Abstract
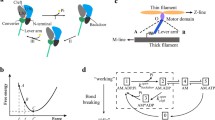
In this article results are reviewed from different experimental approaches to determine the size of the power stroke generated by myosin molecules during their ATPase cycle. While data from fiber studies and protein crystallography predict a stroke size of about 10 nm for skeletal muscle myosins, single molecule studies imply a stroke size for these myosins of only about 5 nm. Single molecule studies also showed the stroke size to be proportional to the length of the light chain binding domain, acting like a lever arm. At the same lever arm length, however, the stroke size of smooth muscle myosin II is found about twice as large and a stroke size of about 14 nm was reported for class-I myosins. It was proposed that such different stroke sizes for molecules with same lever arm length result from different extend of converter domain rotation. Only for class-I myosins, however, an about 30° larger rotation of the converter was found so far by protein crystallography. This, however, is far too small to account for the almost 3-fold larger stroke size reported from single molecule studies. In this contribution we discuss some factors that might account for the apparent discrepancies between single molecule studies on the one hand and protein crystallography as well as some fiber studies on the other hand. In addition, we present some modeling to illustrate that the power stroke very likely is underestimated to a large extent in current single molecule approaches. We further show that differences in the stroke size for various classes of myosins reported from single molecule studies might be related to small differences in the probability to execute the power stroke kinetics. We demonstrate that such small changes in power stroke kinetics can seriously affect the extent to which the ‘true’ power stroke is underestimated by present single molecule approaches.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Anson M, Geeves MA, Kurzawa SE, Manstein DJ, (1996) Myosin motors with artificial lever arms Embo J 15:6069–6074
Brenner B (1988) An experimental approach to determine cross-bridge turnover kinetics during isometric and isotonic steady state contraction using skinned skeletal muscle fibres of the rabbit Pflügers Arch 411: R186
Brenner B (1990) In: Squire J (ed) Molecular Mechanism of Muscular Contraction. (pp. 77–149). Macmillan, London
Burton K (1992) Myosin step size: estimates from motility assays and shortening muscle J Muscle Res Cell Motil 13:90–607
Carragher BO, Cheng N, Wang ZY, Korn ED, Reilein A, Belnap DM, Hammer JA III, Steven AC (1998) Structural invariance of constitutively active and inactive mutants of acanthamoeba myosin IC bound to F-actin in the rigor and ADP-bound states Proc Natl Acad Sci USA 95: 15206–15211
Chen YD, Brenner B (1993) On the regeneration of the actin–myosin power stroke in contracting muscle Proc Natl Acad Sci USA 90: 5148–5152
Cooke R (1986) The mechanism of muscle contraction CRC Crit Rev Biochem 21: 53–118
Corrie JE, Brandmeier BD, Ferguson RE, Trentham DR, Kendrick-Jones J, Hopkins SC, Van Der Heide UA, Goldman YE, Sabido-David C, Dale RE, Criddle S, Irving M (1999) Dynamic measurement of myosin light-chain-domain tilt and twist in muscle contraction Nature 400: 425–430
Finer JT, Simmons RM, Spudich JA (1994) Single myosin molecule mechanics: piconewton forces and nanometre steps Nature 368: 113–119
Ford LE, Huxley AF, Simmons RM (1977) Tension responses to sudden length change in stimulated frog muscle fibres near slack length J Physiol (Lond) 269: 441–515
Geeves MA, Holmes KC (1999) Structural mechanism of muscle contraction Annu Rev Biochem 68: 687–728
Guilford WH, Dupuis DE, Kennedy G, Wu J, Patlak JB, Warshaw DM (1997) Smooth muscle and skeletal muscle myosins produce similar unitary forces and displacements in the laser trap Biophys J 72: 1006–1021
Harada Y, Sakurada K, Aoki T, Thomas DD, Yanagida T (1990) Mechanochemical coupling in actomyosin energy transduction studied by in vitro movement assay J Mol Biol 216: 49–68
Higuchi H, Goldman YE (1991) Sliding distance between actin and myosin filaments per ATP molecule hydrolysed in skinned muscle fibres Nature 352: 352–354
Hill TL (1974) Theoretical formalism for the sliding filament model of contraction of striated muscle. Part I Prog Biophys Mol Biol 28: 267–340
Homsher E, Irving M, Wallner A (1981) High-energy phosphate metabolism and energy liberation associated with rapid shortening in frog skeletal muscle J Physiol (Lond) 321: 423–436
Howard J, Spudich JA (1996) Is the lever arm of myosin a molecular elastic element? Proc Natl Acad Sci USA 93: 4462–4464
Huxley AF (1957) Muscle structure and theories of contraction Prog Biophys Biophys Chem 7: 255–318
Huxley AF, Simmons RM (1971) Proposed mechanism of force generation in striated muscle Nature 233: 533–538
Huxley AF, Simmons RM (1973) Mechanical transients and the origin of muscular force Cold Spring Harb Symp Quant Biol 37: 669–680
Huxley HE, Stewart A, Sosa H, Irving T (1994) X-ray diffraction measurements of the extensibility of actin and myosin filaments in contracting muscle Biophys J 67: 2411–2421
