Abstract
Secondary chemical shift analysis is the main NMR method for detection of transiently formed secondary structure in intrinsically disordered proteins. The quality of the secondary chemical shifts is dependent on an appropriate choice of random coil chemical shifts. We report random coil chemical shifts and sequence correction factors determined for a GGXGG peptide series following the approach of Schwarzinger et al. (J Am Chem Soc 123(13):2970–2978, 2001). The chemical shifts are determined at neutral pH in order to match the conditions of most studies of intrinsically disordered proteins. Temperature has a non-negligible effect on the 13C random coil chemical shifts, so temperature coefficients are reported for the random coil chemical shifts to allow extrapolation to other temperatures. The pH dependence of the histidine random coil chemical shifts is investigated in a titration series, which allows the accurate random coil chemical shifts to be obtained at any pH. By correcting the random coil chemical shifts for the effects of temperature and pH, systematic biases of the secondary chemical shifts are minimized, which will improve the reliability of detection of transient secondary structure in disordered proteins.
Similar content being viewed by others

Avoid common mistakes on your manuscript.
Introduction
Intrinsically disordered proteins (IDPs) are abundant and important in biology (Dyson and Wright 2005). In structural studies of IDPs, NMR spectroscopy has proven a powerful technique that gives high resolution information about transiently formed structures (Dyson and Wright 2002). The simplest way to characterize transiently formed structure in IDPs is secondary chemical shift analysis of the peptide backbone atoms. Chemical shifts are attractive probes of transient structure as they can be measured with very high precision, which is required for detection of marginally populated structural elements in disordered proteins.
The backbone chemical shifts depend on the dihedral angles ψ and φ, and are thus affected by the secondary structure content of the peptide chain (Spera and Bax 1991). In folded proteins, secondary structural elements can be identified by calculating a chemical shift index (Wishart et al. 1992; Wishart and Sykes 1994). In disordered proteins, secondary structure is only formed transiently, and the chemical shift index is thus not appropriate, as it is primarily intended to identify fully formed secondary structure. Instead, IDPs are evaluated in terms of secondary chemical shifts, Δδ, that identify the location and the population of transiently structured regions. Identification of transiently structured regions is of biological interest, as these are frequently involved in molecular recognition (Mohan et al. 2006).
The secondary chemical shift is calculated for each atom as the difference between the experimentally determined shifts and a set of random coil chemical shifts. The choice of an appropriate set of random coil values is pivotal for calculation of reliable secondary chemical shifts, however, it is unclear how the appropriate values should be defined. Numerous datasets have been proposed by different groups using a variety of methods. The procedure for obtaining the random coil chemical shifts can be grouped into two categories, depending on whether the chemical shifts are derived from a protein chemical shift database (Zhang et al. 2003; De Simone et al. 2009; Wishart et al. 1991; Peti et al. 2001) or a set of model peptides (Wishart et al. 1995a; Schwarzinger et al. 2000; Richarz and Wuthrich 1978; Bundi and Wuthrich 1979; Jimenez et al. 1986; Braun et al. 1994; Thanabal et al. 1994; Merutka et al. 1995; Plaxco et al. 1997; Bienkiewicz and Lumb 1999). Database methods collect chemical shift data from a large number of proteins recorded at a variety of different experimental conditions. The large datasets in database methods allow good approximation of the average values, but the averaged chemical shifts are only appropriate if the protein of interest is investigated at conditions similar to the database average, which is usually poorly defined. Alternatively, random coil chemical shifts have also been derived from short unstructured peptides. The datasets used for determination of peptide based random coil chemical shifts are much smaller than the database derived values. However, the experimental conditions can be tightly controlled, which makes small peptides optimal for investigation of the effects of environmental perturbations such as pH, denaturant concentration and temperature. A widely used random coil peptide series has the sequence Ac-GGXGG-NH2, where the conformational freedom of the glycines combined with chemical denaturant ensure that the peptides are unstructured (Schwarzinger et al. 2000).
Sequential neighbors in the peptide chain affect the random coil chemical shifts (Schwarzinger et al. 2001), and if this is not taken into account, the neighbor effect could mask the transiently structured regions. One approach for eliminating the neighbor effects compares the chemical shifts to those of the same protein recorded under highly denaturing conditions (Modig et al. 2007). This procedure gives clean secondary chemical shifts for IDPs (Kjaergaard et al. 2010a). This approach is, however, expensive in terms of instrument time, sample consumption and analysis time and the denaturant may affect the ensemble of the disordered state. Alternatively, the sequence effects can be removed using a set of sequence correction factors. Like the random coil chemical shifts, the sequence correction factors can either be derived from database approaches (Wang and Jardetzky 2002; De Simone et al. 2009; Wang et al. 2007) or from peptide studies (Schwarzinger et al. 2001). An elegant approach for experimentally determining the sequence correction factors was proposed by Schwarzinger et al. (Schwarzinger et al. 2001). This method uses the chemical shifts of the glycines in the Ac-GGXGG-NH2 peptide series to calculate correction factors for the X residue. The underlying assumption of this method is thus that all residue types will experience a similar perturbation of the random coil chemical shift as glycine. The dataset reported by Schwarzinger et al. (Schwarzinger et al. 2000; Schwarzinger et al. 2001), however, was determined at pH 2.3 where Asp, Glu and His are fully protonated. When this dataset is used for analysis of chemical shifts recorded at neutral pH, these residues show large deviations in the secondary chemical shift profile suggesting that these random coil values and correction factors are inappropriate for IDPs investigated at neutral pH.
