric MyHC protein expression in control and FoxO1-infected cells. c Densitometric analyses showed FoxO1 enhanced fast MyHC protein but suppressed slow MyHC protein. *P < 0.05 relative to control
To evaluate whether the transcriptional effects of FoxO1 were translated into protein changes, Western blot analysis was performed. The results indicated that FoxO1 enhanced fast MyHC protein but suppressed MyHC slow protein (Fig. 4b,c), which was consistent with the MyHCs transcript expression (Fig. 4a). Notably, the relative expression of the total sarcomeric MyHC protein was also decreased in FoxO1-expressing cells. These results suggest that FoxO1 plays a critical role in the myofiber specification toward fast-twitch type.
FoxO1 decreases oxidative capacity in C2C12 myotube
To determine whether a decrease in FoxO1 function induces the differentiation of slow fiber type, cells were treated with resveratrol (RSV, 100 μM), an activator of Sirt1, which inhibited the endogenous FoxO1 activity [30–32]. As expected, Gadd45α, which is known to be regulated by FoxO transcription factors [33, 34], was significantly decreased (Fig. 5a). The results showed that treatment of cells with RSV increased the oxidative fibers related gene TnI slow and myoglobin (Fig. 5b) whereas it was not enough to change the ratio of fast glycolytic to slow oxidative-type muscle fibers (Fig. 5c).
Inhibition of FoxO1 increases myotube oxidative capacity. a RSV (100 μM) inhibited the expression level of the FoxO1 target gene Gadd45α. b The real-time PCR showed FoxO1 largely blocked RSV-induced the mRNA abundance of oxidative fiber markers. c Western blots and densitometric analyses showed effects of RSV on muscle fiber transition. Enhanced fast MyHC expression by FoxO1-A3 was inhibited in the presence of RSV whereas suppressed slow MyHC was abrogated. *P < 0.05, **P < 0.01
To test whether RSV’s effects on oxidative fibers function are mediated by FoxO1, the overexpression of FoxO1 largely blocked the RSV-induced increase in TnI slow and myoglobin expression (Fig. 5b). Importantly, in FoxO1-infected C2C12 cells addition of RSV reversed FoxO1-induced muscle fiber switch (Fig. 5c).
FoxO1 inhibits calcineurin phosphatase activity
The calcineurin/NFAT pathway is important for the formation of slow oxidative fibers [20, 35]. The authors examined whether FoxO1-mediated myofiber switch was associated with calcineurin/NFAT signaling. In this study, relative to empty vector–infected control cells, cells expressing FoxO1-A3 showed a significant decrease in endogenous calcineurin phosphatase activity (Fig. 6a). Figure 6b shows that FoxO1 activation significantly decreased the mRNA abundance of modulatory calcineurin interacting protein, exon 4 isoform (MCIP1.4), a direct target of the calcineurin/NFAT pathway [36, 37].
FoxO1 decreases calcineurin phosphatase activity. a Relative phosphatase activity was normalized to protein content. FoxO1 inhibited calcineurin phosphatase activity, and RSV (100 μM) was not sufficient to induce the clearly increased calcineurin activity. b Overexpression of FoxO1 decreased calcineurin/NFAT pathway target gene MCIP1.4, while RSV (100 μM) antagonized the activity of FoxO1 factor. Results are graphed as mean ± SEM of a representative experiment with quadruplicate samples. * P < 0.05, ** P < 0.01
The effect of RSV treatment on calcineurin activity and found no changes (Fig. 6a) was tested. However, using a more sensitive assay, quantitative real-time PCR showed that inhibition of FoxO1 activity by RSV markedly increased the mRNA abundance of MCIP1.4 (Fig. 6b). Furthermore, RSV antagonized FoxO1-induced down-regulation of MCIP1.4 expression (Fig. 6b), coincident with previous study by the authors on increased muscle oxidative capacity (Fig. 5b).
