Abstract
MicroRNAs are master regulators of gene expression in many biological and pathological processes, including mammary gland development and breast cancer. The differentiation program termed the epithelial to mesenchymal transition (EMT) involves changes in a number of microRNAs. Some of these microRNAs have been shown to control cellular plasticity through the suppression of EMT-inducers or to influence cellular phenotype through the suppression of genes involved in defining the epithelial and mesenchymal cell states. This has led to the suggestion that microRNAs maybe a novel therapeutic target for the treatment of breast cancer. In this review, we will discuss microRNAs that are involved in EMT in mammary cells and breast cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
microRNAs
MicroRNAs (miRNAs) are non-coding RNAs that regulate gene expression at the post-transcriptional level by inhibiting translation or initiating mRNA destruction. There is increasing evidence that miRNAs maybe master regulators of many fundamental biological processes, such as embryogenesis [1] and organ development [2]. Individual miRNAs can enforce a developmental switch or tissue-specific gene expression through regulation of a key mRNA target [3, 4] or through many mRNA targets [5]. Similar to other forms of gene regulation, miRNAs have been shown to be involved in human cancers, acting as oncogenes [6, 7] or tumor suppressors [8, 9]. Therefore, understanding this form of gene regulation maybe a key to regulating fundamental biological processes and controlling disease progression.
Some miRNAs are expressed in a cell-specific, tissue-specific and/or developmental stage-specific manner, while others are expressed relatively ubiquitously. As with other genes, this is dependent upon the transcription factors that regulate their expression. MiRNA genes are transcribed by RNA polymerase II as their own dedicated transcript [10] or can be cleaved from introns of protein-coding genes. Cleavage of the primary transcript by the DROSHA/DGCR8 complex [11] generates a 60–70 nucleotide (nt) stem-loop pre-miRNA, which is exported from the nucleus by the exportin 5-RanGTP system [12]. Within the cytoplasm, the RNAse III enzyme Dicer processes the pre-miRNA to yield the 18–24 nt mature miRNA [13, 14]. Through imperfect base pairing, miRNAs bind to target mRNAs in the context of the RNA-induced silencing complex (RISC) [15]. Complementarity between nucleotides 2–8 at the 5′ end of the miRNA (termed the “seed sequence”) is important for efficient targeting. While a single miRNA can be encoded in one primary transcript, often a cluster of multiple miRNAs are encoded in a polycistronic transcript. Differences in the seed sequences of miRNAs within a cluster means that a single cluster of miRNAs has the potential to regulate an enormous range of targets.
miRNA Regulation of EMT
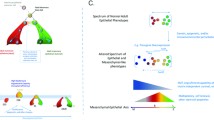
Epithelial to mesenchymal transition (EMT) is involved in embryonic development, wound healing and cancer progression (reviewed in [16]). During an EMT, epithelial cells lose cell-cell contacts and undergo cytoskeletal remodelling and polarity changes, resulting in acquisition of mesenchymal morphology and enhanced migratory ability. Importantly, the process of EMT is reversible (termed a mesenchymal to epithelial transition or MET), so that polarised epithelium can be generated in a new site. Many signalling pathways induce EMT, including Transforming Growth Factor β (TGF-β), Wnt and Notch, in part through regulation of several transcription factors, including Snail1/2, E47, Klf8, ZEB1/2 and Twist (reviewed in [16]). A key target of these EMT-inducing transcriptional repressors is the epithelial-specific junctional protein, E-cadherin. Loss of epithelial specific proteins, such as E-cadherin, and increased expression of mesenchymal specific proteins, such as vimentin, can be used as markers to show that an epithelial cell has undergone an EMT. Here we highlight miRNAs that are regulated by and control EMT (Fig. 1).
Microarray analysis has been used to identify miRNAs that are involved in EMT, comparing cells before and after induction of EMT. When EMT is induced with different stimuli, the most consistently striking change is the reduction in levels of the miR-200 family and miR-205 [4, 17–20]. The miR-200 family include two clusters of miRNAs — miR-200b∼200a∼429 and miR-200c∼141 [4]. The dramatic decrease in miR-200 family and miR-205 expression with EMT is reflected in lower expression of the miR-200 family and miR-205 in mesenchymal cell lines compared to epithelial lines [4, 20–22]. Significantly, these correlations have led to the discovery that the miR-200 family are key regulators of EMT. Through their suppression of the mesenchymal-specific repressors of E-cadherin transcription, ZEB1 and ZEB2, the miR-200 family increase the levels of E-cadherin and are capable of enforcing the epithelial phenotype in mesenchymal cells [4], including mesenchymal-like, MDA-MB-231 breast cancer cells [17, 20]. Additionally, their expression in epithelial cells can inhibit TGF-β1-induced EMT in canine kidney and murine mammary epithelial cell lines, MDCK and NMuMG cells respectively [4, 22]. Conversely, inhibition of the miR-200 family in epithelial cells induces EMT [4, 17, 20]. In part, the reversibility of EMT is determined by the fact that ZEB1 and ZEB2 suppress miR-200 family expression [23]. This double negative feedback loop enables the miR-200 family and ZEBs to respectively maintain the epithelial and mesenchymal cellular states. Additional EMT-related targets of the miR-200 family have also been reported, including TGF-β2 by miR-141 [17] and Beta-catenin (CTNNB1) by miR-200a [24]. Given that miRNAs are capable of regulating hundreds of miRNA targets and the key role the miR-200 family play in enforcing the epithelial phenotype, further miR-200 family targets that are involved in EMT may still be identified.
The key role that the TGF-β signalling pathway plays in EMT and cancer (reviewed in [25]), has prompted studies focussed upon identifying miRNAs involved in TGF-β-induced EMT. In contrast to the miR-200 family, levels of miR-155 are increased during TGF-β-induced EMT in mammary epithelial cell model systems, via transcriptional activation by SMAD4 [19]. While ectopic expression of miR-155 did not induce an EMT, cell polarity and tight junction formations were disrupted and cells responded more rapidly to TGF-β. Significantly, loss of miR-155 suppresses TGF-β induced EMT in NMuMG cells. A key miR-155 target is RhoA, which plays an important role in formation and stabilisation of cell junctions [19]. In addition to miR-155 regulation of RhoA, TGF-β induces the ubiquitination and degradation of RhoA by Smurf1 E3 ligase, in response to activation by Par6 [26, 27]. Accordingly, miRNAs such as miR-155 provide a further layer of regulation that ensures the suppression of specific proteins to control cellular phenotype.
MiR-29a and miR-21 levels increase upon TGF-β-induced EMT in mammary epithelial model systems [18, 19] and are higher in many mesenchymal cell lines compared to epithelial cell lines [20], although the precise relationship between these miRNAs and EMT has not been extensively explored. Significantly, suppression of tristetraprolin (TTP) by miR-29a induces EMT in oncogenic Ras-expressing murine mammary epithelial cells [18]. However, since the overexpression of miR-29a in non-tumorigenic murine mammary epithelial cells did not induce an EMT, the relationship between miR-29a and EMT is clearly context dependent. In contrast, there are no reports of miR-21 directly regulating EMT. However, there have been multiple reports of pathways regulating miR-21 in response to TGF-β. Pre-miR-21 and mature miR-21 levels are increased by TGF-β in MDA-MB-468 breast carcinoma cells via a SMAD4-independent, post-transcriptional mechanism, involving increased processing of the miR-21 primary transcript by the Drosha complex [28, 29]. Additionally, miR-21 transcription can be activated by the TGF-β-regulated transcription factor, AP-1 [30, 31] or by the EMT-inducer, ZEB1 [32, 33]. The functional significance of increased levels of miR-21 with EMT maybe elucidated from its target mRNAs, which have roles in EMT, cell cycle control and apoptosis (TGFβR2, DAXX, Cdc25A, PDCD4, TPM1, PTEN) [34–39]. Further exploration of the role of these miRNAs in EMT is required.
