Abstract
The p7 membrane protein encoded by hepatitis C virus (HCV) assembles into a homo-hexamer that selectively conducts cations. An earlier solution NMR structure of the hexameric complex revealed a funnel-like architecture and suggests that a ring of conserved asparagines near the narrow end of the funnel are important for cation interaction. NMR based drug-binding experiments also suggest that rimantadine can allosterically inhibit ion conduction via a molecular wedge mechanism. These results suggest the presence of dilation and contraction of the funnel tip that are important for channel activity and that the action of the drug is attenuating this motion. Here, we determined the conformational dynamics and solvent accessibility of the p7 channel. The proton exchange measurements show that the cavity-lining residues are largely water accessible, consistent with the overall funnel shape of the channel. Our relaxation dispersion data show that residues Val7 and Leu8 near the asparagine ring are subject to large chemical exchange, suggesting significant intrinsic channel breathing at the tip of the funnel. Moreover, the hinge regions connecting the narrow and wide regions of the funnel show strong relaxation dispersion and these regions are the binding sites for rimantadine. Presence of rimantadine decreases the conformational dynamics near the asparagine ring and the hinge area. Our data provide direct observation of μs–ms dynamics of the p7 channel and support the molecular wedge mechanism of rimantadine inhibition of the HCV p7 channel.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The viroporin p7 encoded by hepatitis C virus (HCV) has been pursued as a potential therapeutic target against HCV infection (Griffin et al. 2008; Luscombe et al. 2010; Steinmann and Pietschmann 2010). The 63-residue p7 is a cleavage product between the structural protein E2 and non-structural protein NS2 and is expressed in the Endoplasmic Reticulum (ER) (Haqshenas et al. 2007; Moradpour et al. 2007). The current consensus in HCV research is that p7 is important for viral infectivity in vivo: clones with deleted or mutated p7 could not produce viral particles in chimpanzees (Sakai et al. 2003). Deleting p7 completely abrogated production of infectious viruses in the cells, but did not affect RNA replication or protein production, suggesting a role post RNA-replication (Jones et al. 2007; Steinmann et al. 2007).
The p7 plays an important role in the viral life cycle by at least two reported functions: protein–protein interaction (Gouklani et al. 2013; Shanmugam and Yi 2013) and cation permeation (Steinmann and Pietschmann 2010). It has been shown that p7 interacts with the NS2 protein to recruit the core protein to the site of the capsid assembly (Popescu et al. 2011). Other proposed cellular functions of p7 include mediating virus budding into the ER lumen (Gentzsch et al. 2013), or modulating the pH of intracellular vesicles during virus egress through its cation permeation function (Wozniak et al. 2010). The channel activity of p7 could potentially depolarize the ER membrane to facilitate membrane curvature formation during virus budding (Agarkova et al. 2008; Nieva et al. 2012). Support for the cation permeation function of p7 comes from observations in lipid bilayers where p7 oligomerizes to form channels and conducts current (Griffin et al. 2003; Montserret et al. 2010; Pavlovic et al. 2003; Premkumar et al. 2004). In addition to assays in lipid bilayers, two-electrode voltage clamp (TEVC) of Xenopus oocytes expressing p7 (OuYang et al. 2013) and proton flux assays in liposomes (Gan et al. 2014) have reported ion conduction functionality of p7. These studies have reported that p7 conducts Na+, K+, H+ and Ca2+. Moreover, the channel activity can be inhibited by rimantadine, long alkylchain iminosugar derivatives and hexamethylene amiloride in vitro, with varying reported efficacies (Griffin et al. 2003; Montserret et al. 2010; Pavlovic et al. 2003; Premkumar et al. 2004).
At the molecular and structural level, not much is known about the channel mechanism, though significant amount of structural information is available for elucidating the mechanism. Earlier NMR studies found that the p7 monomer has three helical segments: two in the N-terminal half of the sequence and one near the C-terminus (Cook and Opella 2011; Montserret et al. 2010). Single-particle electron microscopy (EM) characterization of the p7 oligomer from HCV genotype 2a (JFH-1 strain) showed that the p7 forms a 42 kDa hexamer and adopts a flower-like shape (Luik et al. 2009). Recently, a detailed structure of the p7 hexamer (from genotype 5a) in DPC micelles was determined using a solution NMR system (OuYang et al. 2013). The NMR structure of the p7 hexamer shows a funnel-like architecture with six minimalist chains, where the individual monomers do not have tertiary contacts within themselves, but neighboring monomers are intertwined to form a hexameric channel. Each monomer contains three helical segments, H1, H2 and H3. The H1 and H2 helices of each monomer form the narrow and wide parts of the pore, respectively, and the H3 helix wraps the channel from the outside through interactions with H2 of i + 2 and H1 of i + 3 monomers. Several conserved residues such as Ile6, Asn9, Leu24 and Arg35 line the pore of the channel. Ile6 forms the narrowest part of the channel. A narrow hydrophobic ring at this position likely serves as a hydrophobic constriction, which prevents water from freely passing through. The carboxamide from Asn9 forms a ring just above the Ile6 ring, and could be involved in coordinating cations.
