Abstract
Pediococcus acidilactici NCDC 252 is a Gram-positive lactic acid bacteria (LAB) that is being studied to develop it as probiotic and starter culture for dairy and meat industry. Peptidoglycan hydrolases (PGHs) are also gaining interest because of their antibacterial activity. NCDC 252 genome was searched for PGH encoded genes and product of these genes were in silico characterized for different parameters. Extracellular PGHs of NCDC 252 were precipitated and characterized. PGHs were screened turbidometrically with Bacillus cereus and Staphylococcus albus as substrates. Spectrum of PGH(s) was studied in renaturing SDS-PAGE by varying substrates and renaturing conditions. PGH(s) band intensity decreased in the presence of NaCl whereas some additional secondary lytic bands appeared in the presence of EDTA. Autolysis of LAB in cheese and other fermented food products is associated with release of oligopeptides by proteolysis and other enzyme activities. Autolytic phenotype of P. acidilactici was evaluated and characterized. Lytic activity studies revealed 45–60% activity against Micrococcus lysodeikticus and B. cereus greater than for P. acidilactici. PGH of P. acidilactici exhibited broad growth inhibiting spectrum of antibacterial activity against studied Gram positive and Gram negative bacteria and thus can be promising in controlling pathogenic bacteria. This also confers probiotic attribute to the studied strain.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Lysis of lactic acid bacteria (LAB) holds special interest due to their use as starter culture and results in release of cytoplasmic peptidases, lipases and other enzymes. Pediococcus acidilactici NCDC 252 is an industrially important LAB of dairy origin with probiotic attributes (Attri et al. 2012, 2015). Pediocoocus sp. was also studied for dietary effects as industrial probiotic (Mikulski et al. 2012). Being facultative anaerobe, its use can be extended to broad range of processes such as in food products viz meat, milk and vegetables to improve their quality and safety along with an important role in cheese ripening.
Peptidoglycan, a major component of bacterial cell is composed of alternating N-acetylglucosaminyl and N-acetylmuramyl residues with cross-linking short peptides of different composition (Vollmer et al. 2008). Peptidoglycan Hydrolases (PGHs) cleave peptide bonds of peptidoglycan and depending upon hydrolytic specificity they are classified as N-acetyl-muramidases, N-acetyl-glucosaminidases, N-acetylmuramoyl-l-alanine amidase and endopeptidases (Chapot-Chartier 1996). PGHs perform different functions like daughter cell separation, cell-wall turnover, autolysis, sporulation and germination, biofilm formation, resuscitation of dormant cells and allolysis in genetic transformation (Smith et al. 2000; Valence and Lortal 1995).
Several studies have been carried out to detect and characterize autolytic enzyme activities in different Gram-positive bacteria viz. Staphylococcus aureus (Tomasz 2000), Bacillus subtilis (Smith et al. 2000), Bacillus thuringiensis (Raddadi et al. 2004), Lactococcus lactis (Huard et al. 2003), Lactobacillus pentosus (Cibik and Chapot-Chartier 2004), Enterococcus faecalis (Mesnage et al. 2008), Pediococcus sp. (Garcia-Cano et al. 2011) and Lactobacillus sakei (Najjari et al. 2016). Autolysins of LAB play an important role in organoleptic properties of fermented dairy products by releasing intracellular enzymes (Cibik and Chapot-Chartier 2004). Ability of PGH(s) to hydrolyse cell wall of pathogenic and food associated bacterial species may enhance the efficacy of biopreservation and thus PGH(s) are suggested as replacement of antimicrobials. LAB being GRAS and probiotics will hold even better place in food preservation and therapeutics.
PGHs also confer probiotic properties on bacteria because they are involved in host-microbe interactions and activation of immune response through anti-inflammatory mechanisms (Rolain et al. 2012). PGHs also function as cell adhesion molecules with antimicrobial properties (Beaussart et al. 2013). PGH(s) need to be studied in detail for better understanding of release muramoyl peptides as probiotic immunomodulators and antipathogenic enzymes so as to extend their use for different applications. Membrane bound and intracellular PGH(s) of P. acidilactici have been reported but there is single report of its extracellular PGHs (Llorente-Bousquets et al. 2008). Extracellular PGH(s) are more important with respect to biopreservation and probiotic attributes because enzymes are supplied continuously during the process. NCDC 252 is the first and novel genome of any P. acidilactici that has been characterized (Bansal et al. 2019). In order to understand role of PGH(s), it is important to isolate and characterize extracellular PGHs and also to assess its performance against some bacteria. This paper describes genome analysis for PGHs, precipitation and characterization of extracellular PGH(s) from P. acidilactici NCDC 252.