The quality of a protein NMR spectrum often depends on the sample conditions. Therefore, NMR investigations of proteins often start by finding the optimal experimental conditions in terms of salt, pH and temperature. For IDPs in particular, low temperatures are often attractive as the exchange rates of amide protons with the solvent are lower. In addition, the secondary structure content is higher at low temperature and transiently formed structures are thus easier to detect (Kjaergaard et al. 2010a). High temperatures, in contrast, are often used for molten globules to reduce peak broadening due to intermediate exchange (Eliezer et al. 1997; Ramboarina and Redfield 2003; Kjaergaard et al. 2010b). Random coil chemical shifts depend on the temperature and it is thus necessary to extrapolate random coil chemical shifts to the desired temperature. For this reason temperature coefficients have been reported for both 1H and 15N random coil chemical shifts (Merutka et al. 1995; Lam and Hsu 2003). The intrinsic temperature effect of the 13C chemical shifts has been assumed to be negligible.
Using the approach of Schwarzinger et al., we report a set of random coil chemical shifts and sequence correction factors recorded for the Ac-GGXGG-NH2 peptide series. The present dataset differs from that determined previously by being recorded at neutral pH and at a lower urea concentration. Furthermore, we have explored the effects of experimental conditions that are likely to be relevant for IDPs, namely the temperature dependence of the 13C random coil chemical shifts and the pH dependence of the histidine random coil chemical shifts. This dataset removes systematic errors in secondary chemical shift analyses due to mismatch between the experimental conditions and those used for determining the random coil chemical shifts. Although this dataset was determined with IDPs in mind, it should be equally suitable for folded proteins determined at neutral pH.
Materials and methods
Peptides with the sequence Ac-GGXGG-NH2 were purchased from KJ Ross-Petersen ApS (Klampenborg, Denmark), where X was each of the 20 common amino acids. All peptides were purified to more than 95% purity by reversed phase HPLC and their identities were confirmed by mass spectrometry. NMR samples were prepared by dissolving 2–3 mg of peptide in 500 μl 20 mM sodium phosphate buffer pH 6.5 containing 5% (v/v) D2O, 1 M urea, 3 mM NaN3, and 1 mM DSS. pH was adjusted to 6.5 by addition of small quantities of HCl. The cysteine containing peptide additionally contained 10 mM DTT to keep the thiol group in the reduced state.
All NMR spectra were acquired on a Varian Unity 800 MHz spectrometer equipped with a cold probe. Chemical shifts were referenced to internal DSS as described previously (Wishart et al. 1995b). Temperatures were measured by a thermocouple mounted in an NMR tube and inserted into the spectrometer probe. For each sample the following spectra were acquired at natural isotope abundance: 1H-15N HSQC, 1H-13C HSQC and a newly developed 1Hα-13C′ HSQC. For all peptides, data were recorded at 5, 15, 25, 35 and 45°C. The 1Hα-13C′ HSQC experiment correlates the Hα protons with the carbonyl resonances of the same and the preceding residue and will be discussed fully in a subsequent publication. In short the 1Hα-13C′ HSQC experiment is a modified version of a previously described experiment (Kay et al. 1992), where the INEPT delay has been changed to 37 ms, and the hard pulses in the INEPT period have been replaced by a selective sech/tanh pulse (Silver et al. 1985) centered on carbonyl for 13C and a REBURP pulse (Geen and Freeman 1991) centered in the Hα region for 1H. NMR data were processed using NMRPipe (Delaglio et al. 1995) and analyzed using CCPNMR Analysis (Vranken et al. 2005). Temperature coefficients were determined by least squares fitting of the chemical shifts to a linear function of temperature using OpenOffice 3.2 to the Eq. 1, where “a” is the temperature coefficient:
The sequence corrected random coil chemical shifts of a residue can thus be calculated at any temperature using the equation:
where B(i+1) is the B correction factor of the subsequent residue etc.
For the histidine containing peptide, a titration series from pH 8 to pH 4 in steps of 0.5 were recorded by stepwise addition of HCl. The NMR spectra were recorded at 5°C to minimize exchange of the amide protons with the solvent and to prevent overlap between the histidine Hα and the water signal. The chemical shifts were assumed to be the linear combination of the chemical shift of a fully protonated and a fully deprotonated species and thus follow the equation:
δHA and δA represent the random coil chemical shifts of the fully protonated and fully deprotonated species, respectively. Ka is the acid dissociation constant of the side chain. The chemical shifts were fitted to Eq. 3 using IGOR PRO (WaveMetrics), where Ka was treated as a global fitting parameter.