Channel recording experiments showed that p7 has higher selectivity for Ca2+ than K+/Na+ (Premkumar et al. 2004), i.e., the channel is permeable to K+/Na+, but in the presence of Ca2+, the K+/Na+ conductance is inhibited and the channel instead prefers to conduct Ca2+. This property has been observed for other channels that have selectivity for divalent cations (Hou et al. 2012). The divalent cation channels often have selectivity rings formed by either carboxylates or carboxamides, probably because they are used to bind and replace the water shell. For examples, the CorA Mg2+ channel has a pentameric ring of asparagines (Lunin et al. 2006) and the Ca2+ release-activated Ca2+ (CRAC) channel Orai has a hexameric ring of glutamic acids (Hou et al. 2012). Since Asn9 is highly conserved across the HCV genotypes (Carrere-Kremer et al. 2002), i.e., residue 9 is asparagine in all strains except in genotype 2 viruses, where it is histidine, the ring of carboxamides near the tip of the funnel is a natural suspect for binding Ca2+. Indeed, mutating residue 9 to alanine caused ~70 % reduction in current in the TEVC assay (OuYang et al. 2013). An unresolved issue is that the size of the Asn9 ring (inner diameter ~7 Å) is significantly larger than those in, for e.g., the CRAC and CorA crystal structures (Supplementary Fig. S1). At this size, the carboxamide ring cannot provide coordination of Ca2+. We thus hypothesize that in the solution state, the Asn9 ring size fluctuates significantly due to conformational exchange between the presumed open and closed states of the channel.
An indirect indication of equilibrium exchange between two states comes from earlier NMR titration experiment for investigating binding of the inhibitor rimantadine to the p7 channel. It was shown that in the absence of rimantadine, the Ile6 methyl resonance is split into an intense and weak peak, possibly corresponding to the open and closed state, respectively, and that increasing the drug concentration shifts the equilibrium that made the weak peak stronger (OuYang et al. 2013). Rimantadine does not directly block the tip of the channel. Instead, it binds to six equivalent hydrophobic pockets (due to the sixfold symmetry of the p7 channel) between the pore-forming and peripheral helices far away from the Ile6 ring. In each site, Leu52, Leu53, and Leu56 from H3 of the i monomer, Val25 and Val26 from H2 of the i + 2 monomer, and Phe20 from H2 of the i + 3 monomer together form a deep hydrophobic pocket that wraps around the adamantane cage of the drug. An important property of the drug binding site is that it consists of elements from different helical segments and from different monomers. The rimantadine may thus act as a molecular wedge that prevents the dynamic “breathing” of the channel required for ion conduction.
In order to investigate the presence of intrinsic conformational dynamics of the p7 channel, we carried out 15N Carr–Purcell–Meiboom–Gill (CPMG) relaxation dispersion measurements to identify regions of the channel that are subject to μs–ms time scale chemical shift exchange as well as proton exchange measurement to characterize the solvent accessibility profile of the channel. We also performed CPMG measurements in the presence of rimantadine to probe the dynamics in the presence of an allosteric inhibitor.
Materials and methods
NMR sample preparation
The p7 sequence for this study was from genotype 5a and was mutated at non-conserved sites as described before (OuYang et al. 2013) for the following purposes: Thr1 is replaced with glycine to avoid the side reaction of cyanogen bromide cleavage, Ala12 is replaced with serine to simplify the backbone assignment, and the three cysteines at positions 2, 27, and 44 are replaced with Ala, Thr, and Ser to avoid the sulfide bond formation during the reconstitution, respectively. The p7 was fused to the His9–trpLE sequence in pMM-LR6 vector (a gift from S.C. Blacklow, Harvard Medical School), expressed and purified from E. coli BL21 (DE3) inclusion bodies as described before (OuYang et al. 2013). Briefly, transformed E. coli strain BL21 (DE3) cells were grown at 37 °C to an absorbance of ~0.7 at 600 nm and were induced at 25 °C with 150 μM isopropyl β-d-thiogalatopyranoside. Cells were harvested after overnight growth and lysed by sonication in lysis buffer (50 mM Tris, 200 mM NaCl, pH 8.0). Protein was then extracted from the inclusion bodies in denaturing conditions (1 % Triton X-100, 6 M Guanidine, 50 mM Tris, 200 mM NaCl, pH 8.0), and purified by nickel affinity chromatography. The 14-kDa trpLE peptide was liberated from the fusion protein by cyanogen bromide cleavage in 70 % formic acid, and separated by reverse phase high-performance liquid chromatography (RP-HPLC) in a PROTO 300 C-18 column (Higgins Analytical) with a gradient of 40 % acetonitrile (0.1 % trifluoroacetic acid) to 60 % acetonitrile (0.1 % trifluoroacetic acid) (Fig. 1a). Pure lyophilized p7 peptide was dissolved in 6 M guanidine and dodecylphosphocholine (DPC), and reconstituted by dialyzing against the NMR buffer (25 mM MES, pH 6.5) to remove the denaturant overnight. To remove excess detergent, the sample was passed through fast protein liquid chromatography (FPLC) in a Superdex 200 10/300 GL column (GE Healthcare) using buffer containing 3 mM DPC, 100 mM NaCl, and 25 mM MES (pH 6.5) (Fig. 1b). Protein containing fractions were collected, dialyzed against NMR buffer to remove salt, and concentrated to yield an NMR sample (Fig. 1c). A typical NMR sample that generates high quality NMR spectra contains 0.8 mM p7, 50 mM DPC, 25 mM MES (pH 6.5) (Fig. 1d). The hexameric formation of p7 complex was identified by electron microscopy. Full deuteration of p7 protein required growth in D2O and substituting appropriate reagents in the bacterial media during growth.