Materials and Methodology
Materials
Ammonium sulphate, Bromophenol Blue, bovine serum albumin (BSA), ethylendiamine tetra acetic acid (EDTA), MRS (de Man Rogosa and Sharpe) medium, prestained markers (11–245 KDa), sodium chloride, sodium dodecyl sulphate, sodium phosphate, Triton X-100 were purchased from Himedia. Potassium hydroxide was purchased from Rankem. Methylene blue, Micrococcus lysodeikticus ATCC 4698 were obtained from Sigma Chemical Co., St. Louis, MO, USA.
In Silico Analysis of PGH from NCDC 252 Genome
In silico analysis of genes encoding for PGH was done by using BLAST2GO platform. The gene sequences were translated by Gene translator (https://www.scfbioiitd.res.in/chemgenome/genetranslator.jsp).
Extinction Coefficient, Aliphatic index and Grand Average of hydropathicity (GRAVY) of PGHs was done using Expasy’s Proptaram (https://www.web.expasy.org/protparam/) server. Subcellular location of proteins was identified by PSORTb.
Homology Modeling
MODELLER was used for comparative and homology modeling of protein’s three-dimensional structure. MODELLER9.21 was downloaded and installed from https://saliab.org/modeller/download_installation.html. The user provides a query sequence (protein) to be modeled with identified related structures and calculates model containing all non-hydrogen atoms. The modeled protein was validated by Protein Structure Validation Software Suite (PSVS) (https://psvs-1-5-dev.nesg.org/).
Ligand Binding Site
Ligand binding sites of modelled proteins were predicted by 3DLigand Site (https://www.sbg.bio.ic.ac.uk/3dligandsite/index.cgi) and COFACTOR (https://zhanglab.ccmb.med.umich.edu/COFACTOR/).
Bacterial Strains and Growth Conditions
Pediococcus acidilactici NCDC 252, Bacillus cereus NCDC 66, E. coli NCDC 134, Staphylocoocus aureus NCDC 110 and Stphylococcus albus NCDC 111 were purchased from NCDC (National collection of Dairy Cultures), National Dairy Research Institute (NDRI), Karnal. Bacterial cultures were maintained as 50% glycerol stock and stored at − 20 °C for further use. P. acidilactici was grown in MRS broth at 32 °C and B. cereus, E. coli, S. aureus and S. albus were grown in nutrient broth at 37 °C. Plating media were prepared by adding 2% agar to medium (w/v).
Isolation of Extracellular PGHs
Pediococcus acidilactici was grown at 32 °C in MRS medium for 36 h in orbital shaker at 250 rpm and culture was centrifuged at 4800 rpm for 45 min at 4 °C (Dynamica) to collect supernatant. The supernatant was subjected to 0–80% ammonium sulphate saturation to isolate extracellular PGHs. The precipitated proteins were collected by centrifugation at 4800 rpm, resuspended in 3 ml of Tris–HCl buffer (pH 7.4) and extensively dialysed against same buffer (100 ml). This sample was used as source of PGH(s).
Isolation of Peptidoglycans
Peptidoglycans were obtained by growing cells in their respective broth upto mid log phase and harvested by centrifugation at 10,000 rpm for 10 min. Pellets (cells) were resuspended in 1 ml of buffer (50 mM Tris–HCl, 7.4) and autoclaved and sonicated at 15 kHz for 4 min. Broken cells were collected by centrifugation at 10,000 rpm for 10 min and pellet was resuspended in the same buffer.
Protein Assay
Total protein content was quantified by Lowry’s method using BSA as standard (Lowry et al. 1951).
PGH(s) Assay by Turbidometric Method
PGH(s) were screened according to method of Shugar with some modifications (Shugar 1952). Bacterial suspension (B. cereus and S. albus), 0.3 mg ml−1 in 0.1 M sodium phosphate, pH 6.4 (950 µl), was incubated with partially purified PGH (50 µl) in a 1 ml cuvette at 25 °C. Rate of lysis was determined by recording change in absorbance at 600 nm with time. One unit of enzyme activity (U) was defined as the amount of enzyme that resulted in decrease in A600 by 0.001 min−1.
