Abstract
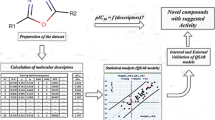
This paper presents the results of an optimization study on biaryl piperidine and 4-amino-2-biarylurea MCH1 receptor antagonists, which was accomplished by using quantitative-structure activity relationships (QSARs), classification and virtual screening techniques. First, a linear QSAR model was developed using Multiple Linear Regression (MLR) Analysis, while the Elimination Selection-Stepwise Regression (ES-SWR) method was adopted for selecting the most suitable input variables. The predictive activity of the model was evaluated using an external validation set and the Y-randomization technique. Based on the selected descriptors, the Support Vector Machines (SVM) classification technique was utilized to classify data into two categories: “actives” or “non-actives”. Several attempts were made to optimize the scaffold of most potent compounds by inducing various structural modifications. Potential derivatives with improved activities were identified, as they were classified “actives” by the SVM classifier. Their activities were estimated using the produced MLR model. A detailed analysis on the model applicability domain defined the compounds, whose estimations can be accepted with confidence.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Melanin-concentrating hormone (MCH) is a cyclic nonadecapeptide which is expressed in the brain of all vertebrates. It has been demonstrated that MCH is involved in feeding and body weight regulation. MCH stimulates food intake in rodents and chronic administration leads to increased body weight. Animals that lack the gene encoding MCH receptor are hypophagic, lean and maintain elevated metabolic rates [1–3].
Two G-protein coupled receptors (GPCRs) have been identified for MCH, namely MCH1R and MCH2R. MCH1R is present in rodents and high mammalian species while MCH2R is expressed only in ferrets, dogs, rhesus monkeys and humans. The pharmacological role of MCH2R in metabolic homeostasis is still undefined whereas the critical role of MCH1R in the regulation of food intake and energy homeostasis has been extensively studied [4–6].
MCH1R has been identified as a key target in MCH regulation, as small molecule antagonists of MCH1R have demonstrated activity in vivo. MCH1R antagonists are potentially interesting agents for treatment of metabolic or obesity-related disorders. This fact has prompted different research groups to design and synthesize MCH1R antagonists which demonstrate in vivo efficiency in the therapy of obesity [7–9]. Alternative techniques could help to decrease the number of animals sacrificed during in vivo testing. In this sense QSAR, virtual screening and pattern recognition could be very useful and promising methodologies. To the best of our knowledge, these techniques have never been attempted so far in the general field of MCH1R. It is thus of particular interest to investigate this possibility.
In this paper we show that QSAR modeling, classification and virtual screening can contribute greatly to the modeling, design and optimization of MCH1R antagonists. The first major result is the development of a QSAR model involving seven descriptors that is able to predict successfully MCH1R binding affinity. The model was generated using a database consisting of a series of 63 MCH1 receptor antagonists including biaryl piperidine- and 4-amino-2-biarylurea-based derivatives [10, 11]. About 69 physicochemical and topological descriptors were examined in terms of their efficacy to determine and predict the activity of the investigated derivatives. The effects of various structural modifications on biological activity were investigated next, within a classification pattern. In particular, the popular SVM classification methodology was utilized to afford novel active patterns. Biological activities of novel structures were estimated using the new QSAR model, while the detection of its domain of applicability defined the compounds whose estimations can be accepted with confidence.
Materials and methods
Data set
The database consists of 63 recently discovered biaryl piperidine and 4-amino-2-biarylbutylureas (Tables 1–3) [10, 11]. In order to model and predict the binding affinity of MCH receptor antagonists, 69 physicochemical constants, topological and structural descriptors (Table 4) were considered as possible input candidates to the model. Before the calculation of the descriptors, all structures were fully optimized using CS Mechanics and more specifically MM2 force fields and the Truncated-Newton-Raphson optimizer, which provide a balance between speed and accuracy (Chemoffice Manual). Before calculating the HOMO and LUMO Energies (eV) all the structures were additionally fully optimized using the AM1 basis set. All the descriptors were calculated using ChemSar and Topix [12, 13].



