Abstract
Under drug selection pressure, emerging mutations render HIV-1 protease drug resistant, leading to the therapy failure in anti-HIV treatment. It is known that nine substrate cleavage site peptides bind to wild type (WT) HIV-1 protease in a conserved pattern. However, how the multidrug-resistant (MDR) HIV-1 protease binds to the substrate cleavage site peptides is yet to be determined. MDR769 HIV-1 protease (resistant mutations at residues 10, 36, 46, 54, 62, 63, 71, 82, 84, and 90) was selected for present study to understand the binding to its natural substrates. MDR769 HIV-1 protease was co-crystallized with nine substrate cleavage site hepta-peptides. Crystallographic studies show that MDR769 HIV-1 protease has an expanded substrate envelope with wide open flaps. Furthermore, ligand binding energy calculations indicate weaker binding in MDR769 HIV-1 protease-substrate complexes. These results help in designing the next generation of HIV-1 protease inhibitors by targeting the MDR HIV-1 protease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
HIV has been a major infectious pathogen since it was discovered in the early 1980s’. In 2009 alone, an estimated 2.6 million new HIV infections occurred worldwide, while about 33.3 million people were living with HIV. In 2009, the global AIDS deaths climbed to 1.8 million (UNAIDS report on the global AIDS epidemic 2010, http://www.unaids.org/globalreport/Global_report.htm).
Highly active antiretroviral therapy (HAART) makes possible the inhibition of in vivo HIV replication to delay the onset of AIDS symptoms [3, 4, 26, 27], while a cure or vaccine of HIV infection is yet unavailable [23, 35]. The usual HAART regimen combines three or more drugs from different classes, such as two nucleoside reverse transcriptase inhibitors (NRTI) and one non-nucleoside reverse transcriptase inhibitor (NNRTI) or protease inhibitor (PI). However, the emerging drug resistance, due to the lack of proofreading ability of the HIV-1 reverse transcriptase [32, 36, 42], requires the development of new and potent drugs to overcome the resistance problem [7, 8, 11, 12].
HIV-1 protease (HIV-1 PR) is an aspartic protease that cleaves newly synthesized HIV-1 polyproteins at nine major cleavage sites to generate the mature protein components of an infectious HIV-1 virion. The HIV-1 gag gene codes for the structural proteins: matrix protein (MA), capsid protein (CA), and nucleocapsid protein (NC), while the gag-pol gene encodes both structural proteins (MA, CA, and NC) and enzymes: protease (PR), reverse transcriptase (RT), and integrase (IN). Without effective HIV-1 PR, HIV-1 virion remains uninfectious, hence making HIV-1 protease inhibitors the most potent anti-AIDS drugs and essential therapeutic components of HAART [4, 12].
Crystallographic studies of WT HIV-1 PR complexed with the natural cleavage site peptides uncover a consensus of binding pattern of the peptides in the protease active site cavity [29, 31]. Based on the wealth of structural information of WT HIV-1 protease substrate complexes, new HIV-1 protease inhibitors mimicking the natural substrates are designed. In addition, it is proposed that inhibitors fitting the substrate envelope are less susceptible to drug resistance [1, 5, 6, 25].
Nonetheless, there is no crystallographic information on the MDR HIV-1 protease substrate complexes. The questions remaining to be addressed are: Do the nine substrates bind to MDR HIV-1 protease active site with a similar conformation? What is the relative binding energy of MDR HIV-1 protease substrate complexes compared to WT HIV-1 protease substrate complexes? What does the substrate envelope look like in the MDR HIV-1 protease?
We chose MDR769 HIV-1 protease as our study model to investigate the MDR HIV-1 protease substrate complex structures. The inactive enzyme (D25N) was chosen to trap the uncleaved peptide in the active site cavity. The high resolution crystal structure of MDR769 HIV-1 protease was solved previously by our group, and it showed an expanded active site cavity with mutations at positions 10, 36, 46, 54, 62, 63, 71, 82, 84, 90 [22]. Nine hepta-peptides corresponding to natural cleavage sites P3 to P4′ were chemically synthesized and successfully co-crystallized with the inactive MDR769 HIV-1 protease. Based on the crystal structures refined to 1.6–2.0 Å resolutions, a multi-drug resistant substrate envelop was determined and the envelope was comparable to that of the WT HIV-1 protease despite a certain degree of variation. The MDR HIV-1 protease substrate envelope was expended relative to WT HIV-1 protease substrate envelope. Furthermore, substrate binding energy calculations showed reduced binding affinity of MDR769 HIV-1 protease to the substrates.
Our observations with the MDR769 HIV-1 protease substrate complexes were consistent with the previous conclusions that the active site cavity was expanded in MDR769 HIV-1 protease. These results suggest that a larger HIV-1 protease inhibitor is needed to inhibit MDR variant of HIV-1.
