Abstract
The goal of this paper is to estimate the number of realistic drug-like molecules which could ever be synthesized. Unlike previous studies based on exhaustive enumeration of molecular graphs or on combinatorial enumeration preselected fragments, we used results of constrained graphs enumeration by Reymond to establish a correlation between the number of generated structures (M) and the number of heavy atoms (N): logM = 0.584 × N × logN + 0.356. The number of atoms limiting drug-like chemical space of molecules which follow Lipinsky’s rules (N = 36) has been obtained from the analysis of the PubChem database. This results in M ≈ 1033 which is in between the numbers estimated by Ertl (1023) and by Bohacek (1060).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Virtual screening of chemical databases is a classical chemoinformatics approach to discover compounds possessing desirable properties, in particularly, new drug molecules. Efficiency of this procedure depends on both performance of the screening tools and the content of the screened database. Nowadays, ensemble of academic, commercial and propriety databases records some 108 structures of existing chemical compounds. Since these collections are limited to already known chemotypes, an effort should be done to generate virtual compounds involving structural moieties which don’t occur in existing structures. Larger library of virtual compounds provides, certainly, with a larger chance to discover new drug-like compounds.
The question arises how large the whole chemical space of realistic drug-like molecules is? Although this question was in the focus of numerous studies [1–3], still there is no consensus in the estimation of the number of potential drug-like molecules (M): depending on the way of its estimation it varies from 1023 to 10180 (Table 1). Efforts were also done to assess the size of sub-spaces covering a given type of chemical compounds: alkanes, substituted heptanes and hexanes, neurological drugs. In these studies, M corresponded either to the number of all graphs containing up to N nodes (exhaustive graphs enumeration [4, 5]), or to the number of graphs resulted from an intersection of several predefined sub-sets of graphs (combinatorial graphs enumeration [6–8]). Each of these approaches has clear drawbacks. Most of structures resulted from an exhaustive graphs enumeration are unrealistic (reactive, strained etc.); thus, some rules should be imposed to select a relatively small portion of molecules which potentially may exist [9]. (Later, we’ll call this “constrained exhaustive graphs enumeration”). Results of the combinatorial graphs enumeration depend on preselected subsets of graphs. Usually they are drawn from already existing molecules, which significantly limits diversity of the resulting structures.
In this context, particular interest represents a project “chemical universe database” initiated by Reymond in 2005 [9]. They performed constrained exhaustive enumeration of structures containing up to 17 C, N, S, O and halogen atoms [10] which resulted to a database of 1.66 × 1011 structures (GDB-17). A lot of potentially reactive or strained structures have been discarded using different filters. Although GDB-17 represents a useful source of molecular diversity to discover new chemotypes, the molecular size (17 heavy atoms) is still too small for typical drug-like molecules. Unlike previous studies, we used the information about the GDB-17 content in order to establish a relationship between the number of structures generated for a given number of heavy atoms using Reymond’s constrains.
Below, we give some information about previous estimations of the number of chemical compounds and potential drug-like molecules followed by description of our approach.
Previous studies
The first attempt to calculate all possible unlabeled 4-valent tree graphs (i.e., the number of alkanes) has been made by Cayley [11]. Later on, numerous publications were devoted to the calculation of the number of molecular graphs corresponding to acyclic non-chiral hydrocarbons [2, 4, 12–18], acyclic chiral monosubstituted hydrocarbons [19], spirits [15], polyenes [20], cyclohexanes [1, 3, 21, 22].
Nowadays, several estimates of the size of chemical universe (M) are reported. As a function of the approach used, this number varies in the range 1023–10180 (Table 1). According to Bohacek et al. [6], the crude number of compounds consisting from thirty C, N, O, S atoms and having up to 4 cycles and 10 branch points is about 1060. Roughly, this corresponds to number of linear molecules with different combinations of atoms (~1023) multiplying by the number of branching/cyclizations for each of them (~1040). In our opinion, this number is overestimated because a large part of structures should be discarded because of steric clashes and strains [10].
Using a set of semantic rules and stereochemistry, D. Weininger concluded that approximately 1033 heptanes and hexanes having molecular weight less than 750 Da and substituted by fragments consisting of H, C, N, O, F atoms [23] may exist. In order to estimate the number of potential neurological drugs, Weaver and Weaver [8] assumed that these compounds should fulfill Lipinski’s rule and fit the 7 Å radius sphere to effectively pass blood–brain barrier. The whole sphere was divided onto 350 functional group volumes. All combinations of up to 5 from 40 possible functional groups correspond to M = 1016 –1021.