EMT can also be induced by the transcription factor Twist [40]. Amongst other genes, Twist activates miR-10b transcription through an E-box proximal to the predicted promoter [41]. In contrast to Twist, miR-10b alone cannot induce an EMT. However, miR-10b increases motility and invasiveness of HMECs (human mammary epithelial cells) and breast cancer cells and contributes to the migratory and invasive properties conferred by Twist. It is still unclear whether miR-10b is specifically related to Twist or is involved in EMT induced by other stimuli. There is some evidence that other E-box binding EMT-inducers may have opposing effects on miR-10b. For example, Snail1 reduced miR-10b expression in HMECs [41]. Microarray data suggest that ZEB1 may increase miR-10b levels in colorectal cancer cells, but decrease miR-10b levels in breast cancer cells [17], suggesting that the link between miR-10b and EMT is context dependent.
Loss of E-cadherin can induce EMT in epithelial cells [42]. Accordingly, miRNAs that affect E-cadherin expression are likely to be regulators of EMT. miR-9 has recently been reported to suppress E-cadherin expression and induce EMT in immortalised human mammary epithelial (HMLE) cells [43]. Interestingly, miR-9 cannot induce EMT in an epithelial breast cancer cell line, SUM149, in vitro, however, there is increased expression of the mesenchymal marker, vimentin, at the tumour-stroma interface in vivo compared to control, which suggests that miR-9 may sensitise cells to signals from the tumour microenvironment that induce EMT.
miRNAs and Mammary Stem Cells
Mammary development and homeostasis is thought to be dependent upon a differentiation hierarchy, with mammary stem cells at the apex (reviewed in [44]). Cell populations enriched in mammary stem cells can be prepared by dissociating mouse mammary tissue into single cell suspensions and fractionating the cells according to cell surface markers, such as CD24med/CD49fhi and CD29hi/CD24+ [45, 46]. Mouse mammary stem cells isolated by these methods are able to reconstitute entire mammary glands in vivo [45, 46]. In an effort to understand molecular pathways regulating stem cell properties, microarray studies were performed to assess differences in mRNAs and miRNAs in mammary stem cells compared to mammary progenitor and mature cells. Higher levels of the mesenchymal marker, vimentin, and lower levels of E-cadherin and the miR-200 family are found in human mammary stem cells (CD49fhiEpCAM-) compared to luminal progenitor and mature luminal cells (CD49f−/+EpCAM+) [47, 48]. Similarly, enriched populations of mouse mammary stem cells (CD24medCD49fhi) have lower levels of the miR-200 family compared to more differentiated mammary epithelial progenitor cells (CD24hiCD49flo) [48]. According to the expression of these epithelial and mesenchymal markers, differentiation of mammary stem cells may follow a program similar to a MET. Importantly, the low levels of the miR-200 family in mammary stem cells are significant. This is most clearly demonstrated when miR-200c is ectopically expressed in murine mammary cells isolated from FVB/NJ mice. In contrast to controls, when miR-200c expressing cells are injected into the cleared mammary fat pad of FVB/NJ mice, normal mammary outgrowth is suppressed and myoepithelial differentiation is induced [48]. This suggests that the miR-200 family may regulate differentiation of mammary stem cells.
In contrast to the miR-200 family, significantly higher expression of miR-205 is observed in enriched populations of [1] basal and myoepithelial stem-cells (CD24+/lo/Sca-1-) compared to luminal (CD24+/hi/Sca-1−/+)cells, [2] self-renewing stem cells (CD29hi/CD24+) compared to non-stem cells and [3] stem cells (CD24med/CD49fhi) or myoepithelial cells (CD24lo/CD49flo) compared to non-stem cells (CD24+/hi/CD49f-) [49]. The high levels of miR-205 in mammary stem cells are functionally significant, because when miR-205 was ectopically expressed in a cell line model of mammary gland progenitor cells, there was a significant expansion of the progenitor population [49, 50]. Despite the similar roles of the miR-200 family and miR-205 in regulating ZEB expression and EMT [4], differences in the response of mammary stem cells to these miRNAs highlights the differences in the range of targets regulated by these miRNAs. Other miR-205 targets have been identified, including PTEN, protein kinase Cε, LRP1 and HER3 [49, 51–53], as well as many other potential targets predicted from a microarray conducted on miR-205-expressing cells [49]. Further experiments are required to fully understand the role of miR-205 in mammary stem cells, development and tissue homeostasis.
miRNAs, EMT and Mammary Gland Development
While there are reports of miRNA involvement in mammary gland development [54, 55], there is very little discussion of EMT in mammary gland development. However, a process reminiscent of wound healing and EMT occurs between lactation and involution. Mammary involution is initiated at the end of lactation, which involves cessation of milk secretion, massive cell death, collapse of the alveoli, clearance of apoptotic cells and remodelling of the epithelial compartment to restore a simple ductal structure (reviewed by [56, 57]). By day 4 of involution, TGF-β signalling pathways increase [58–60], coinciding with decreased levels of E-cadherin [61, 62]. miRNA expression profiles suggest that miRNAs may play a role in this EMT-like process in late involution, with decreased levels of miR-200a, miR-429 and miR-141 and increased expression of miR-29a, miR-21, and miR-10b [54]. Similar to the changes observed in mammary progenitor cells [49, 50], changes in miR-205 levels are in direct contrast to the miR-200 family. However, the precise roles of EMT and associated miRNAs in mammary gland development have yet to be explored.
miRNAs, EMT and Breast Cancer Metastasis
Metastasis is the most common cause of death for breast cancer patients. Evidence for the involvement of miRNAs in the metastatic process is rapidly accumulating, presenting an attractive novel therapeutic possibility. Altered expression of the miRNA processing machinery is observed in more invasive, aggressive breast cancers. For example, levels of Dicer are lower in mesenchymal compared to epithelial breast cancer cell lines and also in the more aggressive basal-like, HER2+ and luminal B type tumours compared to luminal A type breast cancer [63–65]. The latter maybe linked to the frequent hemizygous deletion of Dicer associated with breast tumors [66]. Specific miRNAs have also been directly linked to metastasis. For example, miR-31, miR-373, miR-520c, miR-126 and miR-335 potentially regulate metastasis through a range of mRNA targets [67–70].
One theory to explain how metastases arise from a primary breast tumor is that peripheral epithelial cells receive signals from the surrounding stroma to undergo the EMT program, thus enhancing tumor cell motility and invasiveness (reviewed in [16]). This hypothesis is supported by studies showing a loss E-cadherin and higher levels of mesenchymal markers in invasive ductal, basal-like and metaplastic carcinoma of the breast [71–73]. The links between metastasis and EMT are also reflected in the expression of EMT-related miRNAs. MiR-21, miR-9 and miR-155 levels are increased in malignant breast cancer and breast cancer cell lines compared to normal tissues and human mammary epithelial cells [74–76]. Both basal and metaplastic breast cancers have reduced expression of the miR-200 family compared to ductal breast tumors, which correlates with their invasiveness [4, 17]. This contrasts miR-205 levels, which are not significantly different between ductal and metaplastic primary breast carcinomas [4], once again highlighting differences in the mRNA targets of between the miR-200 family and miR-205. However, direct links between these EMT-related miRNAs and metastasis are most clear upon comparison of primary tumour samples to metastases — higher levels of miR-10b, miR-21 and miR-155 and lower levels of the miR-200 family were observed in metastatic samples compared to matched primary tumours [77]. These observations have led to the hypothesis that manipulation of the EMT program through EMT-regulating miRNAs may limit metastasis.
