Abstract
1H NMR spectroscopy of sera from HIV-1 infected and uninfected individuals was performed on 300 and 600 MHz instruments. The resultant spectra were automatically data reduced to 90 and 180 integral segments of equal length. Analysis of variance identified significant differences between the sample groups, especially for the samples analyzed on 600 MHz and reduced to fewer segments. Linear discriminant analysis correctly classified 100% of the samples analyzed on the 300 MHz NMR (reduced to 180 segments); an increase in instrument sensitivity resulted in lower percentages of correctly classified samples. Multinomial logistic regression (MLR) resulted in 100% correct classification of all samples from both instruments. Thus 1H-NMR metabonomics on either instrument distinguishes HIV-positive individuals using or not using anti retroviral therapy, but the sensitivity of the instrument impacts on data reduction. Furthermore, MLR is a novel multivariate statistical technique for improved classification of biological data analyzed in NMR.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diseases cause disturbances in the concentrations and fluxes of endogenous metabolites involved in cellular pathways and, due to the homeostatic nature of the cellular environment, cells respond to stressors generally by adjustment of their extra-cellular environment. This metabolic adjustment is expressed as a fingerprint of biochemical perturbations, which is characteristic of the nature or site of a disease process (Lindon 2004). Metabonomics, defined as “the quantitative measurement of the dynamic multiparametric metabolic response of living systems to pathophysiological stimuli or genetic modification” (Nicholson et al. 1999) has contributed to the identification and study of human disease, especially in the investigation of toxicological mechanisms, disease diagnosis and response to therapy. NMR-based metabonomics consists of using high-resolution proton NMR (1H-NMR) spectroscopic profiling of biological fluids combined with multivariate analysis, to identify low molecular weight metabolites that correlate with changes in physiological conditions (Nicholson and Wilson 1989). High-resolution 1H NMR spectroscopy datasets are represented as complex matrices with several hundreds of proton signals originating from the various metabolites. This complexity can be simplified by the application of multivariate, statistical, pattern recognition and informatics (chemometric) methods that are used to reduce the dimension of the 1H-NMR data for visualization purposes, and to identify inherent patterns among sets of spectral measurements (Ramadan et al. 2006).
This study compared serum metabolic profiles of healthy HIV-1 negative individuals, HIV-1 positive volunteers not using ART and HIV-1 positive volunteers on ART using both 300 and 600 MHz 1H NMR spectroscopy. The goal was to detect metabolites influenced or disturbed by HIV and/or ART, possibly detecting even minor differences in metabolite profiles when using the more sensitive instrument. The resultant spectra were automatically data reduced to 90 and 180 integral segments (variables) of equal length, in order to investigate how data reduction influenced chemometrics-assisted classification of the groups as either HIV-1 negative or positive on treatment or not. Analysis of variance (ANOVA); linear discriminant analysis (LDA) and multinomial logistic regression (MLR) was used to identify areas of variability, and to find a linear combination of amplitudes which best separate the different samples. MLR is a relatively new multivariate statistical technique that has not been used in metabonomic research, and hence this study investigated whether it could be a novel multivariate statistical technique for classification of this form of biological data.
Analysis of variance identified significant differences between the three sample groups. LDA was able to correctly classify 100% of the samples analyzed on 300 MHz NMR, 93.1% of samples from the 600 MHz analysis and 95.2% correct classification of samples analyzed under optimal 600 MHz conditions (all of the above was for 180 segmented data). The MLR correctly classified 100% of all samples analyzed on both instruments (90-variable data). These data suggests that 1H-NMR metabonomics on either instrument distinguishes between HIV-positive individuals using and not using ART, however, sensitivity of the instrument influences statistical processing of reduced data (e.g., fewer segments for better classification using MLR).
Materials and methods
Sample collection
Blood was drawn from 39 HIV-1 positive patients attending the HIV/AIDS clinic at the Helen Joseph Hospital in Johannesburg, South Africa. Patient recruitment criteria was limited to HIV status, CD4 count, absence of any other signs of AIDS and knowledge of usage of ART since exact side-effects were not at issue here only the ability to distinguish HIV or ART influenced metabolites using NMR.
