Abstract
Meningioma is the one of the most common type of brain tumor, it as arises from the meninges and encloses the spine and the brain inside the skull. It accounts for 30% of all types of brain tumor. Meningioma’s can occur in many parts of the brain and accordingly it is named. In this paper, a mixture model based classification of meningioma brain tumor using MRI image is developed. The proposed method consists of four stages. In the first stage, with respect to the cells’ boundary, it is necessary to further processing, which ensures the boundary of some cells is a discrete region. Mathematical Morphology brings a fancy result during the discrete processing. Accurate cancer cell nucleus segmentation is necessary for automated cytological image analysis. Thresholding is a crucial step in segmentation..An adaptive binarization technique is an important step for medical image analysis.Finally, a novel hybrid Fuzzy SVM is designed in the classification stage meningioma brain tumor. The tumor classification results of proposed feature extraction with SVM is 74.24%, MM with FSVM is 82.67% and MM with RBF is 62.71% and our proposed method MM with Hybrid SVM is 91.64%.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Our brain is a complex integrative network where information is continuously processed and shared between structurally and functionally interconnected regions. Functional magnetic resonance imaging (fMRI) indirectly measures such brain activities by detecting relative alterations in blood oxygen level. For the past three decades, most studies have been carried out focusing on localization of neural activities associated with a variety of cognitive, sensory and motor tasks. Recent developments in functional neuro imaging have shifted the research focus to investigating and interpreting functional interactions between different brain regions [1,2,3].
Meningioma is the most common and aggressive primary malignant central nervous system (CNS) tumors. Initially termed multiform, meningioma belongs to a group of heterogeneous tumors because of their significant morphological, cytological and molecular variability. Because of their frequency and severity, meningioma’s are a major therapeutic issue for neurosurgeons, neuro-oncologists, and radiation therapists. The diagnosis of a meningioma is a sensitive operation where the error can lead to critical results. CT,PET and MRI images are medical imaging modalities used by doctors and researchers to acquire a best visualization for soft tissues anatomy. The naming of the Meningioma Tumors is based on the location it occupies in the brain. The growth rate of Meningioma Tumors are veryslow and therefore often people suffering from Meningioma Tumors are unaware that they are having the tumor in the brain [4,5,6].
In many cases patients come to know about it due to sudden critical seizure and numbness in their hands or legs. Depending on the position of the tumor in the brain certain parts of the body are affected. For example, a person having Parasagittal Meningioma tumor in the left side of the brain will find difficulty on their right hand and right leg movement. They will feel numbness, shivering on their right side and will also find difficult in seeing. When tumor exist on the left side of the brain, part of the brain controlling emotions are also strained [7]. Anatomical magnetic resonance imaging is today the most suitable non-invasive method for fine-resolution observation of biological tissues, and in particular brain tissue. Its sensitivity to small tissue changes is in fact a valuable and powerful ally in the diagnosis of brain tumors. MRI provides digital images that provide the information needed to implement treatment and analysis algorithms to help diagnose brain tumors [8,9,10]. This challenge consists in precisely detecting the tumor zone. Brain tumor segmentation is a major and difficult task in surgical and treatment planning. Segmentation plays a relevant role in the process of medical image analysis and diagnostic assistance. The need to design an automatic segmentation algorithm is indisputable but remains a persistent challenge. In brain imaging, the difficulty of segmentation insists, given the structural complexity of MRI images and the often-insufficient contrast to extract the structure of interest, without any prior knowledge of its shape and location. Segmentation must naturally be reproducible, accurate and reliable.
The segmentation task becomes more complex, the extraction of useful features is a demanding task due to the complex structures of different tissues such as WM, GM and CSF and volume segmentation techniques operates in the spatial domain by classifying voxels. This is done directly instead of applying an intermediate transfer function to their data values. In principle it has a chance to replace the classification step. They are generally either too time-consuming for routine use because they require a massive amount of user interaction, e. g., iterating over all slices and marking the desired structure with high precision, mostly, they do not yield accurate results for volumes with low signal-to-noise ratio [11,12,13].