Separation into a training and a validation set
The separation of the dataset into training and validation sets was performed according to the popular Kennard and Stones algorithm [14]. The algorithm starts by finding two samples that are the farthest apart from each other on the basis of the input variables in terms of some metric, e.g., the Euclidean distance. These two samples are removed from the original data set and placed into the calibration data set. This procedure is repeated until the desired number of samples has been reached in the calibration set. The advantages of this algorithm are that the calibration samples map the measured region of the input variable space completely with respect to the induced metric and that the test samples all fall inside the measured region. According to Tropsha [15] and Wu [16], the Kennard and Stones algorithm is one of the best ways to build training and test sets.
MLR model development-variable selection
Our first objective was to determine the best variables which produce the most significant linear QSAR models linking the structure of compounds with their binding affinity. The ES-SWR algorithm was used on the training data set to select the most appropriate descriptors. ES-SWR is a popular stepwise technique [17] that combines Forward Selection (FS-SWR) and Backward Elimination (BE-SWR).
Model validation
The accuracy of the proposed MLR model was illustrated using validation through an external test set and Y-randomization. The leave-one-out and leave-five-out cross-validation procedures were used to illustrate the robustness of the MLR modeling technique in our particular QSAR study.
Cross-validation test
Cross-validation is popular technique used to explore the reliability of statistical models. Based on this technique, a number of modified data sets are created by deleting in each case one or a small group (leave-some-out) of objects. For each data set, an input–output model is developed, based on the utilized modeling technique. The model is evaluated by measuring its accuracy in predicting the responses of the remaining data (the ones that have not been utilized in the development of the model) [18].
Validation through the external validation set
According to Tropsha’ group [15, 19] a QSAR model is considered predictive, if the following conditions are satisfied:
In Eqs. 2 and 3, R 2 is the coefficient of determination between experimental values and model prediction on the training set. Mathematical definitions of \( R_{\rm{o}}^2 \), \( R_{\rm{o}}^{'2} \), k and k′ are based on regression of the observed activities against predicted activities and the opposite (regression of the predicted activities against observed activities). The definitions are presented clearly in [20] and are not repeated here for brevity.
Y-randomization test
This technique ensures the robustness of a QSAR model [15, 21]. The dependent variable vector (biological action) is randomly shuffled and a new QSAR model is developed, using the given modeling algorithm. The procedure is repeated several times and the new QSAR models are expected to have low R 2 and Q 2 values. If the opposite happens then an acceptable QSAR model cannot be obtained for the specific modeling method and data.
Defining model applicability domain
In order for a QSAR model to be used for screening new compounds, its domain of application [15, 20] must be defined and predictions for only those compounds that fall into this domain may be considered reliable. Extent of Extrapolation [15] is one simple approach to define the applicability of the domain. It is based on the calculation of the leverage h i [22] for each chemical, where the QSAR model is used to predict its activity
In Eq. 4 x i is the row vector containing the k model parameters of the query compound and X is the n × k matrix containing the k model parameters for each one of the n training compounds. A leverage value greater than 3k/n is considered large. It means that the predicted response is the result of a substantial extrapolation of the model and may be not reliable.
Support vector machines
The Support Vector Machine (SVM) method is a new and very promising supervised machine learning technique, originally developed by Vapnik and co-workers while working on Structural Risk Minimization [23–25]. The SVM method is quite different from empirical risk minimization algorithms and is gaining popularity due to the many attractive features it possesses and its promising empirical performance. SVM is a classification approach that has been suggested as being particularly appropriate for chemical applications. In particular, the popular Library for Support Vector Machines (LIBSVM) [26] was utilized in this work. A detailed presentation of the theory behind the SVM technique can be found in several books and tutorials [27]. Here we briefly summarize the main principles of SVMs, when they are used for classification purposes.