Purification and TROSY-HSQC spectra of p7. a The p7 peptide was purified from a mixture of trpLE, p7, trpLE-p7 fusion protein by HPLC using a C18 column and a gradient of acetonitrile. b FPLC elution profile after applying buffer containing 3 mM DPC, 100 mM NaCl, and 25 mM MES (pH 6.5) and running through a Superdex 200 10/300 GL column (GE Healthcare) to remove excess detergent. c SDS-PAGE of the FPLC elution fraction showing the purity of the p7 in the final NMR sample. d Two-dimensional 15N TROSY-HSQC spectrum of 2H–15N-labeled p7 hexamer reconstituted in DPC micelles at pH 6.5 (recorded at 1H frequency of 600 MHz). The labels are residue-specific assignments of the amide resonances
To make protein sample containing drug, rimantadine dissolved in buffer (3 mM DPC, 25 mM MES, pH 6.5) was added to the concentrated p7 sample such that the final rimantadine concentration is 5 mM.
NMR spectroscopy
All NMR experiments were recorded at 30 °C using Agilent 600 MHz, Agilent 700 MHz, or Bruker 900 MHz spectrometer with cryogenic probes.
The water-amide proton exchange rates were measured on the Bruker 900 MHz using a uniformly 2H-, 15N-labeled protein sample. For this measurement, a series of interleaved 2D TROSY-HSQC having the water exchange (WEX) filter element (Mori et al. 1994, 1996a) in the beginning of the TROSY-HSQC pulse sequence were recorded. Moreover, the experiment has been implemented to achieve optimal water suppression for the use with cryogenic probes (Supplementary Fig. S2). The interleaved experiment recorded eight spectra with mixing times (Tm) of 0, 10, 20, 30, 40, 100, 140, 180, and 220 ms. Magnetization transfer from water to the residue-specific amide sites in the protein, which is a direct measure of exchange, was monitored as increasing NMR signals with longer Tm in the 2D TROSY-HSQC spectrum.
A 2D TROSY-HSQC version of a single quantum Carr-Purcell-Meiboom-Gill (CPMG) relaxation dispersion (RD) experiment (Loria et al. 1999; Palmer et al. 2001) was performed using the pulse sequence of Vallurupalli et al. (2008). CPMG RD experiments measure the 15N transverse relaxation as a function of the repetition rates of CPMG refocusing pulses, νCPMG. For p7, the experiments were recorded on an Agilent 600 MHz and an Agilent 700 MHz spectrometer. The CPMG experiments were recorded in an interleaved manner with 9 different CPMG frequencies, 100, 200, 300, 400, 500, 600, 700, 800, 900 Hz and two repeat experiments at νCPMG = 200 and 700 Hz (for error analysis) with a CPMG constant time delay of 20 ms. Spectra were recorded with complex points of 1024 and 128 in the t2 and t1 dimensions, respectively.
Data analysis
Spectra were processed using NMRPipe software system (Delaglio et al. 1995) and visualized using either Sparky (Goddard and Kneller 2008) or CcpNmr (Vranken et al. 2005) software. Residue specific assignment of p7 backbone amide resonances as reported previously (OuYang et al. 2013) was used for data analysis. For water-proton exchange experiments, peak intensities were quantified using Sparky. Peak intensities were normalized as a percentage of intensity of amide resonances from a reference spectrum recorded with no WEX element. Initial rate of change of normalized peak intensity with respect to mixing time was calculated for each residue to derive proton exchange rates. Uncertainty in slope fitting was used to determine error bars.