Given the key role that the miR-200 family plays in regulation of the epithelial phenotype and EMT, it has been hypothesized that enforced expression of these miRNAs will limit metastasis [78]. In support of this hypothesis, knockdown of the key miR-200 target ZEB1 in Panc1 cells decreases primary pancreatic tumour size and inhibits local infiltration and metastasis upon intrapancreatic injection into nude mice [79]. A role for miR-200 in suppressing metastasis is supported by work with a lung adenocarcinoma model, where miR-200 family expression in metastasis-prone cells did not affect primary tumor growth rate but inhibited metastasis from tumors formed by subcutaneous injection of the cells into the posterior flank of 129Sv mice [80]. The potential for the miR-200 family to suppress metastasis is also inferred from other identified miR-200 family targets, leptin receptor and cofilin 2, which are known promoters of metastasis [17]. This suggests that the miR-200 family maybe capable of limiting metastasis through suppression of a range of mRNA targets involved in multiple pathways. Numerous studies have found that miR-200 inhibits migration or invasion of cells in vitro. This includes MDCK cells [4], 344SQ lung adenocarcinoma cells [80], NPC nasopharyngeal carcinoma cells [24], SW480 colorectal cancer cells [17], Hec50 endometrial cancer cells [81], ovarian cancer SKOV-3 cells [82], LNCaP prostate cancer cells [83], TGF-β1 treated MCF10A breast cancer cells [84], MDA-MB-231 cells [17, 20, 85] and 4T07 breast cancer cells [22]. This latter result with 4T07 cells is contradicted by a report that ectopic expression of miR-200c∼141 in 4TO7 cells increased migration, despite the cells having undergone an MET (according to their epithelial-like morphology, loss of ZEB2 and gain of E-cadherin) [86]. One difference between the two studies is that the latter investigated migration through membranes coated with a mixture of basement membrane components whereas the former study used uncoated membranes, although it is not clear that this accounts for the differences in effect of miR-200 on migration, especially since other reports show miR-200 reducing invasion through matrigel [17, 24, 80, 81, 83, 85]. Consistent with the enhanced in vitro migration of miR-200-expressing 4TO7 cells, they produced more metastases from mammary tumors formed by injection into the mammary fat pad of BALB/cJ mice [86]. Clearly more work is required to resolve why miR-200 appears to inhibit invasion and metastasis in some systems but to apparently promote it in the 4T07 model.
Loss of E-cadherin is a key characteristic of EMT and is associated with tumour progression, metastasis and poorer prognosis in breast cancer [72, 87, 88]. Knockdown of E-cadherin in breast cancer cells is sufficient to dramatically increase metastasis when these cells are injected into nude mice [42]. In accordance with its ability to suppress E-cadherin, miR-9 can regulate metastasis [43]. However, the increased migration and invasion of miR-9 expressing HMLE or SUM149 cells in vitro can only partially be rescued by ectopic expression of E-cadherin, suggesting that miR-9 may have other mRNA targets that effect migration and invasion. Importantly, the link between miR-9 and metastasis is likely to be clinically relevant, because significantly higher miR-9 levels were observed in primary breast tumours from patients with metastases compared to patients with no metastases [43]. Since miR-9 levels are lower in metastases compared to matched primary tumour samples [77], miR-9 maybe involved in an early step in the metastatic process.
Suppression of the EMT-inducer, Twist, inhibits metastasis of breast cancer cells injected into the mammary gland of Balb/c mice [40]. A key target of Twist is miR-10b, as loss of miR-10b suppresses the Twist-induced migration and invasion of HMLE cells in vitro [41]. MiR-10b is capable of promoting invasion in vitro and metastasis upon orthotopic injection into NOD/SCID mice, as shown with over-expression of miR-10b in non-invasive, non-metastatic SUM149 breast cancer cells. MiR-10b suppresses homeobox D10 (HOXD10), thus permitting the expression of the pro-metastatic protein RHOC [41]. While it is clear that miR-10b can initiate metastasis, the correlation of miR-10b and breast cancer is complicated. miR-10b levels are reduced in breast cancer samples compared to normal tissue [75, 89] and miR-10b expression does not correlate with distant metastases, recurrence-free survival or distant relapse free survival [89]. To reconcile these observations, a likely hypothesis is that induction of EMT at the invasive front of a breast tumour may lead to transient activation of Twist and miR-10b, thus promoting invasion and metastasis [41, 90]. Further investigation is required to assess the precise role of miR-10b in metastasis and its link to EMT.
The pro-tumorigenic, pro-metastatic role of miR-21 has been firmly established [91]. Increased miR-21 expression is consistently observed in breast cancer, particularly invasive breast cancer [92–95] and high levels of miR-21 correlate with poor disease free survival and high levels of TGF-β1 [96]. MiR-21 directly suppresses known metastasis suppressors, including Tropomyosin 1 (TPM1), PDCD4 and maspin [91]. Further investigation is required to determine whether reversion of mesenchymal, metastatic cells to an epithelial phenotype would lead to a reduction in miR-21 and a suppression of metastasis. Manipulation of EMT, through the differential expression of these EMT-related miRNAs, may present a novel therapeutic strategy for the treatment of advanced breast cancer although many obstacles remain, with delivery being one of them.
miRNAs, EMT and Breast Cancer Stem Cells
The concept of cancer stem cells is controversial and may not apply to all human cancers, but support for the cancer stem cell model comes from the ability to fractionate cancer cells into populations enriched for tumour-initiating cells. Breast cancer stem cells (bCSCs) can be enriched from solid breast tumours or pleural effusions from metastatic breast cancer patients according to cell surface markers (CD44+CD24−/low) and serially passaged in immunocompromised mice, generating tumours each time that have the same heterogeneous mix of cells present in the initial tumour, consistent with the notion of stem cell-like properties [97]. Aldehyde dehydrogenase 1 (ALDH1) activity is also used to enrich for bCSCs and is associated with metastatic ability and poor prognosis of breast cancer [98, 99]. These studies indicate that bCSC frequency varies depending on treatments, grade and sub-type of breast cancer. Regardless of the method of enrichment, bCSCs have increased tumorigenic potential and propensity to metastasise compared to other cancer cells [97, 99]. This has led to the generation of an “invasiveness” signature by expression profiling of CD44+CD24−/low (bCSC-like) cells relative to normal breast epithelium, which is highly predictive of the propensity of a tumour to metastasise [100]. Importantly, bCSCs are resistant to current chemotherapy and radiation therapies [101–104]. The combination of the bCSC properties of self-renewal, ability to reconstitute a tumour and resistance to therapy facilitate recurrence in advanced breast cancer.