Results and discussion
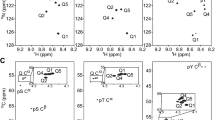
Three spectra were recorded for each peptide: 1H-15N HSQC, 1H-13C HSQC and 1Hα-13C′ HSQC. The amide peaks were well resolved in the 1H-15N HSQC (Fig. 1A) and assignments were readily transferred from the previous study at low pH. The glycine signals were in the same region of the 1H-13C HSQC (Fig. 1B), but they could be separated to allow determination of the chemical shifts of each glycine. The 1Hα-13C′ HSQC spectrum correlates Hα(i) with 13C′(i) and 13C′(i − 1), which allows the glycine Hα resonances to be assigned by a sequential walk through the carbonyl resonances (Fig. 1C). The chemical shifts of glycine peaks are similar to those reported at low pH. The signals were weaker in the 1H-15N HSQC at 35°C and absent at 45°C due to exchange of the amide protons with the solvent. Accordingly, the 1H-15N HSQC spectrum was not recorded at 45°C for most peptides. In a few spectra, the chemical shifts could not be obtained due to overlap with the water signal or due to proton exchange. The temperature dependence could still be analyzed as chemical shifts from at least three temperatures were available for all nuclei with the amide resonances from histidine as the only exception. The chemical shifts depend linearly on temperature (Fig. 2) and were thus analyzed by least squares fitting to a linear function. Random coil chemical shifts and temperature coefficients for all 20 residues are reported in Table 1. Sequence correction terms for residue X were obtained using the method of Schwarzinger et al. (Schwarzinger et al. 2001) by subtracting the chemical shifts of the Ac-GGGGG-NH2 peptide from those of the Ac-GGXGG-NH2 peptide (Tables 2, 3). Glycine 1, 2, 4 and 5 give rise to correction factors A, B, C and D, respectively.
2D NMR spectra used for determination of chemical shifts for the glutamate containing peptide. a 1H-15N HSQC, b 1H-13C HSQC and c 1Hα-13C′ HSQC spectra recorded at pH 6.5 and 25°C on the peptide with the sequence Ac-GGEGG-NH2. * denotes aliased peaks from the N-terminal acetyl group and E3 Cγ-Hγ. The dashed line in (c) illustrates a sequential walk through the sequence, which forms the basis for sequential assignment of the peaks. The peaks from Hα of G4 are split due to prochirality induced by E3
Temperature dependence of the random coil chemical shifts of glutamate at pH 6.5. Random coil chemical shifts from 25°C have been subtracted from all datasets to allow the different nuclei to be represented together. Lines represent the best linear fit from which temperature coefficients were extracted
For all other residues than histidine, aspartate, and glutamate, the random coil chemical shifts obtained at pH 6.5 are very similar to those reported by Schwarzinger et al. (Schwarzinger et al. 2000) at pH 2.3 and 8 M urea. For 13C resonances, the difference in chemical shifts was below 0.2 ppm and in most cases it was much smaller. This shows that for residues that do not change protonation state, the 13C random coil chemical shifts are largely independent of pH and denaturant concentration. For the 15N chemical shifts, the deviations from the values reported by Schwarzinger et al. (2000) are larger. This effect on the random coil chemical shifts is presumably due to the effect of urea rather than pH as most residues do not change protonation state. For the 1H chemical shifts, the random coil chemical shifts are similar to the values determined at low pH and at a high concentration of urea. The sequence correction terms are almost identical to the low pH values, and even the residues deprotonated at pH 6.5 show only minor differences in the sequence correction factors.
The motivation for determining the random coil chemical shifts at a higher pH was to obtain accurate values for histidine, aspartate, and glutamate. Aspartate and glutamate have side chain pKa values of approximately 4, which means that the side chains will be fully protonated at the conditions where Schwarzinger et al. determined their shifts (pH 2.3), 10% protonated at the conditions of Wishart et al. (pH 5) and fully deprotonated at the conditions used here (pH 6.5). Consistent with the pH-values, the 13C and 15N chemical shifts for aspartate and glutamate at 6.5 are significantly higher than those determined by Schwarzinger et al. and slightly higher than those reported by Wishart et al. due to the remaining protonation at pH 5. For the 1H chemical shifts, protonation does not change the chemical shifts in the same direction for all protons. Similarly to the 13C and 15N chemical shifts, the 1H chemical shifts are significantly different from those determined at pH 2.3 and slightly different from those determined at pH 5. In contrast to peptide based random coil chemical shifts, the effect of pH is more difficult to control in database based random coil values, where it is assumed that the chemical shift effect from differences in the pH values is averaged by using a large database. Averaging of the pH effects requires that pH induced changes at higher pH values are canceled by changes of opposite sign at lower pH. However, this will not be the case for the pH dependence of chemical shifts as the chemical shifts only are affected when the pH is near the pKa value. When the average pH value in the database is near 6, the chemical shifts of aspartate and glutamate at pH values above 6 are identical to the value at pH 6, as the side chain is already fully protonated. Accordingly, the contributions from chemical shifts recorded at pH values below 6 are not canceled by averaging with values at higher pH values. Unless care is taken to eliminate datasets recorded at lower pH values, database methods are likely to result in random coil values representing partially protonated side chains of aspartate and glutamate. The distribution of pH values used for random coil databases suggests that this may be a problem for at least some datasets (De Simone et al. 2009). Random coil chemical shift values for aspartate and glutamate determined by database based approaches are generally much more similar to the values determined at pH 6.5 than those determined at pH 2.3, however, the deviations are consistent with a small contribution from low pH datasets.