Brain tumor classification system
Segmentation plays a relevant role in the process of medical image analysis and diagnostic assistance. The need to design an automatic segmentation algorithm is indisputable but remains a persistent challenge. In brain imaging, the difficulty of segmentation insists, given the structural complexity of MRI images and the often-insufficient contrast to extract the structure of interest, without any prior knowledge of its shape and location. Segmentation must naturally be reproducible, accurate and reliable.
Pre-processing
Data preprocessing is a crucial concern in machine learning research. It is performed before the construction of learning models to prepare reliable input data sets. In this work, the experimental images pass through the filter to eliminate the noise then only doing the segmentation and classification. Usually, the background of an image does not contribute any fruitful information at the same time increases the processing time. Hence to improve the processing speed and decrease memory usage, remove background, eyes, scalp, skull, and structures that are not the regions of interest. The Brain Surface Extractor (BSE) algorithm which is used only for MR images can be used for skull removal [15]. The images are filtered using BSE removing irregularities, performing morphological erosions, detecting edges and isolating brain. It also does image masking and surface clean up. In our system, anisotropic filter is used for removing the noise; it eliminated the noise pixel without affect the edge pixel of the image. In this work, skull removed is done by using skull stripping and tumor regions are identified in region props algorithm. Mathematical morphology from the initial processing of binaryimages, and gradually developed to deal with grayscale image. The skull striping processing is given in Eq. (1)
ROI segmentation
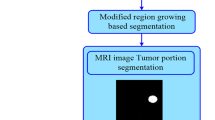
In the past decade, level set method is too cumbersome in computational complexity, it still has a unique advantage: first of all, the coordinate system for the level set method of the given PDE is fixed, so it is a no parameter method, secondly, a topology change may occurs for the curve during the evolution process, for the level set method, the whole curve movement is embedded in the high-dimensional surface motion, so the curve on the topology changes are same embedded in the high-dimensional surface motion, thirdly, the segmentation accuracy of the level set can reach sub-pixel, and the medical images require a class of high precision, so the level set has a wide range of applications [14,15,16].The segmentation results of proposed method is given in Fig. 1.
Feature extraction using modified multi-trend structure descriptor
Feature extraction can be defined as the process of transforming or converting an image into its group of features. The different methods employed for feature extraction includes texture features, co-occurrence matrix, Gabor features, wavelet transform based features, decision boundary feature extraction, minimum noise fraction transform, nonparametric weighted feature extraction and spectral mixture analysis. For feature reduction principal component analysis, linear discriminate analysis and independent component analysis are used. Integration of the feature extraction with the feature reduction algorithms leads to accurate systems that use less number of features that can be extracted with less computational cost [17]. The tumor type and grade are the two main factors that decide the features used for brain tumor segmentation. This is because diverse types and grades of tumor exhibit variations in appearance that may be shape, location, regularity, contrast uptake etc. The commonly used features are the image intensities assuming that different tissues have varying gray levels [18,19,20].
First order intensity statistics are also referred to as voxel intensity based features. First order statistical features express the distribution of grey levels within selected regions of interest (ROI), represented by supervoxels in our case. We use 16 features including the average, standard deviation, variance, mean of the absolute deviation, median absolute deviation, coefficient of variance, skewness, kurtosis, maximum, minimum, median and mode of the intensity values, central moments, range, inter quartile range and entropy. Due to the complexity and heterogeneity of tumour tissue, first order intensity features are generally not sufficient for an accurate segmentation. In the proposed algorithm, The texton mask 2 × 2 is move the image from left to right and top to bottom for identify the texton pattern of entire image. Texton mask are labeled as T1, T2, T3, T4, T5, and T6, respectively, the proposed 2 × 2 grid as shown in Fig. 2 and texton calculation process is given in Fig. 3, it is defined as per Eq. (2)
In block counting, the texton image is converted into a number of grids by drawing an imaginary line of equal space and the imaginary lines were drawn horizontally and vertically over the image. The grids are formed to make accurate measurements. Based on the requirement, number of grids can be made. The number of grids commonly made includes 4, 18 and 24 grids. Then count the number of block of the texton image of each intensity value 1 to 255. Also the original image is converted into number of grids, count number of block of the origin image of each intensity value 1 to 255. The original and orientation image block count values are used for tumor classification.
