Abstract
Saccharina latissima is an ecologically and economically important kelp species native to the coastal regions of the Northern Hemisphere. This species has considerable phylogeographic structure and morphological plasticity, but lack of resolution of available genetic markers prevents a finer characterization of its genetic diversity and structure. Here, we describe 12 microsatellite loci identified in silico in a genomic library, and assess their polymorphism in three distant populations. Allelic richness at the species level was relatively high (5–23 alleles per locus), as was gene diversity within populations (0.42 < H E < 0.62). In addition, individuals readily form distinct genotypic clusters matching their populations of origin. The variation detected confirms the great potential of these markers to investigate the biogeography and population dynamics of S. latissima, and to better characterize its genetic resources for the establishing farming industry.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The sugar kelp Saccharina latissima (Linnaeus) C.E. Lane, C. Mayes, Druehl and G.W. Saunders (=Laminaria saccharina) is a cryptic perennial, canopy-forming, edible brown seaweed (Laminariales, Heterokontophyta) distributed throughout much of the Northern Hemisphere. Its vast range extends from the high Arctic to cold-temperate Iberia (NE) and New Jersey (NW) in the Atlantic, and to Central California (NE) and Korea (NW) in the Pacific (Müller et al. 2009). In addition to its ecological importance as a foundational species in lower intertidal and shallow subtidal environments (Steneck et al. 2002, Gutiérrez et al. 2011), S. latissima is cultivated/harvested for human consumption (Smale et al. 2013). It is prized for its sweet flavor, particularly in Asian markets (where it is sold with other species as “kombu”), but also in Europe and America, where sea farming is currently a growing enterprise (Peteiro et al. 2014).
The species S. latissima is morphologically and genetically polymorphic throughout its range, comprising regional ecotypes (e.g., now synonymized Saccharina longicruris from Canada) and at least three distinct mtDNA phylogeographic lineages (McDevit and Saunders 2010). Where these overlap, such as along the Canadian Arctic and western Greenland, nuclear internal transcribed spacer (ITS) sequences are consistent with ongoing admixture, suggesting that S. latissima may correspond to an incipient species complex resulting from historical cycles of glacial isolation and interglacial admixture. The genetic markers available so far are, however, not suited to adequately assess gene flow across secondary contact zones, or intra-oceanic structure, highlighting the need for independent, more polymorphic, co-dominant markers such as microsatellites. Other applications for such markers include identifying regional hotspots of genetic diversity and endemism (i.e., refugia) to better forecast the consequences of ongoing climatic and environmental change (Müller et al. 2009; Moy and Christie 2012; Martínez et al. 2015), and characterizing available genetic resources, population dynamics, and relevant life-history traits of S. latissima for the establishing farming industry.
Given the above, we describe 12 microsatellite markers for S. latissima identified in silico following 454 pyrosequencing, and characterize their polymorphism and differentiation in three selected populations.
Material and methods
Four hundred fifty-four DNA shotgun pyrosequencing of Saccharina latissima was performed at Biocant (Cantanhede, Portugal). DNA was extracted from 100 mg of dried tissue from four individuals collected at Uummannaq (western Greenland) using a modified protocol of the commercial DNeasy Plant Mini (Qiagen, Hilden) DNA extraction kit (Snirc et al. 2010). Simple di- to hexanucleotide microsatellite loci were searched for, and primers designed, using WebSat (Martins et al. 2009). A subset of 22 promising loci (possessing high number of repeats, adequate anchoring sites for primers and close melting temperatures of primers) was amplified with labeled primers and evaluated for amplification success, ease of scoring, and polymorphism, and 12 were eventually selected (Table 1) to screen three distant populations (n = 20) from the NE Atlantic/Arctic distribution: Uummannaq (70.7 °N), Bergen (southwestern Norway, 60.4 °N), and Roscoff (northwest France, 48.7 °N).
PCR amplifications were performed in 15 μL reaction mixtures containing ±10 ng of template DNA, 1× buffer, 1.5–2.5 mM of MgCl2 (Table 1), 0.28 mM of dNTPs (Bioline), 0.1–0.3 μM of each primer pair with 5′end of forward primer fluorescently labeled, and 1 U of GoTaq G2 Flexi DNA Polymerase (Promega). PCR reactions were performed in a ABI 2720 Thermal Cycler (Applied Biosystems) and included an initial denaturation step (95 °C, 5 min), 35 cycles of 95 °C for 30 s, T a °C for 30 s (Table 1) and 72 °C for 40 s, and a final extension step (72 °C, 20 min; 5 min for SLN314). Amplified fragments were run in an ABI PRISM 3130xl automated DNA sequencer (Applied Biosystems) at CCMAR, Portugal. Alleles were manually scored in STRand (Toonen and Hughes, 2001) using the GeneScan 500 LIZ size standard (Applied Biosystems), and binned using MsatAllele (Alberto 2009).
GENETIX v4 (Belkhir et al. 1996) was used to test for potential linkage disequilibrium (LD) of alleles and to estimate the total number of alleles or allelic richness (A), expected (H E) and observed (H O) heterozygosities and inbreeding (F IS) in each population. Polymorphic information content (PIC) was calculated using Cervus v3 (http://www.fieldgenetics.com/pages/home.jsp). Population differentiation was inspected using a factorial correspondence analysis (FCA) based on individual multilocus genotypes.