PGH(s) Detection by Zymography
Zymography by SDS-PAGE containing peptidoglycan followed by in-situ enzyme renaturation is the most common method of PGHs detection. PGH activity was analysed on renaturing SDS-PAGE (15%) (Buist et al. 1995). Autoclaved cells of M. lysodeikticus ATCC 4698 (0.2% w/v) were incorporated into polyacrylamide running gel. Sample was prepared by mixing 40 µl sample with 30 µl of loading buffer (50 mmol−l Tris–HCl, pH 6.8 containing 2% (w/v) SDS, 2.5% (v/v) glycerol and 0.5% (w/v) bromophenol blue). After electrophoresis, gels were soaked in 200 ml of 1% Triton X-100 for 1–2 h and then in 200 ml of 50 mmol−1 Tris–HCl (pH 7.4) for 12–36 h. Bands of lytic activity were visualized by staining gel with 1% (w/v) methylene blue in 0.01% (w/v) KOH and subsequent destaining in sterilized distilled water.
Molecular weight for peptidoglycan hydrolases was analysed by comparing the PGH(s) band(s) on zymogram with prestained protein markers with molecular weight in the range of 10–245 KDa. A green band of 25 KDa and red of 75 KDa were the reference proteins.
Characterization of Peptidoglycan Hydrolases
Effect of pH
Effect of pH on PGHs was evaluated by zymographic analysis by varying pH of renaturation buffer viz. 50 mM Na-acetate buffer pH 4.0; 50 mM Tris–HCl buffer pH 7.4; 50 mM Glycine–NaOH buffer of pH, 10.5.
Effect of Chemicals
Effect of NaCl and EDTA on PGHs was studied by adding at 1% concentration in renaturation buffer.
Effect of Renaturation Time
Effect of renaturation time on PGHs was studied by incubating gel slices in renaturation buffer (50 mM Tris–HCl, 7.4) for different time intervals.
Substrate Specificity
Substrate specificity of peptidoglycan hydrolase(s) was evaluated by incorporating cells of B. cereus, P. acidilactici and M. lysodeikticus in polyacrylamide gel for zymographic analysis.
Autolytic Phenotype of P. acidilactici
Pediococcus acidilactici cells were grown in MRS and harvested in exponential phase. Cells were washed in Na-Phosphate buffer (50 mM, pH 6.5) and resuspended in same buffer such that O.D.600 nm was between 0.6 and 0.8. The cells were incubated at 32 °C and extent of autolysis was studied and expressed as percentage decrease in O.D.600 nm after 72 h.
Triton X-100 Induced Autolysis
Pediococcus acidilactici cells were grown to exponential phase and harvested by centrifugation at 5000 rpm for 10 min at 4 °C. Cells were then washed with 50 mM Na-Phosphate buffer (pH 6.5) and resuspended in same buffer having 0.05% Triton X-100 and incubated at 32 °C. Autolysis was monitored by recording the O.D.600 of the cell suspensions at different intervals (Rolain et al. 2012).
Lytic Activity
Lysis of bacterial strains was studied by method of Valence and Lortal (1995) with slight modifications. One hundred microliter of PGH was separately added to 1.4 ml of M. lysodeikticus, sonicated, B. cereus and P. acidilactici cells (0.2 mg in Na-phosphate buffer at pH 6.5). The samples were incubated at 40ºC with constant stirring. Autolysis was monitored by recording O.D.600 nm for 8 h. One unit of enzyme activity was defined as the amount of enzyme that resulted in decrease of 0.001 U/min at 40 °C in O.D.600 nm for both substrates.
Antibacterial Activity of PGHs
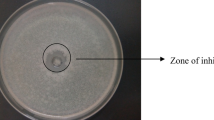
Antibacterial effect of partially purified PGH(s) was studied by agar well diffusion method (Tagg et al. 1976). S. aureus, S. albus, B. cereus and E. coli were used as indicator organisms to check sensitivity to purified PGH of P. acidilactici. Solidified agar was bored with a sterile metal cork borer and filled with 100 μl PGH(s). After diffusion of PGH(s), plates were incubated at 37 °C for 18 h.