A typical application of the SVM technology in chemoinformatics consists of defining two classes of molecules, determining a set of descriptors that characterize each molecule and using the SVM algorithm to develop a classification model. If we assume that a number of training compounds have been classified using experimental data into an active class or an inactive class and that a set of significant descriptors has been calculated for each training compound, the procedure for developing an SVM classification model can be summarized as follows: every molecule has its own image in the multidimensional space, where each dimension corresponds to a different descriptor. The coordinates of the image are obviously the values of the various descriptors. The SVM model seeks to find an optimal hyperplane that best separates the two sets of classes corresponding to the active and non-active compounds in the multidimensional space. There are numerous hyperplanes that may separate the data in this manner. The optimal hyperplane is the one which maximizes the margin, defined as the closest distance from any point to the separating hyperplane. The points used to define the optimal hyperplane are often a small fraction of the entire data and, as such, they allow a SVM model to be less prone to overtraining while maintaining an excellent degree of generalizability. Thus, the produced SVM model can be used to classify other than the training molecules as active or inactive. The predicted class of a molecule that is not included in the training set depends on which side of the separating hyperplane the image of the molecule is located.
Results and discussion
First, the data set of 63 derivatives was partitioned into a training set of 35 compounds, and a validation set of 28 compounds according to the Kennard and Stones [14] algorithm. The algorithm was applied on the complete database consisting of all 69 available descriptors. The validation examples are marked with b in Tables 1–3. The validation data were not involved by any means in the process of selecting the most appropriate descriptors or in the development of the QSAR model. They were considered as a completely unknown external set of data, which was used only to test the accuracy of the produced model.
The MLR QSAR model was thus developed by applying the ES-SWR algorithm on the set of training data. The result was the following 8-parameter (seven descriptors and the intercept) equation:
n = 35; R 2 = 0.78; \( R_{{\rm{adj}}}^{\hbox{2}} = 0.72 \); F = 13.72; RMSE = 0.6256; Q 2 = 0.65; S PRESS = 0.7959.
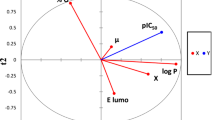
The above equation shows that the most significant descriptors according to the ES-SWR algorithm are Lipophilicity (ClogP), LUMO energy, PSAr, Kier&Hall information index order 2 (KiInf2), Kier&Hall index order 3 (Ki3), Xu3 index and Chicluster4 (ChiCl4). Table 5 presents the correlation matrix and the variance inflation factors (VIF), for the seven descriptors. These statistics indicate that the selected descriptors are not highly correlated. The chemical meaning of the seven descriptors is briefly described next.
Lipophilicity is known to be important for absorption, permeability, and in vivo distribution of organic compounds [28] and has been used as a physicochemical descriptor in QSARs with great success. From the derived QSAR equation we can conclude that lipophilic groups favor the biological action under study.
Polar surface area (PSAr) is defined as the part of the surface area of the module associated with oxygens, nitrogens, sulfurs and the hydrogens bonded to any of these atoms. PSAr has proven to be a very useful parameter for QSAR studies [17].
LUMO energy in particular has been identified as being of significant value to QSAR studies [17, 29]. Molecules with low LUMO values are more able to accept electrons than molecules with high LUMO energy values. The LUMO energy value is increased with the presence of electron donating groups (EDG) such us NMe2, NH2, NHEt and OMe and decreased with the presence of electron withdrawing groups (EWG) such as halogens. From the derived QSAR equation we can conclude that EWGs favor the biological action under study.
In addition to the aforementioned indices, four topological indices were found to significantly influence the activity [17, 30]. Topological indices give information not only about the atomic constitution of a compound but also about the presence and character of chemical bonds by which the atoms are connected to each other.
