To elucidate the role of innate immune responses in celiac disease, we investigated the effect of gliadin on blood monocytes from patients with celiac disease. Gliadin induced substantial TNF-α and IL-8 production by monocytes from patients with active celiac disease, lower levels by monocytes from patients with inactive celiac disease, and even lower levels by monocytes from healthy donors. In healthy donor monocytes gliadin induced IL-8 from monocytes expressing HLA-DQ2 and increased monocyte expression of the costimulatory molecules CD80 and CD86, the dendritic cell marker CD83, and the activation marker CD40. Gliadin also increased DNA binding activity of NF-κB p50 and p65 subunits in monocytes from celiac patients, and NF-κB inhibitors reduced both DNA binding activity and cytokine production. Thus, gliadin activation of HLA-DQ2+ monocytes leading to chemokine and proinflammatory cytokine production may contribute to the host innate immune response in celiac disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Celiac sprue is a chronic inflammatory disorder of the small intestine induced by dietary gluten or related rye and barley proteins in genetically predisposed individuals. More than 90% of patients with celiac disease express the HLA class II molecule HLA-DQ2, encoded by the DQA1*05 and DQB1*02 alleles, compared to 20–30% in the general population in Europe and Northern America. Symptoms and pathological changes in the small intestine, including villous atrophy, crypt hyperplasia, infiltration of inflammatory cells, and activation of the infiltrating inflammatory cells, are effectively treated by the exclusion of gluten from the diet (1–6).
Lamina propria CD4+ T cells that produce interferon (IFN)-γ in response to gliadin and intraepithelial CD8+ lymphocytes cytotoxic for epithelial cells expressing MHC class I polypeptide-related sequence A (MICA) appear to play fundamental roles in the pathogenesis of celiac disease (7–14). The activation of local as well as systemic humoral responses is reflected in the presence of circulating IgG and IgA antibodies to gliadin, endomysium, tissue transglutaminase (tTG), and other autoantigens (15–22).
Recent evidence indicates that innate immune response cells, including monocytes/macrophages and dendritic cells (DCs), also contribute to celiac disease pathogenesis. Gliadin fragments formed after pepsin digestion induce mouse peritoneal macrophages to produce TNF-α, IL-10, and RANTES and to release increased amounts of nitric oxide in the presence of IFN-γ (23–25). In addition, gliadin triggers NF-κB activation, interferon regulatory factor (IRF)-1 release, and STAT-1-mediated signal transduction in mouse monocytes (26). Importantly, gliadin-induced activation of human monocytes appears to be dependent on the stage of cell differentiation (23–25, 27–29). Gliadin fragments also induce phenotypic and functional maturation of human monocyte-derived dendritic cells (30).
To further understand the possible role of innate immune cells in celiac disease, we investigated whether peptic fragments of gliadin activate blood monocytes and the mechanism of that activation, and whether the gliadin-induced response of monocytes from celiac patients with active disease differs from the response of monocytes from patients with inactive disease on a gluten-free diet.
METHODS
Food Proteins
Peptic fragments of gliadin and soya protein were prepared using pepsin-agarose gel (ICN, Biomedicals, Inc., Ohio), as previously described (24, 30). Protein concentrations were measured by Bicinchoninic acid assay (BCA Protein assay, Pierce, Rockford, IL). All reagents were tested by the E-toxate test for lipopolysaccharide (LPS) (Sigma, St. Louis, MO) and shown to be below the limit of detection (2 pg/mL).
Study Subjects
The study population consisted of 54 patients with biopsy-proven celiac disease, including 14 symptomatic, untreated subjects [ESPGAN criteria; (31)] and 40 asymptomatic, treated (gluten-free diet) subjects. The control group consisted of 45 healthy blood donors. Patients were recruited after appropriate local ethics committee approval, and informed consent was obtained from all subjects.
Cells and Their Activation
Peripheral blood mononuclear cells were isolated by Ficoll paque (Amersham Biosciences, Piscataway, NJ) and incubated for 24 h in 24-well plates (Nunc, Roskilde, Denmark). Nonadherent cells were removed by washing, and the adherent monocytes were recultured in RPMI-1640 medium supplemented with 10% fetal calf serum, 2 mM l-glutamine, 100 μg/mL streptomycin, and 100 U/mL penicillin (Sigma). The monocytes were removed by scraping and analyzed by flow cytometry (BD FACSVantage SE, San Jose, CA) after staining with the indicated monoclonal antibodies (mAbs). The monocytes (70–80% CD14+) were cultured at a concentration of 1 × 106 cells/mL for 24 h in complete RPMI-1640 with gliadin (100–200 μg/mL) alone or with gliadin (100 μg/mL) plus IFN-γ (150 U/mL; R&D Biosciences, Minneapolis, MN), as described previously (7, 29, 32). In parallel monocyte cultures, the NF-κB inhibitors l-1-tosylamido-2-phenylethyl chloromethyl ketone (TPCK, 1.0–25.0 μM) or pyrrolidine dithiocarbamate (PDTC, 1.0–10.0 μM) (both Sigma) were added to the cells for 30 min prior to the addition of gliadin. Cultures were maintained at 37°C, 5% CO2 in a humidified incubator.