2 Materials and Methods
2.1 Preparation of Substrate Peptides
The nine substrate hepta-peptides shown in Table 1 were purchase from SynBioSci Corporation, Livermore, CA. All the hepta-peptides were purified by HPLC to purity higher than 98%. Peptide powder was dissolved in DMSO to prepare stock solution of 20 mM concentration and the samples were stored at −20 °C.
2.2 Protein Purification and Co-Crystallization
The MDR769 HIV-1 protease A82T was overexpressed by using a T7 promoter expression vector in conjunction with the E. coli host, BL21 (DE3). In brief, a fresh E. coli transformant of BL21(DE3) with the MDR769 plasmid was cultured in 5 mL LB medium containing 100 μg/mL ampicillin for 7 h, which was used to inoculate a 50 mL LB medium containing 100 μg/mL ampicillin and the culture was grown overnight. Later, two liters of LB medium with 100 μg/mL ampicillin was inoculated with 50 mL seed culture and bacterial growth was monitored by absorbance at A600. After 4–6 h of incubation with shaking at 37 °C, the A600 reached 0.5 and the cells were harvested at 10,000×g for 10 min at 4 °C using a Sorvall RC5B Plus centrifuge [40]. The MDR protease was isolated from inclusion bodies by using a series of buffered washes, followed by denaturation in 6 M urea. For purification of the unfolded protease, an anion exchange column with pH 8.6 (Q Sepharose, Amersham Biosciences) was used that allowed the protease to pass through, while the E. coli contaminants to remained bound to the column. The protease purity was monitored by Coomassie blue-stained SDS–PAGE [16] and the MDR769 HIV-1 protease was purified to greater than 95% homogeneity. The 6 M urea was removed to refold the protease by using a series of dialysis exchanges that were carried out at 4 °C. The first buffer consisted of 0.2 M sodium phosphate monobasic, 0.2 M sodium phosphate dibasic, 1.0 M urea, 0.2% βME, and 10% glycerol. The next step consisted of buffer exchange with 0.2 M sodium phosphate monobasic, 0.2 M sodium phosphate dibasic, 0.2% βME, and 10% glycerol. The last two buffer exchanges were with 10 mM sodium acetate, pH 5.0, 1 mM DTT and 10% glycerol. The protease was concentrated by Amicon to 2 or 3 mg/mL and stored in aliquots at −80 °C.
The substrate hepta-peptides were mixed with 2.5 mg/mL MDR 769 HIV-1 protease and diluted to 0.4 mM final concentration. The mixtures were incubated at 4 °C for 2 h, which were subject to centrifugation to remove the insoluble impurities. The hanging drop vapor diffusion method was used to form the bipyramidal crystals of the MDR 769 protease. Using a grid screen consisting of sodium chloride (0.7–1.4 M) and MES-HEPES buffer (pH 5.5–8.1), the HIV-1 protease substrate complex crystals formed overnight at 22 °C. Routinely, 0.2 mm crystals, in the longest dimension, were obtained after 14 days of incubation. In each well, there were two droplets, containing 1 μL of protease substrate mixture, 1 μL of reservoir solution and 2 μL of protease substrate mixture, 1 μL of reservoir solution, respectively, 07 mL of well solution.
2.3 Data Collection and Crystallographic Refinement
Protease crystals were dipped in 30% glucose for cryoprotection and flash frozen in liquid nitrogen. The diffraction data were collected at 1.00 Å wavelength at the Advanced Photon Source (APS) (LS-CAT 21), Argonne National Laboratory (Argonne, IL). Data were reduced with the program CrystalClear (CrystalClear: An Integrated Program for the Collection and Processing of Area Detector Data, Rigaku Corporation©, 1997–2002). The data statistics are shown in Table 2. In all cases the crystals belong to the same space group P4 1 . Molecular replacement was performed with Molrep-autoMR in CCP4 with the model previously solved in our lab [20].
Initial refinements were performed without substrate hepta-peptides using Refmac5 [24, 38]. The nine hepta-peptides were built into the difference electron density maps as the refinement proceeded in the COOT program [10]. The electron density maps contained two orientations of bound peptide to the MDR HIV-1 protease. The Gag-Pol peptide binding in two orientations was also reported earlier by Prabu-Jeyabalan et al. [31] for the wild type HIV-1 protease. The MDR HIV-1 protease complexes were modeled and refined with the hepta-peptides in two orientations. The two peptide orientations were related by a pseudo-2-fold symmetry and there was minimal asymmetry for the two orientations of the peptide as was the case earlier for the wild type HIV-1 protease peptide complexes. To carry out the substrate envelope analysis in an analogous way as reported for the HIV-1 wild type complexes, we deleted one hepta-peptide orientation from the model. In all nine crystal structures, the N-terminus of the hepta-peptide was oriented close to monomer A of the AB protease dimer to allow for systematic analysis of the substrate envelope.
