Abstract
The focus of the computational structural biology community has taken a dramatic shift over the past one-and-a-half decades from the classical protein structure prediction problem to the possible understanding of intrinsically disordered proteins (IDP) or proteins containing regions of disorder (IDPR). The current interest lies in the unraveling of a disorder-to-order transitioning code embedded in the amino acid sequences of IDPs/IDPRs. Disordered proteins are characterized by an enormous amount of structural plasticity which makes them promiscuous in binding to different partners, multi-functional in cellular activity and atypical in folding energy landscapes resembling partially folded molten globules. Also, their involvement in several deadly human diseases (e.g. cancer, cardiovascular and neurodegenerative diseases) makes them attractive drug targets, and important for a biochemical understanding of the disease(s). The study of the structural ensemble of IDPs is rather difficult, in particular for transient interactions. When bound to a structured partner, an IDPR adapts an ordered conformation in the complex. The residues that undergo this disorder-to-order transition are called protean residues, generally found in short contiguous stretches and the first step in understanding the modus operandi of an IDP/IDPR would be to predict these residues. There are a few available methods which predict these protean segments from their amino acid sequences; however, their performance reported in the literature leaves clear room for improvement. With this background, the current study presents ‘Proteus’, a random forest classifier that predicts the likelihood of a residue undergoing a disorder-to-order transition upon binding to a potential partner protein. The prediction is based on features that can be calculated using the amino acid sequence alone. Proteus compares favorably with existing methods predicting twice as many true positives as the second best method (55 vs. 27%) with a much higher precision on an independent data set. The current study also sheds some light on a possible ‘disorder-to-order’ transitioning consensus, untangled, yet embedded in the amino acid sequence of IDPs. Some guidelines have also been suggested for proceeding with a real-life structural modeling involving an IDPR using Proteus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
After extensive research over one-and-a-half decades, it is evident that many functional proteins lack well-folded 3D structures. These intrinsically disordered proteins (IDPs), could be completely disordered or contain intrinsically disordered protein regions (IDPRs) [1–5]. In contrast to the classical view of protein folding [6], where a nascent cytoplasmic polypeptide chain folds into a stable globule, concomitantly while being synthesized [7, 8], these proteins are born disordered [3] and remain either completely or partially unstructured throughout their entire life span. It is only when they interact with functionally relevant binding partners that they switch to ordered structures [4]. In fact, their existence in a biologically active form without adapting to a unique 3D-structure contradicts the traditional notion of the “one protein–one structure–one function” paradigm [1].
IDPs are highly abundant in nature and have been found to be involved in a number of functions within the living cell, most of which belong to the non-classic (non-enzyme) type [9, 10]. They possess remarkable binding promiscuity [4] in a wide range of intermolecular interactions, complementing the functional repertoire of ordered globular proteins, similar to the phenomena of enthalpy—entropy compensation [11]. The promiscuity is primarily manifested in their ability to interact specifically with structurally diverse molecular partners and obtaining different structures upon binding. It is highly likely that these peculiar characteristics may be attributed to their non-native-like multi-funneled and relatively flat energy landscapes [12, 13], wherein the favored conformations closely resemble to the partially folded molten globules [13] which also enable them to preserve the necessary amount of disorder even in their bound forms [4]. Considering this flexible nature, they have been referred to as part of the ‘edge of chaos’ systems [14], serving as a bridge between well-ordered and chaotic system that is critical in the context of cellular energy balance.
In addition to these peculiar biophysical and folding attributes, IDPs are also of considerable biomedical interest due to their functional importance. In fact, the functions they are involved in (e.g., regulation, signaling, and control) are mostly the ones that require high specificity–low-affinity interactions [15]. Recent studies have highlighted their multifarious activities as molecular rheostats and molecular clocks, in tissue specific and alternative splicing of mRNA, transport of rRNA and proteins and RNA-chaperons [16]. Also, by sustaining enough disorder even in the bound form, IDPs are equipped to participate in both one-to-many and many-to-one signaling [2]. Their promiscuity in binding also suggests that not only misfolding [17], but also misidentification or mis-signaling [2] in biomolecular recognition could serve as the root cause of some extremely complex human diseases [3] including cancer, diabetes, amyloidoses, and cardiovascular and neurodegenerative diseases [18].
Taken together, there is a great need for a deeper understanding of IDPs and their interactors. However, since obtaining information about IDPs from experiments is difficult owing to their inherent disorder, computational modeling provides a realistic way forward. For most IDPs, only a subset of the disordered residues can actually undergo a disorder-to-order transition, upon binding to a folded protein, leading to the concept of ‘folding coupled with binding’ [19]. These segments are called protean borrowed from Greek mythology, meaning ‘ever-changeable’ or ‘mutable’ [19]. To model the 3D structure of an interacting IDP/IDPR, a first aim would be to predict the potential ‘mutable’ protean regions. It is important to note that, due to the intrinsic disorder, these regions in an isolated X-ray structure are presented as ‘missing electron density’ patches (listed in REMARK 465 in the corresponding PDB file [5]), and should only appear structured in their bound forms. This is in fact also the definition of a ‘protean’ segment.
Extensive studies have analyzed the sequence space of IDPs in relation to their intrinsic disorder. These studies reveal their correspondence to low entropy sequences with less complexity [5, 20]. In particular, tandem repeats have often been found to be embedded in these sequences (e.g. polyglutamine stretches in amyloid beta [21]) giving rise to the notion of ‘the more perfect the less structured’ proteins [22]. Thus, in a sense, low sequence entropy can potentially lead to high conformational entropy, characteristic of the IDPs. Some mechanistic insights into the origin of the disorder have also been suggested, for example, the low content of hydrophobic residues with an abundance of charged residues in IDPs [23] disfavoring self-folding [24] by potentially decreasing the number of possible two-body contacts [25]. Furthermore, the charge—hydrophobicity boundary have been envisaged to represent a trade-off between repulsive and attractive interactions reminiscent of globular—disorder transitions [26].
