Abstract
High-temperature stress is a major abiotic stress that affects the yield and quality of Pyropia, which is produced for use as a nutrient. Pyropia haitanensis strain Z-61 is tolerant to high temperature and is widely cultivated in South China. Here, we conducted a comparative proteomic analysis of the blades of Z-61 strain at 29 °C. Changes in the proteome of the blades were analyzed every other day for 8 days at 29 °C. Approximately 1,000 protein species were reproducibly detected at all time points, whereas 87 protein spots were differentially expressed at least at one time point. Fifty-nine protein spots were successfully identified using liquid chromatography–tandem mass spectroscopy and database searching. The proteins were classified into nine functional categories: photosynthesis (27.12 %), energy metabolism (22.03 %), other metabolic pathways (11.86 %), redox homeostasis (11.86 %), response to stimuli (8.47 %), proteins involved in transcription and translation (6.78 %), cytoskeleton-related proteins (5.08 %), proteins that mediate signal transduction (1.69 %), and unknown proteins (5.08 %). These findings indicated that the blades of Z-61 resist high-temperature stress by inhibiting photosynthesis and other nonessential metabolic processes. In contrast, to increase energy metabolism, the expression of proteins related to redox homeostasis and response to stimuli were upregulated, thereby maintaining physiological balance during stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pyropia, a genus of marine red algae, is one of the most economically important mariculture crops, with an annual harvest of more than 120,000 t (dry weight) and a value of over US$ 2 billion per year (Blouin et al. 2010). With the expansion of artificial seeding and development of the floating culture method, farming and processing of Pyropia have generated the largest seaweed industries in East Asian countries, including China, Japan, and South Korea (Sahoo et al. 2002). In China, two major cultivars, Pyropia yezoensis (Ueda) Hwang et Choi and Pyropia haitanensis (Chang et Zheng) Kikuchi et Miyata, are distributed in North and South China, respectively. P. haitanensis, a typical warm temperate zone species originally found in Fujian Province, is widely cultivated along the coasts of South China (Zhang et al. 2005). The production of P. haitanensis now represents 75–80 % of the total production of cultivated Pyropia in China (Xie et al. 2010).
Under natural conditions, the optimum temperature for conchospore release and net seeding by P. haitanensis is 25 to 27 °C. However, in early seeding period (autumn in South China), Pyropia farms often suffer from sustained high temperatures or temperature increases following a dramatic drop to temperatures suitable for conchospore release and germination, which is known as “temperature rebound” (Zhang et al. 2011a, b). The persistence of high temperatures can negatively affect Pyropia blades by inhibiting the survival and division of the conchospores. In addition, the germlings would be prone to disease, premature senility, and eventual decay, leading to a substantial reduction in yield (Yan et al. 2010). In recent years, high temperatures associated with global warming have markedly affected the cultivation of P. haitanensis and reduced its yield along the coasts of Fujian and Zhejiang (China), which comprise one of the two primary areas used for cultivation. Therefore, in addition to screening for cultivars with higher yield and better quality, breeding cultivars for tolerance to high temperature is a major objective.
Chen et al. (2008) and Yan et al. (2010) selected several strains (Z-26, Z-61, and ZS-1) of P. haitanensis exhibiting high-temperature tolerance and high growth rates. These strains are widely cultivated in South China, and their traits of high-temperature tolerance have been validated by field cultivation. To understand the physiological mechanism of high-temperature tolerance, Zhang et al. (2011a, b) compared the physiological response of a high-temperature tolerance strain (Z-61) to that of a wild-type strain and found that Z-61 can induce its antioxidant and osmoregulation systems, whereas the wild-type strain only induced its osmoregulation system. Further, the baseline activity of the antioxidant system differed significantly between the two strains. However, little is known about the molecular changes that occur in the tolerant strains that affect the biochemical pathways involved in their response to high-temperature stress. Although much evidence exists detailing the molecular mechanisms of high-temperature stress tolerance in vascular plants, and many genes related involved in the high-temperature response have been identified (Wahid et al. 2007), much less is known about the molecular mechanism of high-temperature tolerance in algae.