Results
In Silico Analysis of PGH from NCDC 252 Genome
Study of genes encoding for PGH in NCDC 252 genome revealed presence of only two PGHs genes (PAN_956, PAN_2301) that encode for PGHs expected to be located in bacterial cell wall. The estimated molecular weight of PGH1 and PGH2 was 105.5 KDa and 30.9 KDa respectively. In silico characterization of these proteins revealed their computed extinction Coefficient (ε) to be 104,360 and 39,880 M−1 cm−1 respectively. The ε of proteins helps to study protein–ligand and protein–protein interactions in solution. AI determines relative value of aliphatic side chains (A, L, I and V) and thus thermostability of proteins. AI of PGH1 and PGH2 was 65.14 and 87.87 respectively. High values of their AI indicate proteins’ stability over wide temperature range. GRAVY value was calculated as sum of hydropathy (hydrophilic and hydrophobic properties of each amino acid side chain) values of all the amino acid divided by number of residues in sequence. Negative GRAVY indicates non polar nature of protein whereas positive value indicates the polar nature. Both PGH1 and PGH2 are non-polar with GRAVY value of − 0.49 and − 0.160 respectively. 3D structures of PGH1 and PGH2 were elucidated by modeller (Fig. 1) using crystal structure of the mucin-binding domain of Spr1345 from Streptococcus pneumoniae (PDB ID: 3NZ3) and crystal structure of N-acetylmuramoyl-l-alanine amidase from Clostridium difficile 630 (PDB ID: 4RN7) respectively as their best templates. The overall G-factor scores of − 0.06 indicated that the model was acceptable as it was greater than recommended value (− 0.5). Verification of 3D results for PGHs model showed positive scores (cut-off score was ≥ 0.2) which indicated the reliability of proposed model.
Ligand Binding Site
Amino acid residues predicted to be involved in binding were Ser, Thr, Asn, Val, Ala and Lys in the modeled PGH1 and His, Asp, Leu, Gly and Asn were present in PGH2 (Fig. 2).
Screening of PGH Activity
In accordance to in silico study, proteins were extracted from cell wall of NCDC 252. The extract when assayed for PGHs revealed lack of PGH activity. Therefore, PGHs were analyzed in extracellular fluid with the rationale that PGHs reportedly coaggregate and/or lyse and even their constituent polypeptides exhibit PGH activity. Extracellular PGHs might be those that are released from its cell wall.
Protein Content and Fold Purification
Total protein content was 2139 mg and 340 mg in crude extract and ammonium sulphate precipitated fraction respectively. This single step resulted in 1.58 ± 0.03 fold purification and yield of 25.09 ± 0.2% with respect to total protein content. Since assay was zymographic, fold purification was calculated with respect to total protein content only.
Assay of PGH Activity
Turbidometric Method
Decrease in O.D.600 nm was observed for Gram positive substrates viz. B. cereus and S. albus and activity units were calculated (Table 1). This confirmed PGH activity in extracellular fluid.
Zymography
Zymogram with M. lysodeikticus (0.2% w/v) as substrate revealed one major lytic band of ~ 11 KDa and minor lytic bands with molecular weight of ~ 15, ~ 23 and ~ 35 KDa (Fig. 3b). Low-intensity lytic signals are supposed to be minor autolysins or major autolysins with reduced activity under the experimental conditions. They also might be partially degraded autolysins that retained weak activity under experimental conditions. Zymogram only analyzes the polypeptides that retain PGH activity under denaturating conditions. The PGH2 with estimated molecular weight of 30.9 KDa might be similar to minor lytic band of ~ 35 KDa. This difference in molecular weight might be due to differential electrophoretic mobility in the PAGE containing bacterial cells and traditional SDS-PAGE. PGH1 is expected either to be retained within the cell wall or lysed/modified to different forms after its release in extracellular medium (the different bands shown on zymogram).
Peptidoglycan hydrolases profile of P. acidilactici evaluated by renaturing SDS-PAGE containing autoclaved M. lysodeikticus cells under different renaturing conditions, prestained markers (Lane a), control (under standard conditions) (Lane b), renaturation in buffer of pH 4.0 (Lane c) pH 7.4 (Lane d) and pH 10.5 (Lane e), buffer of pH 7.4 containing 1% NaCl (Lane f) and 10 mM EDTA (Lane g)
Characterization of Peptidoglycan Hydrolases
Peptidoglycans are affected by change in pH and presence of chemicals so PGH activity repeatedly varies under different assay conditions. Therefore PGH(s) were characterized zymographically by varying the conditions of renaturation buffer.