For CPMG experiments, peak intensities were quantified using NMRPipe. Effective relaxation rate, R2,eff was calculated from peak intensities according to the equation (Mulder et al. 2001)
where T cp is the CPMG constant time delay, I (0) and I (νCPMG) are peak intensities in the absence and presence of a CPMG field, respectively. The relaxation dispersion data was analyzed by fitting to a Carver-Richards two-site exchange model (Carver and Richards 1972) using an in-house software. A global fit was performed, for the residues that showed non-flat RD curves, assuming uniform exchange rate, kex and population of the excited state, pb, as described previously (Korzhnev et al. 2005).
Results
Water-amide proton exchange experiment
We determined water-amide proton exchange rates for the hexameric p7 complex to identify solvent-accessible regions of the protein, and to characterize fast exchange interactions between the protein and the solvent. The NMR experiment used is a 2D TROSY-HSQC (Pervushin et al. 1998) with the water exchange (WEX) filter (Hwang et al. 1997; Mori et al. 1994, 1996a, b) in the beginning of the experiment (“Materials and methods”; Supplementary Fig. S2). In this experiment, water protons are selectively excited while the protein proton magnetization is purged. The water magnetization is then transferred, for a variable mixing time (Tm), to the protein backbone amide sites through proton exchange and/or nuclear Overhauser effect (NOE) for detection. Completely deuterated protein sample was used to eliminate NOE-related artifacts due to unwanted excitation of the sidechain alpha or beta protons. We recorded the spectrum at 8 different mixing times in an interleaved manner. At Tm = 0, no 1H–15N correlation peaks are seen because no magnetization is transferred from water proton to amide (Fig. 2a). With increasing values of Tm, a subset of amide peaks began to appear (Fig. 2a), and the rates at which these peaks appear were quantified. The normalized peak intensity as a function of Tm was plotted and the slope of the initial linear region was calculated and reported as the proton exchange rate (Fig. 2b, Supplementary Table S1). The initial slope was calculated for Tm range between 0 and 40 ms for most of the residues, with only few residues where it was calculated over the 100 ms range (Supplementary Table S1). We emphasize that within this time range, the proton exchange process is dominant while the NOE contribution becomes significant beyond this time range. The proton exchange rates we get are relative and semi-quantitative, and can be used for comparison between the residues.
Peak intensities and proton exchange rate. a Peak intensity of amide resonances as a function of mixing time (Tm) for selected residues of p7. Black lines represent residues in the helical regions and gray lines represent ones in the loop region. b Relative water-amide proton exchange rates for each residue of the p7 channel (calculated from normalized peak intensities as described in “Materials and methods”). Gray bars represent residues in the loop region and black bars represent residues in the helical regions
The overall solvent exchange profile agrees with the funnel architecture of the p7 hexamer, e.g., most of the residues lining the cavity and near the wide and narrow mouths of the funnel show higher exchange rates (Figs. 2b, 3). Among the helical segments, H1 (residue 5–16) and parts of H2 (residues 20–41) that line the pore or face the solvent on the top wide region of the channel show fast exchange, indicating that these regions are water accessible (Fig. 3). In contrast, H3 (residue 48–58) does not show any proton exchange. This is consistent with the hexameric structure of p7 determined in DPC micelles because in the structure, H1 and H2 form the channel pore that is largely hydrophilic. It is expected from the structure that the H3 helices, which face the detergent, show little to no exchange. In addition to the helical segments, the loop region between H2 and H3 (residues 42–47) shows fast exchange. This is also expected because this loop is at the wide mouth of the funnel. Furthermore, the amide protons in the loop region exchange faster because they are not protected by hydrogen bonding.
If we compare the relative exchange rates of the pore lining residues in H1 and H2 (residues 20–34 are pore lining), we find that residues in H1 have faster exchange rates (Fig. 2b). Although both helical segments are facing the polar cavity, the H2 contains more hydrophobic residues, e.g., Val26, Leu28, Leu36 and Val37 (Supplementary Table S1). Slower exchange in this helical segment thus suggests less hydration around H2. On the other hand, faster exchange in H1 could be due to looser helix–helix packing and thus weaker hydrogen bonding in H1 compared to H2. From simple inspection of the structure, the H2 helices are indeed more tightly packed than the H1 helices.
Micro-milli seconds dynamics of the p7 channel
Conformational fluctuations in proteins can occur over a wide range of time scales. To probe the dynamics of the p7 channel on a biologically relevant time scale we performed backbone 15N RD experiments. These experiments measure dynamics on the μs–ms time scale, and the measured relaxation rate, R2,eff, is depended, assuming conformational exchange between two states, on the populations (pa,b) and the chemical shift difference (Δω) between the two interconverting conformations as well as the rate of interconversion (kex) (Palmer et al. 2001). 2D TROSY-HSQC 15N-CPMG experiments were recorded on a 2H/15N-labeled p7 sample at two different magnetic fields, 600 and 700 MHz. The transverse relaxation rate, R2,eff, was plotted as a function of νCPMG frequency and the resulting RD profiles were fit employing a two state model using the Carver–Richards formalism (Carver and Richards 1972). Several residues such as Asn9 and Val25 were precluded from analysis due to peak overlap.