Feature classification
Classification of MR images is an essential process in resolving the human tissues, especially at the time of clinical analysis. The Brain is explicitly complex and mainly it consists of three healthy tissues named as WM, GM and CSF which can be detected using segmentation of the brain MR image and are very important in surgical planning and in diagnosing neurological diseases [21,22,23].
In non linear data, the input space X can be mapped into higher dimensional space ψ to make the feature space become linearly separable. The mapping function ψ should be in accordance with the Mercer’s theorem. It is given in Eq. (3)
Kernel functions can be chosen from the following functions, it is given in Eq. (4),
Let K1(X, Xi) and K2(X, Xi) be RBF kernels over Ξ × Ξ, Ξ ⊆ Rp, and K3(X, Xi) be a kernel over Rp × Rp.The proposed pair of RBF kernels are designed in Eq. (5)
Experimental results
For performance evaluation of the proposed algorithm, the experimental data set contains 100 meningioma tumor images and 40 normal images. These images are obtained from a hospital. In this case, 50 normal samples and 80 tumor samples are used as the training set. Success of the MRI image identification is measured due to its sensitivity, specificity, accuracy and coverage which can be explained as per Eq. (6)
The sensitivity, specificity and accuracy results of various tumor classification methods are shown in Table 1.The investigational results using k-fold cross validation method of sensitivity, specificity and accuracy are given in Figs. 4, 5 and 6. The overall results of feature extraction with different classifiers are shown in Table 2.
Conclusion
This paper proposed an excellent and innovative classification of brain image is vital in identifying disorders and treatment planning in the field of medicine. The proposed method consists of four stages. In the first stage, with respect to the cells’ boundary, it is necessary to further processing, which ensures the boundary of some cells is a discrete region. Mathematical Morphology brings a fancy result during the discrete processing. Secondly, the brain MRI is segmented into white matter (WM), gray matter (GM), Cerebrospinal fluid (CSF) and outliers by ordering of observations. In classification, novel hybrid Fuzzy SVM is designed in the classification stage meningioma brain tumor. The proposed method is validated using k-fold cross validation method. The classification accuracy of MM with SVM (MMSVM) is 74.24%, MM with FSVM is 82.67% and MM with RBF is 62.71% and our proposed method MM with Hybrid SVM is 91.64%.
References
Ramsay, C. R., Matowe, L., Grilli, R., Grimshaw, J. M., and Thomas, R. E., Interrupted time series designs in health technology assessment: lessons from two systematic reviews of behavior change strategies. Int. J. Technol. Assess. Health Care 19(4):613–623, 2003.
Zikic, B., Glocker, E. K., Criminisi, A., Demiralp, C., Shotton, J., Thomas, O. M., Das, T., Jena, R., and Price, S. J., Decision forests for tissue-specific segmentation of high-grade gliomas in multi-channel. Journal of Medical Image Computing and Computer Assisted Intervention 7512:369–376, 2012.
Dhanasekaran, R., and Jayachandran, A., Severity analysis of brain tumor in MRI images uses modified multi-texton structure descriptor and kernel-SVM. Arab. J. Sci. Eng. 39(10):7073–7086, 2014.
Dubey, M. H., Gupta, S. K., and Gupta, S. K., Semi-automatic Segmentation of MRI Brain Tumor. Journal of Graphics, Vision and Image Processing. 9:33–40, 2009.
Kromer, C., Xu, J., Ostrom, Q. T. et al., Estimating the annual frequency of synchronous brain metastasis in the United States 2010-2013: a population-based study. J. Neuro-Oncol. 134(1):55–64, 2017.
Posner, J. B., and Chernik, N. L., Intracranial metastases from systemic cancer. Adv. Neurol. 19:579–592, 1978.
Vishvaksenan, K. S., Mithra, K., Kalidoss, R., and Karthipan, R., Experimental study on Elliot wave theory for Handoff Prediction. Fluctuation and Noise Letters 15(4):1–11, 2016.