Equation 5 was used to predict the binding affinity for the validation examples. The results are presented in the last columns of Tables 1–3 and correspond to the following statistics: \( R_{{\rm{pred}}}^2 = 0.83 \), RMSE = 0.6549. In Fig. 1, the experimental vs. predicted values are plotted for both the training and validation sets. The results illustrate that the linear MLR technique combined with a successful variable selection procedure are adequate to generate an efficient QSAR model for predicting the binding affinity of different compounds.
The proposed model (Eq. 5) passed all the tests related to the predictive ability (Eqs. 1–3)
For a more exhaustive testing of the model building technique that was followed in this work, the LOO and L5O cross-validation techniques were applied on the training set of compounds. The L5O method was implemented by selecting randomly groups of five compounds from the available training data. Each group was left out and that group was predicted by the model developed from the remaining observations. 3000 random groups of five compounds were selected for the implementation of the L5O cross-validation test. It should be emphasized that the procedure for developing the QSAR models included the selection of the best descriptors. Therefore, each time one (LOO) or five (L5O) compounds were excluded from the training set, the modeling procedure selected the best descriptors and developed an MLR model based only on the remaining observations. The excluded compounds were not involved by any means in the development of the model. It was important that the model was stable to the inclusion–exclusion of compounds. The results produced by the LOO (Q 2 = 0.65) and the L5O ( \( Q_{{\rm{L}}5{\rm{O}}}^{\hbox{2}} = 0.66 \)) cross-validation tests illustrated the validity of the modeling approach.
The model was further validated by applying the Y-randomization. Several random shuffles of the Y vector were performed and the low R 2 and Q 2 values that were obtained show that the good results in our original model are not due to a chance correlation or structural dependency of the training set. It should be noted that for each random permutation of the Y vector, the complete training procedure was followed for developing the new QSAR model, including the selection of the most appropriate descriptors. The results of the Y-randomization test are presented in Table 6.
The extrapolation method was applied to the compounds that constitute the validation set. The leverages for all 28 compounds were computed (Table 7). All 28 compounds in the test set fall inside the domain of the model (the warning leverage limit is 3k/n 3*8/35 = 0.686).
After the pre-selection of the descriptors the next step was to build the classification model by using SVM. The SVM classification approach has been suggested as being particularly appropriate for chemical applications and well-suited for virtual screening purposes. A successful SVM model is expected to efficiently discriminate between important and unimportant features. For constructing the SVM model, the LIBSVM package was used [26] after scaling both the training and validation data in the range [−1,+1]. The Kernel type that was adopted in the present work was the Radial Basis Function (RBF). The first task is the assignment of each compound to one class, namely “active” or “non-active” based on a cut-off value that was set to 60 for binding affinity K i . For classification purposes, active compounds are assigned a +1 value whereas non-active compounds are assigned a −1 value. These classes are defined a priori by groups of objects in the training set belonging to these classes. The cost parameter C and the gamma parameter γ in the kernel function were optimized to achieve the best possible discrimination between classes. The optimized values obtained were C = 10 and γ = 1. The predictive ability of the model was tested on the validation set of compounds. The total accuracy of the SVM model was 91%, meaning that the model assigned the correct class to 91% of the compounds. Accuracy for the training and validations sets was 100 and 82%, respectively. The results for the validation set are listed in Table 8. The misclassified samples (marked with an asterisk) are clearly indicated.
Our final objective was to be able to classify compounds that are not involved in the training procedure. These compounds were derived from virtual optimization of the lead compounds by insertions, substitutions, and deletions of pharmacophoric substituents of the main building block scaffolds. More specifically, based on the produced SVM classification model, a group of new derivatives, previously not tested for the specific biological action, was subjected to virtual screening. The aim was, starting from a primary hit and using both pharmacophore-based and substructure-based modifications to discover a structurally diverse set of potent leads [31, 32].