Flow Cytometric Analysis
To evaluate the effect of gliadin fragments on monocyte surface antigen expression, blood monocytes (5 × 105) were incubated with optimal concentrations of FITC-labeled mouse mAbs to HLA-DR, CD40, CD80, CD83, CD86 (all from BD Biosciences Pharmingen), and CD14 and MEM-18 (kindly provided by V. Hořejší, Charles University, Prague). Cells also were incubated with FITC- or PE-conjugated irrelevant antibodies of the same concentration and isotype. After two washings, cells were resuspended in ice-cold PBS with 0.1% NaN3 or fixed with 2% paraformaldehyde and analyzed by flow cytometry (FACSCalibur, BD Bioscience). Data were evaluated using CellQuest software (BD Biosciences). Staining with propidium iodide was performed to assess cell viability.
Preparation of Nuclear Extracts and Colorimetric NF-κB Assays
Nuclear extracts were prepared from purified monocytes stimulated for 90 min with gliadin digest (100 μg/mL) as mentioned earlier, with or without TPCK (1–25 μM) and PDTC (1–10 μM), using the Nuclear Extract Kit (Active Motif, Carlsbad, CA). NF-κB DNA binding activity was determined using the TransAM NF-κB family transcription factor assay (Active Motif), according to the manufacturer’s protocol. Briefly, microwells coated with a double-stranded oligonucleotide containing the NF-κB consensus sequence were incubated with nuclear extracts for 1 h at room temperature and washed with washing buffer. The wells containing captured active transcription factor were incubated for 1 h with mAb specific for p50 or p65 NF-κB subunits, then for 1 h with anti-rabbit IgG coupled to horseradish peroxidase and, after washing, exposed to developing solution for 10 min. Optical density was measured at 450 nm using a Titertec Multiscan MCC/340 (Flow Lab., Irvine, Scotland).
Measurement of IL-8 and TNF-α Proteins
The amounts of IL-8 and TNF-α in supernatants of monocytes (1 × 106 cells/mL) cultured for 24 h with or without gliadin (100 μg/mL) in the presence or absence of IFN-γ (150 U/mL) were determined by ELISA (R&D Systems, Europe, Germany) according to the manufacturer’s instructions.
Preparation of Genomic DNA
QIAamp spin columns (QIAGEN, GmbH, Hilden, Germany) were used for rapid DNA purification according to the specifications of the manufacturer. Briefly, lysis buffer and proteinase K were added to the monocyte cultures, which were mixed by pulse-vortexing and incubated for 10 min at 56°C. After the addition of 100% ethanol, the mixture was applied to the QIAamp Spin Column, centrifuged, and the filtrate discarded. After the QIAamp Spin Column was washed with washing buffers, elution buffer was applied, and the column was incubated at 15–20°C for 1 min. The DNA filtrate was collected by centrifugation and stored at 4°C.
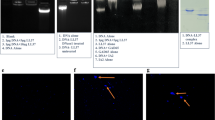
Analysis of the HLA Class II Molecule HLA-DQ2
Genes were typed by PCR with sequence-specific primers (33, 34). Oligonucleotide primers (Genovision, West Chester, PA) were designed to amplify the second exon of the class II genes. PCR amplification was carried out in a final volume of 10 μL containing: 6 ng/μL of genomic DNA, PCR Master Mix (200 μM of each dNTP, PCR buffer: 50 mM KCl, 1.5 mM MgCl2, 10 mM Tris-HCl pH 8.3, 0.001% w/v gelatin, 5% glycerol, 100 μg/mL cresol red) complete with 0.4 U/μL Taq polymerase and specific primer mix. Thermal cycling was accomplished by heating the sample at 94°C for 2 min and then running 10 cycles of 10 s at 94°C (denaturation), 60 s at annealing temperature 65°C, and 20 cycles of 10 s at 94°C (denaturation), 50 s at annealing temperature 61°C, 30 s at 72°C (extension). After 30 cycles, the extension was completed, and the samples were stored at 4°C.
