Abstract
Structure folding dictates protein functions. Recent advances in structural analysis such as NMR and X-ray crystallography enabled to determine protein structures at high resolution, which helps understanding of protein folding and recognition mechanisms. Additionally, solution thermodynamic analysis provides useful information to visualize the real behavior of proteins because they largely fluctuate in solution. In this study, we analyzed the effect of a putative salt-bridge between His-137 and Asp-178 in the c-Myb DNA-binding domain on the thermodynamics of its folding and motion. The minimum unit for specific DNA-binding of c-Myb consists of the two repeats, R2 and R3. The residues, His-137 and Asp-178, are located in R2 and R3, respectively, and are spatial proximity, possibly forming a salt-bridge as revealed by a previous crystal structure analysis. D178N mutation caused slight changes in the tertiary structure of R2R3. The thermal stability of the D178N mutant in the absence of DNA was much lower than that of wild-type R2R3. The decrease in stability was due to unfavorable enthalpy change, partially compensated by the favorable entropy change. The largely decreased enthalpy change indicated that the disruption of the salt-bridge weakens the overall intramolecular interactions in the folded state. Additionally, the increased entropy change indicated that the dynamics of the folded state increase upon mutation. In contrast, the D178N mutation slightly affected the thermal stability of the DNA-bound state, indicating that the salt-bridge is required for proper folding of R2R3 in the DNA-free state.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
High-resolution 3D structures of proteins have been determined using X-ray crystallography, NMR, and electron microscopy. These techniques provide useful information on protein folding and recognition mechanisms. However, because most of the available structures are static or averaged, information on structural dynamics is needed to understand protein motions in solution. Proteins actually fluctuate in solution and express their functions as the result of conformational changes upon ligand binding. The degree of protein fluctuation varies extensively depending on their function. For example, inhibitor proteins are relatively rigid because they need to oppose hydrolyzation by proteases, whereas some DNA-binding proteins are intrinsically disordered probably because of the advantage to find the specific DNA sequence [1, 2]. Limited knowledge of protein structural dynamics caused difficulties in understanding protein structure–activity relationship and inhibited applications such as protein engineering and structure-based drug design. Thermodynamic analysis of proteins in solution could compensate the lack of information arising from static pictures of proteins. Thermodynamic studies unveiling the differences between a protein and its site-specific mutant could show the effect of the mutated site on the protein dynamics and intramolecular interactions such as hydrogen bonds and electrostatic interactions.
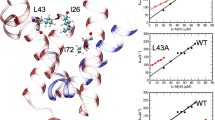
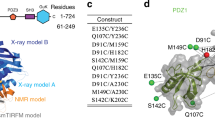
The transcriptional factor c-Myb consists of three functional domains involved in DNA-binding, transcriptional activation, and negative regulation [3]. The DNA-binding domain consists of three repeats, namely R1, R2, and R3, with R2R3 being the minimum unit of specific-DNA binding. We have analyzed the structural and physical properties of c-Myb R2R3 and have shown that its structural dynamics correlate well with its function [4–7]. At physiological temperature, at which c-Myb R2R3 binds to a specific DNA sequence in vivo, our NMR analysis showed that fluctuations were exceptionally large [8]. In the absence of DNA (i.e., DNA-free state), R2R3 exists as “semi-intrinsically disordered state.” The 3D structure of R2R3 in the absence or presence of DNA has been revealed by both NMR and crystallography [9–11]. In the crystal structure of R2R3 in the DNA-free state (PDB code 1GV2), which would be one of the stable structures, Asp-178 is spatially next to His-135, both of which form a surface-exposed salt-bridge with each other (Fig. 1). Upon binding to DNA, the relative positions of these two residues change, resulting in salt-bridge disruption. Based on the available structural information, the putative salt-bridge contributes to the “fragile” 3D structure of R2R3. In this study, we prepared a mutant of R2R3 (i.e., D178N), which lacks the salt-bridge, and analyzed its folding thermodynamics compared to those of wild-type R2R3. The results showed that the salt-bridge actually forms in solution and largely contributes to the 3D structure of R2R3 in the DNA-free state, but it barely contributes to that in the DNA-bound state.