Nevertheless, it remains highly challenging to decipher the root cause of intrinsic disorder from pure sequence-based investigation given the limited structural data. Concerted efforts have been made to untangle a possible disorder code from amino acid sequence alone which includes deciphering the propensity for intrinsic disorder [26], and proposition of statistical mechanical potentials describing sequence-derived elasticity [27]. The nature of the problem is ideal for machine learning algorithms given the availability of annotated sequence data. In fact, quite a few predictors have recently been developed that predict not only the disordered regions [28–32], but also the ‘protean’ segments [32–35]. Still, ‘protean prediction’ is in an early stage, offering much room for improvement. In this background, the current study does not only attempt to shed some light on a possible yet unexplored sequence consensus of such ‘disorder-to-order’ transitions, but also presents ‘Proteus’, a random forest classifier that predicts protean segments solely from the amino acid sequence of an IDP. Proteus compares favorably to the existing predictors. Some guidelines have also been suggested on how and where to use Proteus during the course of a real-life structural modeling involving an IDPR.
Methods
Training dataset
Two databases containing proteins with annotated protean segments, IDEAL [19] and MoRF [34] were pulled together to build the final training dataset. IDEAL (Intrinsically Disordered proteins with Extensive Annotations and Literature) contains 557 proteins with experimentally verified protean segments called ‘ProS’ in the database. However, only 203 of 557 proteins in this database actually contain protean segments. The rest are IDPs where no protean segments have yet been experimentally verified and thus serve as negative examples in training. The MoRF dataset comes from MoRFpred [34], one of the existing classifiers. It contains 840 proteins, and all of them have at least one protean segment. More importantly, all members of MoRF have direct structural evidence from the PDB. Members from IDEAL and MoRF will henceforth be referred to as ‘ProS’ and ‘MoRF’ respectively, and the combined dataset as ‘PnM’. The details of all datasets have been enlisted in Table 1.
Independent benchmark
Nine proteins that were used as independent benchmark in the DISOPRED3 study [32] were used as an independent benchmark set here as well. In the DISOPRED3 study 29 chains having protean segments were initially culled using database annotations and publications, which later had to be reduced to nine proteins, as the other 20 chains were found to be used in the training datasets of the competing methods, ANCHOR [33], MoRFpred [34] or MFSPSSMPred [35]. None of the nine proteins were similar to any protein in the current training dataset.
Target function
The binary status for each amino acid residue in the sequence to be protean or non-protean was used as the target function in training the classifier, denoted by 1 and 0 respectively, for positive and negative examples. It is to be noted that non-protean residues refer to the ordered as well as disordered residues which do not undergo the ‘disorder-to-order transition’ upon binding and hence, remain disordered even in the bound state.
Data clustering and cross-validation benchmark
To avoid training and testing on similar examples, BLASTclust was used to cluster the protein sequences in the combined dataset ‘PnM’. Sequences with a pairwise similarity of at least 30% over at least 50% of the sequence length (-S 30 -L 0.5) were clustered. This resulted in 774 clusters, the largest containing 38 proteins, and 253 clusters containing more than one protein. One-third of all ProS sequences were found to be similar to at least one MoRF sequence and vice-versa.
To prepare the data for fivefold cross-validation, five folds were built by grouping clusters in such a way that the number of target proteins remain consistent among the folds. This resulted in four folds with 280 targets and one fold with 279 targets, containing between 158,651 and 218,870 amino acid residues, and around 1.4–2.2% positive examples. During cross-validated training, four of the folds are used for training and the remaining one is used for testing. This is repeated five times to make predictions for all five folds.
Random forest classifier
The random forest classifier module in scikit-learn Python package [36] was used for training. Every decision tree in the forest classify examples as positive or negative, and a final decision is made according to a majority vote.
Evaluation measures
In binary classification, there are four possible outcomes when classifying an example: (i) True Positive (TP): a positive example correctly classified as positive; (ii) True Negative (TN): a negative example, correctly classified as negative; (iii) False Negative (FN): a positive example incorrectly classified as negative; and (iv) False Positive (FP): a negative example incorrectly classified as positive. By counting these four possible outcomes, the following evaluation measures were calculated.
Precision (PPV)
Precision, also known as specificity or the Positive Predicted Value (PPV), measures how many examples classified as positive were actually positive, calculated by the ratio, TP/(TP + FP).
Recall (TPR)
Recall (or coverage) measures how many positive examples were correctly classified as positives. It is also called the ‘True Positive Rate’ (TPR) and calculated by the ratio, TP/∑P, where ∑P is the total number of positives, i.e., ∑P = TP + FN.
F1-score
F1-score is the harmonic mean between PPV and TPR and could be interpreted as a trade-off between PPV and TPR. It is defined by the following equation: F1 = 2PPV × TPR/(PPV + TPR).
Matthews correlation coefficient
Another direct evaluation measure of classification performance is the Matthews Correlation Coefficient (MCC) ranging from −1 (perfect inverse prediction) to +1 (perfect prediction) and calculated as: MCC = ((TP × TN) − (FP × FN))/((TP + FP)(TP + FN)(TN + FP)(TN + FN))1/2. This was used in conjugation with the F1-score to estimate the overall performance of the predictor.
Tuning training parameters
Decision tree depth
In general the deeper the tree, the more complex patterns it can fit. However, this can easily lead to over-fitting. Thus, finding an optimal tree depth is important. The maximum depth was varied between 1 and 25 (Supplementary Fig. S1) and a depth of 13 yielded the highest MCC and F1 scores.
Number of trees in the forest
Another important parameter is how many decision trees to use. In theory, the more trees the better, but there is a saturation in performance, beyond which the increase in performance is only marginal. Therefore, it is important to find the optimal number of trees to save computational time. As can be seen from the Supplementary Fig. S2, 50 decision trees yield a reasonable performance, which is only slightly increased (by ~5%) using more trees. Therefore, using 50 trees was considered to be enough for the computationally expensive feature selection part. However, for the final selected combination of features, 500 trees were used to achieve maximum performance.
Probability cutoff
The classifier needs a user-defined probability cutoff (Pcut) above which an example is classified as positive. Pcut was varied in the whole range of 0.0–1.0 and based on the performance (Supplementary Fig. S3), was set to 0.5 (majority vote). Therefore, if 50% or more decision trees voted for the particular example to be positive, it was classified as positive.