Ertl [7] estimated the number of combinations of two and three substituents attached to one same scaffold. Both scaffolds and substituents were generated from the in-house database containing about 3 million organic compounds comprising C, N, O, S, P, Se, Si and halogens and containing up to 36 atoms leading to M ≈ 1023. He has noted that more than 10100 compounds (most of which unrealistic) could potentially be constructed if no restrictive filters are used. Ogata et al. [24] split the ligands extracted from 100 PDB complexes onto fragments, replaced atoms by all possible combinations of C, N, S, O or Cl considering bond orders and combined obtained fragments in new structures. Extrapolation of these results resulted in 1026 compounds containing up to 50 atoms. Drew et al. [25] approximated the number of available compounds in ChemSpider and NIST Chemistry WebBook by power function of the number of carbon atoms. Obtained equation clearly underestimates the number of compounds consisting in up to 100 atoms (~109). There are some other estimates of M in the range from 1014 to 10200 [23, 26–29] given without any clear explanation.
GDB-based chemical space of drug-like compounds
In this study, in order to assess M we had to solve two problems: (1) to establish equation linking M with the number of heavy atoms (N), and (2) to estimate limiting value of N for the drug-like chemical space.
At the first stage, we used the information about the number of generated structures M as a function of N (N = 1–17) tabulated in Ref. [10]. Notice that only two filters were applied upon generation of structures containing up to 11 heavy atoms: “smallest atomic volume” one discarding strained structures and functional group filter discarding reactive non-drugable molecules. To generate structures with N = 12–17, several additional filters have been applied in order to avoid combinatorial explosion [10]. Therefore, only information about the structures with N = 1–11 have been used to build a relationship. According to Giménez and Noy [30], the number of connected undirected planar labeled graphs (M) is linked with the number of vertexes (N) by the relationship M ~ N!, hence logM ~ N × logN. Fitting the latter for GDB-17 compounds with N = 1–11 [10] using the R software [31] results in Eq. (1):
In order to estimate value of N which limits drug-like chemical space, a classical Lipinski’s definition of drug-likeness we used. According to Lipinski’s “rule of five”, orally absorbed drug-like molecules should have the following properties: (1) molecular weight MW ≤ 500 Da, (2) the number of H-donor ≤ 5, (3) the number of H-acceptors ≤ 10, and (4) logP ≤ 5 [32]. The last three parameters don’t limit the number of structures that can potentially follow them; whereas molecular weight can be used as a confining parameter. Thus, we suggested that MW ≤ 500 Da can be used as a bound on drug-like chemical space.
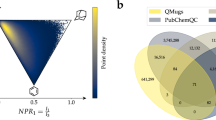
The approximate number of heavy atoms (N) corresponding to molecular weight (MW) of 500 Da has been estimated based on PubChem molecules extracted from the ZINC database (accessed in 2010) [33]. A subset of 23 million compounds containing only C, N, O, S and halogen atoms (as in GDB-17 database) has been selected from the initial set of 31 million compounds. From the linear correlation found between median MW and N (Fig. 1) one can easily assess N ≈ 36 corresponding to MW = 500. Using this number together with Eq. 1 results in M ≈ 1033 (Fig. 2).
The number of 3D structures is even larger if one takes into account all stereoisomers corresponding to one planar molecular graph. According to Ref. [10], GDB-17 compounds contain, in average, 6.4 stereocenters per molecule. Suggesting that the number of stereocenters increases linearly with N, one expects about 12 stereocenters per molecule for the dataset containing compounds up to 36 atoms. This corresponds to 212 = 4096, e.g., the number of stereoisomers is proportional to 103. Thus, the overall number of 3D structures with MW ≤ 500 Da is about 1036.
It seems that remaining three Lipinski’s “rules of five” are valid for most of these molecules. Indeed, Ruddigkeit et al. [10] demonstrated that the vast majority of GDB-17 compounds follow these rules. Thus, it has been shown that the average number of H-bond donors in GDB-17 slowly increases with N and for compounds having 17 atoms this value is equals to 2.5. Average CLog P values remain almost constant and equal to zero independently on the number of heavy atoms in GDB-17 molecules. Thus, we believe that N = 1033 is a reasonable empirical estimation of the size of chemical space of drug-like compounds which follow Lipinski’s “rule of five”.
It should be noted that the idea of extrapolation of the number of drug-like compounds based on fully enumerated compounds in GDB-17 was recently suggested by Shoichet [34]. However, neither mathematical equation for M nor its estimated value were reported in [34].