Of the 20 common amino acids, only histidine has a side chain with a pKa value near neutral pH. The pH values used in protein NMR studies are highly variable, but are generally kept slightly below neutral pH to minimize proton exchange. Due to the variability in pH values, a set of generally applicable random coil chemical shifts needs to take the pH dependence of histidine into account. We have performed a pH titration using the Ac-GGHGG-NH2 peptide to address this problem. The random coil chemical shifts of this peptide were determined from pH 4 to 8, which is the range where protein NMR studies are usually carried out (Fig. 3). As expected, the chemical shifts of all nuclei change with a midpoint at pH 6.9, corresponding to the expected pKa of the histidine side chain. The data are fitted to Eq. 3 using Ka as a common parameter for all nuclei. This procedure extracts the chemical shifts of the fully protonated and the fully deprotonated residue (Table 4) and Ka = 1.2 × 10−7 M. Using these values and Eq. 3, the pH corrected random coil chemical shift for histidine at any desired pH can be calculated. A similar procedure can be applied to aspartate and glutamate residues, using the values reported by Schwarzinger et al. as the fully protonated state and the values reported here as the fully deprotonated state.
pH dependence of the random coil chemical shifts of histidine. A pH titration was carried out for the Ac-GGHGG-NH2 peptide at 5°C. The random coil chemical shifts as a function of pH were fitted to Eq. 3 with pKa as a global fitting parameter. This gives the random coil chemical shifts of the fully protonated, δHA, and the fully deprotonated, δHA, species. Data could not be collect for the amide at pH 8 due to hydrogen exchange with the solvent
Random coil chemical shifts depend on the temperature. If the temperature dependence is not taken into account, a systematic bias is introduced into the secondary chemical shifts. The temperature dependence of 1H and 15N random coil chemical shifts has been addressed by others (Merutka et al. 1995; Lam and Hsu 2003) and our values resemble those reported previously. The temperature dependence of 13C random coil chemical shifts is relatively small and has not been addressed previously. In folded proteins with fully formed secondary structure, the temperature dependence of the 13C random coil chemical shifts is less important due to the small temperature coefficients relative to the secondary chemical shifts. In IDPs, however, the contribution from the temperature dependence of the random coil chemical shifts might be of the same magnitude as the secondary chemical shifts and consequently even minute contributions to the random coil chemical shifts need to be taken into account. Table 1 shows that C′ temperature coefficients are always negative and Cβ temperature coefficients are, with the exception of proline, positive. Histidine is an exception and will be discussed below. The consistent sign of the temperature coefficients means that if the temperature dependence is not taken into consideration, low temperature data will consistently overestimate the α-helix content and underestimate the degree of β-structure and vice versa. If random coil chemical shifts recorded at 25°C are used for analysis of NMR data recorded at 5°C, the magnitude of the error is up to 0.2 ppm, corresponding to a systematic error of 5–10% in the calculated secondary structure content. The Cα temperature coefficients are smaller than those of C′ and Cβ and can be positive or negative. Accordingly, when the temperature dependence of Cα chemical shifts is not corrected for, it is less likely to lead to a systematic error in quantification of transiently formed secondary structure elements. The temperature dependence of the Cα shifts will, however, still add noise to the secondary chemical shifts if the chemical shifts are recorded at a different temperature than the random coil chemical shifts.
With respect to temperature dependence of the chemical shift, histidine is a special case. A change of temperature results in a change in the pH and as a consequence a change in the protonation state and thus the random coil chemical shifts. Small temperature induced pH changes are, however, difficult to avoid. In order to minimize the temperature dependence of the pH value, we have used phosphate buffer whose pH is relatively insensitive to temperature (Dawson et al. 1986). Small pH changes can, however, not be avoided, and accordingly the positive temperature coefficients for C′ and the unusually large Cβ temperature coefficient for histidine may thus represent small temperature induced changes in the pH.