There is much evidence linking bCSCs and EMT. BCSCs enriched from breast tumours and metastatic breast pleural effusions express markers similar to cells that have undergone an EMT [105–107]. Similarly, EMT and stem cell markers are frequently associated with breast cancers that have a propensity to metastasise, such as basal-like [73] and metaplastic [108] breast cancers. Consistent with the expression of EMT markers in bCSCs, there is differential expression of EMT-related miRNAs in bCSCs compared to other breast cancer cells. miRNA expression profiling of bCSCs isolated from human breast tumours compared to the remaining breast cancer cells showed high levels of miR-155 and low levels of the miR-200 family in bCSCs [48]. In addition, low to undetectable levels of the miR-200 family are found in chemotherapy-resistant bCSCs isolated from breast cancer cell lines [104]. However, the most direct association between bCSCs and EMT comes from studies showing that the induction of EMT in vitro in transformed mammary epithelial cells [106, 109] or in vivo in epithelial breast cancer cells in mouse models [110] generates cells with bCSC properties. This suggests that through the EMT program breast epithelial cells can gain bCSC properties. Accordingly, the EMT program is involved in cancer progression. To assess whether reversal of EMT may suppress cancer stem cell properties, several groups have manipulated levels of regulators of EMT. For example, knockdown of ZEB in pancreatic cancer cell lines suppresses cancer stem cell properties [79]. Similarly, miR-200c expression in bCSCs suppresses cancer stem cell properties, as demonstrated by reduced tumorigenicity of miR-200c-expressing CD44+CD24−/low cells (isolated from an early passage human breast xenograft tumour) [48]. This study showed that in addition to enforcing the epithelial phenotype through repression of ZEB, miR-200c acts upon self-renewal pathways through regulation of BMI1. Further miR-200 family targets have been identified that are involved in self-renewal and cancer stem cell properties, such as Sox2 and Klf4 [79]. Therefore, the miR-200 family is able to affect two key properties of cancer stem cells — differentiation and self-renewal.
miRNAs that Affect Breast Cancer Response to Endocrine- or Chemotherapy
There are many determinants of sensitivity and resistance to cancer therapies such as drug metabolizing enzymes, drug transporters, proteins involved in DNA repair, cell division, and apoptosis, and levels of the drug targets themselves (such as estrogen receptor α for breast cancer endocrine therapy). MiRNAs can regulate these determinants and thereby influence sensitivity to therapeutic agents, as evidenced by knockdown of Dicer in breast cancer cells increasing sensitivity to cisplatin [111]. Furthermore, chronic exposure to anti-cancer agents can affect expression of miRNAs and lead to changes in the pharmacokinetic and pharmacodynamic properties of the agents.
The most global study to date on this topic examined the effects of three miRNAs, let7i, miR-16 and miR-21, which target RAS, BCL2, and PTEN respectively, and also used in silico methods to compare drug potency patterns of 3,089 compounds to miRNA expression profiles across the entire NCI-60 human cancer cell line panel [112]. The three miRNAs tested were capable of altering the potency of a number of anti-cancer agents by up to 4 fold and the in silico comparison of drug potencies to miRNA expression profiles across the NCI-60 panel demonstrated that 30 miRNAs, including miR-21, showed significant correlation with the potency of numerous anti-cancer agents, indicating a substantial role for miRNA in determining drug responsiveness [112]. Specific examples of miRNA affecting drug response by directly targeting important determinants of drug sensitivity or resistance are rapidly accumulating, including targeting of BCRP/ABCG2 and CYP1B1 by miR-328 and miR-27b [113, 114].
Increasing evidence links EMT and drug resistance. Epithelial markers are lost in cetuximab-resistant urothelial carcinoma cell lines [115], gemcitabine-resistant pancreatic cancer cells [116] and in erlotinib- and gefitinib-resistant non-small cell lung carcinoma and head and neck squamous cell carcinomas [117, 118], suggesting that these drug-resistant cells have undergone an EMT. Similarly, residual breast cancers after chemotherapy have low expression of E-cadherin and high expression of mesenchymal markers [119]. Additionally, EMT induced by knockdown of E-cadherin in HMLE cells increases resistance to doxorubicin, actinomycin D and paclitaxel [120].
Specifically linking the process of EMT with chemotherapy resistance is the finding that miR-200 not only represses ZEB1 and ZEB2, but also directly represses other mesenchymal and neuronal genes such as fibronectin, moesin, NTRK2 and class III beta-tubulin (TUBB3), not normally expressed in epithelial cells, but aberrantly expressed in high-grade, de-differentiated carcinoma cells that have undergone EMT [81]. Expression of TUBB3 (an isoform of tubulin normally limited to neuronal cells) is a common mechanism of resistance to microtubule-binding chemotherapeutic agents in many types of carcinoma, including breast cancer [121–124]. Enhanced miR-200c expression in carcinoma cells dramatically increases sensitivity to microtubule-targeting agents [81]. The ability of miR-200c to restore chemosensitivity to taxanes is attributable to its ability to directly target TUBB3 since introduction of exogenous non-targetable TUBB3 lacking its 3’UTR miR-200c target site, reverses the effect [125]. Similarly, miR-200b and miR-200c can increase the sensitivity of breast cancer cells to doxorubicin in vitro [85]. Additionally, miR-205 suppresses ERBB3/HER3 (a ligand binding, kinase inactive receptor tyrosine kinase of the epidermal growth factor receptor (EGFR) family). MiR-205 expression increases the sensitivity of breast cancer cells to the epidermal growth factor receptor inhibitors, gefitinib and lapatinib [52].
Nearly, 70% of breast cancers express estrogen receptor alpha (ESR1) and are consequent candidates for endocrine therapy. Tamoxifen, is the most commonly prescribed endocrine therapy for pre-menopausal women (while aromatase inhibitors are commonly used in post-menopausal women) and tamoxifen has also been recommended as a preventative. Nevertheless, 30–40% of patients with ESR1-positive tumours fail adjuvant tamoxifen therapy and nearly all patients with metastatic disease develop tamoxifen resistance [126–128]. De novo and acquired tumor resistance to endocrine therapy remains a poorly understood clinical problem. Recent studies have identified multiple miRNAs that target ER and affect ER-signalling [22, 86, 129–136]. MiR-206 has been demonstrated to directly target ESR1 [135] and may modulate tamoxifen resistance [130] and this miRNA appears to be up-regulated in ESR1-negative tumours [75]. However, the most compelling evidence of the critical role of miRNAs in downregulating ER and contributing to the acquisition of tamoxifen resistance has emerged from studies of miR-221/222, which are highly expressed in ESR1 negative breast tumours and cells lines, directly target ERα and render cells resistant to tamoxifen [132, 133]. While additional miRNAs differentially expressed in tamoxifen-resistant cell lines have been identified, their functional role in tamoxifen-resistance has not been elucidated [133].
Conclusions
Manipulation of EMT-related miRNAs may represent a novel therapeutic strategy for the treatment of advanced breast cancer. The attractiveness of using miRNAs as targeted therapeutics arises from the fact that unlike traditional gene therapy, the miRNA can simultaneously target many key genes/proteins involved in the process of EMT. As an example, expression of the differentiation- and “stemness”-associated miR-200 or miR-203 cannot only suppress transcription factors that repress E-cadherin, but they also target genes normally only expressed in mesenchymal cells or stem cells. Some of these targets may only be elucidated following proteomic analysis. In theory EMT-related miRNAs hold potential as a form of “differentiation therapy” that would drive differentiation, reduce invasion, and enhance chemosensitivity. However, in reality many obstacles remain, with delivery and timing being the largest hurdles to overcome. It remains to be determined how miRNAs introduced to prevent or reverse EMT in breast carcinoma cells will affect normal differentiated epithelium or normal stem cells. Furthermore, recent findings raise the question as to whether it will be detrimental to drive an MET in cancer [137].