The estimated number of molecules is hardly accessible, at least at the current level of computer power. Indeed, simple calculations show that the best modern 500 supercomputers in the world will be able to generate just 1014 compounds per year which corresponds to full enumeration of compounds containing up to 18 atoms. This shows that exhaustive enumeration doesn’t seem to be an effective way to generate “all” useful drug-like compounds. On the other hand, combinatorial generation based on molecular fragments taken from fully enumerated library looks more perspective. For instance, if one uses for this purpose GDB-17 (1.66 × 1011 compounds with up to 17 heavy atoms), 1.66 × 1011 × 1.66 × 1011 = 2.8 × 1022 their possible combinations could be generated. Since each pair of species can be linked by 17 × 17 = 289 different ways, a pairwise linking of GDB-17 molecules results in “only” 2.8 × 1022 × 289 = 8 × 1024 compounds, which is more affordable than N = 1033. Thus, combinatorial generation of drug-like compounds based on fully enumerated libraries of small fragments looks more realistic than fully enumerated compounds libraries. This strategy has also another advantage: the structures can be generated on the fly during virtual screening which allows one to avoid the difficulties with storage and maintenance of such a huge database. The generation of “useful” compounds could always be tuned in a guided enumeration, which fits generated molecules to the target chemical space. Generally, this can be achieved using a fitting function which includes different parameters like target property value, ADME/Tox properties, diversity of the generated library, etc. [35, 36].
References
Pólya G, Read RC (1987) Combinatorial enumeration of groups, graphs, and chemical compounds. Springer-Verlag Inc., New York
Bergeron F, Labelle G, Leroux P (1997) Combinatorial species and tree-like structures, vol 67. Cambridge University Press, Cambridge
Fujita S (1991) Symmetry and combinatorial enumeration in chemistry, vol 8. Springer-Verlag, Berlin, Heidelberg
Henze HR, Blair CM (1931) The number of isomeric hydrocarbons of the methane series. J Am Chem Soc 53(8):3077–3085. doi:10.1021/ja01359a034
Blair CM, Henze HR (1932) The number of stereoisomeric and non-stereoisomeric paraffin hydrocarbons. J Am Chem Soc 54(4):1538–1545. doi:10.1021/ja01343a044
Bohacek RS, McMartin C, Guida WC (1996) The art and practice of structure-based drug design: a molecular modeling perspective. Med Res Rev 16(1):3–50. doi:10.1002/(sici)1098-1128(199601)16:1<3:aid-med1>3.0.co;2-6
Ertl P (2002) Cheminformatics Analysis of Organic Substituents: identification of the most common substituents, calculation of substituent properties, and automatic identification of drug-like bioisosteric groups. J Chem Inf Comput Sci 43(2):374–380. doi:10.1021/ci0255782
Weaver DF, Weaver CA (2011) Exploring neurotherapeutic space: how many neurological drugs exist (or could exist)? J Pharm Pharmacol 63(1):136–139. doi:10.1111/j.2042-7158.2010.01161.x
Fink T, Bruggesser H, Reymond J-L (2005) Virtual exploration of the small-molecule chemical universe below 160 Daltons. Angew Chem Int Ed 44(10):1504–1508. doi:10.1002/anie.200462457
Ruddigkeit L, van Deursen R, Blum LC, Reymond J-L (2012) Enumeration of 166 billion organic small molecules in the chemical universe database GDB-17. J Chem Inf Model 52(11):2864–2875. doi:10.1021/ci300415d
Cayley E (1875) Ueber die analytischen Figuren, welche in der Mathematik Bäume genannt werden und ihre Anwendung auf die Theorie chemischer Verbindungen. Ber Dtsch Chem Ges 8(2):1056–1059. doi:10.1002/cber.18750080252
Herrmann F (1897) Ueber das Problem, die Anzahl der isomeren Paraffine von der Formel CnH2n + 2 zu bestimmen. Ber Dtsch Chem Ges 30(3):2423–2426. doi:10.1002/cber.18970300310
Schiff H (1875) Zur Statistik chemischer Verbindungen. Ber Dtsch Chem Ges 8(2):1542–1547. doi:10.1002/cber.187500802191
Losanitsch SM (1897) Die Isomerie-Arten bei den Homologen der Paraffin-Reihe. Ber Dtsch Chem Ges 30(2):1917–1926. doi:10.1002/cber.189703002144