Frequency analyses of protean and disordered residues
Amino acid propensity
The propensity (Pr) for a particular amino acid, X to occupy a particular ‘class’ (e.g. protean vs. disordered residues) was calculated as the ratio of two probabilities (P) as: Pr(X) = P(X)class/P(X)full = (N(X)class/N(All)full)/(N(X)full/N(All)full) where ‘full’ stands for the entire training dataset and N denotes the raw count of amino acid(s) in the said ‘class’. A propensity value of 1 represents no preference whereas a higher or lower value represent higher or lower preference, respectively, of the amino acid to occupy the given class with respect to the baseline.
Predicted secondary structural content
PSIPRED [37] was used to predict the secondary structure in three classes (H: Helix, E: Strand, C: Coil). For each amino acid, the relative fraction of each of the three main secondary structural classes (H, E, C) were calculated for protean, non-protean, disordered and ordered sequences. The aim was to decipher if there was any preference in disorder versus order sequences that might have propagated to protean segments during the ‘disorder-to-order’ transitions.
Design of the sequence-driven features
Consideration of local and global effects
The origin of disorder is a conjunction of multiple factors. Therefore, ideally the contribution of both, the local sequence (neighboring effect) and that of the global three-dimensional fold of the protein should be considered in the design of features. However, it is highly non-trivial to take into account the global effect of the overall protein fold without actually attempting to build homology models for the predicted ‘structured’ regions, in their bound form. Incorporating such a modeling pipeline will be computationally costly and will also have low confidence associated with the built models due to the lack of enough structural data. One alternative way to indirectly take into account the global constraints is to perform a homology search against all non-redundant sequences [38] and then convert the sequence into a profile. To this end, PSI-BLAST [39] was used to construct sequence profiles. In addition, PSIPRED [37] was used to predict secondary structure and DISOPRED3 [32] was used to predict disorder probability for each amino acid residue. Thus, the plausible global constraints were also accounted for in the designed features, at least implicitly.
To describe the neighboring environment, a sliding window of 15-residues centered around the current residue was considered in the design of most features. This will produce an average property of the feature, taking into account the local sequence dependence associated with disorder-to-order transitions. The size of the window was optimized by trying different lengths in the range of 9–21. The optimal size agrees with the average length of protean segments (Fig. 1).
In total 342 features, in seven different feature groups, were used and are described in detail below (Table 2).
Feature group 1: amino acid mutability (features: 1–300)
Considering the influence of the local sequence on disorder, it is likely that empirical trends (over and under-representations) will be found in the distribution of amino acids in protean compared to non-protean regions. In other words, certain amino acids might preferentially occur in the protean segments but not others. This was represented by Position Specific Scoring Matrices (PSSM) constructed by running three iteration (−j 3) of PSI-BLAST [39] against UniRef90 [38] with an inclusion E-value threshold of 10−3 (−h 0.001). The PSSM contains scores for each of the 20 possible amino acid substitutions in each position, representing the amino acid mutability at any given position. The higher the score, the higher the probability of the corresponding amino acid to occur at that position. To improve convergence, the raw PSSM scores were linearly scaled to [0.0, 1.0] based on the maximum and minimum values observed for each amino acid in the whole training set. To account for the local sequence bias, a 15-residue window of the PSSM was used, centered around the current residue, giving 300 (15 × 20) features in total for each residue.
Feature group 2: amino acid conservation (feature: 301)
The conservation score is derived by PSI-BLAST [39] from the PSSM matrix, and, as the name suggests, conceptually, it is complementary to that of ‘mutability’. Numerically, it is a modified Shannon Entropy [40] term representative of the heterogeneity of amino acid substitutions for a given position in the input sequence. Again, to take care of the neighboring environment, the conservation score was averaged over a 15-residue window. In contrast to all other feature groups, this group consists of only a single value.
Feature group 3: amino acid composition (features: 302–321)
This feature group describes the individual concentration of all amino acids, in a 15-residue long window, i.e. 20 features in all, representing a coarse-grain estimation of the amino acid properties in the local neighborhood around the central residue.
Feature group 4: amino acid properties (features: 322–330)
It is natural to believe that the physiochemical properties of different amino acids hold the key for developing intrinsic disorder and also for the disorder-to-order transitions. In contrast to the ‘amino acid composition group’ described above, Polarity, Charge, Hydrophobicity and Molecular Weight were explicitly described in this feature-group, in a 15-residue sliding window. Polarity was divided into polar, non-polar, acidic-polar or basic-polar, and charge into positive, negative, and neutral [41]. Hydrophobicity was described using the Kyte Doolittle scale [42]. For each of these seven features, the corresponding counts were averaged over the 15-residue window.
Feature group 5: predicted secondary structure (features: 331–333)
Secondary structural propensities of individual amino acids in the close neighborhood of a residue might have major influence on disorder and might serve as a discriminative feature between protean and non-protean fragments. For example, if this likelihood keeps altering between helices to sheets along the sequence, the resultant main-chain trajectory would potentially keep wobbling giving rise to an unstructured region. The other possibility is of course having most residues predicted as ‘random coils’. The probabilities of each amino acid residue in a sequence to form one of the three main secondary structures (Helix, Strand, Coil) were predicted by PSIPRED [37] and averaged over a 15-residue sliding window, serving as three distinct features.
Feature group 6: predicted disorder probability (features: 334–340)
The probability for disorder was predicted using DISOPRED3 [32]. The disorder prediction score from DISOPRED3 is a confidence estimate (or probability) for a residue in a protein sequence to be disordered. It is defined in the range [0, 1] and DISOPRED3 assigns the disordered status to a residue if the score is >0.5. The disorder prediction score, averaged over the 15-residue window centered on the current residue was directly used as the first feature in this group. In addition, to describe the local properties of the disorder prediction, the length of disordered and ordered segments and the start and end positions relative to the total sequence length were also used. In detail, if the score was ≥0.5, the positions on either side of the current residue where the score drops below 0.5 were identified. From this, the length, start and stop positions of the segment could be calculated. This was performed for residues predicted to be disordered (score ≥0.5) and for residues predicted to be ordered (score <0.5), resulting in 7 (1 + 3 + 3) features. Depending on the predicted disorder of the segment, three of the seven features will always remain zero.