Abbreviations
- miRNA:
-
microRNA
- EMT:
-
epithelial to mesenchymal transition
- MET:
-
mesenchymal to epithelial transition
- TGF-β:
-
Transforming Growth Factor β
- HMEC:
-
human mammary epithelial cell
- bCSC:
-
breast cancer stem cell
References
Wienholds E, Koudijs MJ, van Eeden FJ, Cuppen E, Plasterk RH. The microRNA-producing enzyme Dicer1 is essential for zebrafish development. Nat Genet. 2003;35(3):217–8.
Yi R, O’Carroll D, Pasolli HA, Zhang Z, Dietrich FS, Tarakhovsky A, et al. Morphogenesis in skin is governed by discrete sets of differentially expressed microRNAs. Nat Genet. 2006;38(3):356–62.
Chen K, Rajewsky N. Natural selection on human microRNA binding sites inferred from SNP data. Nat Genet. 2006;38(12):1452–6.
Gregory PA, Bert AG, Paterson EL, Barry SC, Tsykin A, Farshid G, et al. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol. 2008;10(5):593–601.
Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature. 2005;433(7027):769–73.
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S, et al. A microRNA polycistron as a potential human oncogene. Nature. 2005;435(7043):828–33.
Voorhoeve PM, le Sage C, Schrier M, Gillis AJ, Stoop H, Nagel R, et al. A genetic screen implicates miRNA-372 and miRNA-373 as oncogenes in testicular germ cell tumors. Cell. 2006;124(6):1169–81.
Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6(11):857–66.
Takamizawa J, Konishi H, Yanagisawa K, Tomida S, Osada H, Endoh H, et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res. 2004;64(11):3753–6.
Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek SH, et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 2004;23(20):4051–60.
Han J, Lee Y, Yeom KH, Kim YK, Jin H, Kim VN. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 2004;18(24):3016–27.
Lund E, Guttinger S, Calado A, Dahlberg JE, Kutay U. Nuclear export of microRNA precursors. Science. 2004;303(5654):95–8.
Hutvagner G, McLachlan J, Pasquinelli AE, Balint E, Tuschl T, Zamore PD. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science. 2001;293(5531):834–8.
Ketting RF, Fischer SE, Bernstein E, Sijen T, Hannon GJ, Plasterk RH. Dicer functions in RNA interference and in synthesis of small RNA involved in developmental timing in C. elegans. Genes Dev. 2001;15(20):2654–9.
Gregory RI, Chendrimada TP, Cooch N, Shiekhattar R. Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell. 2005;123(4):631–40.
Thiery JP, Acloque H, Huang RY, Nieto MA. Epithelial-mesenchymal transitions in development and disease. Cell. 2009;139(5):871–90.
Burk U, Schubert J, Wellner U, Schmalhofer O, Vincan E, Spaderna S, et al. A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep. 2008;9(6):582–9.
Gebeshuber CA, Zatloukal K, Martinez J. miR-29a suppresses tristetraprolin, which is a regulator of epithelial polarity and metastasis. EMBO Rep. 2009;10(4):400–5.
Kong W, Yang H, He L, Zhao JJ, Coppola D, Dalton WS, et al. MicroRNA-155 is regulated by the transforming growth factor beta/Smad pathway and contributes to epithelial cell plasticity by targeting RhoA. Mol Cell Biol. 2008;28(22):6773–84.
Park SM, Gaur AB, Lengyel E, Peter ME. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev. 2008;22(7):894–907.
Hurteau GJ, Carlson JA, Spivack SD, Brock GJ. Overexpression of the microRNA hsa-miR-200c leads to reduced expression of transcription factor 8 and increased expression of E-cadherin. Cancer Res. 2007;67(17):7972–6.
Korpal M, Lee ES, Hu G, Kang Y. The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J Biol Chem. 2008;283(22):14910–4.
Bracken CP, Gregory PA, Kolesnikoff N, Bert AG, Wang J, Shannon MF, et al. A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res. 2008;68(19):7846–54.
Xia H, Ng SS, Jiang S, Cheung WK, Sze J, Bian XW, et al. miR-200a-mediated downregulation of ZEB2 and CTNNB1 differentially inhibits nasopharyngeal carcinoma cell growth, migration and invasion. Biochem Biophys Res Commun. 2010;391(1):535–41.
Massague J. TGFbeta in Cancer. Cell. 2008;134(2):215–30.
Ozdamar B, Bose R, Barrios-Rodiles M, Wang HR, Zhang Y, Wrana JL. Regulation of the polarity protein Par6 by TGFbeta receptors controls epithelial cell plasticity. Science. 2005;307(5715):1603–9.
Wang HR, Zhang Y, Ozdamar B, Ogunjimi AA, Alexandrova E, Thomsen GH, et al. Regulation of cell polarity and protrusion formation by targeting RhoA for degradation. Science. 2003;302(5651):1775–9.
Davis BN, Hilyard AC, Lagna G, Hata A. SMAD proteins control DROSHA-mediated microRNA maturation. Nature. 2008;454(7200):56–61.
Zavadil J, Narasimhan M, Blumenberg M, Schneider RJ. Transforming growth factor-beta and microRNA:mRNA regulatory networks in epithelial plasticity. Cells Tissues Organs. 2007;185(1–3):157–61.
Fujita S, Ito T, Mizutani T, Minoguchi S, Yamamichi N, Sakurai K, et al. miR-21 Gene expression triggered by AP-1 is sustained through a double-negative feedback mechanism. J Mol Biol. 2008;378(3):492–504.
Shih SC, Claffey KP. Role of AP-1 and HIF-1 transcription factors in TGF-beta activation of VEGF expression. Growth Factors. 2001;19(1):19–34.
Du J, Yang S, An D, Hu F, Yuan W, Zhai C, et al. BMP-6 inhibits microRNA-21 expression in breast cancer through repressing deltaEF1 and AP-1. Cell Res. 2009;19(4):487–96.
Yang S, Du J, Wang Z, Yan J, Yuan W, Zhang J, et al. Dual mechanism of deltaEF1 expression regulated by bone morphogenetic protein-6 in breast cancer. Int J Biochem Cell Biol. 2009;41(4):853–61.
Asangani IA, Rasheed SA, Nikolova DA, Leupold JH, Colburn NH, Post S, et al. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene. 2008;27(15):2128–36.
Frankel LB, Christoffersen NR, Jacobsen A, Lindow M, Krogh A, Lund AH. Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J Biol Chem. 2008;283(2):1026–33.
Kim YJ, Hwang SJ, Bae YC, Jung JS. MiR-21 regulates adipogenic differentiation through the modulation of TGF-beta signaling in mesenchymal stem cells derived from human adipose tissue. Stem Cells. 2009;27(12):3093–102.
Meng F, Henson R, Wehbe-Janek H, Ghoshal K, Jacob ST, Patel T. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology. 2007;133(2):647–58.
Papagiannakopoulos T, Shapiro A, Kosik KS. MicroRNA-21 targets a network of key tumor-suppressive pathways in glioblastoma cells. Cancer Res. 2008;68(19):8164–72.