Three residues, Val7, Leu8 and Phe19 have non-flat curves that show significant chemical exchange, with ΔR2 > 3 Hz for 600 MHz RD, where ΔR2 = R2,eff (νCPMG = 100 Hz) − R2,eff (νCPMG = 900 Hz) (Figs. 4a, 5 top panel). The individual kex rates for Val7, Leu8 and Phe19 are 1197 ± 100, 1236 ± 130 and 746 ± 128 s−1 respectively (Supplementary Table S2). For the individual fits of these three residues, the data was insufficient to get reasonable and reliable excited state population percentages and delta-omega values. A global fit was performed on these three residues, which yielded a population of 10 ± 3 % for the excited state and an exchange rate, kex, of about 1031 ± 79 s−1 between the two states (Supplementary Table S2). Interestingly, Val7 and Leu8 are located near the N-terminal tip of the channel (Fig. 4b), which also contains some of the channel elements that are thought to be important for ion conduction. The proton exchange data as described above also suggest that this region of the channel is dynamic, albeit on a different time scale compared to the relaxation dispersion data.
CPMG relaxation dispersion analysis of the p7 channel. a Difference between R2,eff at CPMG frequency 100 and 900 Hz (ΔR2) for each residue of the p7 channel recorded at two different magnetic fields, 600 MHz (red triangles) and 700 MHz (blue squares). Error bars were calculated based on uncertainty in repeat CPMG experiments. b Mapping residues that show significant relaxation dispersion, Val7, Leu8 and Phe19 on the p7 structure (PDB: 2M6X). These three residues are marked by red spheres
CPMG relaxation dispersion curves for p7 in the presence and absence of rimantadine. CPMG relaxation dispersion curves for three residues, Val7, Leu8 and Phe19 at two different magnetic fields, 600 MHz (red) and 700 MHz (blue) in the apo state (top panel) and in the presence of 5 mM rimantadine (bottom panel). Circles represent data points and the dotted line represents the fit for two-state model
Furthermore, Phe19 is located in the flexible hinge region between H1 and H2 (Fig. 4b). This region is important from a structural perspective as near this region, i.e. at Phe20, H3 forms tertiary contacts with H2. In addition, rimantadine binding site is also near this hinge region. As such, interactions in this region are important for maintaining the overall conformation of the channel and any changes in the channel conformation would affect resides at this hinge region. Significant chemical exchange at Phe19 suggests the presence of a conformational exchange. In our data we do not see much conformational exchange in the H2 helix. This is probably because H2 is more rigid and packed (as suggested by the proton exchange data), and this part of the cavity is more stable. Indeed, our experiments probing dynamics at different time scales suggest that H1 segment is more dynamic compared to other segments.
Dynamics of the p7 channel in the presence of rimantadine
We also recorded 2D TROSY-HSQC 15N-CPMG experiments on p7 in the presence of 5 mM rimantadine. Addition of rimantadine causes significant chemical shift changes in the p7 spectra (OuYang et al. 2013). CPMG data was analyzed in a similar manner as the apo state of p7. In the presence of drug, only two residues, Leu8 and Phe19, show significant relaxation dispersion (Fig. 5, bottom panel). Relaxation dispersion curve for Val7, which has significant chemical exchange in the apo state, is completely flat. Individual fits for Leu8 and Phe19 give kex values of 1514 ± 189 and 67 ± 182 s−1 respectively (Supplementary Table S2). Since the kex are in the different time scales, we could not get a global fit for the drug bound state. When we compare the relaxation dispersion curves for Val7 and Phe19 (Fig. 5), we find that in the presence of rimantadine the curves become more flat, which suggests that the binding of rimantadine makes the channel less dynamic.
Discussion
We have shown that the reconstituted p7 channel in DPC micelles has intrinsic conformational exchange and the regions showing sub-millisecond time scale motion are overall consistent with the architecture and solvent accessibility profile of the channel. Interestingly, presence of an allosteric inhibitor (rimantadine) decreases the conformational exchange at some sites. The 15N relaxation dispersion data show that Val7 and Leu8 of H1 undergo significant chemical exchange, and that the rate of conversion between the two states is in the range of 1000 s−1. These two residues are in the packing interface between the H1 helices of the adjacent monomers. Therefore, large chemical exchange in these two residues suggests that the portion of the cavity formed by H1 is intrinsically breathing. In addition, Phe19 at the hinge between H1 and H2 also shows pronounced exchange, and the conformational switching of the hinge should be related to the movement of the H1 helices that cause the exchange in Val7 and Leu8. The hinge region is also near the rimantadine binding site. Addition of rimantadine decreases the relaxation dispersion at this site, in addition to Val7 of H1. Effect of rimantadine binding at Val7 suggests a wedge mechanism where the dynamics is perturbed allosterically.