Feature group 7: disorder topography (features: 341–342)
Disorder topography measures the topography of peaks and valleys in the predicted disorder score graph (Supplementary Fig. S4). Each residue is classified as being part of a peak (1), valley (−1) or neither (0). A residue is part of a peak if on both sides, there exists another residue with a score at least 10% lower than the current residue. Likewise, a residue is part of a valley if there are residues with disorder scores at least 10% higher than the current residue. If a residue is neither at a peak nor in a valley it is classified as neither. In addition, the length of the current peak- or valley- residue is also calculated and used as a separate feature. Thus, the disorder topography feature consists of the peak/valley/neither classification (feature no. 341) and the topographic length of the current peak/valley (feature no. 342).
Results and discussion
Propagation of sequence consensus during disorder-to-order transitions
Sequence-driven properties such as amino acid propensities and predicted secondary structural content might serve as crucial consensus in the information transfer during the ‘disorder-to-order’ transition. A comparative study of these properties in predicted disordered and annotated protean segments will also serve to explore and identify empirical trends in the designed features and thereby act as a guide in determining the features that are more discriminative compared to the features that can act as filters. Taking this into account, the referred properties were investigated in (i) protean versus non-protean residues and (ii) disordered versus ordered residues (as predicted by DISOPRED3 [32]) and compared with each other. The aim was to identify any pattern that might be responsible for the disorder-to-order transitions, implicitly embedded in the protean sequences. To that end, we wanted to collect the most discriminating trends in the disordered versus ordered regions which were either maintained or inverted in the protean versus non-protean segments. These combined trends should be instrumental in both sustaining the intrinsic disorder and also in the information transfer during the ‘disorder-to-order’ transitions. However, since the ‘disorder versus order’ classification is clearer and more distinct, it was expected that the trends for ‘disorder versus order’ should be more prominent than the ‘protean versus non-protean’ trends.
Amino acids preference in protean and disorder residues
The first and most fundamental characteristic investigated was the of amino acid propensity in disordered, ordered and protean/non-protean residues. The predicted disordered regions show drastic under-representations of hydrophobic amino acids compared to predicted ordered regions (Fig. 2a). Even among the distribution of hydrophobic amino acids, there is an unmistakable trend with respect to the size of the hydrophobic side-chain. The gradual increase in the propensity of the hydrophobic side-chains in the predicted ordered regions is found to be directly proportional to their side-chain volume (Ala → Val → Leu → Ile → Phe → Tyr → Trp) (Fig. 2b); whereas in the predicted disordered regions, the relationship appears inversely proportional. This trend is perfectly consistent with the notion of hydrophobic core formation within ordered protein tertiary structures [43], and on the other hand, bulky aromatics (Phe, Tyr, Trp) should be unfavorable in disordered regions, due to their potential incompatibility with regard to side-chain volume and entropy. The other noticeable features include the significant over-representation of cysteines in ordered regions with a concomitant under-representation in disordered regions, consistent with the idea of fold stabilization by disulfide bridges [44], which must be avoided during the natural design of intrinsic disorder. On the other hand, prolines are significantly over-represented in disordered compared to ordered regions, which is consistent with their ability to break regular secondary structures [45], especially helices [46]. Even if found in regular secondary structures (β-sheets for example), proline needs additional structural constraints from pre-prolines (e.g. glycine rescue) to become stabilized [47]. In line with these observations, proline has been identified as the most disorder promoting amino acid residue [48].
Amino acid propensities in the predicted A disordered versus B ordered regions. The black horizontal line (propensity = 1.0) serves as the baseline; meaning no preferential occurrence of the said amino acid in the said class. A propensity greater or lesser than 1.0 represents over and under representations respectively
The other well-known residue, responsible for backbone flexibility, glycine [45] was also found to be over-represented in disordered compared to ordered regions. This is in accord with the well- established idea that prolines and glycines are general indicators of entropic elasticity [27, 48] and hence control self-organization of elastomeric proteins (e.g., amyloid fibrils) [49]. In fact, recent studies have formulated correlation functions of elasticity in terms of coiling propensity based on sequences rich in proline and glycine in disordered proteins [27, 48].
The other noticeable difference was seen for serine, again a small and polar amino acid, significantly over-represented in disordered and under-represented in ordered regions. Indeed, serine-rich proteins in bacterial enzymes like kinases [50] and eukaryotic splicing factors [51] have been reported to be part of intrinsically disordered proteins. The other polar (Thr, Asn, Gln) and charged (Asp, Glu, Lys, Arg) amino acids were found to have similar or slightly higher propensities in disordered compared to ordered sequences, which agrees well with the earlier observations [47, 48].
But as mentioned earlier, the focus of the current work was to identify patterns that were not only discriminative in disorder versus order sequences but were also maintained in protean versus non-protean sequences and therefore might help in establishing a crucial consensus in the understanding of disorder-to-order transitions. However, as expected, the patterns in protean versus non-protean sequences were not as prominent as in disordered versus ordered sequences (Fig. 3). The collection of all non-ProS plus non-MoRF sequences served as the ‘non-protean’ baseline which raised a value of ~1.00 (±0.01) for the baseline propensities of all amino acids (Fig. 3b). This was not surprising since the bulk majority of the training dataset contained negative examples (non-protean sequences). Similar to amino acid propensities obtained for the ordered regions, all large hydrophobic residues (Leu, Ile, Phe, Tyr, Trp) were found to be over-represented in the protean segments (Fig. 3a) and drastically under-represented in the disordered regions (Fig. 2a). This inversion in trends from disordered to protean segments is rather interesting since the protean segments are merely subsets of the originally disordered regions. It strongly indicates that the potential to get ordered by mediating enough hydrophobic interactions is in fact implicitly embedded in the protean sequences, just like that of globular proteins, but masked by neighboring or flanking disordered residues in their unbound forms.