Crystallographic waters were added with the ARP/wARP program [17]. The structures were refined in Refmac5 to 1.6–2.0 Å resolution. The final stereo-chemical parameters were checked using PROCHECK [39]. Images were generated in PyMol.
2.4 Analysis
PISA Server was used to calculate the ligand binding energy in both MDR and WT HIV-1 protease substrate complexes [15]. Dissociation constants were calculated based on the formula ΔGint = RTlnKd, where the ΔGint is the intrinsic binding energy as defined by PISA. Hydrogen bonds with the bound hepta-peptides were analyzed with PISA server. Conserved hydrogen bonds within protease were identified by with the program Ligplot [41]. Temperature factor analysis was performed with the CCP4 program Temperature Factor Analysis. Protease substrate complexes were superimposed based on protease residues 1–99 Cα and the RMSD of Cα was analyzed with the CCP4 program Superpose Molecules [14]. Solvent accessible area was calculated with the CCP4 program Accessible Surface Areas. All structures were visualized using PyMol.
3 Results
3.1 Determination of Crystal Structures of MDR769 HIV-1 Protease-Substrate Complexes and the Calculation of Ligand Binding Energy
We report the crystal structures of nine MDR769 HIV-1 protease substrate complexes. The results support our working hypothesis that the MDR769 HIV-1 protease with wide open flaps binds weaker (20–162,000 times) to the substrates. In addition, the expanded MDR HIV-1 protease substrate envelope requires larger inhibitor to overcome drug resistance.
The inactive MDR769 HIV-1 protease is co-crystallized with nine hepta-peptides representing the nine natural substrate cleavage site sequences in the Gag and Gag-Pol polyproteins. The sequences and names of the hepta-peptides are shown in Table 1. The hepta-peptides are named based on proteins released after cleavage: matrix (MA), capsid (CA), nucleocapsid (NC), trans-frame peptide (TF), protease (PR), reverse transcriptase (RT), RNase H (RH), and integrase (IN).
The structures of the complexes are solved with crystallographic statistics shown in Table 2. These complexes crystallize in space group P4 1 , in which each crystallographic asymmetric unit contains one biologically relevant HIV-1 protease dimer complexed with ligands. The structures are refined to 1.6–2.0 Å.
The ligand binding energy is calculated for all the nine MDR769 HIV-1 protease substrate complexes by using the PISA server [15], and is compared to that of WT HIV-1 protease substrate complexes. Based on the calculated binding energy from crystallographic structure, dissociation constant (Kd) of each substrate cleavage site peptides against MDR and WT HIV-1 proteases is calculated and shown in Table 3. The contribution of hydrogen bonds to the binding free energy and Kd is not considered since there is a high degree of variation in the strength of the various hydrogen bonds based on the distance and angle of the hydrogen bonds.
3.2 The Substrates Bind Weaker to the MDR Protease Relative to the WT Protease, Despite Similar Ligand Conformation as That in WT Protease
All the nine hepta-peptides are bound to the MDR HIV-1 protease active site cavity in an extended conformation. The structure of the MDR HIV-1 protease MA/CA peptide complex, shown in Fig. 1, is representative for all the nine complexes. The hepta-peptides establish extensive contacts with both protease monomers; however, the contacts between monomers are not equally distributed. The asymmetric nature of hepta-peptides breaks the symmetry of protease dimer in complexes by forming more contacts with one monomer than the other. Structural analysis shows that in complexes MA/CA, NC/p1, p1/p6, and TF/PR, RMSD of Cα at residue 79 is significantly different between the two monomers (Liu et al., data not shown). In RH/IN complex the significant difference is present at residues 67 and 68 rather than at residue 79. However, there is no big RMSD variation between monomers in complexes CA/p2, PR/RT and RT/RH (Liu et al., data not shown).
Structure of the MDR 769 HIV-1 protease MA/CA complex. The two protease monomers are shown in ribbon rendering. The MA/CA substrate peptide is shown by stick model. The substrate shows an extended conformation of the MA/CA peptide binding to the MDR HIV-1 protease dimer, which is representative to all the nine complex structures
Although the active site cavity of MDR 769 HIV-1 protease is expanded leaving enough space for an alternate conformation, all the nine hepta-peptides are located at the bottom of the active site cavity with flexible amino and carboxy termini. P1 and P1′ reside between Asn 25/25′. Seven out of nine hepta-peptides (except for p1/p6 and RT/RH), point their carbonyl oxygen of P1 (P1 O) in between Asn25/25′, bridging the two “catalytic site” residues. In the p1/p6 complex, the P1 O forms a hydrogen bond with only Asn25 ND2 atom. In contrast, in complex RT/RH, a water molecule bridges the connection between P1 O and Asn25′ ND2.