IL-8 and TNF-α production by monocytes from patients with celiac disease. Monocytes isolated from patients with active celiac disease (n = 14), asymptomatic patients with celiac disease on a gluten-free diet (n = 40), and healthy donors (n = 45) were cultured for 24 h with gliadin (100 μg/mL) alone or with gliadin plus IFN-γ (150 U/mL). The amounts of (A) IL-8 and (B) TNF-α (mean ± SD) released into the supernatants were determined by ELISA. The level of significance for patients versus healthy donors are indicated as follows: ** P < 0.01, *** P < 0.001, oo P < 0.01, ooo P < 0.001.
Statistical Analysis
The differences among healthy donors and active and treated celiac patients groups were evaluted by the Kruskal–Wallis test and Mann–Whitney test for pairwise comparison. The levels of significance of the Mann–Whitney tests were adjusted according to Holm’s method (35). P values <0.05 were considered significant.
RESULTS
Cytokine Production by Gliadin-Stimulated Monocytes
Since monocytes are a major source of the chemokine IL-8 and the proinflammatory cytokine TNF-α, we evaluated blood monocytes from celiac patients and healthy donors for spontaneous and inducible IL-8 and TNF-α production. Monocytes from patients with active celiac disease and patients with inactive celiac disease on a gluten-free diet spontaneously released low levels of IL-8 (525 ± 404 pg/mL and 965 ± 682 pg/mL, respectively, P > 0.05), which were significantly greater than the amounts spontaneously released by monocytes from healthy control subjects (131 ± 202 pg/mL) (P < 0.001) (Fig. 1A). The addition of IFN-γ (150 U/mL) to the cultures did not significantly enhance IL-8 production by monocytes from patients or healthy donors. In sharp contrast, the addition of gliadin fragments (100 μg/mL) to the cultures induced monocytes from all three groups to secrete markedly higher levels of IL-8; monocytes from patients with active celiac disease produced more IL-8 (3365 ± 2451 pg/mL than monocytes from patients with inactive disease on a gluten-free diet (2645 ± 1723 pg/mL), which in turn produced more IL-8 than monocytes from healthy donors (1046 ± 783 pg/mL) (P<0.001). The production of TNF-α by monocytes from the three groups paralleled that of IL-8 (Fig. 1B). The addition of IFN-γ did not further enhance gliadin-induced IL-8 production, but did enhance TNF-α production. Similar results were obtained for IL-8 and TNF-α production when the monocytes were preincubated with IFN-γ (data not shown). In contrast to gliadin, soya proteins (100 μg/mL) with or without IFN-γ did not induce production of IL-8 or TNF-α by monocytes from the three groups.
Effect of gliadin on monocyte phenotype. Blood monocytes from healthy donors were incubated for 24 h in media or with IFN-γ alone, gliadin alone, or IFN-γ plus gliadin at the indicated concentrations and then analyzed by FACS for the expression of activation and differentiation markers. Staining after exposure to isotype-matched Ig is shown as a shaded histogram. Values are the percent monocytes that expressed the indicated marker from a representative donor (n = 3).
Gliadin Fragments Induce Differentiational Changes in Monocyte Surface Phenotype
Systemic immune activation is well documented in patients with celiac disease (36–39). In related studies (30), we have shown that gliadin fragments induce maturation of monocyte-derived dendritic cells. Therefore, we determined whether incubation of blood monocytes from healthy persons with gliadin fragments (100 μg/mL) induced changes in monocyte phenotype. Incubation of blood monocytes with gliadin, and to a lesser extent with IFN-γ (150 U/mL) alone, induced a slight increase in the percentage of cells expressing dendritic cell (DC) markers of maturation (CD83) and activation (CD80, CD40), compared to monocytes incubated in medium (Fig. 2). Importantly, incubation of monocytes with gliadin fragments plus IFN-γ induced a marked upregulation in the surface density and percentage of cells expressing CD80, CD86, CD83, and CD40 (Fig. 2). In contrast to the ability of gliadin to induce phenotypic changes, soya proteins (in the presence or absence of IFN-γ) did not alter monocyte surface protein expression (data not shown). This altered phenotype suggests a synergistic effect of gliadin and IFN-γ on monocyte activation.
Effect of HLA-DQ2 on Monocyte Cytokine Production
Since the majority of celiac patients are HLA-DQ2+, we studied the relationship between HLA genotype and monocyte IL-8 production in healthy control subjects. Monocytes from HLA-DQ2+ healthy donors secreted IL-8 spontaneously. However, in the presence of gliadin or IFN-γ, the monocytes released two- to three-fold more IL-8 than monocytes from HLA-DQ2− donors (Table I). These findings suggest that the HLA-DQ2+ genotype predisposes monocytes to increased IL-8 secretion.