Amino acid propensities in the annotated A protean versus B non-protean segments. The black horizontal line (propensity = 1.0) serves as the baseline; meaning no preferential occurrence of the said amino acid in the said class. A propensity greater and lesser or 1.0 represents over and under representations respectively
At the same time, the charged residues (Glu, Asp, Lys, Arg) also acquired much larger propensities compared to what they had in disordered sequences, and also noticeably higher propensities compared to ordered sequences in general (Figs. 2b, 3a). The results clearly indicate that both large, hydrophobic and charged residues are preferentially selected during the ‘disorder-to-order’ transitions (via binding). In other words, not all disordered regions undergo the same transition; rather, there is a preferential selection of sequences containing large hydrophobic and charged residues leading to stabilization through hydrophobic and salt-bridge interactions at the protein–protein interface. This is in accord with the general notion of stability upon binding in protein–protein interfaces where both shape and electrostatic complementaries are crucial for binding [52, 53].
Finally, as for disorder residues, cysteines are clearly under-represented in protean residues as well, reflecting the fact that the stability of protean residues should not involve disulfide bridges (at the cost of massive loss of plasticity). However, in contrast to disordered residues, both proline and glycine are under-represented in protean residues, indicating that these residues do not undergo disorder-to-order transition; instead, they remain disordered.
Secondary structure preference in protean and disordered residues
It is also important to conceptualize the secondary structural trends during the course of disorder-to-order transitions. The relative content of coil (C), including loops and turns is higher than helix (H) and strands (E) in all classes of sequences ranging from disorder to order and from protean to non-protean. But when comparing between two opposite class (e.g. disordered vs. ordered), it is the relative increment in (H + E)/C that is of interest. On that note, ordered sequences naturally have far greater regular secondary structures (H + E) amounting to ~50% of the whole population than disordered sequences (H + E: ~15%; C: ~85%) (Fig. 4). As expected, the relatively low proportion (~15%) of helices and strands in disordered residues definitely increases the disorder-to-order transitions in protean segments (H + E:~40%), which is roughly the same as in non-protean sequences (Fig. 5). Recall that the large majority of the non-protean sequences are in fact the usual ordered sequences and the subset of disordered sequences that gets ordered only constitute a small (leftover) fraction. Among the regular secondary structures, helices appear to be more prevalent in protean (~32%) than non-protean segments (~27%) whereas beta-strands seem to be slightly more preferred in non-protean (~10%) compared to protean segments (~5%).
Indecisiveness in adapting a particular secondary structure class from sequence
Another property investigated based on secondary structure is the indecisiveness of an amino acid sequence in adapting a particular secondary structure. The assumption was that protean segments, when disordered in isolation, might be indecisive in their choice to adapt a particular secondary structure (H, E or C) along the main-chain trajectory and thereby end up being unstructured. Given the current lack of structural data for these sequences, PSIPRED [37] was used to predict secondary structure and to test the above hypothesis. A measure for the indecisiveness or randomness in secondary structure prediction called Altscore was defined as the average number of transitions (H → C, C → E etc.) for each protean and non-protean segment. Regions with an Altscore value of ‘zero’ were omitted for both protean and non-protean regions, since they would only add noise to any potential signal. Focusing on the regions with Altscore >0, the frequency distribution (Fig. 6) clearly discriminated between protean and non-protean classes with a wider spread being obtained for the protean class in addition to a peak-shift towards higher values (0.1 compared to 0.05 for non-protean). The results indicate that the intrinsic disorder associated with the unbound protean segments potentially suffers from the indecisiveness of the main-chain trajectory to adapt a particular secondary structure.
Both the above observations, (i) the reappearance of large hydrophobic and charged amino acids into the protean segments, as well as (ii) the indecisiveness associated with their predicted secondary structures should serve constructively in unraveling a hidden consensus in promoting disorder-to-order transitions.
Training a classifier to predict protean residues
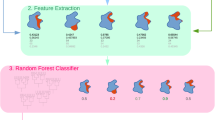
To be able to predict protean residues from sequence, a random forest classifier was trained on the features described in the “Methods” section. Most features were calculated using a 15-residue sliding window, optimized by trying different window sizes in the range of 9–21 (Supplementary Fig. S5). The chosen window size was optimal in the sense that it fell right at the center of the distribution of the protean segment-lengths (Fig. 1). An identical sliding window size was also used to determine protein-binding residues embedded within disordered regions previously [32]. Note that for all feature groups except Feature Group 1 (Amino Acid Mutability), the number of features will remain the same even with a different window size. Among all features, some features might be non-informative, others might be redundant. Indeed, some features are similar in their physiochemical descriptions and therefore might be excluded without loss in performance. But sometimes it is advantageous for the classifier to learn from explicit rather than implicit features. To find the best combination of the seven feature groups, all 127 possible combinations were exhaustively examined by measuring the final cross-validated performance using MCC and F1-scores for each feature group combination.
The 20 best feature group combinations according to the MCC and F1-scores have been shown in Supplementary Figs. S6 and S7 respectively. The difference is small between the top feature group combinations. Also, the top-combinations as evaluated by MCC and F1 are not identical, whereas, using all features resulted in good scores being attained in both evaluations. Therefore, the combination of all feature groups was chosen judiciously. The absolute MCC and F1 score values are relatively small (~0.13), owing to a large number of false positives and negatives. However, the magnitude of the scores are comparable to other studies [32–35], and reflect the difficulty of predicting residues that will be ordered upon binding, from information in one of the binding partners only. This is further illustrated in the recall versus precision (PPV) curves for the best combination (Fig. 7). The recall versus precision curves were constructed by varying the cutoff (Pcut) and calculating precision and recall for each cutoff. The random base line precision is 1.9% and the curve for the best combination is clearly above that. It can also be seen that 500 trees perform slightly better than 50. But the question remains whether the rather modest 10% precision at 23% recall (Pcut > 0.5) is useful at all. Considering that it is still five times better than a random prediction, it is arguably useful given the state-of-the-art. But there is of course plenty of room for improvement which might be brought about in future studies by incorporating additional information not directly obtainable from the sequence alone. In principle, one might perform structure prediction with these sequences, and, during the course, filter out residues that are actually ordered by themselves; and also predict the surrounding residues. The predicted and demarcated structural units can also be used as starting templates in molecular dynamic simulations or docking studies exploring a reduced conformational space. In addition, advanced structural validation tools [54–56] could also be incorporated as filters in an iterative prediction pipeline to improve the sequence-based prediction.