The substrates bind weaker to the MDR HIV-1 protease relative to the WT HIV-1 protease. Based on the ligand binding energy (ΔG) calculated by PISA server, the binding energy is reduced dramatically (34–78%) in MDR HIV-1 protease substrate complexes compared to that in WT HIV-1 protease substrate complexes (Table 3). Correspondingly, Kd of MDR HIV-1 protease substrate complexes is much higher (20–126,000 folds), which indicates lower binding affinity of the substrate hepta-peptides (Table 3).
3.3 Conserved Substrate Conformation in MDR HIV-1 Protease Substrate Complexes Gives Rise to an Unevenly Expanded Substrate Envelope Compared to That of WT HIV-1 Protease Substrate
Substrate envelope is defined as the overall space occupied by the nine substrate peptides in the superimposed nine complexes based on Cα of the protease dimers. As a result, this substrate envelope demonstrates the maximal flexibility of substrates inside of the activity site cavity of the MDR HIV-1 protease.
The substrate peptides are taken out from the MDR HIV-1 protease active site cavity upon dimer superposition and listed individually in Fig. 2. Moreover, these substrate peptides are shown in superposition mode based on the protease dimer Cα to display the spatial overlay in the MDR HIV-1 protease active site cavity (Fig. 3a). However, when compared to those in WT HIV-1 protease, these cleavage site peptides are more flexible in the active site cavity according to the B factor calculation (data not shown). The flexibility may be partially due to the expanded MDR HIV-1 protease active site cavity. Detailed investigation shows that the enlargement of the substrate envelope is not uniform through the different sites. For example, P1/P1′ is expanded by 12%, while the overall expansion of the substrate envelope is 7%. The comparison of the WT and MDR HIV-1 protease substrate envelope is shown in Fig. 3b–d.
Orientation of nine substrate peptides in the MDR 769 HIV-1 protease active site cavity. The nine MDR HIV-1 protease substrate peptide complexes are superposed according to the Cα of the protease dimer structure. The substrate peptides are displayed individually after the dimer superposition, showing conserved conformation of these peptides in MDR HIV-1 protease active site cavity
Panel A Superposition of the nine substrate peptides in the MDR HIV-1 protease active site cavity. The superposition shows not only the conserved conformation, but also the conserved position of these substrate peptides in the MDR HIV-1 protease active site cavity. Panel B The substrate envelope of the MDR HIV-1 protease. The transparent shading represents the MDR HIV-1 protease substrate envelope. The stick model inside the substrate envelope is the MA/CA substrate peptide which is representative of all the substrate peptides studied here. Panel C Substrate envelope of the WT HIV-1 protease. To facilitate direct comparison of the WT and MDR HIV-1 protease complexes, the nano-peptides from the WT studies are truncated to P3 to P4′ hepta-peptides used in this study. Panel D Superposition of the WT and MDR HIV-1 protease substrate envelope. The protruding substrate envelope indicates the expanded substrate envelope of the MDR HIV-1 protease
3.4 Altered MDR Protease Binding Pockets Result in Weaker Binding of the Substrates
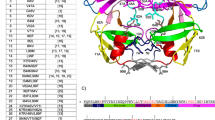
Detailed investigation of the interaction between protease and nine hepta-peptides reveals altered binding pockets compared to WT HIV-1 protease. Five conserved binding pockets are found in MDR HIV-1 protease, namely S1/S3, S2, S1′/S3′, S2′, and S4′ as shown in Fig. 4. Fewer residues relative to WT HIV-1 protease contribute to the pockets S1/S3, S2, S1′/S3′, S2′, meanwhile a conserved S4′ pocket, missing in WT HIV-1 protease, is found in the MDR HIV-1 protease. The reduced number of amino acid residues lining the binding pocket lead to weaker binding between the active site cavity and the nine substrates. First, the S1/S3 binding pocket involves Gly27′, Asp 29′, Arg8, Thr82, and Asn25 in at least four complexes. Leu23 is involved in NC/p1, MA/CA, and RT/IN complexes, whereas Val84 is involved in NC/p1, RT/RH, and CA/p2 complexes. Side chains of Gly27′, Asp29′, Arg8, Asp25, and Val84 adopt the same conformation in all complexes; whereas side chains of Thr82 and Leu23 have variable conformations. Second, in the S1′/S3′ pocket, Leu23′, Asn25′, Val84′, Arg8, Gly27, and Asp29 are present in at least four complexes; Thr82′ participates in only three complexes p2/NC, RT/TH, and NC/p1. Surprisingly, Gly48 in the flap region is involved in the S1′/S3′ binding pocket in complexes RT/RH, TF/PR, and MA/CA. Except for Thr82′, all residues in S1′/S3′ binding pocket are conserved in the same conformation. Third, side chains from Ala28′, Asp29′ and Asp30′ constitute the binding pocket of the S2 site. Fourth, side chains from Gly27, Ala28, Asp29, and Asp30 constitute the binding pocket at the S2′ site. All residues involved in S2/S2′ binding are highly conserved in conformation, forming a rigid binding pocket. The S2 binding pocket shows fewer contacts relative to that in the S2′ pocket. Finally, the S4′ binding pocket contains two segments: one is flexible and the other is rigid. The flexible segment consists of residues Lys45, Leu46, Ile47 and Gly48. The rigid segment is comprised of Asp29 and Asp30.