Gliadin Induces Monocyte Cytokine Production via NF-κB Activation
We have shown perviously that NF-κB activation is involved in gliadin-induced cytokine production by monocyte-derived DCs and THP-1 cells (30). Therefore, we next determined whether the NF-κB signal transduction pathway is also involved in gliadin-induced cytokine production by monocytes. Stimulation of monocytes from celiac patients with gliadin digest (200 μg/mL) resulted in a marked increase in the binding activity of the NF-κB p50 and p65 subunits (P < 0.05). Incubation of monocytes from celiac patients with the NF-κB inhibitor TPCK reduced p50 and p65 binding to 20% and 54%, respectively (Fig. 3). Supporting the role of NF-κB in gliadin-induced cytokine production by monocytes from patients with active celiac disease, NF-κB inhibitors also reduced gliadin-induced IL-8 and TNF-α production (Fig. 4). In contrast, stimulation of monocytes from healthy subjects with gliadin caused weak NF-κB binding, which was completely inhibited by TPCK (25 μM/mL), and reduced cytokine production to background levels.
Gliadin induction of monocyte NF-κB p50 and p65 subunit binding. Monocytes from patients with active celiac disease and healthy donors were analyzed for NF-κB DNA binding activity after a 90 min incubation with gliadin (in the presence or absence of the NF-κB inhibitor TPCK). Data are presented as the fold-increase in DNA binding activity by gliadin-stimulated versus nonstimulated monocytes (mean ± SD for three separate experiments). * P < 0.05 corresponds to the binding activity of monocytes from celiac patients versus that of monocytes from healthy donors.
NF-κB inhibitors block gliadin-induced monocyte IL-8 and TNF-α production. Monocytes from patients with celiac disease and healthy donors were cultured for 24 h with gliadin in the presence or absence of the NF-κB inhibitors TPCK (1–25 μM) or PDTC (1–10 μM), and the amount of IL-8 and TNF-α released into the culture supernatants were measured by ELISA. Results are expressed as mean ± SD of cytokine production of monocytes from three patients with active celiac disease and six healthy donors.
DISCUSSION
We report for the first time that gliadin fragments stimulate cytokine production by blood monocytes from patients with celiac disease. The level of cytokine production varied with disease activity, as monocytes from patients with active celiac disease produced more gliadin-induced TNF-α and IL-8 than monocytes from patients with inactive disease, which in turn produced more inducible TNF-α and IL-8 than monocytes from healthy subjects. We also investigated whether IFN-γ further enhanced gliadin-induced TNF-α and IL-8 production by blood monocytes, since gliadin-induced T cells from patients with celiac disease produce increased levels of IFN-γ. IFN-γ plus gliadin upregulated TNF-α, but not IL-8, production. These data suggest that monocytes from celiac patients are activated in vivo, consistent with the increased levels of other cytokines, i.e., IL-6, that have been detected in patients with celiac disease (38, 39). Coincident with gliadin-induced monocyte activation, gliadin also induced a DC phenotype in blood monocytes, consistent with our earlier observation that gliadin promoted maturation of monocyte-derived DCs (30). Moreover, coincubation of gliadin with IFN-γ further enhanced the upregulation in the surface density and percentage of cells expressing CD80, CD86, CD83, and CD40, suggesting a synergistic effect of gliadin and IFN-γ on monocyte activation. Since our data indicates that gliadin enhances monocyte IL-8 release, and the majority of celiac patients are HLA-DQ2+ (1–6), we examined the ability of monocytes from HLA-DQ2+ and HLA-DQ2− healthy subjects to release IL-8 following exposure to gliadin. Monocytes from HLA-DQ2+, but not HLA-DQ2−, healthy subjects spontaneously released IL-8, and monocytes from HLA-DQ2+ persons released two- to threefold more IL-8 than monocytes from HLA-DQ2− persons. These findings suggest that the enhanced cytokine production by monocytes from patients with celiac disease is due, at least in part, to the activated HLA-DQ2+ population of monocytes.