Discussion
FoxO1 is a transcription factor that has been shown to be a negative regulator of muscle growth [38, 39] and involved in metabolism and mitochondrial biogenesis [9]. In this study, the authors describe that FoxO1 induces a slow-to-fast fiber-type switch and suppresses muscle oxidative capacity at least in part through inhibition of the calcineurin pathway.
These specialized myofibers are highly plastic, and under several physiological and pathological circumstances, the metabolic and enzymatic nature of the fiber can be altered [6]. The most notable is exercise training, and an endurance exercise protocol could increase muscle oxidative capacity and trigger transformation of glycolytic fibers into oxidative ones [5, 7]. Previous study discovered FoxO1 gene increased by 5.2-fold at 3 h, but returned to baseline by 48 h during recovery from acute training in human skeletal [40], consistent with other studies [41]. In addition, acute exercise improved insulin signaling, increasing insulin-stimulated Akt, and FoxO1 phosphorylation [42]. However, FoxO1 expression pattern after endurance exercise is still unclarified. Here, Swimming training to mimic endurance exercise-induced adaptation was adopted [25] and FoxO1 expression change in this process was detected. Concomitant with the increase in slow fiber type (Fig. 1), the authors also found a significant reduction of FoxO1 mRNA contents both in slow and fast muscles compared to untrained mice (Fig. 2b,c). Surprisingly, total FoxO1 protein has no change; however, the level of FoxO1 phosphorylation at Ser256 was increased (Fig. 2b, d). These results reflect that FoxO1 inactive appears to influence muscle fiber-type composition.
Myogenic regulation factors are important regulators of myogenesis and, recently have been shown to have a role in the regulation of MyHC gene expression. MyoD and myogenin mRNAs are preferentially expressed in adult fast-glycolytic and slow-oxidative muscle fibers, respectively [29]. Overexpression MyoD or Myf-5 significantly increased MyHC IIb promoter activity [43]. In this study, the authors have clearly shown that overexpression FoxO1 leads to decreased myogenin transcription factor (Fig. 3f, h). It is noteworthy that FoxO1 activation also up-regulates MyHCIIx but down-regulates the other three type MyHC genes, viz., MyHC I, MyHC IIa, and MyHC IIb (Fig. 4a). Thus, in FoxO1-infected myotubes, the changes in MyHC isoforms tend to follow a general scheme of sequential and reversible transitions: I↔IIa↔IIx↔IIb [1, 6]. Furthermore, the rise in fast MyHC protein and the reduction of MyHC slow protein were revealed. These findings suggest that FoxO1 promotes the formation of fast-twitch fiber at the expense of slow-twitch fiber. Interestingly, the authors use MF-20 monoclonal antibody specific for sarcomeric MyHC and find that the expression of the total MyHC protein was also suppressed in FoxO1-expressing myotubes (Fig. 4b), which probably connect with FoxO1-caused muscle atrophy [38, 39].
Even though an increase in fast twitch fiber's composition was noted in FoxO1-infected cells, inhibition of endogenous FoxO activity by RSV was not sufficient to change the ratio of oxidative to glycolytic type (Figs. 4a, 5c). However, the expression of oxidative fibers-related gene troponin I and myoglobin was increased by RSV inhibition of FoxO1 (Fig. 5b). Moreover, the active form of FoxO1-mutant mediated the transformation from slow-to-fast fiber type was partially reversed by addition of RSV (Figs. 4a, 5c). Therefore, FoxO1 functions as a negative factor in oxidative metabolism. These findings further support the notion that FoxO1 induces slow-to-fast fiber transition. On the other hand, RSV increases the activities of AMP-activated protein kinase (AMPK) and peroxisome proliferator-activated receptor-γ coactivator 1α (PGC-1α) [44, 45], both of which are required for mitochondrial biogenesis and could improve the oxidative activity of skeletal muscle [46, 47]. Also, PGC-1α is a principal factor regulating muscle fiber-type determination, which is known to be preferentially expressed in slow muscle and drives the formation of slow-twitch muscle fibers [48]. As the FoxO1 protein can interact with the PGC-1α protein [49] and transfection of PGC-1α into adult fibers reduced the capacity of FoxO3 [50], FoxO1 may counteract the function of PGC-1α in this process. Thus, further studies are reqiured to examine this possibility.