3.5 Limited Induced Fit of the MDR Protease Results in Weaker Binding of the Substrates
There is a limited induced fit in the MDR HIV-1 protease when compared to the WT HIV-1 protease substrate complexes, which may be a key reason to the weaker ligand binding. First, flap distance, measured from Cα Ile 50 to Cα Ile 50′, is a good indicator of binding. Analysis of the flap distance shows variation among the nine complexes, ranging from 9.9 to 12.3 Å. When compared to flap distances in WT HIV-1 protease substrate complexes, which range from 5.9 to 6.3 Å [29–31, 37], the MDR HIV-1 substrate complexes are still wide open, i.e. 4–6 Å wider compared to the WT crystal structures. This partial induced fit may contribute to the weaker biding due to fewer contacts between the HIV-1 protease flap area and the substrates. However, when compared to the flap distance in uncomplexed MDR 769 HIV-1 protease (12.2 Å) [22], most complexes show significant reduction in flap distance, except for two complexes: CA/p2 and RH/IN, 11.7 and 12.3 Å, respectively (Table 3). Second, the dimer interface area is also reduced in the MDR HIV-1 protease substrate complexes (1,469.7–1,485.5 Å2) relative to that of WT complexes (1,680.3–1,768.8 Å2) [29–31, 37]. This induced fit in the MDR HIV-1 protease weakens the protease dimer, which may lead to the relative instability of the MDR HIV-1 protease substrate complexes. Surprisingly, the binding of substrate to MDR HIV-1 protease reduces the dimer interface (1,508.5 Å2 in uncomplexed MDR HIV-1 protease), whereas the substrate binding increases the dimer interface area in the WT HIV-1 protease (1,487.7 Å2 in uncomplexed WT HIV-1 protease). The factor that ligand binding decreases the stability of MDR HIV-1 protease makes it possible to develop new dimerization inhibitors. Third, one conserved high RMSD pattern is found after the substrates bind to MDR HIV-1 protease. All the nine complexes were superposed with an uncomplexed wide-open MDR769 HIV-1 protease structure, based on Cα. High RMSD regions are located at residues 15 and 16 (loop 2 between β strands 2, 3), residue 21 (β strand 3), residues 40–63 (flap), residues 66–68 (loop between β strands), residues 72 and 73 (loop between β strands), and residues 78–82 (80′ loop interacting with ligands), except for the RH/IN complex which shows high RMSDs at residues 15, 34, 65, and 66. The introduction of these high RMSD regions may contribute to the weaker binding.
3.6 Substrate Conformation Difference Between MDR and WT HIV-1 Protease Complexes
As mentioned before, the overall substrate conformation is conserved between the MDR and WT HIV-1 protease substrate complexes; however, some subtle variation in the substrate conformation makes the MDR HIV-1 protease substrate complexes less stable (Fig. 5). Details of these substrate conformation differences will be discussed below.
Compared to the side chains, both position and conformation of the substrate backbone are relatively conserved between the WT and MDR HIV-1 protease substrate complexes. The WT and MDR HIV-1 proteases complex with the same substrate peptides are superimposed onto each other according to Cα of the protease dimer. The deviation of corresponding Cα in substrate peptide is shown in Table 4. Except for the substrate peptide in complex RT/IN, the Cα shows more deviation in the C-terminus than in the N-terminus. In addition, the deviation of P1 O is shown in the last column of Table 4 of corresponding substrate peptides from both WT and MDR HIV-1 protease substrate complexes.
In contrast to backbone atoms, the side chain atoms are much more flexible in the MDR complexes. Although no apparent pattern is identified in side chain deviation, it is found that the deviation is larger at both termini.
The relative position of P1 O is totally different from the MDR HIV-1 substrate complexes to the WT HIV-1 protease substrate complexes. The distances between the P1 O atom to Asn25 and Asn25′ in both MDR and WT HIV-1 protease substrate complexes are listed in Table 5. In the MDR HIV-1 protease substrate complexes, the P1 O, Asn25 ND2 and Asn25′ ND2 constitute an isosceles triangle with equal distance between P1 O and Asn25/25′. The only exception to the observation is p1/p6, which is similar to substrates in the WT complexes with unequal distance to Asn25 and Asn25′. The symmetric conformation in the MDR HIV-1 protease substrate complexes may indicate that a symmetric inhibitor might be more suitable to overcome the drug resistance problem.