At the time of collection, 19 patients were undergoing treatment with combination antiretroviral therapy while the other 20 patients were not on treatment. All patients on treatment started ART when their CD4 counts were below 200 cells/μl blood and had been on treatment for less than 1 year at the time of sample collection. Patients not on treatment had CD4 cell counts in excess of 200 cells/μl blood. Hospital records confirmed the status, anti-retroviral treatment use and general well-being of the patients. HIV positive individuals were selected for the study based on the fact that they were not on treatment for any secondary infections at the time of blood collection.
Blood samples were allowed to clot in non-anti-coagulant vacutainers (Greiner Bio-One, Germany) for 2 h at room temperature (RT) and sera was collected after centrifugation at 1028×g. Aliquots (500 μl) of all serum samples were stored at −80°C until assayed on either 300 or 600 MHz MNR instrumentation. Samples were thawed only once on the day of analysis when it was also diluted with Deuterium oxide (D2O). Aliquots of the same 9 ART and 10 non-ART patient sera were analyzed on both 600 and 300 MHz NMR using the same protocol. An additional 10 ART and 10 non-ART samples were analyzed using the optimized 600 MHz NMR protocol.
HIV negative sera was collected from 21 HIV-1 negative laboratory personnel (status confirmed with HIV 1/2 Determine™ rapid tests, Abbott Laboratories, Abbott Park, IL.). Samples were aliquoted as explained before; aliquots of the same 10 samples were analyzed on both instruments using the standard protocol and aliquots of 11 additional HIV negative samples analyzed with the optimized 600 MHz protocol. Informed consent was obtained from all volunteers and patient anonymity was maintained throughout.
1H NMR spectroscopy
Aliquots of the serum samples (500 μl) were diluted with 200 μl of 52% D2O (Deuterium oxide, 99.9% atom % D, Aldrich Chemicals Company, South Africa) and placed in 5 mm high quality NMR tubes (Sigma Aldrich, RSA). Conventional 1H NMR spectra of each serum sample was measured at 300 MHz on a Varian Unity Inova using an IDG300–5EHT indirect detection gradient probe. The following pulse sequence was used: 90°–90°-RD–satdly–t m-90°- acquire free induction decay (FID). RD represents a relaxation delay of 2 s, saturation delay (satdly, also called saturation pulse length) was 1.5 s, during which the water resonance was selectively irradiated, and the mixing time (t m) was 0.4 ms. For each sample, 128 FIDS were collected with a spectral width (also called a spectral window) of 4001.6 Hz and an acquisition time of 3.744 s. The acquired NMR spectra were collected for phase and baseline distortions and referenced to alanine at 1.30 ppm. Different aliquots of the same samples were then run on a Varian Unity Inova (Department of Chemistry and Polymer Science, University of Stellenbosch, South Africa) using a 5 mm HCN indirect detection PFG probe operating at 600 MHz for 1H, with exactly the same conditions as the 300 MHz analysis.
Due to the fact that these settings were optimized for the 300, 600 MHz analysis was repeated with the following optimized settings: pulse angle of 90°, relaxation delay of 3 s, and a saturation delay of 1.5 s. For each sample, 64 FIDS were collected with a spectral width of 8000 Hz, and an acquisition time of 4 s.
Data reduction of NMR data
The 1H-NMR spectra obtained for each serum sample were automatically data reduced to 90 integral segments (variables) of equal width (δ 0.04 ppm), corresponding to the δ range 4.0–0.4 ppm. In order to evaluate whether data reduction would influence the results, the original 1H-NMR spectra were also reduced to 180 variables of equal length (δ 0.02 ppm). Each segment consisted of the integral of the NMR region with which it was associated. Although the area in the suppression of the water resonance was not included, the region δ 6.0–4.4 was set to zero integral.
Statistical analysis
All statistical analysis was performed by the Statistics Department of the University of Pretoria, using SPSS (version 14.0) software.