Intrinsically disordered proteins often undergo temperature dependent structural changes (Uversky 2009; Kjaergaard et al. 2010a; Kim et al. 2007; Wu et al. 2008; Hsu et al. 2009). The temperature dependence of chemical shifts in disordered proteins consists of the sum of two components, the chemical shift change induced by structural transitions, and the change caused by the temperature dependence of the random coil chemical shifts. The temperature dependence of the random coil chemical shifts thus complicates analysis of temperature induced structural changes in disordered proteins. Intrinsic random coil referencing solves this problem as the chemical shifts can be recorded at the same temperatures as the reference chemical shifts (Kjaergaard et al. 2010a), however, this approach requires reassignment of the protein under denaturing conditions at several temperatures. The tabulated temperature coefficients reported here allow analysis of temperature dependent structural changes by removing the intrinsic temperature dependence of the random coil chemical shifts. To compare the temperature coefficients to intrinsic statistical coil referencing, the chemical shift differences calculated for a 40°C temperature difference using Eq. 2 were compared to the differences recorded on the urea denatured state of ACTR between 5 and 45°C (Fig. 4). The chemical shifts of the urea denatured state vary more than the tabulated values, which is likely to result from the effects of the neighboring residues on the temperature coefficients. There is not a complete agreement between the two methods for correcting for the intrinsic temperature dependence of the random coil chemical shifts, but they suggest that the intrinsic temperature effect has the same sign and magnitude. This shows that the tabulated temperature coefficients remove the systematic error from secondary chemical shift analyses. However, intrinsic random coil referencing is expected to perform better due to the contributions of neighboring residues to the temperature coefficients.
Comparison of the difference in random coil chemical shifts between 5°C and 45°C obtained using intrinsic random coil referencing and peptide based temperature coefficients at pH 6.5. The temperature induced change in random chemical shifts between 5 and 45°C were calculated for the activation domain of ACTR based on the temperature coefficients in Table 1 (dashed line) for Cα (black) and C′ (red). For comparison, the temperature induced changes in the intrinsic random coil chemical shifts reported previously are displayed as a full line. The intrinsic random coil chemical shifts are measured in the presence of 6 M urea
The residue preceding a proline experiences an unusually large sequence effect (Schwarzinger et al. 2001). This most likely represent the unusual φ/ψ distributions of these residues, where the population of the α-helical region of the Ramachandran plot is unusually low (Ting et al. 2010) due to steric clashes between the proline Cδ and the Cβ of the X-residue (MacArthur and Thornton 1991). Glycine does not have a Cβ atom and glycine derived correction factors are thus not able to correct for this phenomenon. The neighbor effect varies considerably depending on the nature of the X residue (Wishart et al. 1995a), reflecting the strong residue-type dependence of the φ/ψ distribution of the residue preceding proline (Ting et al. 2010). For this reason, we recommend using the amino acid specific values correction factors determined previously for proline (Wishart et al. 1995a) instead of the glycine derived value presented here.
The effect of the neighboring residues on the random coil chemical shifts is often corrected using a set of correction factors derived from glycine rich peptides (Schwarzinger et al. 2001). The sequence correction factors for residue X are derived from the chemical shifts of the glycines in the Ac-GGXGG-NH2 peptide relative to those of the Ac-GGGGG-NH2 peptide. The underlying assumption of this approach is thus that all residues experience the same neighbor perturbation as glycine. Since the dataset reported here is determined under similar conditions to that reported previously by Wishart et al. (Wishart et al. 1995a), we can test this assumption. Figure 5 compares chemical shifts for the Ac-GGXAGG-NH2 peptide series obtained by experiment or calculated from the Ac-GGXGG-NH2 peptide series. In all cases, the correction factors improve agreement between the two peptide series. The predicted random coil chemical shifts are, however, consistently too low, consistent with the correction factors being too small. This suggests that the glycine-derived correction factors are able to correct only a part of the neighbor effect. Residue-type specific correction factors for the residue preceding proline suggest that glycine is an outlier with smaller correction factors than all other residues. Two different mechanisms can be envisioned for the neighbor effect. First, the different chemical structures of the amino acids affect the polarization of the chemical bonds, and thus the electron density around the nuclei. Presumably, this effect is similar for all residue types, and may thus be corrected using glycine-derived correction factors. Second, the neighboring residues may affect the φ/ψ distribution in the disordered state as was demonstrated using residual dipolar couplings of a set of disordered peptides (Dames et al. 2006). Glycine has an unusual φ/ψ distribution due to the absence of a side chain, and is thus not perturbed by the neighboring residues in the same way as other residue types. Correction factors derived from glycine residues are thus only able to correct for certain aspects of the neighbor effects.