4 Discussion
4.1 The Hepta-Peptides Bind with Reduced Binding Energy and Increased Kd Values
Although the nine hepta-peptides adopt a similar conformation to bind to MDR HIV-1 protease, the binding energy reveals varied affinities of the hepta-peptides to the protease (Table 3). The binding energy is calculated by using the PISA server [15]. Based on the semi-empirical calculation used in PISA server, the RH/IN complex holds the strongest binding energy between the ligand and protease, although it has the longest inter-flap distance. When compared to the binding energy derived from WT HIV-1 protease substrate complexes, the binding energy for MDR HIV-1 protease substrate complexes is by 34–78%. The Kd value is calculated for both MDR and WT HIV-1 protease substrate complexes, with 20–162,000 fold increase in MDR HIV-1 protease substrate complexes. Among all MDR HIV-1 protease substrate complexes, CA/p2 has the largest decrease in binding energy relative to WT HIV-1 protease CA/p2 complex (22%) and a 162,000 times larger Kd (Table 3).
4.2 The MDR HIV-1 Protease Substrate Envelope Is Enlarged by 12% at P1/P1′ Site
Hepta-peptides bound to MDR769 HIV-1 protease yield a larger substrate envelope relative to that in the WT HIV-1 protease substrate complexes. The volume of substrate envelope determines the size of HIV-1 protease inhibitor required to overcome the drug resistance issue. The substrate envelope is enlarged overall by 7% in MDR HIV-1 protease substrate complexes. The superposition of the MDR with WT HIV-1 protease substrate envelope is shown in Fig. 3d. Further examination shows that the enlargement of MDR HIV-1 protease substrate envelope is not uniform at all binding sites. The MDR HIV-1 protease substrate envelope at P1/P1′ site is enlarged by 12% compared to the P1/P1′ site in the WT HIV-1 protease substrate envelope, while together the P1/P1′ and P2/P2′ increase in surface area by 9.54%. This result suggests that a larger P1/P1′ site is needed to overcome the multi-drug resistance.
4.3 Compensatory Mutations May Restore Enzymatic Activity
With the reduced protease catalytic activity restraining the HIV-1 maturation and replication, HIV-1 virion develops compensatory mutations in and outside the substrate cleavage sites (i.e. mutations at p1/p6 and NC/p1) to restore the protease catalytic activity [2, 9, 30, 33, 43]. Work is currently in progress with the p1/p6and NC/p1 mutant substrate complexes to understand their structural and functional impact on MDR769 HIV-1 protease. New structural information of these two protease-substrate complexes may reveal additional information about the MDR HIV-1 protease substrate envelope and replication of the MDR HIV-1 viron.
4.4 Mutations in CA/p2 Lead to Drug Resistance to the Maturation Inhibitor PA457, Which Is Represented in the Complexes
PA457 (Bevirimat) targeting the CA/p2 cleavage site, blocking the cleavage of CA/p2 by HIV-1 protease, is currently under phase II clinical trials to address the drug resistance problem [13, 18, 19, 21, 34]. It is found that the introduction of single residue mutation to the flanking sequence of CA/p2 cleavage site results in a drug resistant phenotype, which requires no mutations from other parts of the virus, including protease [28, 44, 46]. Among these single mutations, Ala to Phe at P1′ develops drug resistance to PA457 [18, 45]. In our complex structure of CA/p2 with MDR769 HIV-1 protease, at the P1′ position, there is a Phe instead of Ala, and consequently molecular dynamics calculation may elucidate the mechanism of drug resistance to PA457.
4.5 The MDR Protease Forms a Weaker Dimer Compared to the WT Protease Dimer Raising the Possibility of Designing Inhibitors That Disrupt the Dimer Interface
Unlike the WT protease which substrate binding increase dimerization interfaces, MDR HIV-1 protease has a reduced dimerization interface after substrate binding. The dimerization interface area is 16% less in MDR HIV-1 protease substrate complexes than that of WT HIV-1 protease substrate complexes, while the dimerization interface area is almost identical between apo MDR HIV-1 protease and WT HIV-1 protease (Table 3). This partial disrupted dimerization interface in MDR HIV-1 protease substrate complexes raises the possibility of designing inhibitors that disrupt the dimer interface during substrate binding.