3D structure of c-Myb R2R3 determined by X-ray crystallography (PDB code 1GV2) [11]. The side-chains of His-137 and Asp-178 are shown in sticks
Experimental
Sample preparations
Prior to the current mutational analyses, the Cys130 in R2, which is the only cysteine residue in the c-Myb R2R3 and is located at a position equivalent to an isoleucine in R3, was replaced with Ile, to facilitate protein purification and characterization. It has been shown that the affinity and the specificity of mutant C130I are similar to those of the wild-type [12, 13]. Therefore, the C130I protein was used as the standard R2R3 and is denoted as R2R3*. The mutant, C130I/D178N is simply denoted as D178N.
R2R3* and D178N were overexpressed in E. coli BL21 (DE3) and purified. The expression and purification methods were described previously [5, 14, 15] and slightly modified. Briefly, the freshly precultivated cells were inoculated into LB medium containing 100 μg mL−1 ampicillin and were grown at 37 °C. After the culture reached OD600 = 0.8, isopropyl-1-thio-β-d-galactopyranoside was added to a final concentration of 0.5 mM and incubated at 37 °C for 4 h. The harvested cells were lysed by the sonication, and the cell debris was removed by the centrifugation. The supernatant was purified by using ammonium sulfate precipitation, and phosphocellulose (P11, Whatman) and gel filtration (Sephacryl S-100, GE healthcare) columns. The purified proteins were concentrated, and the buffer was exchanged to phosphate-buffered saline (PBS, pH 7.4) or NMR sample buffer using Amicon Ultra-4 (Merck). The purity of each protein was determined to be about 95% by SDS/PAGE analysis. The protein concentrations were determined from UV absorption at 280 nm and were calculated by using the molar absorption coefficient of 3.7 × 104 M−1 cm−1 [13], determined by the amino acid composition analysis.
The 22-mer oligonucleotide CACCCTAACTGACACACATTCT, containing the Myb-binding site in the simian virus 40 enhancer sequence (MBS-I) was chemically synthesized and purified as described previously [13].
CD measurements
Far-UV (195-250 nm) and Near-UV (250-350 nm) CD spectra were recorded on a Jasco J-825 spectropolarimeter at 20 °C equipped with Peltier-type temperature control system. The spectra of R2R3* and D178 N were obtained in PBS (pH 7.4), using quartz cell with 0.1 cm path-length. The protein concentrations were 0.2 and 2.3 mg mL−1 for far- and near-UV, respectively. For the analysis of stability in the presence of DNA, MBS-I, the DNA concentration was equimolar to the protein, 16 μM. CD spectra were obtained using scanning speed of 20 nm min−1, a time response of 1 s, a bandwidth of 1 nm, and an average over 4 or 16 scans, for the far- and near-UV range, respectively.
The melting curves were recorded in temperature mode at 222 nm, from 20 to 80 °C with a heating rate of 1.0 °C min−1. The protein concentrations were 0.02 mg mL−1, and the quartz cell with 1 cm path-length was used. The analysis of the transition curves obtained by temperature-scanning CD measurements was performed on the basis of two-state transition model, as described previously [7, 14, 16].
The molar ellipticity data of temperature-scanning CD (Y) were fitted with the weighted-mean combination of two linear functions for the folded and the unfolded states, that is,
and
from Eqs. (1) and (2), as shown below:
Here, f D represents the molar fraction of the unfolded state and be expressed with the function of ΔG 01, the free energy of the transition:
ΔG 01 is also the function of the temperature involving three parameters, which are heat capacity change (ΔC p), transition temperature (T m), and van’t Hoff enthalpy change (ΔH vH) at T m:
In this study, we calculated the equation on the assumption that ΔC p is 0 for either R2R3* or D178N in the DNA-free and DNA-bound states. Substituting Eqs. (4) and (5) into (3), Y results in the function containing six parameters, that is A n, A d, B n, B d, T m, and ΔH vH, to be determined. The curve fitting was carried out by the nonlinear least-square method on Origin 5.0 software.