Results and discussion
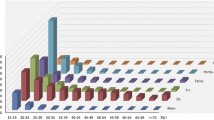
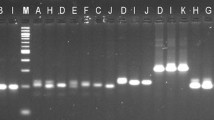
Four hundred fifty-four pyrosequencing produced a total of 214,127 quality reads between 40 and 640 bp, of which 22,454 (10.5 %) contained at least one simple di- to hexanucleotide microsatellite with at least 4 repeats. All 12 microsatellite loci selected successfully amplified in the three surveyed populations. Consistent LD across populations was not detected in any loci. Globally, the number of alleles per locus (A) ranged from 5 to 23 (averaging 12.42), and within populations, it ranged from 1 to 13. Polymorphic information content (PIC) was rather high for all loci, ranging from 0.452 to 0.939 in the pooled dataset. Polymorphism within populations was also appreciable, with mean expected heterozygosity (H E) ranging between 0.48 and 0.62 and mean A ranging between 4 and 6.25 alleles per locus. Significant homozygote excess was observed in all populations in at least some loci (Table 2). These disequilibria were locus- and population-specific, however, and thus were more likely to reflect sampling artifacts or null alleles rather than local differences in mating systems or population dynamics. Indeed, MICRO-CHECKER (Van Oosterhout et al. 2004) identified null alleles at loci SLN34 (Uummannaq, Bergen) and SLN35 (Uummannaq). Allelic compositions across populations were sufficiently different so that individual multilocus genotypes formed clearly discontinuous and coherent clusters according to geography (Fig. 1).
The levels of intra- and interpopulation polymorphism in this preliminary survey confirm the usefulness of these markers to characterize population diversity and structure of S. latissima at multiple spatial scales, and their potential use to identify and manage genetic resources (e.g., selection of germplasm banks) for cultivation purposes.
References
Alberto F (2009) MsatAllele-1.0: an R package to visualize the binning of microsatellite alleles. J Hered 100:394–397
Belkhir K, Borsa P, Chikhi L, Raufaste N, Bonhomme F (1996) GENETIX 4.05, logiciel sous Windows TM pour la génétique des populations. Laboratoire Génome, Populations, Interactions. Université de Montpellier II, Montpellier.
Gutiérrez JL, Jones CG, Byers JE, Arkema KK, Berkenbusch K, Commito JA, Duarte CM, Hacker SD, Lambrinos JG, Hendriks IE, Hogarth PJ, Palomo MG, Wild C (2011) Physical ecosystem engineers and the functioning of estuaries and coasts. In: Wolanski E, McLusky D (eds.) Treatise on Estuarine and Coastal Science, Vol 7, Elsevier pp. 53–81.
Martínez B, Afonso-Carrillo J, Anadón R, Araújo R, Arenas F, Arrontes J, Bárbara I, Borja A, Díez I, Duarte L, Fernández C, Tasende MG, Gorostiaga JM, Juanes JA, Peteiro C, Puente A, Rico JM, Sangil C, Sansón M, Tuya F, Viejo RM (2015) Regresión de las algas marinas en la costa atlántica de la Península Ibérica y en las Islas Canarias por efecto del cambio climático. Algas 49:5–12
Martins WS, Lucas DCS, Neves KFDS, Bertioli DJ (2009) WebSat—a web software for microsatellite marker development. Bioinformation 3:282–283
McDevit DC, Saunders GW (2010) A DNA barcode examination of the Laminariaceae (Phaeophyceae) in Canada reveals novel biogeographical and evolutionary insights. Phycologia 49:235–248
Moy FE, Christie H (2012) Large-scale shift from sugar kelp (Saccharina latissima) to ephemeral algae along the south and west coast of Norway. Mar Biol Res 8:309–321
Müller R, Laepple T, Bartsch I, Wiencke C (2009) Impact of oceanic warming on the distribution of seaweeds in polar and cold-temperate waters. Bot Mar 52:617–638
Peteiro C, Sánchez N, Dueñas-Liaño C, Martínez B (2014) Open-sea cultivation by transplanting young fronds of the kelp Saccharina latissima. J Appl Phycol 26:519–528
Smale DA, Burrows MT, Moore P, O’Connor N, Hawkins SJ (2013) Threats and knowledge gaps for ecosystem services provided by kelp forests: a northeast Atlantic perspective. Ecol Evol 3:4016–4038
Snirc A, Silberfeld T, Bonnet J, Tillier A, Tuffet S, Sun J-S (2010) Optimization of DNA extraction from brown algae (Phaeophyceae) based on a commercial kit. J Phycol 46:616–621
Steneck RS, Graham MH, Bourq BJ, Corbett D, Erlandson JM (2002) Kelp forest ecosystems, biodiversity, stability resilience and future. Environ Conserv 29:436–459
Toonen RJ, Hughes S (2001) Increased throughput for fragment analysis on an ABI Prism® 377 automated sequencer using a membrane comb and STRand software. BioTechniques 31:1320–1324
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Acknowledgments
This work was funded by Portuguese Fundação para a Ciência e Tecnologia (FCT) through the project EXTANT (EXCL/AAG-GLO/0661/2012) and postdoctoral fellowship to JN (SFRH/BPD/88935/2012), and through the European Union’s Seventh Framework Programme, grant agreement no. 226248—Arctic Tipping Points (ATP).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Paulino, C., Neiva, J., Coelho, N.C. et al. Characterization of 12 polymorphic microsatellite markers in the sugar kelp Saccharina latissima . J Appl Phycol 28, 3071–3074 (2016). https://doi.org/10.1007/s10811-016-0811-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-016-0811-x