Effect of pH on PGH Activity
Differential PGH activity was observed at different pH (Fig. 3c–e). No PGH band was observed at pH 10.5. An intense band of ~ 11 KDa appeared at pH 7.4 along with minor lytic bands of ~ 15, ~ 23 and ~ 35 KDa while at pH 4.0, intensity of even ~ 11 KDa PGH was very low.
Effect of NaCl and EDTA
Main lytic band of ~ 11 kDa appeared in all conditions. Band intensity decreased in general. Only band of ~ 11 kDa with low intensity appeared in comparison to control whereas other bands disappeared in the presence of 1% NaCl (Fig. 3f). In presence of EDTA, an additional band of 50 KDa appeared (Fig. 3g) along with all other PGH(s) bands in control (Fig. 3b). Bands intensity increased in presence of EDTA.
Effect of Time of Renaturation
Incubation in renaturation buffer was done from 6 h upto 20 h. One major band of ~ 11 KDa was seen along with other minor lytic bands. Intensity of bands was relatively very low when incubated for 6 h (Fig. 4b). Additional bands of ~ 35 and ~ 23 and ~ 15 KDa along with ~ 11 KDa band (high intensity) appeared with increase in incubation time in renaturation buffer (Fig. 4c).
Peptidoglycan hydrolases profile of P. acidilactici on renaturing SDS-PAGE with M. lysodeikticus as substrate (0.2% w/v): prestained markers (Lane a), Fewer low intensity bands were obtained after renaturation for 6 h (Lane b), bands of 15, 23 and 35 KDa along with major band of 11 KDa were observed after renaturaion for 20 h (Lane c)
Substrate Specificity
Spectrum of PGH activity was determined with different substrates viz. Bacillus cereus, P. acidilactici and M. lysodeikticus. With M. lysodeikticus a major band of ~ 11 KDa was seen along with minor lytic bands of ~ 15, ~ 23 and ~ 35 KDa (Fig. 5a). Hydrolytic bands of ~ 15–20 KDa appeared for B. cereus (Fig. 5b) and that of ~ 20–25 KDa with P. acidilactici (Fig. 5c). PGHs reported with all substrates are of low molecular weight.
Characterization of Autolytic Phenotype of P. acidilactici
The autolytic rate of P. acidilactici at 32 °C was evaluated, at pH 6.5 after 72 h. Autolysis of 30% was reported (Fig. 6). Carbon source played an effective role on autolytic phenotype and autolysis increased from 30 to 41% in the presence of glucose.
Effect of Triton X-100
Autolysis rate also increased from 13 to 36% in the presence of 0.05% Triton X-100 upto 15 h respectively (Fig. 7).
Lytic Enzyme Activity Against Different Substrates
Lysis was studied by monitoring the decrease in O.D.600 nm of M. lysodeikticus, B. cereus and P acidilactici. M. lysodeikticus and B. cereus were lysed in range of 45–60% with calculated enzyme activity units of 0.843 and 0.920 U/min respectively whereas activity units of P. acidilactici was lysed to a lesser extent with 0.181 U/min (Fig. 8).
Application Studies of PGH
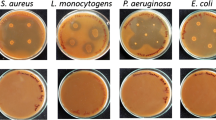
In order to explore the potential of PGHs on lysis of mixed bacterial community their effect was tested on Gram positive (S. albus, S. aureus and B. cereus) and Gram negative bacteria (E. coli) by agar well diffusion method. Partially purified PGHs showed antibacterial activity against all the studied bacteria (Fig. 9). Highest antibacterial activity was observed against E. coli and B. cereus with zone of inhibition of 18 mm and 17 mm respectively.