Discovering and understanding the functions of genes whose expression responds to high-temperature stress is essential. To this end, a global transcriptomic study of the response to high temperature identified more than 1,000 differentially expressed genes (Xie et al., unpublished). It is important, however, to confirm studies of transcription with those that address protein expression. In fact, several studies have shown that changes in transcription often do not correspond with changes in protein expression, and only direct measurements of proteins will reveal authentic functional changes (Gygi et al. 1999; Hack 2004; Ghazalpour et al. 2011). Proteins are the direct effectors of the response of plants to stress. They not only include enzymes that mediate changes in the levels of metabolites but also serve as components of the transcription and translation machinery. Proteomic studies can thus lead to identification of proteins involved in the stress response and contribute significantly to our understanding of the molecular mechanisms underlying tolerance to stress. It is imperative to determine the pattern of proteins expressed by P. haitanensis in response to high-temperature stress to gain a better understanding of the molecular basis of high-temperature stress, as well as identifying proteins that mediate this process.
With the development and improvement of protein separation and identification techniques, comparative proteomics has evolved into a powerful tool for identifying previously unrecognized proteins and those expressed differentially in response to abiotic stress in higher plants (Neilson et al. 2010; Kosová et al. 2011). However, to the best of our knowledge, few proteomic studies have focused on macroalgae. Examples include reports describing the extraction and identification of proteins obtained from Gracilaria changii (Wong et al. 2006), Ecklonia kurome (Nagai et al. 2008), and Saccharina japonica (Kim et al. 2011); the analysis of seasonal variation in protein expression in the sporophyte of S. japonica (Yotsukura et al. 2010); and the identification of 19 proteins involved in copper tolerance in Scytosiphon gracilis (Contreras et al. 2010). One reason for this is that extracting proteins from macroalgae is difficult because of the presence of secondary metabolites, mainly polysaccharides and polyphenols, which cause proteins to precipitate or streak or generate artifactual spots in two-dimensional electrophoresis (2-DE) gels (Yotsukura et al. 2010; Kim et al. 2011). We have developed an effective method for protein isolation from P. haitanensis and the separation these proteins by 2-DE to facilitate their identification (Hang et al. 2011). The aim of the present study was to analyze the proteomic profiles of P. haitanensis Z-61 in response to high-temperature stress and to identify proteins that are differentially expressed using a combination of 2-D PAGE gels and mass spectrometry analysis, coupled with database searching.
Materials and methods
Plants and stress treatment
We used P. haitanensis strain Z-61, which is tolerant to high temperatures and grows with a high yield. It was selected and purified by researchers working in the Laboratory of Germplasm Improvements and Applications of Pyropia in Jimei University, Fujian, China. Fifty 20 ± 2-cm long blades of Z-61 were randomly selected and cultured in ten aerated flasks (2,000 mL) with five blades each, at 21 °C (control temperature) and 29 °C (high temperature) and illuminated by 50–60 μmol photons m−2 s−1 (10:14, L/D) for 0, 2, 4, 6, and 8 days. The culture medium was natural seawater enriched with MES medium and was refreshed every 2 days.
Chlorophyll fluorescence measurements
To monitor the intensity of high-temperature stress on Z-61 blades of P. haitanensis, the optimum quantum yield, a chlorophyll fluorescence marker for photosynthetic efficiency, was measured with a portable pulse amplitude-modulated fluorometer (DIVING-PAM, Walz, Germany). Intrinsic fluorescence (F 0) was determined after maintaining the tissue in darkness for 30 min. A saturating actinic light pulse (5,000 μmol photons m−2 s−1, 800 ms) was applied to obtain maximum fluorescence (F m) in the dark-adapted samples. Variable fluorescence (F v) was determined as the difference between F m and F 0, and optimum quantum yield (F v/F m) was calculated (Schreiber et al. 1994).