Wang P, Zou F, Zhang X, Li H, Dulak A, Tomko Jr RJ, et al. microRNA-21 negatively regulates Cdc25A and cell cycle progression in colon cancer cells. Cancer Res. 2009;69(20):8157–65.
Yang J, Mani SA, Donaher JL, Ramaswamy S, Itzykson RA, Come C, et al. Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell. 2004;117(7):927–39.
Ma L, Teruya-Feldstein J, Weinberg RA. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature. 2007;449(7163):682–8.
Onder TT, Gupta PB, Mani SA, Yang J, Lander ES, Weinberg RA. Loss of E-cadherin promotes metastasis via multiple downstream transcriptional pathways. Cancer Res. 2008;68(10):3645–54.
Ma L, Young J, Prabhala H, Pan E, Mestdagh P, Muth D, et al. miR-9, a MYC/MYCN-activated microRNA, regulates E-cadherin and cancer metastasis. Nat Cell Biol. 2010;12(3):209–11.
Visvader JE. Keeping abreast of the mammary epithelial hierarchy and breast tumorigenesis. Genes Dev. 2009;23(22):2563–77.
Shackleton M, Vaillant F, Simpson KJ, Stingl J, Smyth GK, Asselin-Labat ML, et al. Generation of a functional mammary gland from a single stem cell. Nature. 2006;439(7072):84–8.
Stingl J, Eirew P, Ricketson I, Shackleton M, Vaillant F, Choi D, et al. Purification and unique properties of mammary epithelial stem cells. Nature. 2006;439(7079):993–7.
Lim E, Vaillant F, Wu D, Forrest NC, Pal B, Hart AH, et al. Aberrant luminal progenitors as the candidate target population for basal tumor development in BRCA1 mutation carriers. Nat Med. 2009;15(8):907–13.
Shimono Y, Zabala M, Cho RW, Lobo N, Dalerba P, Qian D, et al. Downregulation of miRNA-200c links breast cancer stem cells with normal stem cells. Cell. 2009;138(3):592–603.
Greene SB, Gunaratne PH, Hammond SM, Rosen JM. A putative role for microRNA-205 in mammary epithelial cell progenitors. J Cell Sci. 2010;123(Pt 4):606–18.
Ibarra I, Erlich Y, Muthuswamy SK, Sachidanandam R, Hannon GJ. A role for microRNAs in maintenance of mouse mammary epithelial progenitor cells. Genes Dev. 2007;21(24):3238–43.
Gandellini P, Folini M, Longoni N, Pennati M, Binda M, Colecchia M, et al. miR-205 Exerts tumor-suppressive functions in human prostate through down-regulation of protein kinase Cepsilon. Cancer Res. 2009;69(6):2287–95.
Iorio MV, Casalini P, Piovan C, Di Leva G, Merlo A, Triulzi T, et al. microRNA-205 regulates HER3 in human breast cancer. Cancer Res. 2009;69(6):2195–200.
Song H, Bu G. MicroRNA-205 inhibits tumor cell migration through down-regulating the expression of the LDL receptor-related protein 1. Biochem Biophys Res Commun. 2009;388(2):400–5.
Avril-Sassen S, Goldstein LD, Stingl J, Blenkiron C, Le Quesne J, Spiteri I, et al. Characterisation of microRNA expression in post-natal mouse mammary gland development. BMC Genomics. 2009;10:548.
Tanaka T, Haneda S, Imakawa K, Sakai S, Nagaoka K. A microRNA, miR-101a, controls mammary gland development by regulating cyclooxygenase-2 expression. Differentiation. 2009;77(2):181–7.
Flanders KC, Wakefield LM. Transforming growth factor-(beta)s and mammary gland involution; functional roles and implications for cancer progression. J Mammary Gland Biol Neoplasia. 2009;14(2):131–44.
Monks J, Henson PM. Differentiation of the mammary epithelial cell during involution: implications for breast cancer. J Mammary Gland Biol Neoplasia. 2009;14(2):159–70.
Andrechek ER, Mori S, Rempel RE, Chang JT, Nevins JR. Patterns of cell signaling pathway activation that characterize mammary development. Development. 2008;135(14):2403–13.
Faure E, Heisterkamp N, Groffen J, Kaartinen V. Differential expression of TGF-beta isoforms during postlactational mammary gland involution. Cell Tissue Res. 2000;300(1):89–95.
Strange R, Li F, Saurer S, Burkhardt A, Friis RR. Apoptotic cell death and tissue remodelling during mouse mammary gland involution. Development. 1992;115(1):49–58.
Boussadia O, Kutsch S, Hierholzer A, Delmas V, Kemler R. E-cadherin is a survival factor for the lactating mouse mammary gland. Mech Dev. 2002;115(1–2):53–62.
Vallorosi CJ, Day KC, Zhao X, Rashid MG, Rubin MA, Johnson KR, et al. Truncation of the beta-catenin binding domain of E-cadherin precedes epithelial apoptosis during prostate and mammary involution. J Biol Chem. 2000;275(5):3328–34.
Blenkiron C, Goldstein LD, Thorne NP, Spiteri I, Chin SF, Dunning MJ, et al. MicroRNA expression profiling of human breast cancer identifies new markers of tumor subtype. Genome Biol. 2007;8(10):R214.
Cheng C, Fu X, Alves P, Gerstein M. mRNA expression profiles show differential regulatory effects of microRNAs between estrogen receptor-positive and estrogen receptor-negative breast cancer. Genome Biol. 2009;10(9):R90.
Grelier G, Voirin N, Ay AS, Cox DG, Chabaud S, Treilleux I, et al. Prognostic value of Dicer expression in human breast cancers and association with the mesenchymal phenotype. Br J Cancer. 2009;101(4):673–83.
Kumar MS, Pester RE, Chen CY, Lane K, Chin C, Lu J, et al. Dicer1 functions as a haploinsufficient tumor suppressor. Genes Dev. 2009;23(23):2700–4.
Bracken CP, Gregory PA, Khew-Goodall Y, Goodall GJ. The role of microRNAs in metastasis and epithelial-mesenchymal transition. Cell Mol Life Sci. 2009;66(10):1682–99.
Huang Q, Gumireddy K, Schrier M, le Sage C, Nagel R, Nair S, et al. The microRNAs miR-373 and miR-520c promote tumour invasion and metastasis. Nat Cell Biol. 2008;10(2):202–10.
Tavazoie SF, Alarcon C, Oskarsson T, Padua D, Wang Q, Bos PD, et al. Endogenous human microRNAs that suppress breast cancer metastasis. Nature. 2008;451(7175):147–52.
Valastyan S, Reinhardt F, Benaich N, Calogrias D, Szasz AM, Wang ZC, et al. A pleiotropically acting microRNA, miR-31, inhibits breast cancer metastasis. Cell. 2009;137(6):1032–46.
Lien HC, Hsiao YH, Lin YS, Yao YT, Juan HF, Kuo WH, et al. Molecular signatures of metaplastic carcinoma of the breast by large-scale transcriptional profiling: identification of genes potentially related to epithelial-mesenchymal transition. Oncogene. 2007;26(57):7859–71.
Prasad CP, Rath G, Mathur S, Bhatnagar D, Parshad R, Ralhan R. Expression analysis of E-cadherin, Slug and GSK3beta in invasive ductal carcinoma of breast. BMC Cancer. 2009;9:325.