Discussion
PGH activity was studied by both turbidometric and zymographic method. Five lytic bands (~ 11, ~ 15, ~ 23, ~ 35, ~ 50 KDa) were observed in zymography as major and minor autolysins. Different peptidoglycan hydrolases were active under different conditions. M. lysodeikticus is standard substrate for zymographic analysis of PGH and also used for other experiments. Peptidoglycan hydrolases of P. acidilactici were active against all studied substrates including P. acidilactici itself. Reduced intensity of PGHs bands was observed under acidic and basic conditions which was also observed earlier (Cibik and Chapot-Chartier, 2000). However cytoplasmic PGH(s) of Pediococcus strain were active over a broad pH range of 5.0–9.5 though band intensity varied (Mora et al. 2003). Increased band intensity in presence of EDTA might be due to chelation of bacterial cell wall Ca2+ by EDTA. However, a slight decrease in band intensity was observed with EDTA in L. sakei (Najjari et al. 2016). Presence of a major lytic band of ~ 11KDa and secondary lytic bands active in low salt concentration and neutral pH suggest their role in defense/stress management. Appearance of additional bands due to incubation in renaturation buffer might be due to renaturation of some additional PGHs.
Lytic studies also support the substrate specificity studied by zymographic method. Lytic bands were observed with all studied substrates. Our studies of substrate specificity are in contrast to other reports as high molecular weight lytic activities were reported earlier in different strains of P. acidilactici viz a high molecular weight PGH of 110 KDa (Llorente-Bousquets et al. 2008), an intracellular lytic band of 116 KDa with M. lysodeikticus substrate (Mora et al. 2003). Two membrane bound lytic activities of 110 KDa and 99 KDa with high N-acetyl muramidase activity and harbouring catalytic sites for N-acetyl muramoyl-l-alanine amidase and N-acetyl glucosaminidase activity were also reported from P. acidilactici (Garcia-Cano et al. 2015). Post translational modification may also result in several different form of same enzyme activity. Even degradation by proteases also produce the degradation products that retained the enzyme activity. Minor lytic bands may be result of partial proteolytic degradation of larger proteins that produce fragments retaining enzymatic activity for many Gram-positive bacteria (Govindasamy-Lucey et al. 2000; Mora et al. 2003). Same lytic bands may have different activities with different substrates under varied renaturation conditions (Foster, 1992). Thus PGHs profile visualized by renaturing SDS-PAGE is a reflection of only those enzymes which could renature even after SDS treatment and not all enzymes.
Autolysis is a strain dependent character that is usually influenced by carbon source in medium. Autolysis of P. acidilactici NCDC252 increased in presence of glucose. These results are in agreement to earlier reports of Pediococcus pentosaceus and P. acidilactici which showed 40–90% autolysis after 48 h at 37 °C (Mora et al. 2003). This can be attributed to weaker cell wall in presence of glucose as reported for other LAB such as Lactococcus lactis (Vegarud et al. 1983), S. thermophilus (Sandholm and Sarimo 1981), Leuconostoc (Cibik and Chapot-Chartier 2000) and L. sakei (Najjari et al. 2016). Difference in lysis in different carbon sources was explained due to modification of cell wall structure (Buist et al. 2008; Steen et al. 2008). Autolytic rate was reported in range of 38–80% for all the studied B. cereus/B. thuringiensis/B. weihenstephanensis strains with only exception for B. cereus DISTAM 345 which showed a rate of 25.4% with higher autolysis in buffer of pH 6.5 (Raddadi et al. 2005). Detergent of non-ionic nature like Triton X-100 forms micelles with lipoteichoic acid present in cell wall peptidoglycans and induce autolysis (Neuhaus and Baddiley 2003).
Further experiments to study effect of peptidoglycan hydrolase(s) on different bacteria confirmed antibacterial activity against Gram positive bacteria and food related pathogens. Antibacterial activity against Gram positive as well as Gram negative bacteria is quite encouraging. There are sporadic reports of antibacterial activity by Gram positive bacteriocin producers against Gram negative strains (Elgado et al. 2004). Antibacterial activity extends potential biotechnological use of PGHs in food safety and preservation as well as in pharmaceuticals. This can be used as starter culture strain in food to control food borne pathogens. Activity against Bacillus cereus and E. coli is of great relevence. B. cereus is frequently the causal agent of gastroenteric diseases associated with food consumption. Resistance of bacteria to antibiotics have reduced their efficacy and there is a hypothesis of using PGHs and antimicrobial peptides as substitute for antibiotics. This will be promising in overcoming the challenge of antibiotic resistance strains. P. acidilactici reportedly produce pediocins effective against several Gram-positive and food-related pathogens such as L. monocytogenes (Mora et al. 2003). PGHs have strong potential to control and treat infections by pathogenic bacteria and also strengthen bactericidal or bacteriostatic effect of antibiotics (Szweda et al. 2012). These aspects are being studied further.