Protein extraction and purification
The blade proteins of Z-61 were extracted according to a previously published method (Hang et al. 2011). Briefly, blades were ground in a precooled mortar in the presence of liquid nitrogen. Approximately 0.2 g of the homogenate was precipitated for 2 h with 10 mL acetone, 10 % trichloroacetic acid, 0.07 % β-mercaptoethanol (BME), and 0.2 % polyvinylpyrrolidone at −20 °C. After centrifugation at 15,000×g for 20 min at 4 °C, the supernatant was removed and the pellet was rinsed three times in ice-cold acetone (including 0.07 % BME). Between the rinsing steps, the sample was incubated for 30 min at 25 °C. The pellet was air-dried, resuspended in 5 mL lysis buffer (7 M urea, 2 M thiourea, 40 mM Tris base, 4 % 3-[(3-cholamidopropyl)-dimethylammonio]-propanesulfonate, 2 % immobilized pH gradient (IPG) buffer (pH 3∼10), 65 mM dithiothreitol (DTT)), and vortexed for 1 h at room temperature. The homogenate was centrifuged at 15,000×g for 20 min at 4 °C, and the supernatant was collected for 2-DE analysis.
2-DE and data analysis
For 2-DE, a 250-μg protein sample was loaded onto immobilized IPG strips (GE Healthcare; pH 4–7, 24 cm linear) and rehydrated passively with 450 μL protein solution for 12 h at 20 °C using an Ettan IPGphor system (GE Healthcare). Isoelectric focusing was conducted in six steps as follows: 100 V for 1 h, 200 V for 1 h, 500 V for 1 h, 1,000 V for 1 h, 8,000 V for 2.5 h, and 10 h at 8,000 V (total ∼80 kV h). The focused strips were then equilibrated twice in equilibration solution for 15 min. The first equilibration was performed in 75 mM Tris–HCl (pH 8.8), 6 M urea, 30 % glycerol, 2 % sodium dodecyl sulfate (SDS), 0.01 % bromophenol blue, and 1 % DTT. The second equilibration was performed in 75 mM Tris–HCl (pH 8.8), 6 M urea, 30 % glycerol, 2 % SDS, 0.01 % bromophenol blue, and 4 % iodoacetamide. For the second dimension, the proteins were separated on 12 % SDS–polyacrylamide gels using an Ettan DALTsix electrophoresis system (GE Healthcare). The proteins in the gels were visualized by silver staining (Yan et al. 2000) and scanned at 300 dpi using a GE image scanner. Image analysis was performed using ImageMaster 2D Platinum software version 6.0 (GE Healthcare). The relative abundance of each protein spot was estimated by the volume percentage (vol%) of total spot densities in the gel. The levels of protein expression were calculated with three biological replicates of each sample, and the results were analyzed using ANOVA. Proteins were considered differentially expressed only if they exhibited reproducible twofold changes in vol% that were statistically significant (p < 0.05).
Protein excision and digestion
Protein spots of interest were manually excised from 2-D gels with a scalpel and washed two times with 70 μL deionized water. Gel slices were destained twice with 100 mM NH4HCO3 in 30 % acetonitrile (ACN) and lyophilized overnight before digestion (16 h) with trypsin (Promega, USA) in 25 mM NH4HCO3 containing 10 % ACN at 37 °C. The peptides were extracted three times with 0.1 % trifluoroacetic acid (TFA) in 60 % ACN, and the extracts were pooled and lyophilized for mass spectrometry (MS) analysis.
MS analysis and database searching
The lyophilized peptide samples were dissolved in 0.1 % TFA, and MS analysis was conducted using a matrix-assisted laser desorption/ionization–time-of-flight mass spectrometer 4800 Plus proteomics analyzer (Applied Biosystems, USA). Peptide mass maps were generated in positive ion reflector mode (accelerating voltage 20 kV) with 1,000 laser shots per spectrum, with a signal-to-noise ratio minimum set to 10 and a local noise window width of m/z 250. Up to 10 of the most intense ions were selected as precursors for MS/MS data acquisition. Further processing and interpretation of the MS/MS spectra were performed using GPS Explorer (V3.6, ABI). MS/MS data from each individual spot were merged into a single file for database searching. The MS/MS data were searched for matches to the NCBI and Swiss-Prot databases using MASCOT V2.1 software (Matrix Science, UK). The search parameters were set as follows: trypsin was used to digest the proteins, up to two missed cleavages were allowed, carbamidomethyl (Cys) was the fixed modification and oxidation (Met) was the variable modification, maximum peptide mass tolerance was 70 ppm, and the general fragment mass tolerance was ±0.25 Da. According to the search engine parameters, scores greater than 65 (p < 0.05) were considered positive.