Celiac disease is characterized by increased infiltration of lymphocytes into the lamina propria and epithelium. Increased production of IL-8 and TNF-α by cells of the innate immune system (monocytes, macrophages, DCs) within the lamina propria could contribute to the pathogenesis of celiac disease through their ability to recruit lymphocytes to the mucosa. In this regard, IL-8 is reported chemotactic for T lymphocytes (40, 41) at concentrations equivalent to those reported in our study (40). Moreover, TNF-α induces the upregulation of vascular ICAM-1, a receptor for the adhesion molecule LFA-1 on memory T cells (42–44). Thus, the production of TNF-α by gliadin-stimulated monocytes in the lamina propria would increase adhesion properties on mucosal vessels and thereby promote T-lymphocyte infiltration into the adjacent mucosa (in celiac disease). In this regard, increased mucosal expression of ICAM-1 has been reported in the intestinal mucosa of patients with celiac disease (42, 43). In addition, IFN-γ, which is abundant in the mucosa of patients with celiac disease (44, 45) and has been implicated in T-lymphocyte recruitment (42–44) (via ICAM-1 upregulation) also has an additive effect on TNF-α-induced T-cell migration (43, 46). Thus, IFN-γ alone, and together with TNF-α, could contribute to T-lymphocyte recruitment in celiac disease. Our findings, therefore, suggest an important potential mechanism whereby gliadin-stimulated release of IL-8 and TNF-α from mucosal innate immune cells contribute to the recruitment of T cells to the mucosa.
While our report has focused on the potential inflammatory effects of gliadin on innate immune cells of the monocytic lineage, other cells within the mucosa, such as epithelial cells, also likely contribute to the innate mucosal response to gliadin in celiac disease. To our knowledge, there are no data available concerning gliadin-induced IL-8 production by primary intestinal epithelial cells, but gliadin is known to alter intercellular tight junction and barrier function in intestinal epithelial cell lines (47, 48), possibly through the induction of zonulin from gliadin-stimulated epithelial cells (47) and macrophages (49).
Members of the NF-κB/Rel family, including the p50 and p65 subunits, regulate inflammatory and immune responses by inducing the expression of specific genes (50). We show here that exposure of monocytes to gliadin increased the binding activity of the p50 and p65 subunits. Increased p50 and p65 binding was more pronounced in monocytes from celiac patients with active disease than in monocytes from healthy subjects. This finding is consistent with our previous observations that gliadin activates the NF-κB p50/p65 complex in human monocyte-derived DCs (30) and THP-1 monocytes (29) and with observations by others that gliadin activation of macrophages is MyD88- and NF-κB-dependent (26, 49). In this regard, both NF-κB/DNA binding activity and p50/p65 nuclear levels are reported to be elevated in the inflamed mucosa of celiac patients (51). The involvement of the NF-κB/IκB complex in gliadin-induced activation of monocytes was confirmed using the NF-κB inhibitors TPCK, which prevents degradation of IκB inhibitor, and PDTC, which blocks dissociation of the NF-κB/IκB complex (52, 53). Both NF-κB inhibitors substantially reduced gliadin-induced NF-κB binding as well as TNF-α and IL-8 secretion by monocytes. Taken together, these findings underscore the likely involvement of monocyte/macrophage-derived innate immune reponses in celiac disease and suggest that NF-κB regulation should be considered in designing future therapeutic strategies.
In normal persons, wheat gluten, the major source of dietary gliadin, does not induce the T-cell activation or intestinal inflammation characteristic of celiac disease. In addition, intestinal macrophages isolated from healthy subjects do not respond to gliadin in vitro (Smythies, unpublished observation), which is consistent with the profound inflammation anergy characteristic of intestinal macrophages (54). Since intestinal macrophages are derived from blood monocytes (55), our findings suggest that local mucosal factors likely downregulate the ability of monocytes newly recruited to the mucosa to respond to gliadin. Since the “normal” downregulation of gliadin-induced monocyte activation and differentiation may be impaired in celiac disease, indentification of these mucosa-derived factors could have important therapeutic implications.
CONCLUSIONS
In summary, gliadin-derived fragments activate monocytes from celiac patients and HLA-DQ2+ monocytes from healthy donors. The higher activity of monocytes in celiac patients could augment gliadin-specific immune responses and thus contribute to the pathogenesis of celiac disease.