While we only see exchange in three residues (Val7, Leu8 and Phe19), it is possible that other residues or regions may also have conformational exchange. The 15N CPMG experiment can only detect exchange if the chemical shift difference in 15N between the two states is sufficiently large. Residues or segments that undergo conformational exchange without significant chemical shift changes will not show significant relaxation dispersion. Additionally, the CPMG experiment assumes a particular model of exchange, e.g., a two-state model system, which may not fully account for all the conformational exchanges the protein is undergoing.
The observed chemical exchanges in the N-terminal end of H1 are nonetheless anticipated if the narrow point or the tip of the funnel is subject to breathing motion. Previous functional assay performed in Xenopus oocytes showed that the Asn9 ring is important for cation conduction—mutating to alanine significantly decreased ion conduction (OuYang et al. 2013). The Asn9 is also highly conserved throughout the HCV genotypes and subtypes. Therefore a reasonable model for channel function is that the Asn9 ring may selectively coordinate and dehydrate the cation near the tip of the funnel, whereas the Ile6 ring is a hydrophobic constriction that would prevent water from freely passing through. The proposed channel breathing may involve reorientation of the H1 helices that widens or narrows the funnel tip, analogous to the dynamic C-terminal helix of KcsA (Cuello et al. 2010). Furthermore, the movements in the H1 segment must be concerted with structural change of the hinge (residues 17–19) between H1 and H2, and this is consistent with the strong chemical exchange observed at Phe19.
The breathing of the portion of the channel formed by the H1 helices is also in qualitative agreement with the significant proton exchange observed for this region of the channel. First, the solvent accessibility profile of p7 reconstituted in DPC micelles show that only the pore forming H1 and H2 helices have proton exchange and H3, which faces the detergent, does not show any exchange. This result is in good agreement with the NMR structure of the p7 hexamer. More importantly, the residues of H1 on average have significantly faster proton exchange rates than those of H2, suggesting that in addition to the presence of water in the cavity, the H1s undergo structural changes that render their secondary structures less stable. The proton exchange rates report not only water accessibility but also the strength of hydrogen bonding in the protein that protect the backbone amide protons from exchanging with water protons. For instance, helical segments that have weaker hydrogen bonding can show faster exchange rates compared to the ones that have stronger bonding even if they have the same water accessibility. A plausible explanation for the less stable helix-specific hydrogen bonds of H1, as suggested by the proton exchange rates, is that the channel is in equilibrium exchange between the closed state in which the H1s are strongly packed and more stable and the open state in which the H1s are loosely packed and less stable. We believe that this mode of channel breathing at the moment provides the most likely explanation for the observed 15N relaxation dispersion of the p7 channel.
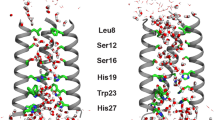
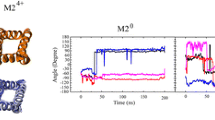
Viroporins are small and usually are very dynamic, and this dynamic nature in some cases has functional relevance. For instance, in the case of M2 channel from influenza A virus, several transmembrane peaks severely broaden when pH is lowered to activate the channel (Schnell and Chou 2008). Indeed for M2, Trp41, which is a pore-lining residue that is important for function and serves as the gate, shows increased dynamics upon activation (Schnell and Chou 2008). Dynamics could also play an important role in the function of viroporins other than M2.
Finally, the observed dynamics of the p7 channel could be important for channel activity and for opening and closing of the channel (Fig. 6), especially because the regions showing strong dynamics contain the functional elements of the channel. In addition, the observed dynamics can have implications in the mechanism of inhibition by the drug rimantadine. Previous NMR study has shown that the drug binds at the region where the three helices interact (Fig. 6). In this region, the drug is not inside the pore but is in a peripheral pocket between H1/H2 and H3 helices. Therefore the mechanism by which the drug is inhibiting is probably allosteric. We see significant dynamics at this region as shown by the relaxation dispersion data. Binding of the drug to this site makes the channel less dynamic, which could thereby prevent the opening and closing motion of the channel that is required for activity.