Jontes JD, Milligan RA (1997) Brush border myosin-I structure and ADP-dependent conformational changes revealed by cryoelectron microscopy and image analysis J Cell Biol 139: 683–693
Jontes JD, Ostap EM, Pollard TD, Milligan RA (1998) Three-dimensional structure of Acanthamoeba castellanii myosin-IB (MIB) determined by cryoelectron microscopy of decorated actin filaments J Cell Biol 141: 155–162
Jontes JD, Wilson-Kubalek EM, Milligan RA (1995) A 32 degree tail swing in brush border myosin I on ADP release Nature 378: 751–753
Kliche W, Fujita-Becker S, Kollmar M, Manstein DJ, Kull FJ (2001) Structure of a genetically engineered molecular motor Embo J 20: 40–46
Kohler D, Ruff C, Meyhofer E, Bahler M (2003) Different degrees of lever arm rotation control myosin step size J Cell Biol 161: 237–241
Köhler J, Winkler G, Schulte I, Scholz T, Mckenna W, Brenner B, Kraft T (2002) Mutation of the myosin converter domain alters cross-bridge elasticity Proc Natl Acad Sci USA 99: 3557–3562
Kollmar M, Durrwang U, Kliche W, Manstein DJ, Kull FJ (2002) Crystal structure of the motor domain of a class-I myosin Embo J 21: 2517–2525
Molloy JE, Burns JE, Kendrick-Jones J, Tregear RT, White DC (1995) Movement and force produced by a single myosin head Nature 378: 209–212
Pate E, White H, Cooke R (1993) Determination of the myosin step size from mechanical and kinetic data Proc Natl Acad Sci USA 90: 2451–2455
Piazzesi G, Lucii L, Lombardi V (2002) The size and the speed of the working stroke of muscle myosin and its dependence on the force J Physiol 545: 145–151
Purcell TJ, Morris C, Spudich JA, Sweeney HL (2002) Role of the lever arm in the processive stepping of myosin V Proc Natl Acad Sci USA 99: 14159–14164
Rayment I, Holden HM, Whittaker M, Yohn CB, Lorenz M, Holmes KC, Milligan RA (1993a) Structure of the actin–myosin complex and its implications for muscle contraction Science 261: 58–65
Rayment I, Rypniewski WR, Schmidt-Base K, Smith R, Tomchick DR, Benning MM, Winkelmann DA, Wesenberg G, Holden HM (1993b) Three-dimensional structure of myosin subfragment−1: a molecular motor Science 261: 50–58
Reconditi M, Linari M, Lucii L, Stewart A, Sun YB, Boesecke P, Narayanan T, Fischetti RF, Irving T, Piazzesi G, Irving M, Lombardi V (2004) The myosin motor in muscle generates a smaller and slower working stroke at higher load Nature 428: 578–581
Ruff C, Brenner B, And Meyhöfer E (2001a) The step size of myosin II is orientationally independent Biophys J 80: 78a
Ruff C, Furch M, Brenner B, Manstein DJ, Meyhofer E (2001b) Single-molecule tracking of myosins with genetically engineered amplifier domains Nat Struct Biol 8: 226–229
Schroder RR, Manstein DJ, Jahn W, Holden H, Rayment I, Holmes KC, Spudich JA (1993) Three-dimensional atomic model of F-actin decorated with Dictyostelium myosin S1 Nature 364: 171–174
Sleep J, Lewalle A, Smith DA. (2005) Displacements in the optical trap underestimate the myosin working stroke by a factor of two Biophys J 88: 3110 (Abstr.)
Smith CA, Rayment I (1996) X-ray structure of the magnesium(II).ADP.vanadate complex of the Dictyostelium discoideum myosin motor domain to 1.9 A resolution Biochemistry 35: 5404–5417
Steffen W, Smith D, Simmons R, Sleep J (2001) Mapping the actin filament with myosin Proc Natl Acad Sci USA 98: 14949–14954 Epub 12001 Dec 14944
Tanaka H, Homma K, Iwane AH, Katayama E, Ikebe R, Saito J, Yanagida T, Ikebe M (2002) The motor domain determines the large step of myosin-V Nature 415: 192–195
Tanaka H, Ishijima A, Honda M, Saito K, Yanagida T (1998) Orientation dependence of displacements by a single one-headed myosin relative to the actin filament Biophys J 75: 1886–1894
Uyeda TQ, Abramson PD, Spudich JA (1996) The neck region of the myosin motor domain acts as a lever arm to generate movement Proc Natl Acad Sci USA 93: 4459–4464
Veigel C, Coluccio LM, Jontes JD, Sparrow JC, Milligan RA, Molloy JE (1999) The motor protein myosin-I produces its working stroke in two steps Nature 398: 530–533
Wakabayashi K, Sugimoto Y, Tanaka H, Ueno Y, Takezawa Y, Amemiya Y (1994) X-ray diffraction evidence for the extensibility of actin and myosin filaments during muscle contraction [published erratum appears in Biophys J 1995 Mar;68(3): 1196–1197]. Biophys J 67: 2422–2435
Warshaw DM, Guilford WH, Freyzon Y, Krementsova E, Palmiter KA, Tyska MJ, Baker JE, Trybus KM (2000) The light chain binding domain of expressed smooth muscle heavy meromyosin acts as a mechanical lever J Biol Chem 275: 37167–37172
Yanagida T, Arata T, Oosawa F (1985) Sliding distance of actin filament induced by a myosin crossbridge during one ATP hydrolysis cycle Nature 316: 366–369
Acknowledgement
This work was supported by the DFG-Priority Program ‘Molecular Motors’ (SPP 1068) grants Br 849/21–1,2,3.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Brenner, B. The stroke size of myosins: a reevaluation. J Muscle Res Cell Motil 27, 173–187 (2006). https://doi.org/10.1007/s10974-006-9056-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10974-006-9056-7