Several signaling pathways are involved in mediating fiber type-specific gene expression. To the best of knowledge, calcineurin is a chief regulatory pathway of slow fiber-selective gene expression [21]. Overexpression of FoxO1 provokes a remarkable decrease in calcineurin phosphatase activity (Fig. 6a) and reduces the basal mRNA level of the MCIP1.4 (Fig. 6b), a direct target of the calcineurin pathway, consistent with previous reports on FoxO1 in heart [22, 23]. Interestingly, the authors detected no changes in calcineurin phosphatase activity triggered by RSV (Fig. 6a). However, RSV treatment significantly increased the mRNA abundance of MCIP1.4 in FoxO1-infected cells (Fig. 6b), consistent with the enhanced expression of oxidative-related gene (Fig. 5b). The data so far available show inhibition of FoxO1 could promote muscle oxidative capacity at least in part through inhibition of the calcineurin pathway.
In summary, the authors have established the relationship between skeletal muscle fiber-type specification and a function of FoxO1 expression. FoxO1 promotes slow-to-fast fiber-type switch and attenuates muscle oxidative metabolism at least, in part, through inhibiting calcineurin pathway.
References
Bassel-Duby R, Olson EN (2006) Signaling pathways in skeletal muscle remodeling. Annu Rev Biochem 75:19–37
Pette D, Staron RS (2000) Myosin isoforms, muscle fiber types, and transitions. Microsc Res Tech 50(6):500–509
Schiaffino S, Reggiani C (1996) Molecular diversity of myofibrillar proteins: gene regulation and functional significance. Physiol Rev 76(2):371–423
Schiaffino S, Sandri M, Murgia M (2007) Activity-dependent signaling pathways controlling muscle diversity and plasticity. Physiology (Bethesda) 22:269–278
Wu H, Rothermel B, Kanatous S, Rosenberg P, Naya FJ, Shelton JM, Hutcheson KA, DiMaio JM, Olson EN, Bassel-Duby R, Williams RS (2001) Activation of mef2 by muscle activity is mediated through a calcineurin-dependent pathway. EMBO J 20(22):6414–6423
Pette D, Staron RS (2001) Transitions of muscle fiber phenotypic profiles. Histochem Cell Biol 115(5):359–372
Demirel HA, Powers SK, Naito H, Hughes M, Coombes JS (1999) Exercise-induced alterations in skeletal muscle myosin heavy chain phenotype: dose-response relationship. J Appl Physiol 86(3):1002–1008
Birkenkamp KU, Coffer PJ (2003) Regulation of cell survival and proliferation by the FOXO (forkhead box, class o) subfamily of forkhead transcription factors. Biochem Soc Trans 31(Pt 1):292–297
Barthel A, Schmoll D, Kruger KD, Bahrenberg G, Walther R, Roth RA, Joost HG (2001) Differential regulation of endogenous glucose-6-phosphatase and phosphoenolpyruvate carboxykinase gene expression by the forkhead transcription factor FKHR in H4IIE-hepatoma cells. Biochem Biophys Res Commun 285(4):897–902
Dijkers PF, Medema RH, Pals C, Banerji L, Thomas NS, Lam EW, Burgering BM, Raaijmakers JA, Lammers JW, Koenderman L, Coffer PJ (2000) Forkhead transcription factor FKHR-L1 modulates cytokine-dependent transcriptional regulation of p27(KIP1). Mol Cell Biol 20(24):9138–9148
Medema RH, Kops GJ, Bos JL, Burgering BM (2000) AFX-like forkhead transcription factors mediate cell-cycle regulation by Ras and PKB through p27kip1. Nature 404(6779):782–787