Results
Effects of high-temperature stress on photosynthesis
Optimum quantum yield is a chlorophyll fluorescence marker for photosynthetic efficiency during abiotic stress (Schreiber et al. 1994). Figure 1 shows the change of optimum quantum yield of Z-61 blades under high temperature (29 °C) and control temperature (21 °C) over time. The quantum yield did not change significantly at the control temperature; however, it decreased significantly during high-temperature stress to 67.9 % of the control after 8 days. The result indicated that the high temperature inhibits photosynthesis in the Z-61 blades significantly.
2-DE analysis of P. haitanensis proteins expressed under high-temperature stress
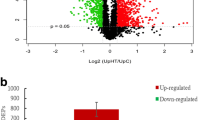
To investigate the dynamic changes in the response of proteins to high-temperature stress, the soluble proteins of Z-61 blades cultured at 21 °C (control group) and 29 °C (high-temperature stress group) for 0, 2, 4, 6, and 8 days were extracted and the proteomic dynamics were analyzed by high-resolution 2-DE. Protein spots displaying reproducible patterns were identified and their expression patterns were analyzed. Representative 2-DE gel maps of each sample are shown in Fig. 2. Between the two groups, 1,263 reproducible protein spots were detected, of which 937 coincided in all gels. Among the 937 coincident spots, the abundance (vol%) of 126 changed by more than twofold (p < 0.05) in the high-temperature stress groups. When they were compared with the control group, 39 were excluded because they also displayed significant changes in the control group. The remaining 87 protein spots were treated as differentially expressed proteins (DEPs) related to high-temperature stress (Fig. 3).
Identification of high temperature-responsive proteins
Among the 87 DEPs, 59 DEPs were successfully identified using LC-MS/MS and database searching (Table 1). Among them, four isoforms were identified, each with two spots located at a different position on the same gel. Spots 60 and 61 were identified as ferredoxin-NADP reductase, spots 2 and 110 as the beta subunit of allophycocyanin, spots 43 and 47 as enoyl-(acyl-carrier-protein) reductase (NADH), and spots 49 and 64 as membrane-associated 30-kDa protein. Further examination of the relative molecular weights (M r) and isoelectric points (pI) showed that the observed and theoretical values of many spots are different (Table 1). This phenomenon might be caused by posttranslational modification or degradation. Posttranslational modifications such as glycosylation and phosphorylation can change the molecular weight, charge, or both.
Among the 59 identified DEPs, three main types of protein expression patterns were detected under high-temperature stress. The first group (group I) includes 24 DEPs whose abundances were consistently upregulated during high-temperature stress, such as spots 2, 16, and 100. The second group (group II) includes 19 DEPs whose abundances were consistently downregulated during high-temperature stress, such as spots 14, 64, and 122. The third group (group III) includes 15 DEPs whose abundances fluctuated, such as spots 11, 54, and 124. During the early stage of high-temperature stress, their abundance was upregulated and then diminished. The abundance of the remaining spot 65 was downregulated during the early stage of high-temperature stress and increased at later times. The dynamic changes in the level of each protein are shown in Table 1.
We were not able to identify 28 protein spots, perhaps because the entire genomic sequence of P. haitanensis has not been determined.
Functional classification of the stress-responsive proteins
The proteins were classified into nine categories according to their functions (Fig. 4). The proteins related to photosynthesis and energy metabolism form the largest two categories, which accounted for 27.12 and 22.03 % of the total. These two categories account for half of the proteins, indicating that high-temperature stress exerts a significant effect on the photosynthesis and energy metabolism. The other categories are as follows: other metabolic pathways (11.86 %), redox homeostasis (11.86 %), response to stimuli (8.47 %), proteins involved in transcription and translation (6.78 %), cytoskeleton-related proteins (5.08 %), proteins that mediate signal transduction (1.69 %), and unknowns (5.08 %).