Standardization of the data
The original data was standardized by calculating a mean and standard deviation per individual, then subtracting the mean from each individual’s amplitude and dividing the result by the standard deviation:
where a fi is the amplitude at frequency f for the ith individual; \( \bar{a}_{i} \) is the mean value of the 90/180 amplitudes for the ith individual; s i is the standard deviation of the 90/180 amplitudes for the ith individual; z fi is the standardized amplitude at frequency f for the ith individual.
The result was that the standardized spectra for each individual had a mean of 0 and a standard deviation of 1.
Analysis of variance
Comparison of the mean standardized spectra of the three groups (HIV negative individuals, HIV patients not using ART and HIV patients using ART) was done by means of Analysis of Variance F-tests. This was done in order to establish whether or not there were significant differences between the groups. Only cases where the significance level (P-value) was less than 0.05 were considered.
Discriminant analysis
Linear discriminant analysis (LDA) was used to classify a case as HIV positive using ART, HIV positive not using ART or HIV negative, by finding the linear combination of amplitudes which best separate the classes, given the NMR spectrum (standardized, as described above). Stepwise discriminant analysis was used to select the best set of frequencies for the classification. The resultant classification was based on 6 of the spectrum amplitudes, in the 300 MHz data, and 5 of the spectrum amplitudes, in the 600 MHz data, which showed the greatest variation between the three groups. The optimized 600 MHz analysis resulted in classification based on 3 amplitudes. A cross-validation was done in which each case was omitted in turn from the analysis and then classified using the analysis based on the other cases. The two discriminant functions, produced by this analysis, were plotted on a scatter diagram.
Logistic regression
Logistic regression is a model for the probability that an individual case belongs to a particular group. In the present study a stepwise technique was used to select the spectrum amplitudes, which would result in the best classification of the three groups. In both cases (300 and 600 MHz) 3 spectrum amplitudes were selected for the classification. In the optimized 600 MHz analysis, 4 and 3 spectrum amplitudes were selected for the 90 and 180 variable data, respectively.
Results and discussion
Clinical and metabolic complications, associated with HIV infection and the use of ART require constant monitoring for the successful management of HIV/AIDS. 1H-NMR on a 300 MHz spectrometer distinguished HIV and ART induced metabolic profiles (Hewer et al. 2006) where sample groups clustered separately following pattern recognition statistical evaluation. Because recent biomedical applications of metabonomics are mostly performed at increased levels of sensitivity (Nicholson et al. 1999, 2002; Brindle et al. 2002), better distinction between the experimental groups may be obtained with higher NMR frequencies and/or minor differences between experimental groups highlighted by more sensitive instrumentation.
The purpose of this study was to determine if there was any effect on the clustering and classification of results (as HIV positive on treatment or not and HIV negative) when the samples were analyzed using different spectrometer observation frequencies and/or when different data reduction and statistical analysis was performed.
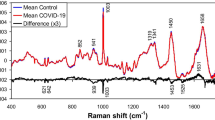
The application of multivariate statistical, pattern recognition and informatic methods were needed in order to establish significant differences in the profiles of the NMR spectra. NMR spectra of human sera are visually similar due to the tight homeostatic control of serum. The original data was standardized (as explained in Section “Standardization of the data”) and Fig. 1 shows the differences between the mean spectra of the three sample groups for the different instruments. The resonances produced by the various metabolites can be assigned directly based on their chemical shifts and signal multiplicities. The main differences observed between the three groups were in the following regions of the spectra: 1.3–1.7 and 3.2–3.9 ppm. These regions represent various lipids and glycerol, glucose and amino acids, respectively (Nicholson et al. 1995; Ala-Korpela 1995; Liu et al. 1997).