DSC measurements
DSC experiments were carried out on a Nano-DSC calorimeter (TA instruments), as described previously [14]. The data were collected by heating the solution from 5 to 90 °C at a rate of 1 °C min−1 and the sample concentrations were 2.0 mg mL−1 in PBS (pH 7.4). The outer buffer solution recovered from final exchanged experiment was used in the reference cell for each case. The data were analyzed using CpCalc software supplied by the manufacture, in order to subtract the baseline from heat capacity data and convert to molar heat capacity. The fitting analysis to obtain the denaturation temperature (T d) where the fraction folded is equal to that unfolded, heat capacity change (ΔC p), and calorimetric enthalpy change (ΔH cal) at T d were performed by the least-squares methods on Origin 5.0 software. The ΔC p was calculated from the difference in the C p values between unfolded and folded states at T d. ΔH cal was calculated by integrating the area in each heat capacity curve. The ΔH vH was calculated by the next equation for assuming the two-state transition.
where R is the gas constant. Using the ΔC p values with the ΔH cal and T d values, the thermodynamic parameters of unfolding as a function of temperature, ΔG (T), ΔH (T), and ΔS (T), could be calculated from the following equation, on the assumption that the ΔC p is constant within the temperature range analyzed, as described previously [7, 14, 15].
Results
Secondary and tertiary structure analysis
R2R3* and D178N were overexpressed in E. coli with similar yields. To investigate the effect of the mutation on the protein structure, far- and near-UV CD spectra were measured. The far-UV CD spectrum of D178N shows an α-helical-rich structure, similar to that of R2R3*, whereas the near-UV CD spectrum is slightly different especially at 290-310 nm (Fig. 2a, b). These results indicated that the D178N mutation affects the tertiary structure with little changes in the secondary structure.
Thermal stability analysis by CD
Thermal transition curves were monitored by measuring the intensity changes in the CD spectrum at 222 nm between 20 and 80 °C. The apparent fractions in the unfolded states were plotted as a function of temperature (Fig. 3a). The transition temperature (T m) and van’t Hoff enthalpy change (ΔH vH) were determined by fitting the obtained curves with a two-state transition model (Table 1). The T m value of D178N is 41.1 ± 0.3 °C, which is 7.4 ± 0.4 °C lower than that of R2R3*. The thermal stability of D178N considerably decreased, indicating that Asp-178 is important for the stability of the c-Myb R2R3 structure. The ΔH vH value of D178N is also smaller than that of R2R3*. The analysis of the thermal stabilities of R2R3* and D178N in the presence of the cognate-DNA MBS-I showed that they increased upon DNA binding. Interestingly, the T m value of the D178N-DNA complex is similar to that of R2R3*-DNA complex (Table 1), whereas the ΔH vH value of D178N is higher than that of R2R3*.
Thermal denaturation analysis. a The transition curves of R2R3* (black) and D178N (gray) in the absence (thick) or presence (thin) of cognate-DNA. The apparent fraction of folded protein by monitoring the CD values at 222 nm is shown as a function of the temperature. b The heat capacity curves of R2R3* (black) and D178N (gray)
Thermal stability analysis by DSC
To determine the thermodynamic parameters of the thermal unfolding in further detail, the thermal stability of D178N was also studied using DSC. Figure 3b shows the heat capacity curves of the protein analyzed at heating rate of 1.0 °C min−1. The DSC measurement showed that the thermal unfolding process was quantitatively reversible, similar to those of other R2R3 mutants [7, 14]. The T d, ΔH cal, and ΔC p values were determined using the two-state transition model, as described previously [14]. The data were well fitted on the assumption that the ratio ΔH cal/ΔH vH is 1 (Table 2). The T d value of D178N is 44.3 °C, which is lower than that of R2R3* and similar to the T m value determined using CD. The ΔH cal and ΔC p values of D178N are 172 kJ mol−1 and 7.79 kJ mol−1 K−1, respectively, both of which are smaller than those of R2R3*.
Discussion
The salt-bridge between His-137 in R2 and Asp-178 in R3 is important for the conformational stability of c-Myb R2R3. The disruption of the salt-bridge changes the cooperativity between R2 and R3, similar to when the linker connecting R2 and R3 is mutated [7]. Salt-bridge formation was expected by observing the crystal structure of the protein (PDB code 1GV2), and its presence was confirmed by the solution thermodynamic analysis of this study.