A variety of modifications of the initial compounds were introduced and the representative modifications that led to “active” compounds are shown in Tables 9–14. Biological activities of the compounds characterized as “actives” were estimated using the developed MLR equation. The activity values together with the leverages are shown in Tables 9–14. More precisely, the last column in Tables 9–14 shows the difference between the warning limit 0.686 and the leverage calculated for each compound. A negative value means that the respective compound falls outside the domain of applicability of the model. The initial study focused on the substitution of the biaryl moiety and indicated that the biphenyl analogs (id 1v–4v, bearing either 2° or 3° alkylamine side chains while falling well within the domain of applicability were not predicted to be significantly active compounds. The fused analogs (id 5v–9v however, gave good predicted activity but the most active structures (id 7v, 9v) fell outside (or marginally inside) of the applicability domain. The tetrahydroquinoline structure 6v was considered for further modification and the nitrogen heterocyclic ring was converted to the isomeric tetrahydroisoquinolines (id 10v and 11v). Structure id 10v (pred. 1.2616, domain 0.2071) which gave improved predicted activity and remained within the applicability domain was further modified by investigating the nature of the N-substitution (id 12v–18v). Generally the addition of larger branched alkyl chains helped improve the activity but several structures fell outside or marginally inside of the domain of applicability. This was in agreement with the model which emphasizes the lipophilicity descriptor. Interestingly, moving the alkyl groups from the ring nitrogen to the neighboring peri position (id 19v–27v) greatly improved the structures fit within the applicability domain and structure id 25v bearing an iso-butyl substituent showed good activity (1.6211) well within the domain (0.3642). Structure id 25v was therefore considered for further modification. N-Alkylation (id 28v) greatly improved the activity (3.7658) but the structure fell outside the applicability domain (−1.0082). Introducing a further degree of unsaturation (id 29v) however, significantly improved activity (2.3716) within the applicability domain (0.2389). The model predicts that lipophilicity and LUMO are important descriptors and as such structure id 29v was further modified by introducing fluorine substituents on the iso-butyl side chain (id 30v–32v). Structures id 31v and 32v were predicted to have very good activities (2.8676 and 2.6800, respectively) and were within the applicability domain. The urea moiety of structure id 32v was modified to afford N-methylurea, carbamate, carbonate or thiocarbamate structures (id 33v–37v) but this did not afford structures with improved activity within the desired domain of applicability. The phenyl urea substituent of structure id 29v was also investigated (id 38v–50v). The 3,4-difluorobenzene substitution pattern gave the best activity comfortably within the applicability domain. The phenyl substituent of the dihydroisoquinoline moiety of this structure (id 44v) was then modified (id 51v–57v) Structure id 51v showed excellent predicted activity (3.1284) within the domain of applicability (0.1105) and introduction of an additional fluorine on the iso-butyl group (id 58v) forced the structure outside the acceptable domain. In general it has been demonstrated that several potentially active structures can be predicted via virtual screening which fall within the models domain of applicability. The introduction of branched alkyl chains and also the use of fluorine substitution are in agreement with the descriptor model which showed a preference for increased lipophilicity and a more negative LUMO energy. Compounds id 31v, 32v, 33v, 47v, 51v, 53v are predicted to exhibit an increased biological activity and simultaneously fall inside the domain of applicability of the model.


Finally a cautionary note should be included dealing with the biological activity scales. While the data from the experimental and virtual studies has been recorded with the same units it must be noted that the predicted activities produced by the virtual model are significantly higher. It would be truly remarkable if the model was able to accurately predict such activities quantitatively but this is unlikely. The synthesis and study of these compounds would be required to truly validate the virtual model and as such is a worthy pursuit but this is outside the scope of this present paper. It must therefore be noted that the virtual screening study acts only as an aid in proposing structural modifications to assist ongoing SAR studies. The high biological activities predicted are only indicative of which structures should be targeted for synthesis on the basis that they meet or approach the optimal values for the chosen descriptors for the given model.