Analysis of variance
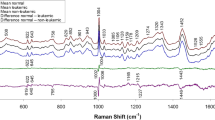
Data reduction simplifies statistical analysis, and reduces the impact of small variations in chemical shift (Keun et al. 2002). In the 300 MHz analysis, 37 of the total 90 segments (41%) and 72 of the 180 segments (40%) were identified through ANOVA as significantly dissimilar (P < 0.05, Fig. 2), with many of them being adjacent to one another. However, in the 600 MHz analysis performed under the same conditions, only 8 of the 90 segments (9%) and 19 of the 180 segments (11%) were identified as significantly different, P < 0.05. When the 600 MHz analysis was performed under optimized conditions, 42 of the 90 segments (47%), and 38 of the 180 segments (21%) were identified as being significantly different, P < 0.05 (Fig. 2). The chemical components that these significantly different segments represent are mainly lipids, including LDL and VLDL, which correlate with known metabolic irregularities of HIV/AIDS patients using ART, such as lipodystrophy and hyperlipidaemia (Calza et al. 2003). ANOVA data suggests that distinction between HIV negative and HIV positive with or without ART treatment can be determined by 1H-NMR metabonomics on either instrument, 300 or 600 MHz. Optimization of the 600 MHz protocol increased the number of segments that differ significantly between groups (compared to standard 600 MHz), especially for the lower number of segments.
Comparison of significantly different integral amplitudes from analysis of variance (ANOVA) between HIV negative individuals, HIV-1 positive/AIDS patients and HIV-1 positive/AIDS patients utilizing anti-retroviral therapy. The data obtained from the optimized 600 MHz analysis segmented into 90 integrals identified the most differences in the data, followed by the 180-segmented data observed for the 300 MHz analysis. A decrease in the number of segments (and therefore an increase in the size of the integral segment) is required when the sensitivity of the instrument increases
Discriminant analysis
Linear discriminant analysis (LDA) easily handles cases where the ‘within-class’ frequencies are unequal and their performances have been examined on randomly generated test data. This method maximizes the ratio of ‘between-class’ variance to the ‘within-class’ variance in any particular data set thereby guaranteeing maximal separability (Fukunaga 1990). Table 1 shows the LDA classification of samples as HIV-1 positive on ART, HIV-1 positive not on ART or HIV-1 negative, with original grouped cases and cross-validated grouped cases. LDA resulted in 96.6% of samples correctly classified from the original grouped cases for both the 300 and 600 MHz analyses (90-segmented data). The cross-validated grouped cases (labeled as ‘naïve cross-validation in Table 1) revealed a slightly higher classification for the 600 MHz data (89.7%) than the 300 MHz analysis (86.2%). Better classification was observed for the 300 MHz 180-segmented data with 100% of the original grouped cases and 96.6% of the cross-validated cases correctly classified. Increasing the number of variables did not improve the 600 MHz analysis with 93.1% of the original grouped cases and 82.8% of the cross-validated cases correctly classified. Optimizing the 600 MHz protocol showed more variables (180) giving the better classification percentage for the original and cross-validated cases. Full cross validation lowered the % correct classification for both instruments and protocols.
Table 2 provides a detailed performance record of main discriminant analyses and classification results showing exactly which groups are being misclassified into which other groups. From these records it’s also possible to calculate LDA error rates. The error rates in the 600 MHz LDA models were higher than in the 300 MHz models, which do not necessarily imply that the error rates were high in general. In the case of 300 MHz the error rates were 3.4% (90 bins) and 0% (180 bins). In the case of the original 600 MHz analysis the error rates were 3.4% (90 bins) and 6.9% (180 bins) which is higher, but not high overall (3.4% means 1 case, 6.9% means 2 cases incorrectly classified). In the “enhanced” (optimized) data set the error rates were a lot higher. Table 3 provides the P-values of variables used in LDA. Error bar graphs (19 charts) of the main variables featuring in the discriminant analysis are provided as an appendix (electronic supplementary material), these give an indication of the degree of differential regulation between groups.