Conclusion
PGHs of P. acidilactici are active under different conditions with different substrates. Autolytic activity of strain release cytoplasmic proteolytic enzymes which release small peptides and free amino acids important in fermentation and cheese ripening. They can be explored for industrial applications but their role in bacterial physiology need further studies. Antibacterial activity of PGH is also a probiotic attribute. Ability of PGH to hydrolyse cell wall of pathogenic and food associated bacterial species can be effective in biopreservation. Autolytic enzymes offer the possibility to develop more robust starter strains. This information will help to select this strain for its use as starter culture for formation of safe food products and probiotic formulations. Further studies will be focused on study of gene(s) constructs and their mutants. This will help to explore role of PGH in autolysis and cell physiology.
References
Attri P, Jodha D, Gandhi D, Dhanda S (2012) Screening of intracellular, extracellular and membrane bound exopeptidases in Lactic Acid Bacteria (LAB). Milchwissenschaft 67:421–424
Attri P, Jodha D, Gandhi D, Chanalia P, Dhanda S (2015) acidilactici national collection of dairy cultures 252 for its probiotic attributes. Int J Dairy Technol 68:533–542
Bansal P, Kumar R, Singh J, Dhanda S (2019) Next generation sequencing, biochemical characterization, metabolic pathway analysis of novel probiotic Pediococcus acidilactici NCDC 252 and it’s evolutionary relationship with other lactic acid bacteria. Mol Biol Rep. https://doi.org/10.1007/s11033-019-05022-z
Beaussart A, Rolain T, Duchene M-C, EI-Kirat-Chatel S, Andre J, Hols P, Dufrene YF (2013) Binding mechanism of the peptidoglycan hydrolase Acm2: low affinity, broad specificity. Biophys J 105:620–629
Buist G, Kok J, Leenhouts KJ, Dabrowska M, Venema G, Haandrikman J (1995) lactis a muramidase needed for cell separation. J Bacteriol 177:1554–1563
Buist G, Steen A, Kok J, Kuipers OP (2008) LysM, a widely distributed protein motif for binding to (peptide) glycans. Mol Microbiol 68:838–847
Chapot-Chartier MP (1996) Les autolysines des bactéries lactiques. Lait 76:91–109
Cibik R, Chapot-Chartier M-P (2000) Autolysis of dairy Leuconostocs and detection of peptidoglycan hydrolases by renaturing SDS-PAGE. J Appl Microbiol 89:862–869
Cibik R, Chapot-Chartier M-P (2004) Characterization of autolytic enzymes in Lactobacillus pentosus. Lett Appl Microbiol 38:459–463
Elgado FB, Guerra MARV, Macayan RA, Mendoza HA, Lirazan MB (2004) Spectrum of bacteriocin activity of Lactobacillus plantarum BS and fingerprinting by RAPD-PCR. Int J Food Microbiol 95:11–18
Foster SJ (1992) Analysis of the autolysins of Bacillus subtilis 168 during vegatative growth and differentiation by using renaturating polyacrylamide gel electrophoresis. J Bacteriol 174:464–470
García-Cano I, Velasco-Pérez L, Rodríguez-Sanoja R, Sánchez S, Mendoza-Hernández G, Llorente-Bousquets A, Farrés A (2011) acidilactici ATCC 8042. J Appl Microbiol 111:607–615
García-Cano I, Campos-Gómez M, Contreras-Cruz M, Serrano-Maldonado CE, González-Canto A, Peña-Montes C, Rodríguez-Sanoja R, Farrés A (2015) acidilactici ATCC 8042. Appl Microbiol Biotechnol 99:8563–8573
Govindasamy-Lucey S, Gopal PK, Sullivan PA, Pillidge CJ (2000) lactis cheese starter bacteria grown in milk. J Dairy Res 67:585–596
Huard C, Miranda G, Wessner F, Bolotin A, Hansen J, Foster SJ, Chapot-Chartier M-P (2003) lactis. Microbiology 149:695–705
Llorente-Bousquets A, Perez-Munguia S, Farres A (2008) Novel extracellular proteolytic activity in P. acidilactici ATCC 8042. Can J Microbiol 54:694–699
Lowry OH, Rossenbrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 93:265–275