Relative importance of features
In an effort to learn what features contributed to the overall prediction, the relative importance of each feature group as calculated by the random forest prediction module was used. To take account of the inherent randomness associated with the classifications, this relative importance was averaged over predictions of 500 decision trees. As we can see, there are three features that stand out above the rest (Fig. 8): Topographic length (group 6) is by far the most important feature and describes the length of the topographic region where the current residue is located. Interestingly, the second most important feature is also a length descriptor, namely the more coarse-grained length of the ordered region corresponding to the current residue (group 5). Note that this feature will be ‘zero’ for all residues predicted to be disordered. The third most important feature is the predicted disorder score averaged over the current window size (group 5).
The other seven features in the top ten were the following. Rank 4: the relative distance to the ordered residue before the current one (group 6), rank 5: length of the disordered region the current residue resides in (group 6), rank 6: the topography score (group 7), rank 7: probability of the current residue to form a coil (group 5), rank 8: probability of the current residue to form a helix (group 5), rank 9: the relative distance to the ordered residue after the current one (group 6), and rank 10: charge-neutrality of the current amino acid (group 4).
True positive enrichment by analyzing the Proteus score
A common test of machine learning predictors is to analyze the true positive enrichment by constructing score plots, which is more detailed compared to recall versus precision curves. Score plots are conventionally defined as the overlay of two independent evaluation measures, Positive Predicted Value (PPV) and recall (true positive rate) as two distinct functions of the predicted Proteus score. Ideally, both the PPV and recall should be high but there is a conflict in finding as many true positives as possible (high recall) and at the same time having a high PPV (few false positives). In reality there will always be at a trade-off between the two, which is also the main reason to use the combined measure F1. In the current case (Fig. 9a), F1 peaks at around the score of 0.5, which is also the cutoff chosen for positive prediction in the final predictor (Pcut = 0.5); corresponding to 10% PPV and 23% recall as discussed above. It can be noted that after that point the PPV increases quite rapidly; and scores >0.7 have PPV >40%. Unfortunately there are rather few examples that obtain this high score resulting in a rather modest recall overall. Still, if the score is high we can certainly trust it to be a relatively accurate prediction. This is also reflected by analyzing the distribution of scores for protean and non-protean residues (Fig. 9b), where the score was found to be much higher for predicted protean than non-protean residues with median values of 0.4 and 0.24 respectively, and with roughly equivalent median absolute deviations. It can also be seen that there are quite a large number of high scoring outliers in the non-protean residues. These might of course be completely wrong, but there is also a possibility that these predictions are actually sites for yet unknown interactions. Since the study of transient interaction is difficult, and the focus of the structural biology community so far has been on stable interactions that can even form crystals. There is still a lot more to be discovered if the dynamics is also taken into account.
Analysis of Proteus score for the cross-validated predictions. a Proteus score versus PPV (solid, blue), recall (dashed, red), and F1 (dotted, orange) for the cross-validated predictions. b Box plots showing the distribution of predicted Proteus scores for protean and non-protean residues. The median of the two distributions is shown by the horizontal red line in the middle of the two boxes
Benchmark on independent data set
In any machine learning scheme it is an advantage if the final classifier can benchmarked on independent data, and against other classifiers. In the recent DISOPRED3 paper [32] the upgraded DISOPRED version was benchmarked with ANCHOR [33] MoRFpred [34], MFSPSSMpred [35], and DISOPRED3 [32] using a set of 2209 residues out of which 163 were protean (i.e., positive examples) from nine proteins (see “Methods” section). None of the examples in the independent set were similar to any example used in training Proteus, thus before classifying, Proteus was retrained on the full non-cross validated training set. The predictions for the other methods were generously made available by the authors of DISOPRED3 through the following link: http://bioinfadmin.cs.ucl.ac.uk/downloads/DISOPRED/suppl_data/. The evaluation measures precision, recall, F1, and MCC were calculated for all methods using the binary classification of each method (Fig. 10) and compared among them, as recall versus precision curves using the raw scores from each method (Supplementary Fig. S8). Overall, Proteus is better in all measures. Proteus has the highest precision (0.26 compared to 0.22 for DISOPRED3, the second best), for a much larger recall (0.56 compared to 0.28 by ANCHOR, the next best). This combined improvement in both precision and recall is also naturally reflected in a concomitant increase in the F1-score (0.35 compared to 0.18 by DISOPRED3, the next best). It also attained a higher MCC value than the other methods (0.30 compared to 0.13 by DISOPRED3). Even though the independent set is small, the high recall is particularly encouraging if Proteus is to be used as an initial step before implementing more elaborate approaches (as discussed earlier). It is crucial not to miss any true positives at an early stage.
Comparison of Proteus with other classifiers using the standard evaluation measures. All methods were tested on the same validation set of nine proteins containing 2209 residues (total number of examples) with 163 protean (positive examples). Precision, Recall, F1-score and MCC tabulated for each method. Proteus predicts twice as many true positives as the second best method (55 vs. 27%) with a much higher precision
Conclusions
With the realization that protein disorder is involved in a range of human diseases, including cancer, cardiovascular and neurodegenerative diseases, it is important to compile more and more structural information for these proteins to understand their modus operandi. A first step in this direction is the classification and prediction of protean segments. The literature shows that there is indeed much room for improvement for the existing predictors [32]. Proteus seems to perform better than the existing predictors on the available independent dataset. Of course this has to be re-evaluated when more data becomes available. It is also possible to combine different individual methods to build hybrid methods to increase the performance even further. Given the current state-of-the-art, the predicted ‘protean’ segments should be considered ‘potential’ binding sites for proteins in general, whereas, for a specific interaction with known partners, the predicted segments should serve as ‘different’ starting points for model building. The built models then need to undergo stringent validation filters in an iterative cycle for screening and selection. It is also important to conceptualize the multiple sequence driven factors and realize that it is their complex coordination which holds the key consensus in promoting the ‘disorder-to-order’ transitions. The consensus is yet untangled and needs other exclusive studies to eventually be resolved, however, the current work explores certain empirically observed trends which appears to be instrumental in the transition from disorder to order. These factors include the reappearance of large hydrophobic and charged amino acids in the protean segments, which are significantly under-represented in the originally ‘disordered’ regions. The study also reflects that there is an inherent indecisiveness to adapt to a specific secondary structure (helices, strands or loops) associated with the protean segments. In other words, the protean segments remain indecisive in their choice to adapt a particular secondary structure. This is consistent with the notion of sustaining enough ‘disorder’ even in the bound form [4] which potentially helps the proteins to sustain their binding promiscuity. To conclude, the study has both a basic and an applied content, both of which should serve the IDP as well as the broad biological community.