LDA classification of samples in a two-population comparison between individuals on ART who show no AIDS related symptoms (other than HIV status) and healthy HIV negative individuals showed 100% correct classification for both instruments and protocols (Table 4, ii). Correct LDA classifications were also possible for a comparison between HIV positive on ART and positive patients not on treatment for both instruments but not the optimized protocol (Table 4, iii). When comparing HIV negative samples to HIV positive patients (not on ART, Table 4, i) and negative compared to positive regardless of ART (Table 4, iv), the highest correct classifications was for the data collected on the 300 MHz instrument. Two-population comparison data processing was done to demonstrate that all sample groups evaluated in this study were statistically different/well-defined. Metabolites that correlate to the variables selected in (Table 4, i) are leads for biomarkers of HIV status while variables selected for other classifications are indicative of metabolites/biomarkers prone to interference by ART.
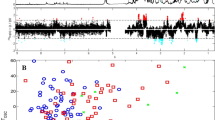
The LDA distinction between groups in the three-population study is better seen graphically in Fig. 3, the resulting scatter diagram of the discriminant functions. The 300 MHz analysis clearly clustered the samples into three different groups (Fig. 3a), whereas in the 600 MHz analysis, the HIV-1 positive with ART cluster is more dispersed and overlaps slightly with the HIV-1 negative group (Fig. 3b). When the study was performed using conditions optimized for the 600 MHz NMR, the resulting scatter diagram (Fig. 3c) shows that most samples in the groups cluster separately but there’s overlap of some samples in all three groups. It is possible that due to higher sensitivity, the 600 MHz spectrometer detects minor changes in metabolite profiles which may mask large differences between the groups (detected by 300 MHz). Higher numbers of minor differences recorded may produce fewer significant variables between the groups hampering accurate classifications.
Scatter plots produced from linear discriminant analysis (LDA) of the 180-segmented data showing distinction between HIV negative, HIV positive patients not on anti-retroviral therapy and HIV-1 positive patients utilizing anti-retroviral therapy. The 300 MHz analysis (a) shows better cluster formation of samples when compared to 600 MHz analysis (b). Optimization of the 600 MHz analysis resulted in a decrease in cluster formation (c). Group centroids indicated by a square sometimes obscures sample symbols and two or more samples sometimes occupy the same space on the graph
Logistic regression
Logistic regression is a relatively new statistical technique and unlike LDA, the logistic regression model does not rely on an assumption of multivariate normality. Furthermore, categorical variables such as gender or population group, if available, may be factored into the model, which is not the case with LDA. The logistic regression model is a linear model in the log-odds {ln(P/(1 − P))} where p is the probability being modeled, resulting in an s-shaped curve for the model of P. The MLR accurately classified 100% of all samples from both 300 and 600 MHz analyses with both sets of data (90 and 180 segments, Table 5). A decrease in classification was seen when MLR was performed on the 180-segmented data obtained from the optimized 600 MHz analysis, however, the 90-variable data resulted in 100% classification for all samples. This finding is consistent with the ANOVA results, whereby the fewer number of variables resulted in the best results for the more sensitive instrument.
This stepwise technique was able to predict classification of samples into the three groups based on three spectral amplitudes: 3.03, 3.11, and 3.33 from the 300 MHz data, and 1.43, 2.17, and 3.03 from the 600 MHz data (180 segmented data, Table 6). The SPSS software chose these amplitudes, as they were the most significantly different amplitudes between the three groups. The amplitude 3.03 (P-value, 0.000003), used to classify both the 300 and 600 MHz data corresponds to the neural metabolite choline. Previous studies have shown that patients with AIDS often show varying degrees of cognitive, motor, and behavioral impairment, including dementia (AIDS dementia complex, ADC; Navia et al. 1986), where an increase in choline is one biomarker used in a predictive model of ADC (Meyerhoff et al. 1999). This preliminary finding (detecting choline) needs to be verified and is worthy of further investigation (e.g., through quantitative NMR to measure choline concentration fluctuations and correlating these with ADC status, using higher sample numbers etc.) because the amplitude for this metabolite was also selected for LDA in two- and three-population analyses. Although none of the patients demonstrated neural impairment, the data collected confirms the sensitivity of biofluid NMR for use as a predictor of disease prior to other symptoms being visible.