Comparison between chemical shifts for the Ac-GGXAGG-NH2 peptide series obtained experimentally (Wishart et al. 1995a) at pH 5 and from Ac-GGXGG-NH2 at pH 6.5 series with (grey) and without (black) glycine derived correction factors. In all cases except for the carbonyl of glycine, glycine derived sequence correction factors improved the agreement between experimental and predicted chemical shifts. The Cα and C′ chemical shifts based on the Ac-GGXAGG-NH2 peptide series show a small, but consistent, systematic deviation from the experimental chemical shifts
To compare the performance of the present data set to other random coil datasets, we compared the random coil values to a dataset consisting of chemical shift assignments of 14 IDPs. The dataset was assembled recently to determine random coil chemical shifts suitable for IDPs (Tamiola et al. 2010). As these proteins are highly disordered, the deviation from the experimental chemical shifts can be used to probe the suitability of random coil datasets for these proteins. Figure 6 shows the RMSD between the experimental chemical shifts and predicted random coil values for seven datasets. The datasets made specifically for IDPs have the smallest RMSD values suggesting the best predictive power. Compared to the present dataset, the database assembled by Tamiola et al. has smaller RMSDs for the amide nuclei. For the Hα, Cα, Cβand C′ nuclei that are more frequently used for secondary chemical shift analysis, these two datasets perform similarly.
Comparison of random coil dataset predictions and experimental chemical shifts reported for 14 IDPs. A dataset consisting of 14 IDPs with available NMR assignments were manually compiled as described previously (Tamiola et al. 2010). The RMSD between predicted and experimental chemical shifts were determined for each random coil dataset. The compared random coil values are Zhang et al. (2003), Schwarzinger et al. (2001), Wang and Jardetzky (2002), Wishart et al. (1995a), De Simone et al. (2009) and Tamiola et al. (2010)
Secondary chemical shift were calculated for the activation domain (residues 1018–1088) of the intrinsically disordered ACTR using the dataset reported here and the six other random coil datasets compared above (Fig. 7). Chemical shifts recorded at 5°C were deliberately chosen to allow the effect of temperature correction to be seen. When the random coil chemical shifts recorded at pH 2.3 are used, spikes are seen at the positions of aspartate and glutamate residues. In datasets that do not correct for the effects of neighbors (Zhang et al. 2003) spikes are seen in the residues preceding proline. Both of these types of spikes disappear when sequence corrected random coil chemical shifts recorded at pH 6.5 are used as the reference. Relative to several of the other datasets, the values reported here appear to have smaller fluctuations in the secondary chemical shifts consistent with the RMSD analysis discussed above. A transiently formed α-helix is observed from residue 1044–1054 using all random coil datasets. This region corresponds to the first α-helix in the complex between ACTR and its ligand, the nuclear coactivator binding domain from CBP (Demarest et al. 2002). This agrees with previous studies on ACTR (Kjaergaard et al. 2010a; Ebert et al. 2008) even though secondary chemical shifts data are less noisy when intrinsic random coil referencing is used (Kjaergaard et al. 2010a). Due to the small magnitude of the secondary chemical shifts, it is primarily Cα and C′ that detect this transiently formed helix. For this reason, these two nuclei are usually the only nuclei for which secondary chemical shifts are reported for IDPs. The effect of temperature correction can be seen most clearly for the C′ and HN secondary chemical shifts, where the values are systematically lower when the temperature correction values are applied. This suggests that temperature correction of the random coil chemical shifts, removes a systematic effect from the secondary chemical shifts. If this effect was not removed, it would cause a systematic bias resulting in overestimation of α-helical populations and underestimation of extended regions.
Comparison of secondary chemical shifts for the intrinsically disordered activation domain of ACTR (residues 1018–1088) using 7 different reference random coil datasets. The spikes in the Zhang ′03 dataset are from the residues preceding proline whereas the spikes in the Schwarzinger dataset are due to side chain protonation at low pH. The six nuclei represented are a Cα, b Cβ, c C′, d N, e HN, and f Hα
Conclusion
We have measured a set of random coil chemical shifts for a Ac-GGXGG-NH2 peptide series including the 20 common amino acid residues of proteins at pH 6.5 and in the temperature range from 5 to 45°C. The dataset overcomes a problem inherent in previous peptide based random coil dataset recorded at acidic pH. The pH dependence problem is prone to persist in database derived random coil datasets due to incomplete averaging of pH dependent chemical shift differences amongst the datasets in the database. Temperature coefficients for the random coil chemical shifts were extracted allowing extrapolation of the random coil values to the temperature at which the chemical shifts are determined. Temperature correction of random coil datasets will thus remove a systematic error from secondary chemical shifts caused by mismatch between the temperature of the random coil study and the protein sample of interest. The dataset reported here has been measured in order to improve the precision of secondary structure analysis of disordered proteins in particular, but it can also be used as a reference for secondary chemical shifts of folded proteins. Future studies will reveal whether peptide- or database derived random coil chemical shifts performs better for IDPs. Even if the database values are eventually found to be better than peptide based values, experimental studies are still useful for the understanding the effects of experimental conditions such as pH and temperature.