REFERENCES
Trier JS: Celiac sprue. N Engl J Med 325:1709–1719, 1991
Marsh MN: Gluten, major histocompatibility complex, and the small intestine. A molecular and immunbiologic approach to the spectrum of gluten sensitivity (celiac sprue). Gastroenterology 102:330–354, 1992
Maki M, Collin P: Coeliac disease. Lancet 349:1755–1759, 1997
Schuppan D: Current concepts of celiac disease pathogenesis. Gastroenterology 119:234–242, 2000
Sollid LM, Markussen G, Ek J, Gjerde H, Vartdal F, Thorsby E: Evidence for a primary association of celiac disease to a particular HLA DQ alpha/beta heterodimer. J Exp Med 169:345–350, 1989
Sollid LM: Coeliac disease: Dissecting a complex inflammatory disorder. Nat Rev Immunol 2:647–655, 2002
Nilsen EM, Jahnsen FL, Lundin KE, Johansen FE, Fausa O, Solid LM, Jahnsen J, Scott H, Brandtzaeg P: Gluten induced an intestinal cytokine response strongly dominated by interferon gamma in patients with celiac disease. Gastroenterology 115:551–563, 1998
Strober W, Fuss IJ: Gluten-sensitive enteropathy and other immunologically mediated enteropaties. In Mucosal Immunology, PL Ogra, J Mestecky, ME Lamm (eds). San Diego, CA, Academic, 1999, pp 1101–1128
Eiras P, Leon F, Camarero C, Lombardia M, Roldan E, Bootello A, Roy G: Intestinal intraepithelial lymphocytes contain a CD3-CD7+ subset expressing natural killer markers and a singular pattern of adhesion molecules. Scand J Immunol 52:1–6, 2000
Hue S, Mention JJ, Monteiro RC, Zhang S, Cellier C, Schmitz J, Verkarre V, Fodil N, Bahram S, Cerf-Bensussan N, Caillat-Zucman S: A direct role for NKG2D/MICA interaction in villous atrophy during celiac disease. Immunity 21:367–377, 2004
Meresse B, Chen Z, Ciszewski C, Tretiakova M, Bhagat G, Krausz TN, Raulet DH, Laner LL, Groh V, Spies T, Ebert EC, Green PH, Jabri B: Coordinated introduction by IL-15 of TCR-independent NKG2D signaling pathway converts CTL into lymphokine-activated killer cells in celiac disease. Immunity 21:357–366, 2004
Sollid LM: Intraepithelial lymphocytes in celiac disease: Licence to kill revealed. Immunity 21:303–304, 2004
Gianfrani C, Auricchio S, Troncone R: Adaptive and innate immune response in celiac disease. Immunol Lett 99:141–145, 2005
Ciccocioppo R, Di Sabatino A, Corazza GR: The immune recognition of gluten in celiac disease. Clin Exp Immunol 140:408–416, 2005
Valeski JE, Kumar V, Beutner EH, Lerner A, Chorzelski TP: Immunology of coeliac disease: Tissue and species specificity of endomysial and reticulin antibodies. Int Arch Allergy Appl Immunol 93:1–7, 1990
Karpati S, Burgin-Wolff A, Krieg T, Meurer M, Stolz W, Braun-Flaco O: Binding to human jejunum of serum IgA antibody from children with coeliac disease. Lancet 336:1335–1338, 1990
Volta U, Molinaro N, Fratangelo D, Bianchi FB: IgA antibodies to jejunum. Specific immunity directed against target organ of gluten-sensitive enteropathy. Dig Dis Sci 39:1924–1929, 1994
Ascher H, Hahn-Zoric M, Hanson LA, Kilander AF, Nilsson LA, Tlaskalova H: Value of serological markers for clinic diagnosis and population studies of coeliac disease. Scand J Gastroenterol 31:61–67, 1996
Karska K, Tuckova L, Steiner L, Tlaskalova-Hogenova H, Michalak M: Calreticulin—the potential autoantigen in celiac disease. Biochem Biophys Res Commun 209:597–605, 1995
Dieterich W, Laag E, Schopper H, Volta U, Ferguson A, Gillett H, Riecken EO, Schuppan D: Autoantibodies to tissue transglutaminase as predictor of coeliac disease. Gastroenterology 115:1317–1321, 1998
Stulik J, Hernychova L, Porkertova S, Pozler O, Tuckova L, Sanchez D, Bures J: Identification of new celiac disease autoantigens using proteomic analysis. Proteomics 3:951–956, 2003
Sollid LM, Jabri B: Is celiac disease an autoimmune disorder? Curr Opin Immunol 17:595–600, 2005
Tuckova L, Flegelova Z, Tlaskalova-Hogenova H, Zidek Z: Activation of macrophages by food antigens: Enhancing effect of gluten on nitric oxide and cytokine production. J Leukoc Biol 67:312–318, 2000
Tuckova L, Novotna J, Novak P, Flegelova Z, Kveton T, Jelinkova L, Zidek Z, Man P, Tlaskalova-Hogenova H, Bezouska K, Havlicek V: Activation of macrophages by gliadin fragments: Isolation and characterization of active peptide. J Leukoc Biol 71:625–631, 2002
Novak P, Man P, Tuckova L, Tlaskalova-Hoqenova H, Bezouska K, Havlicek V: Monitoring of in vitro deamidation of gliadin peptic fragments by mass spectrometry may reflect one of the molecular mechanisms taking place in celiac disease development. J Mass Spectrom 37:507–511, 2002.