References
Wright PE, Dyson HJ (1999) Intrinsically unstructured proteins: re-assessing the protein structure-function paradigm. J Mol Biol 293:321–331. doi:10.1006/jmbi.1999.3110
Dunker AK, Garner E, Guilliot S, Romero P, Albrecht K, Hart J et al (1998) Protein disorder and the evolution of molecular recognition: theory, predictions and observations. Pac Symp Biocomput. Pac Symp Biocomput 3:473–484
Kulkarni P, Rajagopalan K, Yeater D, Getzenberg RH (2011) Protein folding and the order/disorder paradox. J Cell Biochem 112:1949–1952. doi:10.1002/jcb.23115
Uversky VN (2013) Unusual biophysics of intrinsically disordered proteins. Biochim Biophys Acta 1834:932–951. doi:10.1016/j.bbapap.2012.12.008
Baruah A, Rani P, Biswas P (2015) Conformational entropy of intrinsically disordered proteins from amino acid triads. Sci Rep. doi:10.1038/srep11740
Anfinsen CB (1973) Principles that govern the folding of protein chains. Science 181:223–230
Harding HP, Zhang Y, Ron D (1999) Protein translation and folding are coupled by an endoplasmic-reticulum-resident kinase. Nature 397:271–274. doi:10.1038/16729
Pestova TV, Hellen CUT (2003) Coupled folding during translation initiation. Cell 115:650–652. doi:10.1016/S0092-8674(03)00981-4
Lau AY, Chasman DI (2004) Functional classification of proteins and protein variants. Proc Natl Acad Sci USA 101:6576–6581. doi:10.1073/pnas.0305043101
Brun C, Chevenet F, Martin D, Wojcik J, Guénoche A, Jacq B (2004) Functional classification of proteins for the prediction of cellular function from a protein-protein interaction network. Genome Biol 5:R6
Starikov EB, Norden B (2012) Entropy-enthalpy compensation as a fundamental concept and analysis tool for systematical experimental data. Chem Phys Lett 538:118–120. doi:10.1016/j.cplett.2012.04.028
Fisher CK, Stultz CM (2011) Constructing ensembles for intrinsically disordered proteins. Curr Opin Struct Biol 21:426–431. doi:10.1016/j.sbi.2011.04.001
Chebaro Y, Ballard AJ, Chakraborty D, Wales DJ (2015) Intrinsically disordered energy landscapes. Sci Rep 5:10386. doi:10.1038/srep10386
Baranger M (2000) Chaos, complexity, and entropy. New England Complex Systems Institute, Cambridge
Multitude of binding modes attainable by intrinsically disordered proteins: a portrait gallery of disorder-based complexes - Chemical Society Reviews (RSC Publishing). Accessed 7 May 2016. http://pubs.rsc.org/en/content/articlelanding/2011/cs/c0cs00057d#!divAbstract
Wright PE, Dyson HJ (2015) Intrinsically disordered proteins in cellular signalling and regulation. Nat Rev Mol Cell Biol 16:18–29. doi:10.1038/nrm3920
Uversky VN (2011) Intrinsically disordered proteins may escape unwanted interactions via functional misfolding. Biochim Biophys Acta 1814:693–712. doi:10.1016/j.bbapap.2011.03.010
Uversky VN, Oldfield CJ, Dunker AK (2008) Intrinsically disordered proteins in human diseases: introducing the D2 concept. Annu Rev Biophys 37:215–246. doi:10.1146/annurev.biophys.37.032807.125924
Fukuchi S, Sakamoto S, Nobe Y, Murakami SD, Amemiya T, Hosoda K et al (2012) IDEAL: intrinsically disordered proteins with extensive annotations and literature. Nucleic Acids Res 40:D507–D511. doi:10.1093/nar/gkr884
Romero P, Obradovic Z, Li X, Garner EC, Brown CJ, Dunker AK (2001) Sequence complexity of disordered protein. Proteins 42:38–48
Chen S, Berthelier V, Hamilton JB, O’Nuallain B, Wetzel R (2002) Amyloid-like features of polyglutamine aggregates and their assembly kinetics. BioChemistry 41:7391–7399
Jorda J, Xue B, Uversky VN, Kajava AV (2010) Protein tandem repeats: the more perfect the less structured. Febs J 277:2673–2682. doi:10.1111/j.1742-464X.2010.07684.x
Mao AH, Crick SL, Vitalis A, Chicoine CL, Pappu RV (2010) Net charge per residue modulates conformational ensembles of intrinsically disordered proteins. Proc Natl Acad Sci USA 107:8183–8188. doi:10.1073/pnas.0911107107
Uversky VN, Gillespie JR, Fink AL (2000) Why are “natively unfolded” proteins unstructured under physiologic conditions? Proteins 41:415–427
Schlessinger A, Punta M, Rost B (2007) Natively unstructured regions in proteins identified from contact predictions. Bioinforma Oxf Engl 23:2376–2384. doi:10.1093/bioinformatics/btm349
Baruah A, Biswas P (2016) Globular–disorder transition in proteins: a compromise between hydrophobic and electrostatic interactions?. Phys Chem Chem Phys 18:23207–23214. doi:10.1039/C6CP03185D
Cheng S, Cetinkaya M, Gräter F (2010) How sequence determines elasticity of disordered proteins. Biophys J 99:3863–3869. doi:10.1016/j.bpj.2010.10.011
Linding R, Jensen LJ, Diella F, Bork P, Gibson TJ, Russell RB (2003) Protein disorder prediction: implications for structural proteomics. Structure 11:1453–1459. doi:10.1016/j.str.2003.10.002
Dosztányi Z, Csizmok V, Tompa P, Simon I (2005) IUPred: web server for the prediction of intrinsically unstructured regions of proteins based on estimated energy content. Bioinformatics 21:3433–3434. doi:10.1093/bioinformatics/bti541
Peng K, Radivojac P, Vucetic S, Dunker AK, Obradovic Z (2006) Length-dependent prediction of protein intrinsic disorder. BMC Bioinform 7:208. doi:10.1186/1471-2105-7-208