To investigate the thermodynamic origin of structural difference, the thermodynamic parameters of D178N at 50.3 °C, the denaturation temperature of R2R3*, were calculated using the ΔC p value, as described previously [14]. Within a narrow range of temperatures around the T d value, the errors on the calculated ΔG and ΔH values at the reference temperature are small, though the ΔC p values obtained have an error. The thermodynamic parameters are summarized in Table 3. The results clearly indicate that the thermal stability of D178N decreases due to unfavorable ΔH, which is partially compensated by favorable ΔS. The lack of a salt-bridge between His-137 in R2 and Asp-178 in R3 in the D178N mutant causes the unfavorable and favorable ΔH and ΔS, respectively. Assuming that the D178N mutation has little effects on the enthalpy and entropy levels in the unfolded state, the decreased ΔH of D178N is due to the increased enthalpy level in the folded state, relative to that of R2R3*. It should be noted that the small ΔH value of D178N (48 kJ mol−1) compared to that of R2R3* is much larger than that could derive from a salt-bridge, suggesting that the disruption of the salt-bridge destabilizes the overall R2R3 structure, similar to what happened in the case of the linker mutation in R2R3 [14]. In addition, the disruption of the salt-bridge increased the dynamics of the folded state, resulting in favorable ΔS value. Previous studies have shown that R2 is very dynamic, but it is stabilized by connecting to R3 [8]. In contrast to R2R3*, the results of well-fitting to the DSC data of D178N on the assumption that the ratio ΔH cal/ΔH vH is 1 indicate that the disruption of the salt-bridge weakens the cooperative interaction between R2 and R3 [14]. Although the ΔH vH determined by the fitting procedure for the change in secondary structure might be different from the accurate ΔH (Fig. 3a), it still suggests the existence of cooperative interaction. Taken together, the salt-bridge between His-137 in R2 and Asp-178 in R3 contributes to the fine-tuned R2R3 architecture.
The thermal stability of D178N was significantly increased by the presence of cognate-DNA. As described in Table S1, the differences in the T m values were determined. The ΔT m (bound, D178N) is approximately twice larger than ΔT m (bound, WT), indicating that the conformational stability of D178N is remarkably affected by DNA binding. Likewise, ΔT m (bound, mut) is significantly smaller than ΔT m (R2R3, mut), and the sum of ΔT m (bound, WT) and ΔT m (R2R3, mut) is nearly equal to ΔT m (bound, D178N). These results suggest that the salt-bridge between His-137 in R2 and Asp-178 in R3 only occurs in the DNA-free state (Fig. 4), which is confirmed by the NMR and crystal structures of R2R3 bound to DNA [9, 11]. Although Asp-178 is not directly involved in DNA recognition, its positive charge would contribute to the overall electrostatic interaction with DNA. The unique dynamic properties of R2R3, which result in fine-tuned folding architecture to which the salt-bridge largely contributes, are required for its specific DNA-binding function [8]. R2R3 largely fluctuates like a “semi-intrinsically disordered” protein to find the specific binding site on the target DNA. When R2R3 binds to the specific-site, it is stabilized by inter-molecular interactions with DNA, resulting in the disruption of the salt-bridge between His-137 and Asp-178.
The DNA-binding domains of Myb-family or Myb-like proteins are classified into two groups: One is composed of single repeats (group 1); the other, two repeats that bind DNA cooperatively (group 2). Group 1 includes many telomere-binding proteins such as hTRF1, hTFR2, TRFL, TBP, and YlTay1 [17–22]. These proteins have no aspartic acid residues corresponding to Asp-178 in c-Myb R3. Instead, group 2 includes a variety of DNA-binding proteins containing helix-turn-helix motifs such as A- and B-Myb, v-Myb, tvMyb1-3, and VvMyb series proteins, which have highly conserved aspartic acid residues corresponding to Asp-178 in c-Myb R3 [11, 23–27]. The solution NMR structure of tvMyb1 showed that several distinctive salt-bridges formed between the two repeats contributed to the stabilization of the relative orientation of the repeats themselves, resulting in high stability toward thermal denaturation [24]. In this study, we showed that Asp-178 largely contributes to the stability of R2R3 due to salt-bridge formation, which seems to be the only interaction between the two repeats based on the crystal structure [11].