References
Bienkiewicz EA, Lumb KJ (1999) Random-coil chemical shifts of phosphorylated amino acids. J Biomol NMR 15:203–206
Braun D, Wider G, Wuthrich K (1994) Sequence-corrected N-15 random coil chemical-shifts. J Am Chem Soc 116:8466–8469
Bundi A, Wuthrich K (1979) H-1-Nmr parameters of the common amino-acid residues measured in aqueous-solutions of the linear tetrapeptides H-Gly-Gly-X-L-Ala-Oh. Biopolymers 18:285–297
Dames SA, Aregger R, Vajpai N, Bernado P, Blackledge M, Grzesiek S (2006) Residual dipolar couplings in short peptides reveal systematic conformational preferences of individual amino acids. J Am Chem Soc 128:13508–13514
Dawson RMC, Elliot DC, Elliot WH, Jones KM (1986) Data for biochemical research, 3rd edn. Oxford University Press, Oxford
De Simone A, Cavalli A, Hsu ST, Vranken W, Vendruscolo M (2009) Accurate random coil chemical shifts from an analysis of loop regions in native states of proteins. J Am Chem Soc 131:16332–16333
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) Nmrpipe—a multidimensional spectral processing system based on Unix pipes. J Biomol NMR 6:277–293
Demarest SJ, Martinez-Yamout M, Chung J, Chen HW, Xu W, Dyson HJ, Evans RM, Wright PE (2002) Mutual synergistic folding in recruitment of CBP/p300 by p160 nuclear receptor coactivators. Nature 415:549–553
Dyson HJ, Wright PE (2002) Insights into the structure and dynamics of unfolded proteins from nuclear magnetic resonance. Adv Protein Chem 62:311–340
Dyson HJ, Wright PE (2005) Intrinsically unstructured proteins and their functions. Nat Rev Mol Cell Bio 6:197–208
Ebert MO, Bae SH, Dyson HJ, Wright PE (2008) NMR relaxation study of the complex formed between CBP and the activation domain of the nuclear hormone receptor coactivator ACTR. Biochemistry 47:1299–1308
Eliezer D, Jennings PA, Dyson HJ, Wright PE (1997) Populating the equilibrium molten globule state of apomyoglobin under conditions suitable for structural characterization by NMR. FEBS Lett 417:92–96
Geen H, Freeman R (1991) Band-selective radiofrequency pulses. J Magn Reson 93:93–141
Hsu ST, Bertoncini CW, Dobson CM (2009) Use of protonless NMR spectroscopy to alleviate the loss of information resulting from exchange-broadening. J Am Chem Soc 131:7222–7223
Jimenez MA, Nieto JL, Rico M, Santoro J, Herranz J, Bermejo FJ (1986) A study of the Nh Nmr signals of Gly-Gly-X-Ala tetrapeptides in H2O at low-temperature. J Mol Struct 143:435–438
Kay LE, Keifer P, Saarinen T (1992) Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J Am Chem Soc 114:10663–10665
Kim HY, Heise H, Fernandez CO, Baldus M, Zweckstetter M (2007) Correlation of amyloid fibril beta-structure with the unfolded state of alpha-synuclein. Chembiochem 8:1671–1674
Kjaergaard M, Norholm AB, Hendus-Altenburger R, Pedersen SF, Poulsen FM, Kragelund BB (2010a) Temperature-dependent structural changes in intrinsically disordered proteins: formation of alpha-helices or loss of polyproline II? Protein Sci 19:1555–1564
Kjaergaard M, Teilum K, Poulsen FM (2010b) Conformational selection in the molten globule state of the nuclear coactivator binding domain of CBP. Proc Natl Acad Sci USA 107:12535–12540
Lam SL, Hsu VL (2003) NMR identification of left-handed polyproline type II helices. Biopolymers 69:270–281
MacArthur MW, Thornton JM (1991) Influence of proline residues on protein conformation. J Mol Biol 218:397–412
Merutka G, Dyson HJ, Wright PE (1995) ‘Random coil’ 1H chemical shifts obtained as a function of temperature and trifluoroethanol concentration for the peptide series GGXGG. J Biomol NMR 5:14–24
Modig K, Jurgensen VW, Lindorff-Larsen K, Fieber W, Bohr HG, Poulsen FM (2007) Detection of initiation sites in protein folding of the four helix bundle ACBP by chemical shift analysis. FEBS Lett 581:4965–4971
Mohan A, Oldfield CJ, Radivojac P, Vacic V, Cortese MS, Dunker AK, Uversky VN (2006) Analysis of molecular recognition features (MoRFs). J Mol Biol 362:1043–1059
Peti W, Smith LJ, Redfield C, Schwalbe H (2001) Chemical shifts in denatured proteins: resonance assignments for denatured ubiquitin and comparisons with other denatured proteins. J Biomol NMR 19:153–165
Plaxco KW, Morton CJ, Grimshaw SB, Jones JA, Pitkeathly M, Campbell ID, Dobson CM (1997) The effects of guanidine hydrochloride on the ‘random coil’ conformations and NMR chemical shifts of the peptide series GGXGG. J Biomol NMR 10:221–230