Leger B, Cartoni R, Praz M, Lamon S, Deriaz O, Crettenand A, Gobelet C, Rohmer P, Konzelmann M, Luthi F, Russell AP (2006) Akt signalling through GSK-3β, mTOR and Foxo1 is involved in human skeletal muscle hypertrophy and atrophy. J Physiol 576(Pt 3):923–933
Southgate RJ, Neill B, Prelovsek O, El-Osta A, Kamei Y, Miura S, Ezaki O, McLoughlin TJ, Zhang W, Unterman TG, Febbraio MA (2007) FOXO1 regulates the expression of 4E-BP1 and inhibits mTOR signaling in mammalian skeletal muscle. J Biol Chem 282(29):21176–21186
Van Der Heide LP, Hoekman MF, Smidt MP (2004) The ins and outs of Foxo shuttling: mechanisms of Foxo translocation and transcriptional regulation. Biochem J 380(Pt 2):297–309
Brunet A, Bonni A, Zigmond MJ, Lin MZ, Juo P, Hu LS, Anderson MJ, Arden KC, Blenis J, Greenberg ME (1999) Akt promotes cell survival by phosphorylating and inhibiting a forkhead transcription factor. Cell 96(6):857–868
Hribal ML, Nakae J, Kitamura T, Shutter JR, Accili D (2003) Regulation of insulin-like growth factor-dependent myoblast differentiation by Foxo forkhead transcription factors. J Cell Biol 162(4):535–541
Bois PR, Grosveld GC (2003) FKHR (FOXO1a) is required for myotube fusion of primary mouse myoblasts. EMBO J 22(5):1147–1157
Kamei Y, Miura S, Suzuki M, Kai Y, Mizukami J, Taniguchi T, Mochida K, Hata T, Matsuda J, Aburatani H, Nishino I, Ezaki O (2004) Skeletal muscle FOXO1 (FKHR) transgenic mice have less skeletal muscle mass, down-regulated type I (slow twitch/red muscle) fiber genes, and impaired glycemic control. J Biol Chem 279(39):41114–41123
Kitamura T, Kitamura YI, Funahashi Y, Shawber CJ, Castrillon DH, Kollipara R, DePinho RA, Kitajewski J, Accili D (2007) A Foxo/Notch pathway controls myogenic differentiation and fiber type specification. J Clin Invest 117(9):2477–2485
Bigard X, Sanchez H, Zoll J, Mateo P, Rousseau V, Veksler V, Ventura-Clapier R (2000) Calcineurin co-regulates contractile and metabolic components of slow muscle phenotype. J Biol Chem 275(26):19653–19660
Olson EN, Williams RS (2000) Remodeling muscles with calcineurin. Bioessays 22(6):510–519
Ni YG, Wang N, Cao DJ, Sachan N, Morris DJ, Gerard RD, Kuro OM, Rothermel BA, Hill JA (2007) Foxo transcription factors activate Akt and attenuate insulin signaling in heart by inhibiting protein phosphatases. Proc Natl Acad Sci USA 104(51):20517–20522
Ni YG, Berenji K, Wang N, Oh M, Sachan N, Dey A, Cheng J, Lu G, Morris DJ, Castrillon DH, Gerard RD, Rothermel BA, Hill JA (2006) Foxo transcription factors blunt cardiac hypertrophy by inhibiting calcineurin signaling. Circulation 114(11):1159–1168
Lara-Pezzi E, Winn N, Paul A, McCullagh K, Slominsky E, Santini MP, Mourkioti F, Sarathchandra P, Fukushima S, Suzuki K, Rosenthal N (2007) A naturally occurring calcineurin variant inhibits Foxo activity and enhances skeletal muscle regeneration. J Cell Biol 179(6):1205–1218
Nakao C, Ookawara T, Kizaki T, Oh-Ishi S, Miyazaki H, Haga S, Sato Y, Ji LL, Ohno H (2000) Effects of swimming training on three superoxide dismutase isoenzymes in mouse tissues. J Appl Physiol 88(2):649–654