Conclusion
In the present study seven descriptors (namely LUMO energy, PSAr, ClogP and four topological descriptors: Ki3, KiInf2, ChiCl4, Xu3) were found to be important for describing biological activity of potent MCH receptor antagonists. The seven-descriptor set contains electronic, topological and physicochemical information about molecules, and describes and models successfully the binding affinity of these small molecules.
An SVM classifier was developed based on the partition of the initial dataset into training and validation compounds. The SVM model was then used to classify novel compounds that were derived by inducing structural modification to the initial compounds of the database. Biological activities of novel compounds were estimated by the produced MLR model. The detailed validation procedure that was followed (separation of the data into two independent sets, cross-validation, Y-randomization) illustrated the accuracy and robustness of the produced model not only by calculating its fitness on the training data, but also by testing its predicting ability. The applicability domain served as a valuable tool to filter out “dissimilar” compounds.
Due to its predictive ability, the proposed model could be a useful aid to the costly and time consuming experiments for determining binding affinity of the MCH receptor antagonists.
References
Kowalski TJ, Spar BD, Weig B, Farley C, Cook J, Ghibaudi L, Fried S, O’Neill K, Del Vecchio RA, McBriar M, Guzik H, Clader J, Hawes BE, Hwa J (2006) Eur J Pharmacol 535:182
McBriar M, Guzik H, Shapiro S, Paruchova J, Xu R, Palani A, Clader JW, Cox K, Greenlee WJ, Hawes BE, Kowalski TJ, O’Neill K, Spar BD, Weig B, Weston DJ, Farley C, Cook J (2006) J Med Chem 49:2294
(a) Palani A, Shapiro S, McBriar MD, Clader JW, Greenlee WJ, Spar B, Kowalski TJ, Farley C, Cook J, van Heek M, Weig B, O’Neill K, Graziano M, Hawes B (2005) J Med Chem 48:4746. (b) McBriar MD, Guzik H, Xu R, Paruchova J, Li S, Palani A, Clader JW, Greenlee WJ, Hawes BE, Kowalski TJ, O’Neill K, Spar B, Weig B (2005) J Med Chem 48:2274
Receveur JM, Bjurling E, Ulven T, Little PB, Norregaard PK, Hogberg T (2004) Bioorg Med Chem Lett 14:5075
Rowbottom MW, Vickers TD, Dyck B, Taminiya J, Zhang M, Zhao L, Grey J, Provencal D, Schwarz D, Heise CE, Mistry M, Fisher A, Dong T, Hu T, Saunders J, Goodfellow VS (2005) Bioorg Med Chem Lett 15:3439
Vasudevan A, Wodka D, Verzal MK, Souers AJ, Gao J, Brodjian S, Fry D, Dayton B, Marsh KC, Hernandez LE, Ogiela CA, Collins CA, Kym PR (2004) Bioorg Med Chem Lett 14:4879
(a) Xu R, Li S, Paruchova J, McBriar MD, Guzik H, Palani A, Clader JW, Cox K, Greenlee WJ, Hawes BE, Kowalski TJ, O’Neill K, Spar BD, Weig B, Weston DJ (2006) Bioorg Med Chem 14:3285. (b) Su J, McKittrick BA, Tang H, Czarniecki M, Greenlee WJ, Hawes BE, O’Neill K (2005) Bioorg Med Chem 13:1829