Mesnage S, Chau F, Dubost L, Arthur M (2008) Role of N-acetylglucosaminidase and N-acetylmuramidase activities in Enterococcus faecalis peptidoglycan metabolism. J Biol Chem 283:19845–19853
Mikulski D, Jankowski J, Naczmanski J, Mikulska M, Demey V (2012) acidilactici) supplementation on performance, nutrient digestibility, egg traits, egg yolk cholesterol, and fatty acid profile in laying hens. Poult Sci 91:2691–2700
Mora D, Musacchio F, Fortina MG, Senini L, Manachini PL (2003) pentocaseus strains. J Appl Microbiol 94:561–570
Najjari A, Amairi H, Chaillou S, Mora D, Boudabaus A, Zagorec M, Ouzari H (2016) Phenotypic and genotypic characterization of peptidoglycan hydrolases of Lactobacillus sakei. Adv Res. https://doi.org/10.1016/j.jare.2015.04.004
Neuhaus FC, Baddiley JA (2003) continuum of anionic charge: structures and functions of D-alanyl-teichoic acids in gram-positive bacteria. Microbiol Mol Biol Rev 67:686–723
Raddadi N, Cherif A, Mora D, Ouzari H, Boudabous A, Molinari F (2004) The autolytic phenotype of Bacillus thuringiensis. J Appl Microbiol 97:158–168
Raddadi N, Cherif A, Mora D, Brusetti L, Borin S, Boudabous A, Daffonchio DJ (2005) The autolytic phenotype of the Bacillus cereus group. J Appl Microbiol 99:1070–1081
Rolain T, Bernard E, Courtin P, Bron PA, Kleerebezem M, Chapot-Chartier M-P, Hols P (2012) Identification of key peptidoglycan hydrolases for morphogenesis, autolysis, and peptidoglycan composition of Lactobacillus plantarum WCFS1. Microb Cell Fact 11:137. https://doi.org/10.1186/1475-2859-11-137
Sandholm E, Sarimo SS (1981) Autolysis of Streptococcus thermophiles. FEMS Microbiol Lett 11:125–129
Shugar D (1952) The measurement of lysozyme activity and the ultraviolet inactivation of lysozyme. Biochim Biophys Acta 8:302–309
Smith TJ, Blackman SA, Foster SJ (2000) Autolysins of Bacillus subtilis: multiple enzymes with multiple functions. Microbiol 146:249–262
Steen A, Buist G, Kramer NE, Jalving R, Benus GF, Venema G (2008) lactis on galactose is a consequence of decreased binding of the autolysin AcmA. Appl Environ Microbiol 74:4671–4679
Szweda P, Schielmann M, Kotlowski R, Gorczyca G, Zalewska M, Milewski S (2012) Peptidoglycan hydrolases-potential weaopens against Staphylococcus aureus. Appl Microbiol Biotechnol 96:1157–1174
Tagg JR, Adjoin AS, Watchmaker LW (1976) Bacteriocins of Gram positive bacteria. Bacteriol Rev 40:722–756
Tomasz A (2000) The staphylococcal cell wall. In: Fischetti VA (ed) Gram-positive pathogens. ASM Press, Washington DC, pp 351–360
Valence F, Lortal S (1995) Zymogram and Preliminary Characterization of Lactobacillus helveticus Autolysins. Appl Environ Microbiol 61:3391–3399
Vegarud G, Castberg HB, Langsrud T (1983) Autolysis of group N streptococci: effect of media compositions, modifications and temperatures. J Dairy Sci 66:2294–2302
Vollmer W, Joris B, Charlier P, Foster S (2008) Bacterial peptidoglycan (murein) hydrolases. FEMS Microbiol Rev 32:259–286
Acknowledgements
Authors are thankful to University Grants Commission, New Delhi, India for funding the research in the form of Major research project. Authors also acknowledge Prof. Sunita Siroha, Dept of English, Kurukshetra University, Kurukshetra for editing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
This article also does not contain studies with animal or human participants.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gandhi, D., Chanalia, P., Bansal, P. et al. Peptidoglycan Hydrolases of Probiotic Pediococcus acidilactici NCDC 252: Isolation, Physicochemical and In Silico Characterization. Int J Pept Res Ther 26, 2119–2127 (2020). https://doi.org/10.1007/s10989-019-10008-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-019-10008-3