Sarrio D, Rodriguez-Pinilla SM, Hardisson D, Cano A, Moreno-Bueno G, Palacios J. Epithelial-mesenchymal transition in breast cancer relates to the basal-like phenotype. Cancer Res. 2008;68(4):989–97.
Hui AB, Shi W, Boutros PC, Miller N, Pintilie M, Fyles T, et al. Robust global micro-RNA profiling with formalin-fixed paraffin-embedded breast cancer tissues. Lab Invest. 2009;89(5):597–606.
Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S, et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res. 2005;65(16):7065–70.
Yan LX, Huang XF, Shao Q, Huang MY, Deng L, Wu QL, et al. MicroRNA miR-21 overexpression in human breast cancer is associated with advanced clinical stage, lymph node metastasis and patient poor prognosis. RNA. 2008;14(11):2348–60.
Baffa R, Fassan M, Volinia S, O’Hara B, Liu CG, Palazzo JP, et al. MicroRNA expression profiling of human metastatic cancers identifies cancer gene targets. J Pathol. 2009;219(2):214–21.
Gregory PA, Bracken CP, Bert AG, Goodall GJ. MicroRNAs as regulators of epithelial-mesenchymal transition. Cell Cycle. 2008;7(20):3112–8.
Wellner U, Schubert J, Burk UC, Schmalhofer O, Zhu F, Sonntag A, et al. The EMT-activator ZEB1 promotes tumorigenicity by repressing stemness-inhibiting microRNAs. Nat Cell Biol. 2009;11(12):1487–95.
Gibbons DL, Lin W, Creighton CJ, Rizvi ZH, Gregory PA, Goodall GJ, et al. Contextual extracellular cues promote tumor cell EMT and metastasis by regulating miR-200 family expression. Genes Dev. 2009;23(18):2140–51.
Cochrane DR, Spoelstra NS, Howe EN, Nordeen SK, Richer JK. MicroRNA-200c mitigates invasiveness and restores sensitivity to microtubule-targeting chemotherapeutic agents. Mol Cancer Ther. 2009;8(5):1055–66.
Hu X, Macdonald DM, Huettner PC, Feng Z, El Naqa IM, Schwarz JK, et al. A miR-200 microRNA cluster as prognostic marker in advanced ovarian cancer. Gynecol Oncol. 2009;114(3):457–64.
Sossey-Alaoui K, Bialkowska K, Plow EF. The miR200 family of microRNAs regulates WAVE3-dependent cancer cell invasion. J Biol Chem. 2009;284(48):33019–29.
Iliopoulos D, Polytarchou C, Hatziapostolou M, Kottakis F, Maroulakou IG, Struhl K, et al. MicroRNAs differentially regulated by Akt isoforms control EMT and stem cell renewal in cancer cells. Sci Signal. 2009;2(92):ra62.
Tryndyak VP, Beland FA, Pogribny IP. E-cadherin transcriptional down-regulation by epigenetic and microRNA-200 family alterations is related to mesenchymal and drug-resistant phenotypes in human breast cancer cells. Int J Cancer. 2010;126(11):2575–83.
Dykxhoorn DM, Wu Y, Xie H, Yu F, Lal A, Petrocca F, et al. miR-200 enhances mouse breast cancer cell colonization to form distant metastases. PLoS ONE. 2009;4(9):e7181.
Gould Rothberg BE, Bracken MB. E-cadherin immunohistochemical expression as a prognostic factor in infiltrating ductal carcinoma of the breast: a systematic review and meta-analysis. Breast Cancer Res Treat. 2006;100(2):139–48.
Kowalski PJ, Rubin MA, Kleer CG. E-cadherin expression in primary carcinomas of the breast and its distant metastases. Breast Cancer Res. 2003;5(6):R217–22.
Gee HE, Camps C, Buffa FM, Colella S, Sheldon H, Gleadle JM, et al. MicroRNA-10b and breast cancer metastasis. Nature. 2008;455(7216):E8–9. author reply E9.
Ma L, Teruya-Feldstein J, Weinberg RA. Nature. 2008;455(7216):E9.
Zhu S, Wu H, Wu F, Nie D, Sheng S, Mo YY. MicroRNA-21 targets tumor suppressor genes in invasion and metastasis. Cell Res. 2008;18(3):350–9.
Huang GL, Zhang XH, Guo GL, Huang KT, Yang KY, Shen X, et al. Clinical significance of miR-21 expression in breast cancer: SYBR-Green I-based real-time RT-PCR study of invasive ductal carcinoma. Oncol Rep. 2009;21(3):673–9.
Qi L, Bart J, Tan LP, Platteel I, Sluis T, Huitema S, et al. Expression of miR-21 and its targets (PTEN, PDCD4, TM1) in flat epithelial atypia of the breast in relation to ductal carcinoma in situ and invasive carcinoma. BMC Cancer. 2009;9:163.
Sempere LF, Christensen M, Silahtaroglu A, Bak M, Heath CV, Schwartz G, et al. Altered MicroRNA expression confined to specific epithelial cell subpopulations in breast cancer. Cancer Res. 2007;67(24):11612–20.
Si ML, Zhu S, Wu H, Lu Z, Wu F, Mo YY. miR-21-mediated tumor growth. Oncogene. 2007;26(19):2799–803.
Qian B, Katsaros D, Lu L, Preti M, Durando A, Arisio R, et al. High miR-21 expression in breast cancer associated with poor disease-free survival in early stage disease and high TGF-beta1. Breast Cancer Res Treat. 2009;117(1):131–40.
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF. Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci USA. 2003;100(7):3983–8.
Charafe-Jauffret E, Ginestier C, Iovino F, Tarpin C, Diebel M, Esterni B, et al. Aldehyde dehydrogenase 1-positive cancer stem cells mediate metastasis and poor clinical outcome in inflammatory breast cancer. Clin Cancer Res. 2010;16(1):45–55.
Ginestier C, Hur MH, Charafe-Jauffret E, Monville F, Dutcher J, Brown M, et al. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell. 2007;1(5):555–67.
Liu R, Wang X, Chen GY, Dalerba P, Gurney A, Hoey T, et al. The prognostic role of a gene signature from tumorigenic breast-cancer cells. N Engl J Med. 2007;356(3):217–26.
Diehn M, Cho RW, Lobo NA, Kalisky T, Dorie MJ, Kulp AN, et al. Association of reactive oxygen species levels and radioresistance in cancer stem cells. Nature. 2009;458(7239):780–3.
Li X, Lewis MT, Huang J, Gutierrez C, Osborne CK, Wu MF, et al. Intrinsic resistance of tumorigenic breast cancer cells to chemotherapy. J Natl Cancer Inst. 2008;100(9):672–9.
Phillips TM, McBride WH, Pajonk F. The response of CD24(-/low)/CD44+ breast cancer-initiating cells to radiation. J Natl Cancer Inst. 2006;98(24):1777–85.
Yu F, Yao H, Zhu P, Zhang X, Pan Q, Gong C, et al. let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell. 2007;131(6):1109–23.
Aktas B, Tewes M, Fehm T, Hauch S, Kimmig R, Kasimir-Bauer S. Stem cell and epithelial-mesenchymal transition markers are frequently overexpressed in circulating tumor cells of metastatic breast cancer patients. Breast Cancer Res. 2009;11(4):R46.
Mani SA, Guo W, Liao MJ, Eaton EN, Ayyanan A, Zhou AY, et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell. 2008;133(4):704–15.