Ueno N, Oh-ishi S, Kizaki T, Nishida M, Ohno H (1997) Effects of swimming training on brown-adipose-tissue activity in obese ob/ob mice: GDP binding and UCP m-RNA expression. Res Commun Mol Pathol Pharmacol 95(1):92–104
Huang H, Muddiman DC, Tindall DJ (2004) Androgens negatively regulate forkhead transcription factor FKHR (FOXO1) through a proteolytic mechanism in prostate cancer cells. J Biol Chem 279(14):13866–13877
Chow LS, Greenlund LJ, Asmann YW, Short KR, McCrady SK, Levine JA, Nair KS (2007) Impact of endurance training on murine spontaneous activity, muscle mitochondrial DNA abundance, gene transcripts, and function. J Appl Physiol 102(3):1078–1089
Hughes SM, Taylor JM, Tapscott SJ, Gurley CM, Carter WJ, Peterson CA (1993) Selective accumulation of MyoD and myogenin mRNAs in fast and slow adult skeletal muscle is controlled by innervation and hormones. Development 118(4):1137–1147
Howitz KT, Bitterman KJ, Cohen HY, Lamming DW, Lavu S, Wood JG, Zipkin RE, Chung P, Kisielewski A, Zhang LL, Scherer B, Sinclair DA (2003) Small molecule activators of sirtuins extend saccharomyces cerevisiae lifespan. Nature 425(6954):191–196
Motta MC, Divecha N, Lemieux M, Kamel C, Chen D, Gu W, Bultsma Y, McBurney M, Guarente L (2004) Mammalian SIRT1 represses forkhead transcription factors. Cell 116(4):551–563
Bai L, Pang WJ, Yang YJ, Yang GS (2008) Modulation of Sirt1 by resveratrol and nicotinamide alters proliferation and differentiation of pig preadipocytes. Mol Cell Biochem 307(1–2):129–140
Tran H, Brunet A, Grenier JM, Datta SR, Fornace AJ Jr, DiStefano PS, Chiang LW, Greenberg ME (2002) DNA repair pathway stimulated by the forkhead transcription factor FOXO3a through the Gadd45 protein. Science 296(5567):530–534
Furukawa-Hibi Y, Yoshida-Araki K, Ohta T, Ikeda K, Motoyama N (2002) Foxo forkhead transcription factors induce G(2)-M checkpoint in response to oxidative stress. J Biol Chem 277(30):26729–26732
Naya FJ, Mercer B, Shelton J, Richardson JA, Williams RS, Olson EN (2000) Stimulation of slow skeletal muscle fiber gene expression by calcineurin in vivo. J Biol Chem 275(7):4545–4548
Rothermel BA, Vega RB, Williams RS (2003) The role of modulatory calcineurin-interacting proteins in calcineurin signaling. Trends Cardiovasc Med 13(1):15–21
Yang J, Rothermel B, Vega RB, Frey N, McKinsey TA, Olson EN, Bassel-Duby R, Williams RS (2000) Independent signals control expression of the calcineurin inhibitory proteins MCIP1 and MCIP2 in striated muscles. Circ Res 87(12):E61–E68
Sandri M, Sandri C, Gilbert A, Skurk C, Calabria E, Picard A, Walsh K, Schiaffino S, Lecker SH, Goldberg AL (2004) Foxo transcription factors induce the atrophy-related ubiquitin ligase atrogin-1 and cause skeletal muscle atrophy. Cell 117(3):399–412
Stitt TN, Drujan D, Clarke BA, Panaro F, Timofeyva Y, Kline WO, Gonzalez M, Yancopoulos GD, Glass DJ (2004) The IGF-1/PI3K/Akt pathway prevents expression of muscle atrophy-induced ubiquitin ligases by inhibiting FOXO transcription factors. Mol Cell 14(3):395–403
Mahoney DJ, Parise G, Melov S, Safdar A, Tarnopolsky MA (2005) Analysis of global mRNA expression in human skeletal muscle during recovery from endurance exercise. FASEB J 19(11):1498–1500