Shimizu K, Hirose S, Noguchi T (2007) POODLE-S: web application for predicting protein disorder by using physicochemical features and reduced amino acid set of a position-specific scoring matrix. Bioinformatics 23:2337–2338. doi:10.1093/bioinformatics/btm330
Jones DT, Cozzetto D (2015) DISOPRED3: precise disordered region predictions with annotated protein-binding activity. Bioinform Oxf Engl 31:857–863. doi:10.1093/bioinformatics/btu744
Mészáros B, Simon I, Dosztányi Z (2009) Prediction of protein binding regions in disordered proteins. PLOS Comput Biol 5:e1000376. doi:10.1371/journal.pcbi.1000376
Disfani FM, Hsu W-L, Mizianty MJ, Oldfield CJ, Xue B, Dunker AK et al (2012) MoRFpred, a computational tool for sequence-based prediction and characterization of short disorder-to-order transitioning binding regions in proteins. Bioinformatics 28:i75–i83. doi:10.1093/bioinformatics/bts209
Fang C, Noguchi T, Tominaga D, Yamana H (2013) MFSPSSMpred: identifying short disorder-to-order binding regions in disordered proteins based on contextual local evolutionary conservation. BMC Bioinform 14:300. doi:10.1186/1471-2105-14-300
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O et al (2011) Scikit-learn: machine learning in Python. J Mach Learn Res 12:2825–2830
Jones DT (1999) Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 292:195–202. doi:10.1006/jmbi.1999.3091
Suzek BE, Huang H, McGarvey P, Mazumder R, Wu CH (2007) UniRef: comprehensive and non-redundant UniProt reference clusters. Bioinformatics 23:1282–1288. doi:10.1093/bioinformatics/btm098
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W et al (1997) Gapped Blast and PsiBlast: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Shannon CE (1948) A mathematical theory of communication. Bell Syst Tech J 27:379–423. doi:10.1002/j.1538-7305.1948.tb01338.x
Cooper GM (2000) The cell, 2nd edn. Sinauer Associates, Sunderland
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132. doi:10.1016/0022-2836(82)90515-0
Munson M, Balasubramanian S, Fleming KG, Nagi AD, O’Brien R, Sturtevant JM et al (1996) What makes a protein a protein? Hydrophobic core designs that specify stability and structural properties. Protein Sci Publ Protein Soc 5:1584–1593
Betz SF (1993) Disulfide bonds and the stability of globular proteins. Protein Sci Publ Protein Soc 2:1551–1558
Chou PY, Fasman GD (1978) Empirical predictions of protein conformation. Annu Rev Biochem 47:251–276. doi:10.1146/annurev.bi.47.070178.001343
Visiers I, Braunheim BB, Weinstein H (2000) Prokink: a protocol for numerical evaluation of helix distortions by proline. Protein Eng 13:603–606. doi:10.1093/protein/13.9.603
Das M, Basu G (2012) Glycine rescue of β-sheets from cis-Proline. J Am Chem Soc 134:16536–16539. doi:10.1021/ja308110t
Theillet F-X, Kalmar L, Tompa P, Han K-H, Selenko P, Dunker AK et al (2013) The alphabet of intrinsic disorder. Intrinsically Disord Proteins 1:e24360. doi:10.4161/idp.24360
Rauscher S, Baud S, Miao M, Keeley FW, Pomès R (1993) Proline and glycine control protein self-organization into elastomeric or amyloid fibrils. Struct Lond Engl 2006;14:1667–1676. doi:10.1016/j.str.2006.09.008
Singh G (2015) Association between intrinsic disorder and serine/threonine phosphorylation in Mycobacterium tuberculosis. FASEB J 29:563.4
Haynes C, Iakoucheva LM (2006) Serine/arginine-rich splicing factors belong to a class of intrinsically disordered proteins. Nucleic Acids Res 34:305–312. doi:10.1093/nar/gkj424
Basu S, Bhattacharyya D, Wallner B (2014) SARAMAint: the complementarity plot for protein–protein interface. J Bioinform Intell Control 3:309–314. doi:10.1166/jbic.2014.1103
Basu S, Wallner B (2016) Finding correct protein–protein docking models using ProQDock. Bioinformatics 32:i262–i270. doi:10.1093/bioinformatics/btw257
Basu S, Bhattacharyya D, Banerjee R (2014) Applications of complementarity plot in error detection and structure validation of proteins. Indian J Biochem Biophys 51:188–200
Uziela K, Shu N, Wallner B, Elofsson A (2016) ProQ3: improved model quality assessments using Rosetta energy terms. Sci Rep 6:33509. doi:10.1038/srep33509
Uziela K, Menéndez Hurtado D, Shu N, Wallner B, Elofsson A (2017) ProQ3D: improved model quality assessments using deep learning. Bioinform Oxf Engl. doi:10.1093/bioinformatics/btw819
Acknowledgements
This work was supported by grants from the Swedish Research Council (VR-NT 2012-5270), the Swedish e-Science Research Center (SeRC) and the Department of Science and Technology – Science and Engineering Research Board, India (DST-SERB research grant PDF/2015/001079). Computational resources were provided by the Swedish National Infrastructure for Computing (SNIC) at the National Supercomputing Center (NSC) in Linköping, Sweden and DST-SERB, India.
Author information
Authors and Affiliations
Corresponding author
Additional information
Software Availability: https://github.com/bjornwallner/proteus.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Basu, S., Söderquist, F. & Wallner, B. Proteus: a random forest classifier to predict disorder-to-order transitioning binding regions in intrinsically disordered proteins. J Comput Aided Mol Des 31, 453–466 (2017). https://doi.org/10.1007/s10822-017-0020-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-017-0020-y