Shipitsin M, Campbell LL, Argani P, Weremowicz S, Bloushtain-Qimron N, Yao J, et al. Molecular definition of breast tumor heterogeneity. Cancer Cell. 2007;11(3):259–73.
Hennessy BT, Gonzalez-Angulo AM, Stemke-Hale K, Gilcrease MZ, Krishnamurthy S, Lee JS, et al. Characterization of a naturally occurring breast cancer subset enriched in epithelial-to-mesenchymal transition and stem cell characteristics. Cancer Res. 2009;69(10):4116–24.
Morel AP, Lievre M, Thomas C, Hinkal G, Ansieau S, Puisieux A. Generation of breast cancer stem cells through epithelial-mesenchymal transition. PLoS ONE. 2008;3(8):e2888.
Santisteban M, Reiman JM, Asiedu MK, Behrens MD, Nassar A, Kalli KR, et al. Immune-induced epithelial to mesenchymal transition in vivo generates breast cancer stem cells. Cancer Res. 2009;69(7):2887–95.
Bu Y, Lu C, Bian C, Wang J, Li J, Zhang B, et al. Knockdown of Dicer in MCF-7 human breast carcinoma cells results in G1 arrest and increased sensitivity to cisplatin. Oncol Rep. 2009;21(1):13–7.
Blower PE, Chung JH, Verducci JS, Lin S, Park JK, Dai Z, et al. MicroRNAs modulate the chemosensitivity of tumor cells. Mol Cancer Ther. 2008;7(1):1–9.
Pan YZ, Morris ME, Yu AM. MicroRNA-328 negatively regulates the expression of breast cancer resistance protein (BCRP/ABCG2) in human cancer cells. Mol Pharmacol. 2009;75(6):1374–9.
Tsuchiya Y, Nakajima M, Takagi S, Taniya T, Yokoi T. MicroRNA regulates the expression of human cytochrome P450 1B1. Cancer Res. 2006;66(18):9090–8.
Black PC, Brown GA, Inamoto T, Shrader M, Arora A, Siefker-Radtke AO, et al. Sensitivity to epidermal growth factor receptor inhibitor requires E-cadherin expression in urothelial carcinoma cells. Clin Cancer Res. 2008;14(5):1478–86.
Shah AN, Summy JM, Zhang J, Park SI, Parikh NU, Gallick GE. Development and characterization of gemcitabine-resistant pancreatic tumor cells. Ann Surg Oncol. 2007;14(12):3629–37.
Frederick BA, Helfrich BA, Coldren CD, Zheng D, Chan D, Bunn Jr PA, et al. Epithelial to mesenchymal transition predicts gefitinib resistance in cell lines of head and neck squamous cell carcinoma and non-small cell lung carcinoma. Mol Cancer Ther. 2007;6(6):1683–91.
Thomson S, Buck E, Petti F, Griffin G, Brown E, Ramnarine N, et al. Epithelial to mesenchymal transition is a determinant of sensitivity of non-small-cell lung carcinoma cell lines and xenografts to epidermal growth factor receptor inhibition. Cancer Res. 2005;65(20):9455–62.
Creighton CJ, Li X, Landis M, Dixon JM, Neumeister VM, Sjolund A, et al. Residual breast cancers after conventional therapy display mesenchymal as well as tumor-initiating features. Proc Natl Acad Sci USA. 2009;106(33):13820–5.
Gupta PB, Onder TT, Jiang G, Tao K, Kuperwasser C, Weinberg RA, et al. Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell. 2009;138(4):645–59.
Pusztai L. Markers predicting clinical benefit in breast cancer from microtubule-targeting agents. Ann Oncol. 2007;18 Suppl 12:xii15–20.
Seve P, Dumontet C. Is class III beta-tubulin a predictive factor in patients receiving tubulin-binding agents? Lancet Oncol. 2008;9(2):168–75.
Paradiso A, Mangia A, Chiriatti A, Tommasi S, Zito A, Latorre A, et al. Biomarkers predictive for clinical efficacy of taxol-based chemotherapy in advanced breast cancer. Ann Oncol. 2005;16 Suppl 4:iv14–19.
Tommasi S, Mangia A, Lacalamita R, Bellizzi A, Fedele V, Chiriatti A, et al. Cytoskeleton and paclitaxel sensitivity in breast cancer: the role of beta-tubulins. Int J Cancer. 2007;120(10):2078–85.
Cochrane DR, Howe EN, Spoelstra NS, Richer JK. Loss of miR-200c: a marker of aggressiveness and chemoresistance in female reproductive cancers. J Oncol. 2010;2010:821717.
Tamoxifen for early breast cancer: an overview of the randomised trials. Lancet. 1998;351(9114):1451–67.
Normanno N, Di Maio M, De Maio E, De Luca A, de Matteis A, Giordano A, et al. Mechanisms of endocrine resistance and novel therapeutic strategies in breast cancer. Endocr Relat Cancer. 2005;12(4):721–47.
Osborne CK, Schiff R. Estrogen-receptor biology: continuing progress and therapeutic implications. J Clin Oncol. 2005;23(8):1616–22.
Bhat-Nakshatri P, Wang G, Collins NR, Thomson MJ, Geistlinger TR, Carroll JS, et al. Estradiol-regulated microRNAs control estradiol response in breast cancer cells. Nucleic Acids Res. 2009;37(14):4850–61.
Adams BD, Cowee DM, White BA. The role of miR-206 in the epidermal growth factor (EGF) induced repression of estrogen receptor-alpha (ERalpha) signaling and a luminal phenotype in MCF-7 breast cancer cells. Mol Endocrinol. 2009;23(8):1215–30.
Pandey DP, Picard D. miR-22 inhibits estrogen signaling by directly targeting the estrogen receptor alpha mRNA. Mol Cell Biol. 2009;29(13):3783–90.
Zhao JJ, Lin J, Yang H, Kong W, He L, Ma X, et al. MicroRNA-221/222 negatively regulates estrogen receptor alpha and is associated with tamoxifen resistance in breast cancer. J Biol Chem. 2008;283(45):31079–86.
Miller TE, Ghoshal K, Ramaswamy B, Roy S, Datta J, Shapiro CL, et al. MicroRNA-221/222 confers tamoxifen resistance in breast cancer by targeting p27Kip1. J Biol Chem. 2008;283(44):29897–903.
Kondo N, Toyama T, Sugiura H, Fujii Y, Yamashita H. miR-206 Expression is down-regulated in estrogen receptor alpha-positive human breast cancer. Cancer Res. 2008;68(13):5004–8.
Adams BD, Furneaux H, White BA. The micro-ribonucleic acid (miRNA) miR-206 targets the human estrogen receptor-alpha (ERalpha) and represses ERalpha messenger RNA and protein expression in breast cancer cell lines. Mol Endocrinol. 2007;21(5):1132–47.
Hossain A, Kuo MT, Saunders GF. Mir-17-5p regulates breast cancer cell proliferation by inhibiting translation of AIB1 mRNA. Mol Cell Biol. 2006;26(21):8191–201.
Tsuji T, Ibaragi S, Hu GF. Epithelial-mesenchymal transition and cell cooperativity in metastasis. Cancer Res. 2009;69(18):7135–9.
No financial or material support
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wright, J.A., Richer, J.K. & Goodall, G.J. microRNAs and EMT in Mammary Cells and Breast Cancer. J Mammary Gland Biol Neoplasia 15, 213–223 (2010). https://doi.org/10.1007/s10911-010-9183-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10911-010-9183-z