4.6 The Catalytic Activity of MDR769 HIV-1 Protease May be Reduced
The catalytic activity may be reduced in MDR 769 HIV-1 protease. Although no direct functional data are available to assess MDR 769 HIV-1 protease catalytic activity, current structural studies, theoretical binding energy calculations, and molecular dynamics simulations indicate the reduced efficiency in peptide cleavage of MDR 769 HIV-1 protease. First, the flap water molecule positioned between Ile50 and Ile50′, bridging the protease flaps and substrate peptides, is missing, and this flap water is crucial in substrate cleavage (Liu et al. unpublished results). Therefore, the MDR HIV-1 protease may adopt other mechanisms to achieve the cleavage of the substrate. As a consequence of the expanded protease active site cavity, both the distance between the two flaps and the distance between flaps and the substrates are considerably increased in the MDR769 HIV-1 protease. As a result, the MDR HIV-1 protease does not bind the substrates tightly in optimal position, hence reducing catalytic cleavage. Second, calculations of substrate binding energies and Kd confirm the results of structural studies of MDR HIV-1 protease substrate complexes. There is a significant decrease of binding energy (34–78% loss) and an increase of Kd (20–162,000 fold gains) in MDR HIV-1 protease substrate complexes. Without tight binding, the substrates flop in and out the active site cavity more frequently, reducing the protease catalytic activity. Third, molecular dynamics calculations with a WT HIV-1 protease substrate complex show a rigid body movement of substrate after a 5 ns simulation, while this movement required for efficient cleavage is missing in MDR substrate complexes molecular dynamics simulations (Yedidi et al. unpublished results).
Despite all the reasoning pointing to the reduced catalytic activity of MDR HIV-1 protease, the Kcat itself may not necessarily be reduced. It is possible that the wide open nature of MDR HIV-1 protease flaps leads to a weaker interaction between the cleavage products and MDR HIV-1 protease. As a result, the release of the cleavage products may be faster in MDR HIV-1 protease substrate complexes compared to that in WT complexes, which may partially compensate for the reduced catalytic activity.
Abbreviations
- MDR:
-
Multi-drug resistance
- HIV:
-
Human immunodeficiency virus
- WT:
-
Wild type
- HAART:
-
Highly active antiretroviral therapy
- PR:
-
Protease
- MA:
-
Matrix
- CA:
-
Capsid
- NC:
-
Nucleocapsid
- RT:
-
Reverse transcriptase
- IN:
-
Integrase
- RH:
-
RNase H
- RMSD:
-
Root mean square deviation
References
Altman MD, Ali A, Reddy GS, Nalam MN, Anjum SG, Cao H, Chellappan S, Kairys V, Fernandes MX, Gilson MK, Schiffer CA, Rana TM, Tidor B (2008) J Am Chem Soc 130:6099–6113
Bally F, Martinez R, Peters S, Sudre P, Telenti A (2000) AIDS Res Hum Retroviruses 16:1209–1213
Barbaro G, Lucchini A, Barbarini G (2005) Minerva Cardioangiol 53:153–154
Bartlett JA, DeMasi R, Quinn J, Moxham C, Rousseau F (2001) AIDS 15:1369–1377
Chellappan S, Kairys V, Fernandes MX, Schiffer C, Gilson MK (2007) Proteins 68:561–567
Chellappan S, Kiran Kumar Reddy GS, Ali A, Nalam MN, Anjum SG, Cao H, Kairys V, Fernandes MX, Altman MD, Tidor B, Rana TM, Schiffer CA, Gilson MK (2007) Chem Biol Drug Des 69:298–313
Clavel F, Hance AJ (2004) N Engl J Med 350:1023–1035
Condra JH, Schleif WA, Blahy OM, Gabryelski LJ, Graham DJ, Quintero JC, Rhodes A, Robbins HL, Roth E, Shivaprakash M (1995) Nature 374:569–571
Croteau G, Doyon L, Thibeault D, McKercher G, Pilote L, Lamarre D (1997) J Virol 71:1089–1096
Emsley P, Cowtan K (2004) Acta Crystallogr D Biol Crystallogr 60:2126–2132
Grabar S, Pradier C, Le Corfec E, Lancar R, Allavena C, Bentata M, Berlureau P, Dupont C, Fabbro-Peray P, Poizot-Martin I, Costagliola D (2000) AIDS 14:141–149