Conclusions
The minimum DNA-binding unit of c-Myb, R2R3, largely fluctuates in solution. The crystal structure of R2R3 has shown that Asp-178 in R3 is spatially next to His-135 in R3, both of which form a surface-exposed salt-bridge with each other. The present mutational and thermodynamic analyses showed that the thermal stability of D178N mutant in the absence of DNA considerably decreased, indicating that Asp-178 is involved in the salt-bridge formation with His-135. The inter-repeat salt-bridge maintains the “semi-intrinsically disordered state” of R2R3, allowing specific DNA-binding. The large fluctuations of R2R3 are crucial for in vivo specific-site recognition of c-Myb.
References
Oda M, Tamura A, Kanaori K, Kojima S, Miura K, Momma K, Tonomura B, Akasaka K. Functional tolerance of Streptomyces subtilisin inhibitor toward conformational and stability changes caused by single-point mutations in the hydrophobic core. J Biochem. 2002;132:991–5.
Dyson HJ. Roles of intrinsic disorder in protein-nucleic acid interactions. Mol Biosyst. 2012;8:97–104.
Ness SA. Myb binding proteins: regulators and cohorts in transformation. Oncogene. 1999;18:3039–46.
Oda M, Furukawa K, Ogata K, Sarai A, Ishii S, Nishimura Y, Nakamura H. Identification of indispensable residues for specific DNA-binding in the imperfect tandem repeats of c-Myb R2R3. Protein Eng. 1997;10:1407–14.
Oda M, Furukawa K, Ogata K, Sarai A, Nakamura H. Thermodynamics of specific and non-specific DNA binding by the c-Myb DNA-binding domain. J Mol Biol. 1998;276:571–90.
Oda M, Furukawa K, Sarai A, Nakamura H. Construction of an artificial tandem protein of the c-Myb DNA-binding domain and analysis of its DNA binding specificity. Biochem Biophys Res Commun. 1999;262:94–7.
Inaba S, Fukada H, Oda M. Thermodynamic effects of a linker region between two repeats of a protein, c-Myb R2R3, on its stability and structural dynamics. J Therm Anal Calorim. 2016;123:1763–7.
Inaba S, Maeno A, Sakurai K, Puthenpurackal SN, Ikegami T, Akasaka K, Oda M. Functional conformer of c-Myb DNA-binding domain revealed by variable temperature studies. FEBS J. 2015;282:4497–514.
Ogata K, Morikawa S, Nakamura H, Sekikawa A, Inoue T, Kanai H, Sarai A, Ishii S, Nishimura Y. Solution structure of a specific DNA complex of the Myb DNA-binding domain with cooperative recognition helices. Cell. 1994;79:639–48.
Ogata K, Morikawa S, Nakamura H, Hojo H, Yoshimura S, Zhang R, Aimoto S, Ametani Y, Hirata Z, Sarai A, Ishii S, Nishimura Y. Comparison of the free and DNA-complexed forms of the DNA-binding domain from c-Myb. Nat Struct Biol. 1995;2:309–20.
Tahirov TH, Sato K, Ichikawa-Iwata E, Sasaki M, Inoue-Bungo T, Shiina M, Kimura K, Takata S, Fujikawa A, Morii H, Kumasaka T, Yamamoto M, Ishii S, Ogata K. Mechanism of c-Myb-C/EBPβ cooperation from separated sites on a promoter. Cell. 2002;108:57–70.
Myrset AH, Bostad A, Jamin N, Lirsac PN, Toma F, Gabrielsen OS. DNA and redox state induced conformational changes in the DNA-binding domain of the Myb oncoprotein. EMBO J. 1993;12:4625–33.
Oda M, Furukawa K, Ogata K, Sarai A, Ishii S, Nishimura Y, Nakamura H. Investigation of the pyrimidine preference by the c-Myb DNA-binding domain at the initial base of the consensus sequence. J Biol Chem. 1997;272:17966–71.
Inaba S, Fukada H, Ikegami T, Oda M. Thermodynamic effects of multiple protein conformations on stability and DNA binding. Arch Biochem Biophys. 2013;537:225–32.