Proposed model for channel function and inhibition. The residues that show conformational exchange are shown with green arrows. The breathing of H1 helix (blue arrows), which is supported by our results, could facilitate opening and closing of the channel. In the open state, Asn9 is able to bind and release Ca2+ ions. Conformational dynamics at the hinge region between H1 and H2 could explain the mechanism of drug inhibition, as the drug binds at this region. Drug binding could act as a molecular wedge to prevent breathing of the complex, and therefore closing the channel
References
Agarkova I, Dunigan D, Gurnon J, Greiner T, Barres J, Thiel G, Van Etten JL (2008) Chlorovirus-mediated membrane depolarization of Chlorella alters secondary active transport of solutes. J Virol 82:12181–12190
Carrere-Kremer S, Montpellier-Pala C, Cocquerel L, Wychowski C, Penin F, Dubuisson J (2002) Subcellular localization and topology of the p7 polypeptide of hepatitis C virus. J Virol 76:3720–3730
Carver JP, Richards RE (1972) A general two-site solution for the chemical exchange produced dependence of T2 upon the Carr–Purcell pulse separation. J Magn Reson 6:89–105
Cook GA, Opella SJ (2011) Secondary structure, dynamics, and architecture of the p7 membrane protein from hepatitis C virus by NMR spectroscopy. Biochim Biophys Acta 1808:1448–1453
Cuello LG, Jogini V, Cortes DM, Perozo E (2010) Structural mechanism of C-type inactivation in K(+) channels. Nature 466:203–208
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) Nmrpipe—a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Gan SW, Surya W, Vararattanavech A, Torres J (2014) Two different conformations in hepatitis C virus p7 protein account for proton transport and dye release. PLoS One 9:e78494
Gentzsch J, Brohm C, Steinmann E, Friesland M, Menzel N, Vieyres G, Perin PM, Frentzen A, Kaderali L, Pietschmann T (2013) hepatitis c Virus p7 is critical for capsid assembly and envelopment. PLoS Pathog 9:e1003355
Goddard T, Kneller D (2008) SPARKY 3. University of California, San Francisco
Gouklani H, Beyer C, Drummer H, Gowans EJ, Netter HJ, Haqshenas G (2013) Identification of specific regions in hepatitis C virus core, NS2 and NS5A that genetically interact with p7 and co-ordinate infectious virus production. J Viral Hepat 20:e66–e71
Griffin SD, Beales LP, Clarke DS, Worsfold O, Evans SD, Jaeger J, Harris MP, Rowlands DJ (2003) The p7 protein of hepatitis C virus forms an ion channel that is blocked by the antiviral drug, Amantadine. FEBS Lett 535:34–38
Griffin S, Stgelais C, Owsianka AM, Patel AH, Rowlands D, Harris M (2008) Genotype-dependent sensitivity of hepatitis C virus to inhibitors of the p7 ion channel. Hepatology 48:1779–1790
Haqshenas G, Mackenzie JM, Dong X, Gowans EJ (2007) Hepatitis C v irus p7 protein is localized in the endoplasmic reticulum when it is encoded by a replication-competent genome. J Gen Virol 88:134–142
Hou X, Pedi L, Diver MM, Long SB (2012) Crystal structure of the calcium release-activated calcium channel Orai. Science 338:1308–1313
Hwang TL, Mori S, Shaka AJ, vanZijl PCM (1997) Application of phase-modulated CLEAN chemical EXchange spectroscopy (CLEANEX-PM) to detect water-protein proton exchange and intermolecular NOEs. J Am Chem Soc 119:6203–6204
Jones CT, Murray CL, Eastman DK, Tassello J, Rice CM (2007) Hepatitis C virus p7 and NS2 proteins are essential for production of infectious virus. J Virol 81:8374–8383
Korzhnev DM, Neudecker P, Mittermaier A, Orekhov VY, Kay LE (2005) Multiple-site exchange in proteins studied with a suite of six NMR relaxation dispersion experiments: an application to the folding of a Fyn SH3 domain mutant. J Am Chem Soc 127:15602–15611
Loria JP, Rance M, Palmer AG 3rd (1999) A TROSY CPMG sequence for characterizing chemical exchange in large proteins. J Biomol NMR 15:151–155
Luik P, Chew C, Aittoniemi J, Chang J, Wentworth P Jr, Dwek RA, Biggin PC, Venien-Bryan C, Zitzmann N (2009) The 3-dimensional structure of a hepatitis C virus p7 ion channel by electron microscopy. Proc Natl Acad Sci USA 106:12712–12716
Lunin VV, Dobrovetsky E, Khutoreskaya G, Zhang R, Joachimiak A, Doyle DA, Bochkarev A, Maguire ME, Edwards AM, Koth CM (2006) Crystal structure of the CorA Mg2+ transporter. Nature 440:833–837
Luscombe CA, Huang Z, Murray MG, Miller M, Wilkinson J, Ewart GD (2010) A novel Hepatitis C virus p7 ion channel inhibitor, BIT225, inhibits bovine viral diarrhea virus in vitro and shows synergism with recombinant interferon-alpha-2b and nucleoside analogues. Antivir Res 86:144–153