Ramboarina S, Redfield C (2003) Structural characterisation of the human alpha-lactalbumin molten globule at high temperature. J Mol Biol 330:1177–1188
Richarz R, Wuthrich K (1978) High-field 13C nuclear magnetic resonance studies at 90.5 MHz of the basic pancreatic trypsin inhibitor. Biochemistry 17:2263–2269
Schwarzinger S, Kroon GJ, Foss TR, Wright PE, Dyson HJ (2000) Random coil chemical shifts in acidic 8 M urea: implementation of random coil shift data in NMRView. J Biomol NMR 18:43–48
Schwarzinger S, Kroon GJ, Foss TR, Chung J, Wright PE, Dyson HJ (2001) Sequence-dependent correction of random coil NMR chemical shifts. J Am Chem Soc 123:2970–2978
Silver MS, Joseph RI, Hoult DI (1985) Selective spin inversion in nuclear magnetic resonance and coherent optics through an exact solution of the Bloch-Riccati equation. Phys Rev A 31:2753–2755
Spera S, Bax A (1991) Empirical correlation between protein backbone conformation and C.alpha. and C.beta. 13C nuclear magnetic resonance chemical shifts. J Am Chem Soc 113:5490–5492
Tamiola K, Acar Bi, Mulder FAA (2010) Sequence-specific random coil chemical shifts of intrinsically disordered proteins. J Am Chem Soc 132:18000–18003
Thanabal V, Omecinsky DO, Reily MD, Cody WL (1994) The C-13 chemical-shifts of amino-acids in aqueous-solution containing organic-solvents—application to the secondary structure characterization of peptides in aqueous trifluoroethanol solution. J Biomol NMR 4:47–59
Ting D, Wang G, Shapovalov M, Mitra R, Jordan MI, Dunbrack RL Jr (2010) Neighbor-dependent Ramachandran probability distributions of amino acids developed from a hierarchical Dirichlet process model. PLoS Comput Biol 6:e1000763
Uversky VN (2009) Intrinsically disordered proteins and their environment: effects of strong denaturants, temperature, pH, counter ions, membranes, binding partners, osmolytes, and macromolecular crowding. Protein J 28:305–325
Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas P, Ulrich EL, Markley JL, Ionides J, Laue ED (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696
Wang Y, Jardetzky O (2002) Investigation of the neighboring residue effects on protein chemical shifts. J Am Chem Soc 124:14075–14084
Wang L, Eghbalnia HR, Markley JL (2007) Nearest-neighbor effects on backbone alpha and beta carbon chemical shifts in proteins. J Biomol NMR 39:247–257
Wishart DS, Sykes BD (1994) Chemical shifts as a tool for structure determination. Methods Enzymol 239:363–392
Wishart DS, Sykes BD, Richards FM (1991) Relationship between nuclear magnetic resonance chemical shift and protein secondary structure. J Mol Biol 222:311–333
Wishart DS, Sykes BD, Richards FM (1992) The chemical shift index: a fast and simple method for the assignment of protein secondary structure through NMR spectroscopy. Biochemistry 31:1647–1651
Wishart DS, Bigam CG, Holm A, Hodges RS, Sykes BD (1995a) 1H, 13C and 15 N random coil NMR chemical shifts of the common amino acids. I. Investigations of nearest-neighbor effects. J Biomol NMR 5:67–81
Wishart DS, Bigam CG, Yao J, Abildgaard F, Dyson HJ, Oldfield E, Markley JL, Sykes BD (1995b) 1H, 13C and 15 N chemical shift referencing in biomolecular NMR. J Biomol NMR 6:135–140
Wu KP, Kim S, Fela DA, Baum J (2008) Characterization of conformational and dynamic properties of natively unfolded human and mouse alpha-synuclein ensembles by NMR: implication for aggregation. J Mol Biol 378:1104–1115
Zhang H, Neal S, Wishart DS (2003) RefDB: a database of uniformly referenced protein chemical shifts. J Biomol NMR 25:173–195
Acknowledgments
This work was supported by the EliteForsk programme (M.K.), The John and Birthe Meyer Foundation, the Carlsberg Foundation grant number 2008-01-0368 and The Danish Natural Research Council grant numbers 272-08-0500 (F.M.P). We thank Kamil Tamiola and Frans Mulder (University of Groningen) for sharing their data for comparison of the random coil data sets and Gitte Wolfsberg Haxholm and Birthe B. Kragelund for valuable discussions and critical comments to the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kjaergaard, M., Brander, S. & Poulsen, F.M. Random coil chemical shift for intrinsically disordered proteins: effects of temperature and pH. J Biomol NMR 49, 139–149 (2011). https://doi.org/10.1007/s10858-011-9472-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-011-9472-x