Taheri, S., Ong, S. H., and Chong, V. F. H., Level-set segmentation of brain tumors using a threshold-based speed function. J. Image Vision Comput. 28:26–37, 2010.
DeAngelis, L. M., and Posner, J. B., Neurologic Complications of Cancer. 2nd edition. New York: Oxford University Press, 2009.
Silberstein, S. D., Practice parameter: evidence-based guidelines for migraine headache (an evidence-based review): report of the Quality Standards Subcommittee of the American Academy of Neurology. Neurology 55(6):754–762, 2000.
Krumholz, A., Wiebe, S., Gronseth, G. et al., Practice Parameter: evaluat- ing an apparent unprovoked first seizure in adults (an evidence-based review): report of the Quality Standards Subcommittee of the Ameri- can Academy of Neurology and the American Epilepsy Society. Neurology 69(21):1996–2007, 2007.
Jayachandran, A., and Dhanasekaran, R., Brain tumor severity analysis using modified multi-texton histogram and hybrid kernel SVM. Int. J. Imaging Syst. Technol. 24(1):72–82, 2014.
Glantz, M. J., Cole, B. F., Glantz, L. K. et al., Cerebrospinal fluid cytology in patients with cancer: minimizing false-negative results. Cancer 82(4):733–739, 1998.
Del Principe, M. I., Buccisano, F., Cefalo, M. et al., High sensitivity of flow cytometry improves detection of occult leptomeningeal disease in acute lymphoblastic leukemia and lymphoblastic lymphoma. Ann. Hematol. 93(9):1509–1513, 2014.
Abdel-Maksoud, E., Elmogy, M., and Al-Awadi, R., Brain tumor segmentation based on a hybrid clustering technique. Egypt. Informatics J. 16:71–81, 2015.
Jayachandran, A., and Dhanasekaran, R., Automatic detection of brain tumor in magnetic resonance images using multi-texton histogram and support vector machine. Int. J. Imaging Syst. Technol. 23:97–103, 2013.
Jayachandran, A., and Dhanasekaran, R., Abnormality segmentation and Classification of multi model brain tumor in MR images using Fuzzy based hybrid kernel SVM. International Journal of Fuzzy Systems 17(3):434–443, 2015.
Patchell, R. A., Tibbs, P. A., Walsh, J. W. et al., A randomized trial of surgery in the treatment of single metastases to the brain. N. Engl. J. Med. 322(8):494–500, 1990.
Hariharan, G., and Jayachandran, A., Color, textures and shape descriptor based cervical cancer classification system of pap smear images. J. Comput. Theor. Nanosci. 14(7):3609–3614, 2017.
Vishvaksenan, K. S., Kalaiarasan, R., Kalidoss, R., and Karthipan, R., Real time experimental study and analysis of Elliott wave theory in signal strength prediction. Proceedings of National Academy of Sciences, Springer 88(1):107–119, 2018.
Luts, J., Laudadio, T., Idema, A. J., Simonetti, A. W., Heerschap, A., Vandermeulen, D., Suykens, J. A. K., and Huffel, S. V., Nosologic imaging: segmentation and classification using MRI and MRSI. Journal of NMR in Biomedicine 22:374–390, 2009.
Borgelt, B., Gelber, R., Kramer, S. et al., The palliation of brain metasta- ses: final results of the first two studies by the Radiation Therapy Oncology Group. Int J RadiatOncolBiol Phys. 6(1):1–9, 1980.
Jayachandran, A., and Dhanasekaran, R., Multi Class Brain Tumor Classification Of MRI Images using Hybrid Structure Descriptor and Fuzzy Logic Based RBF Kernel SVM. Iranian Journal of Fuzzy system 14(3):41–54, 2017.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
No potential conflict of interest was reported by the authors.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
This article is part of the Topical Collection on Image & Signal Processing
Rights and permissions
About this article
Cite this article
Arokia Jesu Prabhu, L., Jayachandran, A. Mixture Model Segmentation System for Parasagittal Meningioma brain Tumor Classification based on Hybrid Feature Vector. J Med Syst 42, 251 (2018). https://doi.org/10.1007/s10916-018-1094-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10916-018-1094-3