Perry D (1932) The number of structural isomers of certain homologs of methane and methanol. J Am Chem Soc 54(7):2918–2920. doi:10.1021/ja01346a035
Polya G (1936) Algebraische Berechnung der Anzahl der Isomeren einiger organischer Verbindungen, Zeit. f. Kristall
Harary F, Norman RZ (1960) Dissimilarity characteristic theorems for graphs. Proc Am Math Soc 11(2):332–334
Read R (1976) The enumeration of acyclic chemical compounds. Academic Press, New York
Robinson RW, Harry F, Balaban AT (1976) The numbers of chiral and achiral alkanes and monosubstituted alkanes. Tetrahedron 32(3):355–361. doi:10.1016/0040-4020(76)80049-X
Cyvin SJ, Brunvoll J, Cyvin BN (1995) Enumeration of constitutional isomers of polyenes. J Mol Struct THEOCHEM 357(3):255–261. doi:10.1016/0166-1280(95)04329-6
Sloane NJA, Sloane N (1973) A handbook of integer sequences, vol 65. Academic Press, New York
Leonard JE, Hammond GS, Simmons HE (1975) Apparent symmetry of cyclohexane. J Am Chem Soc 97(18):5052–5054. doi:10.1021/ja00851a003
Weininger D (2002) Combinatorics of small molecular structures. In: Encyclopedia of computational chemistry. John Wiley & Sons, Ltd. doi:10.1002/0470845015.cna014m
Ogata K, Isomura T, Yamashita H, Kubodera H (2007) A quantitative approach to the estimation of chemical space from a given geometry by the combination of atomic species. QSAR Comb Sci 26(5):596–607. doi:10.1002/qsar.200630037
Drew KLM, Baiman H, Khwaounjoo P, Yu B, Reynisson J (2012) Size estimation of chemical space: how big is it? J Pharm Pharmacol 64(4):490–495. doi:10.1111/j.2042-7158.2011.01424.x
Walters WP, Stahl MT, Murcko MA (1998) Virtual screening—an overview. Drug Discov Today 3(4):160–178. doi:10.1016/S1359-6446(97)01163-X
Gorse A-D (2006) Diversity in medicinal chemistry space. Curr Trends Med Chem 6(1):3–18
Mario Geysen H, Schoenen F, Wagner D, Wagner R (2003) Combinatorial compound libraries for drug discovery: an ongoing challenge. Nat Rev Drug Discov 2(3):222–230
Valler MJ, Green D (2000) Diversity screening versus focussed screening in drug discovery. Drug Discov Today 5(7):286–293. doi:10.1016/S1359-6446(00)01517-8
Giménez O, Noy M (2005) The number of planar graphs and properties of random planar graphs. In: International conference on analysis of algorithms DMTCS proc. AD, Barcelona, Spain, 6-10 June 2005. Discrete Mathematics and Theoretical Computer Science (DMTCS), Nancy, France. p 147–156
R: A Language and Environment for Statistical Computing (2012) R Foundation for Statistical Computing, Vienna, Austria
Lipinski C (1995) Computational alerts for potential absorption problems: profiles of clinically tested drugs. Paper presented at the tools for oral absorption. Part II. Predicting human absorption. BIOTEC. PDD symposium, AAPS, Miami
Irwin JJ, Sterling T, Mysinger MM, Bolstad ES, Coleman RG (2012) ZINC: a free tool to discover chemistry for biology. J Chem Inf Model 52(7):1757–1768. doi:10.1021/ci3001277
Shoichet BK (2013) Drug discovery: nature’s pieces. Nat Chem 5(1):9–10
Gillet VJ, Khatib W, Willett P, Fleming PJ, Green DVS (2002) Combinatorial library design using a multiobjective genetic algorithm. J Chem Inf Comput Sci 42(2):375–385. doi:10.1021/ci010375j
van Deursen R, Reymond J-L (2007) Chemical space travel. ChemMedChem 2(5):636–640. doi:10.1002/cmdc.200700021
Acknowledgments
Authors thank Dr. I. Baskin, Prof. I. Antipin and Dr. G. Marcou for valuable comments. PP thanks the French Embassy in Ukraine for the support of his stay at the University of Strasbourg in 2012. TM acknowledges Kazan Federal University for the support of his stay at the University of Strasbourg in 2012.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Polishchuk, P.G., Madzhidov, T.I. & Varnek, A. Estimation of the size of drug-like chemical space based on GDB-17 data. J Comput Aided Mol Des 27, 675–679 (2013). https://doi.org/10.1007/s10822-013-9672-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-013-9672-4