Gulick RM, Mellors JW, Havlir D, Eron JJ, Meibohm A, Condra JH, Valentine FT, McMahon D, Gonzalez C, Jonas L, Emini EA, Chodakewitz JA, Isaacs R, Richman DD (2000) Ann Intern Med 133:35–39
Holzgrabe U (2004) Pharm Unserer Zeit 33:160
Krissinel E, Henrick K (2004) Acta Crystallogr D Biol Crystallogr 60:2256–2268
Krissinel E, Henrick K (2007) J Mol Biol 372:774–797
Laemmli UK (1970) Nature 227:680–685
Lamzin VS, Wilson KS (1993) Acta Crystallogr D Biol Crystallogr 49:129–147
Li F, Goila-Gaur R, Salzwedel K, Kilgore NR, Reddick M, Matallana C, Castillo A, Zoumplis D, Martin DE, Orenstein JM, Allaway GP, Freed EO, Wild CT (2003) Proc Natl Acad Sci USA 100:13555–13560
Li F, Zoumplis D, Matallana C, Kilgore NR, Reddick M, Yunus AS, Adamson CS, Salzwedel K, Martin DE, Allaway GP, Freed EO, Wild CT (2006) Virology 356:217–224
Logsdon BC, Vickrey JF, Martin P, Proteasa G, Koepke JI, Terlecky SR, Wawrzak Z, Winters MA, Merigan TC, Kovari LC (2004) J Virol 78:3123–3132
Martin DE, Blum R, Wilton J, Doto J, Galbraith H, Burgess GL, Smith PC, Ballow C (2007) Antimicrob Agents Chemother 51:3063–3066
Martin P, Vickrey JF, Proteasa G, Jimenez YL, Wawrzak Z, Winters MA, Merigan TC, Kovari LC (2005) Structure 13:1887–1895
McMichael AJ, Hanke T (2003) Nat Med 9:874–880
Murshudov GN, Vagin AA, Dodson EJ (1997) Acta Crystallogr D Biol Crystallogr 53:240–255
Nalam MN, Ali A, Altman MD, Reddy GS, Chellappan S, Kairys V, Ozen A, Cao H, Gilson MK, Tidor B, Rana TM, Schiffer CA (2010) J Virol 84:5368–5378
Natarajan V, Bosche M, Metcalf JA, Ward DJ, Lane HC, Kovacs JA (1999) Lancet 353:119–120
Palella FJ, Delaney KM, Moorman AC, Loveless MO, Fuhrer J, Satten GA, Aschman DJ, Holmberg SD (1998) N Engl J Med 338:853–860
Pettit SC, Everitt LE, Choudhury S, Dunn BM, Kaplan AH (2004) J Virol 78:8477–8485
Prabu-Jeyabalan M, King NM, Nalivaika EA, Heilek-Snyder G, Cammack N, Schiffer CA (2006) Antimicrob Agents Chemother 50:1518–1521
Prabu-Jeyabalan M, Nalivaika EA, King NM, Schiffer CA (2004) J Virol 78:12446–12454
Prabu-Jeyabalan M, Nalivaika EA, Romano K, Schiffer CA (2006) J Virol 80:3607–3616
Prabu-Jeyabalan M, Nalivaika E, Schiffer CA (2000) J Mol Biol 301:1207–1220
Prabu-Jeyabalan M, Nalivaika E, Schiffer CA (2002) Structure 10:369–381
Roberts JD, Preston BD, Johnston LA, Soni A, Loeb LA, Kunkel TA (1989) Mol Cell Biol 9:469–476
Robinson LH, Myers RE, Snowden BW, Tisdale M, Blair ED (2000) AIDS Res Hum Retroviruses 16:1149–1156
Schooley RT, Mellors JW (2007) J Infect Dis 195:770–772
Sukasem C, Churdboonchart V, Sukeepaisarncharoen W, Piroj W, Inwisai T, Tiensuwan M, Chantratita W (2008) Int J Antimicrob Agents 31:277–281
Takeuchi Y, Nagumo T, Hoshino H (1988) J Virol 62:3900–3902
Tie Y, Boross PI, Wang YF, Gaddis L, Liu F, Chen X, Tozser J, Harrison RW, Weber IT (2005) FEBS J 272:5265–5277
Vagin AA, Steiner RA, Lebedev AA, Potterton L, McNicholas S, Long F, Murshudov GN (2004) Acta Crystallogr D Biol Crystallogr 60:2184–2195
Vaguine AA, Richelle J, Wodak SJ (1999) Acta Crystallogr D Biol Crystallogr 55:191–205
Vickrey JF, Logsdon BC, Proteasa G, Palmer S, Winters MA, Merigan TC, Kovari LC (2003) Protein Expr Purif 28:165–172
Wallace AC, Laskowski RA, Thornton JM (1995) Protein Eng 8:127–134
Weber J, Grosse F (1989) Nucleic Acids Res 17:1379–1393
Zhang YM, Imamichi H, Imamichi T, Lane HC, Falloon J, Vasudevachari MB, Salzman NP (1997) J Virol 71:6662–6670
Zhou J, Chen CH, Aiken C (2004) Retrovirology 1:15
Acknowledgments
This research was supported by the National Institutes of Health grant AI65294 and a grant from the American Foundation for AIDS Research (106457-34-RGGN).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, Z., Wang, Y., Brunzelle, J. et al. Nine Crystal Structures Determine the Substrate Envelope of the MDR HIV-1 Protease. Protein J 30, 173–183 (2011). https://doi.org/10.1007/s10930-011-9316-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-011-9316-2