Montserret R, Saint N, Vanbelle C, Salvay AG, Simorre JP, Ebel C, Sapay N, Renisio JG, Bockmann A, Steinmann E et al (2010) NMR structure and ion channel activity of the p7 protein from hepatitis C virus. J Biol Chem 285:31446–31461
Moradpour D, Penin F, Rice CM (2007) Replication of hepatitis C virus. Nat Rev Microbiol 5:453–463
Mori S, Johnson MO, Berg JM, Vanzijl PCM (1994) Water exchange filter (Wex Filter) for nuclear-magnetic-resonance studies of macromolecules. J Am Chem Soc 116:11982–11984
Mori S, Abeygunawardana C, vanZijl PCM, Berg JM (1996a) Water exchange filter with improved sensitivity (WEX II) to study solvent-exchangeable protons. Application to the consensus zinc finger peptide CP-I. J Magn Reson Ser B 110:96–101
Mori S, Berg JM, vanZijl PCM (1996b) Separation of intramolecular NOE and exchange peaks in water exchange spectroscopy using spin-echo filters. J Biomol NMR 7:77–82
Mulder FA, Skrynnikov NR, Hon B, Dahlquist FW, Kay LE (2001) Measurement of slow (micros-ms) time scale dynamics in protein side chains by (15)N relaxation dispersion NMR spectroscopy: application to Asn and Gln residues in a cavity mutant of T4 lysozyme. J Am Chem Soc 123:967–975
Nieva JL, Madan V, Carrasco L (2012) Viroporins: structure and biological functions. Nat Rev Microbiol 10:563–574
OuYang B, Xie S, Berardi MJ, Zhao X, Dev J, Yu W, Sun B, Chou JJ (2013) Unusual architecture of the p7 channel from hepatitis C virus. Nature 498:521–525
Palmer AG 3rd, Kroenke CD, Loria JP (2001) Nuclear magnetic resonance methods for quantifying microsecond-to-millisecond motions in biological macromolecules. Methods Enzymol 339:204–238
Pavlovic D, Neville DC, Argaud O, Blumberg B, Dwek RA, Fischer WB, Zitzmann N (2003) The hepatitis C virus p7 protein forms an ion channel that is inhibited by long-alkyl-chain iminosugar derivatives. Proc Natl Acad Sci USA 100:6104–6108
Pervushin K, Riek R, Wider G, Wuthrich K (1998) Transverse relaxation-optimized spectroscopy (TROSY) for NMR studies of aromatic spin systems in C-13-labeled proteins. J Am Chem Soc 120:6394–6400
Popescu CI, Callens N, Trinel D, Roingeard P, Moradpour D, Descamps V, Duverlie G, Penin F, Heliot L, Rouille Y et al (2011) NS2 protein of hepatitis C virus interacts with structural and non-structural proteins towards virus assembly. PLoS Pathog 7:e1001278
Premkumar A, Wilson L, Ewart GD, Gage PW (2004) Cation-selective ion channels formed by p7 of hepatitis C virus are blocked by hexamethylene amiloride. FEBS Lett 557:99–103
Sakai A, Claire MS, Faulk K, Govindarajan S, Emerson SU, Purcell RH, Bukh J (2003) The p7 polypeptide of hepatitis C virus is critical for infectivity and contains functionally important genotype-specific sequences. Proc Natl Acad Sci USA 100:11646–11651
Schnell JR, Chou JJ (2008) Structure and mechanism of the M2 proton channel of influenza A virus. Nature 451:591–595
Shanmugam S, Yi MY (2013) Efficiency of E2-p7 processing modulates production of infectious hepatitis C virus. J Virol 87:11255–11266
Steinmann E, Pietschmann T (2010) Hepatitis C virus p7—a viroporin crucial for virus assembly and an emerging target for antiviral therapy. Viruses 2:2078–2095
Steinmann E, Penin F, Kallis S, Patel AH, Bartenschlager R, Pietschmann T (2007) Hepatitis C virus p7 protein is crucial for assembly and release of infectious virions. PLoS Pathog 3:e103
Vallurupalli P, Hansen DF, Kay LE (2008) Structures of invisible, excited protein states by relaxation dispersion NMR spectroscopy. Proc Natl Acad Sci USA 105:11766–11771
Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas M, Ulrich EL, Markley JL, Ionides J, Laue ED (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696
Wozniak AL, Griffin S, Rowlands D, Harris M, Yi M, Lemon SM, Weinman SA (2010) Intracellular proton conductance of the hepatitis C virus p7 protein and its contribution to infectious virus production. PLoS Pathog 6:e1001087
Acknowledgments
We thank Dr. Qin Yang and Dr. Kirill Oxenoid for helpful discussions, and Dr. Tanxing Cui for help with Sparky. S. B. is a recipient of an Erwin Schrödinger postdoctoral fellowship of the Austrian Science Fund (FWF, J3251). This work was supported by the NIH Grant GM094608 (to J. J. C.).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dev, J., Brüschweiler, S., Ouyang, B. et al. Transverse relaxation dispersion of the p7 membrane channel from hepatitis C virus reveals conformational breathing. J Biomol NMR 61, 369–378 (2015). https://doi.org/10.1007/s10858-015-9912-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-015-9912-0