Russell AP, Hesselink MK, Lo SK, Schrauwen P (2005) Regulation of metabolic transcriptional co-activators and transcription factors with acute exercise. FASEB J 19(8):986–988
Ropelle ER, Pauli JR, Cintra DE, Frederico MJ, de Pinho RA, Velloso LA, De Souza CT (2009) Acute exercise modulates the Foxo1/PGC-1α pathway in the liver of diet-induced obesity rats. J Physiol 587(Pt 9):2069–2076
Allen DL, Sartorius CA, Sycuro LK, Leinwand LA (2001) Different pathways regulate expression of the skeletal myosin heavy chain genes. J Biol Chem 276(47):43524–43533
Baur JA, Pearson KJ, Price NL, Jamieson HA, Lerin C, Kalra A, Prabhu VV, Allard JS, Lopez-Lluch G, Lewis K, Pistell PJ, Poosala S, Becker KG, Boss O, Gwinn D, Wang M, Ramaswamy S, Fishbein KW, Spencer RG, Lakatta EG, Le Couteur D, Shaw RJ, Navas P, Puigserver P, Ingram DK, de Cabo R, Sinclair DA (2006) Resveratrol improves health and survival of mice on a high-calorie diet. Nature 444(7117):337–342
Lagouge M, Argmann C, Gerhart-Hines Z, Meziane H, Lerin C, Daussin F, Messadeq N, Milne J, Lambert P, Elliott P, Geny B, Laakso M, Puigserver P, Auwerx J (2006) Resveratrol improves mitochondrial function and protects against metabolic disease by activating SIRT1 and PGC-1alpha. Cell 127(6):1109–1122
Zong H, Ren JM, Young LH, Pypaert M, Mu J, Birnbaum MJ, Shulman GI (2002) AMP kinase is required for mitochondrial biogenesis in skeletal muscle in response to chronic energy deprivation. Proc Natl Acad Sci USA 99(25):15983–15987
Wu Z, Puigserver P, Andersson U, Zhang C, Adelmant G, Mootha V, Troy A, Cinti S, Lowell B, Scarpulla RC, Spiegelman BM (1999) Mechanisms controlling mitochondrial biogenesis and respiration through the thermogenic coactivator PGC-1. Cell 98(1):115–124
Lin J, Wu H, Tarr PT, Zhang CY, Wu Z, Boss O, Michael LF, Puigserver P, Isotani E, Olson EN, Lowell BB, Bassel-Duby R, Spiegelman BM (2002) Transcriptional co-activator PGC-1 alpha drives the formation of slow-twitch muscle fibres. Nature 418(6899):797–801
Puigserver P, Rhee J, Donovan J, Walkey CJ, Yoon JC, Oriente F, Kitamura Y, Altomonte J, Dong H, Accili D, Spiegelman BM (2003) Insulin-regulated hepatic gluconeogenesis through FOXO1-PGC-1alpha interaction. Nature 423(6939):550–555
Sandri M, Lin J, Handschin C, Yang W, Arany ZP, Lecker SH, Goldberg AL, Spiegelman BM (2006) PGC-1alpha protects skeletal muscle from atrophy by suppressing Foxo3 action and atrophy-specific gene transcription. Proc Natl Acad Sci USA 103(44):16260–16265
Acknowledgments
This study was supported by Major Projects for Genetically Modified Organisms Breeding (No. 2009ZX08009-157B). We thank Professor Haojie Huang (Minnesota University, USA) for the kind donation of FoxO1-WT and FoxO1-A3 plasmids. We also thank Post Ph.D Qingwu Shen for his kind help in the revision of the original manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yuan, Y., Shi, Xe., Liu, Yg. et al. FoxO1 regulates muscle fiber-type specification and inhibits calcineurin signaling during C2C12 myoblast differentiation. Mol Cell Biochem 348, 77–87 (2011). https://doi.org/10.1007/s11010-010-0640-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-010-0640-1