Concluding remarks
Metabolic changes induced by HIV and/or ART can be monitored using either 300 or 600 MHz NMR spectrometry, as the statistical analysis post-data acquisition is able to classify samples into separate distinct groups based on their respective metabolites. The best results are obtained when the data reduction and the sensitivity of the instruments are both considered. This study also illustrated that MLR can be an effective tool in metabonomic research.
References
Ala-Korpela M (1995) 1H NMR spectroscopy of human blood plasma. Prog NMR Spectrosc 27:475–554
Brindle JT, Antti H, Holmes E, Tranter G, Nicholson JK, Bethell HW, Clarke S, Schofield PM, McKilligin E, Mosedale DE, Grainger DJ (2002) Application of chemometrics to 1H NMR spectroscopic data to investigate a relationship between human serum emtabolic profiles and hypertension. Nat Med 8:1439–1444
Calza L, Manfredi R, Chiodi F (2003) Hyperlipidaemia in patients with HIV-1 infection receiving highly active antiretroviral therapy: epidemiology, pathogenesis, clinical course and management Int. J Antimicrob Agents 22:89–99
Fukunaga K (1990) Introduction to statistical pattern recognition. Academic Press, San Diego
Hewer R, Voster J, Steffens FE, Meyer D (2006) Applying biofluid 1H NMR-based metabonomic techniques to distinguish between HIV-1 positive/AIDS patients on antiretroviral treatment and HIV-1 negative individuals. J Pharm Biomed Anal 41(4):1442–1446
Keun HC, Ebbels TM, Antti H, Bollard ME, Beckonert O, Schlotterbeck G, Senn H, Niederhauser U, Holmes E, Lindon JC, Nicholson JK (2002) Analytical reproducibility in 1H NMR-based metabonomic urinalysis. Chem Res Toxicol 15:1380–1386
Lindon JC (2004) In: Cooper E (ed) Business briefing: future drug discovery. World Markets Research Center Ltd, London, pp 1–6
Liu M, Nicholson JK, Parkinson JA, Lindon JC (1997) Measurement of biomolecular diffusion coefficients in blood plasma using two-dimensional 1H-1H diffusion-edited total-correlation NMR spectroscopy. Anal Chem 69:1504–1509
Meyerhoff DJ, Bloomer C, Cardenas V, Norman D, Weiner MW, Fein G (1999) Elevated subcortical choline metabolites in cognitively and clinically asymptomatic HIV+ patients. Neurology 52:995–1003
Navia BA, Cho ES, Petito CK, Price RW (1986) The AIDS dementia complex: II. Neuropathology. Ann Neurol 19:525–535
Nicholson JK, Wilson ID (1989) High resolution proton magnetic resonance spectroscopy of biological fluids. Prog NMR Spectrosc 21:449–501
Nicholson JK, Foxall PJD, Spraul M, Farrant DR, Lindon JC (1995) 750 MHz and 1H-13C NMR spectroscopy of human blood plasma. Anal Chem 67:793–811
Nicholson JK, Lindon JC, Holmes E (1999) ‘Metabonomics’: understanding the metabolic responses of living systems to pathophysiological stimuli via multivariate statistical analysis of biological NMR data. Xenobiotica 29:1181–1189
Nicholson JK, Connelly J, Lindon JC, Holmes E (2002) Metabonomics: a platform for studying drug toxicity and gene function. Nat Rev Drug Discov 1:153–161
Ramadan Z, Jacobs D, Grigorov M, Kochhar S (2006) Metabolic profiling using principal component analysis, discriminant partial least squares and genetic algorithms. Talanta 68:1683–1691
Acknowledgments
This research was funded by University of Johannesburg Faculty of Science.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Philippeos, C., Steffens, F.E. & Meyer, D. Comparative 1H NMR-based metabonomic analysis of HIV-1 sera. J Biomol NMR 44, 127–137 (2009). https://doi.org/10.1007/s10858-009-9329-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-009-9329-8