Inaba S, Fukada H, Oda M. Folding thermodynamics of c-Myb DNA-binding domain in correlation with its α-helical contents. Int J Biol Macromol. 2016;82:725–32.
Morii H, Uedaira H, Ogata K, Ishii S, Sarai A. Shape and energetic of a cavity in c-Myb probed by natural and non-natural amino acid mutations. J Mol Biol. 1999;292:909–20.
Nishikawa T, Okamura H, Nagadoi A, Konig P, Rhodes D, Nishimura Y. Solution structure of a telomeric DNA complex of human TRF1. Structure. 2001;9:1237–51.
Karamysheva ZN, Surovtseva YV, Vespa L, Shakirov EV, Shippen DE. A C-terminal Myb extension domain defines a novel family of double-strand telomeric DNA-binding proteins in Arabidopsis. J Biol Chem. 2004;279:47799–807.
Court R, Chapman L, Fairall L, Phodes D. How the human telomeric protein TRF1 and TRF2 recognize telomeric DNA: a view from high-resolution crystal structure. EMBO Rep. 2005;6:39–45.
Hanaoka S, Nagadoi A, Nishimura Y. Comparison between TRF2 and TRF1 of their telomeric DNA-bound structures and DNA-binding activities. Protein Sci. 2005;14:119–30.
Kramara J, Willcox S, Gunisova S, Kinsky S, Neosek J, Griffith JD, Tomoska L. Tay1 protein, a novel telomere binding factor from Yarrowia lipolytica. J Biol Chem. 2010;285:38078–92.
Visacka K, Hofr C, Willcox S, Necasova I, Pavloskova J, Sepsiova R, Wimmerova M, Simonicova L, Nosek J, Fajkus J, Griffith JD, Tomoska L. Synergism of the two Myb domains of Tay1 protein results in high affinity binding to telomeres. J Biol Chem. 2012;287:32206–15.
MoInotosh PB, Frenkiel TA, Wollborn U, McCormick JE, Klempnauer K-H, Feeney J, Carr MD. Solution structure of the B-Myb DNA-binding domain: a possible role for conformational instability of the protein in DNA binding and control of gene expression. Biochemistry. 1998;37:9619–29.
Lou Y-C, Wei S-Y, Rajasekaran M, Chuo C-C, Hsu H-M, Tai J-H, Chen C. NMR structural analysis of DNA recognition by a novel Myb1 DNA-binding domain in the protozoan parasite Trichomonas vaginalis. Nucleic Acids Res. 2009;37:2381–94.
Hichri I, Deluc L, Barrieu F, Bogs J, Mahjoub A, Regad F, Gallois B, Granier T, Trossat-Magnin C, Gomès E, Lauvergeat V. A single amino acid change within the R2 domain of the VvMYB5b transcription factor modulates affinity for protein partners and target promoters selectivity. BMC Plant Biol. 2011;11:117.
Jiang I, Tsai C-K, Chen S-C, Wang S-H, Amiraslanov I, Chang C-F, Wu W-J, Tai J-H, Liaw Y-C, Huang TH. Molecular basis of the recognition of the ap65-1 gene transcription promoter elements by a Myb protein from the protozoan parasite Trichomonas vaginalis. Nucleic Acids Res. 2011;39:8992–9008.
Wei S-Y, Lou Y-C, Tsai J-Y, Ho M-R, Chou C-C, Rajasekaran M, Hsu H-M, Tai J-H, Hsiao C-D, Chen C. Structure of the Trichomonas vaginalis Myb3 DNA-binding domain bound to a promoter sequence reveals a unique C-terminal β-hairpin conformation. Nucleic Acids Res. 2012;40:449–60.
Acknowledgements
This work was supported by Grant-in-Aid for Japan Society for the Promotion of Science Fellow (Grant No. 15J03567). The analyses of this study were performed in part at the Okinawa Institute of Science and Technology Graduate University (OIST); MO is an OIST visiting researcher.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Inaba, S., Fukada, H. & Oda, M. Effect of a salt-bridge between inter-repeats on the 3D structure of the c-Myb DNA-binding domain revealed by thermodynamic analysis. J Therm Anal Calorim 131, 335–341 (2018). https://doi.org/10.1007/s10973-017-6524-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10973-017-6524-9