De Stefano D, Maiuri MC, Iovine B, Ialenti A, Bevilacqua MA, Carnuccio R: The role of NF-kappa B, IRF-1, and STAT-1 alpha transcription factors in the iNOS gene induction by gliadin and IFN-gamma in RAW 264.7 macrophages. J Mol Med 84:65–74, 2006
Maiuri L, Ciacci C, Ricciardelli I, Vacca L, Raia V, Auricchio S, Picard J, Osman M, Quaratinos S, Londei M: Association between innate response to gliadin and activation of pathogenic T cells in coeliac disease. Lancet 362:30–37, 2003
Maiuri MC, De Stefano D, Mele G, Iovine B, Bevilacqua MA, Greco L, Auricchio S, Carnuccio R: Gliadin increases iNOS gene expression in interferon-gamma-stimulated RAW 264.7 cells through a mechanism involving NF-kappa B. Naunyn Schmiedebergs Arch Pharmacol 368:63–71, 2003
Jelinkova L, Tuckova L, Cinova J, Flegelova Z, Tlaskalova-Hogenova H: Gliadin stimulates human monocytes to production of IL-8 and TNF-alpha through a mechanism involving NF-kappa B. FEBS Lett 571:81–85, 2004
Palová-Jelínková L, Roková D, Pecharová B, Bartova J, Sediva A, Tlaskalova-Hogenova H, Spisek R, Tuckova L: Gliadin fragments induce phenotypic and functional maturation of human dendritic cells. J Immunol 175:7038–7045, 2005
Walker-Smith JA, Guandalini S, Schmitz J, Working Group of European Society of Paediatric Gastroenterology and Nutrition: Revised criteria of diagnosis of coeliac disease. Arch Dis Child 65:909–911, 1990
Mazzarella G, Maglio M, Paparo F, Nardone G, Stefanile R, Greco L, van de Wal Y, Kooy Y, Koning F, Auricchio S: An immunodominant DQ-8 restricted gliadin peptide activates small intestinal immune response in in vitro cultured mucosa from HLA DQ-8 positive but not HLA-DQ8 negative coeliac patients. Gut 52:57–62, 2003
Olerup O, Zatterquist H: HLA DR typing by PCR amplification with sequence-specific primers (PCR-SSP) in 2 h: An alternative to serological DR typing in clinical practice including donor-recipient matching in cadaveric transplantation. Tissue Antigens 39:225–235, 1992
Zetterquist H, Olerup O: Identification of HLA-DRB1*04, DRB1*07 and DRB1*09 alleles by PCR amplification with sequence-specific primers (PCR-SSP) in 2 h. Hum Immunol 34:64–74, 1992
Holm S: A simple sequentially rejective multiple test procedure. Scand J Stat 6:65–70, 1979
Jelinkova L, Tuckova L, Sanchez D, Krupickova S, Pozler O, Nevoral J, Kotalova R, Tlaskalova-Hogenova H: Increased levels of cirulating ICAM-1, E-selectin and IL-2 receptors in celiac disease. Dig Dis Sci 45:398–402, 2000
Merendino RA, Di Pasquale G, Sturniolo GC, Ruello A, Albanese V, Minciullo PL, Di Mauro S, Gangemi S: Relationship between IL-18 and sICAM-1 serum levels in patients affected by coeliac disease: Preliminary considerations. Immunol Lett 85:257–260, 2003
Romaldini CC, Barbieri D, Okay TS, Raiz R Jr, Cancado EL: Serum soluble interleukin-2 receptor, interleukin-6, and tumor necrosis factor-alpha levels in children with coeliac disease: Response to treatment. J Pediatr Gastroenterol Nutr 35:513–517, 2002
Cataldo F, Lio D, Marino V, Scola L, Crivello A, Corazza GR, Working Group of the SIGEP, Working Group of “Club del Tenue”: Plasma cytokine profiles in patients with celiac disease and selective IgA deficiency. Pediatr Allergy Immunol 14:320–324, 2003
Pace E, Gjomarkaj M, Melis M, Profita M, Spatafora M, Vignola A, Bonsignore G, Mody CH: Interleukin-8 induces lymphocyte chemotaxis into the pleural space. Role of pleural macrophages. Am J Respir Crit Care Med 159:1592–1599, 1999
Roth SJ, Carr MW, Springer TA: C-C chemokines, but not the C-X-C chemokines interleukin-8 and interferon-g inducible protein-10, stimulate transendothelial chemotaxis of T lymphocytes. Eur J Immunol 25:3482–3488, 1995