(a) Kanuma K, Omodera K, Nishiguchi M, Funakoshi T, Chaki S, Semple G, Tran T-A, Kramer B, Hsu D, Casper M, Thomsen B, Beeley N, Sekiguchi Y (2005) Bioorg Med Chem Lett 15:2565. (b) Kanuma K, Omodera K, Nishiguchi M, Funakoshi T, Chaki S, Semple G, Tran T-A, Kramer B, Hsu D, Casper M, Thomsen B, Sekiguchi Y (2005) Bioorg Med Chem Lett 15:3853. (c) Kanuma K, Omodera K, Nishiguchi M, Funakoshi T, Chaki S, Nagase Y, Iida I, Yamaguchi J-I, Semple G, Tran T-A, Sekiguchi Y (2006) Bioorg Med Chem 14:3307
Vasudevan A, Wodka D, Verzal MK, Souers AJ, Gao J, Brodjian S, Fry D, Dayton B, Marsh KC, Hernandez LE, Ogiela CA, Collins CA, Kym PR (2004) Bioorg Med Chem Lett 14:4879
Guo T, Hunter RC, Gu H, Shao Y, Rokosz LL, Stauffer TM, Hobbs DW (2005) Bioorg Med Chem Lett 15:3691
Guo T, Shao Y, Qian G, Rokosz LL, Stauffer TM, Hunter RC, Babu SD, Gu H, Hobbs DW (2005) Bioorg Med Chem Lett 15:3696
CambridgeSoft Corporation http://www.cambridgesoft.com
http://www.lohninger.com/topix.html
Kennard RW, Stone LA (1969) Technometrics 11:137
Tropsha A, Gramatica P, Gombar VK (2003) QSAR Comb Sci 22:69
Wu W, Walczak B, Massart DL, Heuerding S, Erni F, Last IR, Prebble KA (1996) Chemometr Intell Lab Syst 33:35
Todeschini R, Consonni V, Mannhold R, Kubinyi H, Timmerman H (Series Editor) (2000) Handbook of molecular descriptors. Wiley-VCH, Weinheim
(a) Efron B (1983) J Am Stat Assoc 78:316. (b) Osten DW (1998) J Chemom 2:39
Shen M, Beguin C, Golbraikh A, Stables J, Kohn H, Tropsha A (2004) J Med Chem 47:2356
Golbraikh A, Tropsha A (2002) J Mol Graph Mod 20:269
Wold S, Eriksson L (1995) In: Van de Waterbeemd H (ed) Chemometrics methods in molecular design, VCH Weinheim, Germany
Atkinson A (1985) Plots, transformations and regression. Clarendon Press, Oxford (UK)
Cortes C, Vapnik V (1995) Mach Learning 20:273
Jorissen RN, Gilson MK (2005) J Chem Inf Model 45:549
Wilton D, Willet P, Lawson K, Mullier G (2003) J Chem Inf Comput Sci 43:469
Chang CC, Lin CJ LIBSVM: http://www.csie.ntu.edu.tw/∼cjlin/libsvm
Burges CJC (1998) Data Min Knowl Discov 2:127
Walters WPA, Murcko MA (1999) Curr Opin Chem Biol 3:384
Karelson M (2000) Molecular descriptors in QSAR/QSPR. Wiley, NY
Kier LB, Hall LB (1986) Molecular connectivity in structure activity analysis. Wiley, Chichester
Afantitis A, Melagraki G, Sarimveis H, Koutentis PA, Markopoulos J, Igglessi-Markopoulou O (2006) J Comput Aid Design 20:83
Afantitis A, Melagraki G, Sarimveis H, Koutentis PA, Markopoulos J, Igglessi-Markopoulou O (2006) Mol Div. 10:405
Acknowledgments
G.M. thanks the Greek State Scholarship Foundation for a doctoral assistantship. A.A. wishes to thank Cyprus Research Promotion Foundation (Grant No. PENEK/ENISX/0603/05) and the Committee of Research of the National Technical University of Athens, Greece for a doctoral assistantship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Melagraki, G., Afantitis, A., Sarimveis, H. et al. Optimization of biaryl piperidine and 4-amino-2-biarylurea MCH1 receptor antagonists using QSAR modeling, classification techniques and virtual screening. J Comput Aided Mol Des 21, 251–267 (2007). https://doi.org/10.1007/s10822-007-9112-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-007-9112-4