Issekutz AC, Meager A, Otterness I, Issekutz TB: The role of tumour necrosis factor-a and IL-1 in polymorphonuclear leucocyte and T lymphocyte recruitment to joint inflammation in adjuvant arthritis. Clin Exp Immunol 97:26–32, 1994
Issekutz TB: Effects of six different cytokines on lymphocyte adherence to microvascular endothelium and in vivo lymphocyte migration in the rat. J Immunol 144:2140–2146, 1990
Ding Z, Xiong K, Issekutz TB: Chemokines stimulate human T lymphocyte transendothelial migration to utilize VLA-4 in addition to LFA-1. J Leukoc Biol 69:458–466, 2001
Sturgess RP, Macartney, JC, Makgoba MW, Hung CH, Haskard DO, Ciclitira PJ: Differential upregulation of intercellular adhesion molecule-1 in coeliac disease. Clin Exp Immunol 82:489–492, 1990
Pober JS, Gimbrone MA Jr, Lapierre LA, Mendrick DL, Fiers W, Rothlein R, Springer TA: Overlapping patterns of activation of human endothelial cells by interleukin-1, tumor necrosis factor, and immune interferon. J Immunol 137:1893–1896, 1986
Drago S, El Asmar R, Di Pierro M, Grazia Clemente M, Tripathi A, Sapone A, Tacar M, Iacono G, Carroccio A, D’Agate C, Not T, Zampini L, Catáis C, Fasano A: Gliadin, zonulin and gut permeability: Effects on celiac and non-celiac intestinal mucosa and intestinal cell lines. Scand J Gastroenterol 41:408–419, 2006
Sander GR, Cummins AG, Henshall T, Powell BC: Rapid disruption of intestinal barrier function by gliadin involves altered expression of apical junctional proteins. FEBS Lett 579:4851–4855, 2005
Thomas KE, Sapone A, Fasano A, Vogel SN: Gliadin stimulation of murine macrophage inflammatory gene expression and intestinal permeability are MyD88-dependent: role of the innate immune response in celiac disease. J Immunol 176:2512–2521, 2006
Ghosh S, May MJ, Kopp EB: NF-kappa B and Rel proteins: Evolutionarily conserved mediators of immune responses. Ann Rev Immunol 16:225–260, 1998
Maiuri MC, De Stefano D, Mele G, Fecarotta S, Greco L, Troncone R, Carnuccio R: Nuclear factor kappa B is activated in small intestinal mucosa of celiac patients. J Mol Med 81:373–379, 2003
Baldwin AS: The NF-kappa B and I kappa B proteins: New discoveries and insights. Ann Rev Immunol 14:649–683, 1996
Snyder JG, Prewitt R, Campsen J, Britt LD: PDTC and Mg132, inhibitors of NF-kappa B, block endotoxin induced vasodilatation of isolated rat skeletal muscle arterioles. Shock 17:304–307, 2002
Smythies LE, Sellers M, Clements RH, Mosteller-Barnum M, Meng G, Benjamin WH, Orenstein JM, Smith PD: Human intestinal macrophages display profound inflammatory anergy despite avid phagocytic and bacteriocidal activity. J Clin Invest 115:66–75, 2005
Smythies LE, Maheshwari A, Clements R, Eckhoff D, Novak L, Vu HL, Mosteller-Barnum M, Sellers M, Smith PD: Mucosal IL-8 and TGF-b recruit blood monocytes: Evidence for cross-talk between the lamina propria stroma and myeloid cells. J Leuk Biol 80:492–499, 2006
ACKNOWLEDGMENTS
This work was supported by the Grant Agency of the Czech Republic (Grants 310/05/2245 and 310/03/H147); the Grant Agency of the Czech Academy of Sciences (Grants AA5020210, AA5020205, 1QS500200572, and B5020407); the Grant Agency of the Ministry of Education (MSM 0021620814, GA UK 30/2006), Mininstry of Agriculture 1B53002 and Institutional Research Concept (Grant AV0Z50200510); and the National Institutes of Health (U.S.) (DK-74033, DK-47322, DK-54495, HD-41361, and DK-064400); the Crohn’s and Colitis Foundation of America; and the Research Service of the Veterans Administration
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
CINOVA, J., PALOVÁ-JELÍNKOVÁ, L., SMYTHIES, L. et al. Gliadin Peptides Activate Blood Monocytes from Patients with Celiac Disease. J Clin Immunol 27, 201–209 (2007). https://doi.org/10.1007/s10875-006-9061-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10875-006-9061-z