Discussion
Proteomics and transcriptomics are becoming indispensable techniques for understanding how organisms adapt to abiotic stresses, and they have promoted the identification of numerous stress-related genes and proteins and the pathways in which they function (Hirayama and Shinozaki 2010; Kosová et al. 2011). Changes in protein accumulation under stress are directly related to the phenotypic response of plants to stress, and proteomic analysis can provide more direct information regarding abiotic stresses than transcriptomic analysis. High-temperature stress exerts a major impact on the yield and quality of Pyropia (Yan et al. 2010; Zhang et al. 2011a, b). The mean seawater temperature is increasing because of global warming; therefore, the study of how Pyropia responds to high-temperature stress at the protein level is important for developing high temperature-tolerant strains. The proteomic responses to high-temperature stress are known for several higher plants, such as Arabidopsis thaliana (Palmblad et al. 2008), rice (Lee et al. 2007; Lin et al. 2005), wheat (Majoul et al. 2003, 2004; Skylas et al. 2002), barley (Sule et al. 2004), Populus euphratica (Ferreira et al. 2006), and Carissa spinarum (Zhang et al. 2010a, b).
In the present study, we analyzed the blade proteome of P. haitanensis under conditions of high-temperature stress for different times. Eighty-seven proteins were observed to be differentially expressed. Only limited genomic data are available for P. haitanensis; therefore, only 59 DEPs (67.82 %) could be identified (Table 1). These proteins were classified into nine categories based on function (Fig. 4, Table 1).
Photosynthesis-related proteins
Photosynthesis is one of the systems that are most sensitive to high-temperature stress. Changes in environmental temperature are primarily reflected by photosynthesis, which triggers a response aimed at reaching the best possible performance under new conditions. For this, a balance is sought between the energy of absorbed light, carbon assimilation, and consumption in metabolic sinks. Several studies have shown that high-temperature stress can significantly inhibit the rate of photosynthesis (Salvucci and Crafts-Brandner 2004). In the present study, 16 proteins (27.12 %) related to photosynthesis were differentially expressed.
The process of photosynthesis is initiated by the absorption of light. Pyropia uses multimeric protein complexes, the phycobilisomes (PBSs), as their major light-harvesting complexes (Liu et al. 2005). PBSs comprise three basic types of biliprotein (phycoerythrin (PE), phycocyanin (PC), allophycocyanin (APC)) and linker polypeptides. Light energy that is absorbed by PE migrates first to PC, then to APC, and finally to chlorophyll a (Samsonoff and MacColl 2001). Linker polypeptides function to stabilize the PBS structure and optimize absorbance characteristics and energy transfer (Liu et al. 2005). In the present study, under high-temperature stress, the downregulation of the phycobilisome linker polypeptide (spot 71) would significantly disturb the stability of PBSs and must, therefore, influence light absorbance and energy transfer of P. haitanensis. In such a case, to maintain the balance of light absorbance and energy transfer, the level of each biliprotein must be upregulated. Thus, the upregulations of the PC alpha subunit (spot 124), APC alpha subunit (spot 99), and APC beta subunit (spots 2 and 110) were understandable.
In seaweeds, carbonic anhydrase plays key roles in the CO2-concentrating mechanism by accelerating the conversion of the external HCO3 − pool into CO2 or by mobilizing the internal pool of HCO3 − to supply the carboxylation reaction of RuBisCo (Zhang et al. 2010a, b). RuBisCo catalyzes the first major step of carbon fixation in photosynthesis. Jordan and Ogren (1984) found that the affinity of RuBisCo for CO2 decreased with increased temperature. In the present study, although the level of carbonic anhydrase (spot 68) was upregulated, the level of expression of the RuBisCo small subunit was downregulated. The diminished capacity of RuBisCo and its low affinity for CO2 clearly indicate that carbon fixation or assimilation is highly inhibited when blades of P. haitanensis are exposed to high-temperature stress. Heat-induced changes of RuBisCo subunits have also been monitored in rice seedlings, and the data indicate that high temperature (over 40 °C), for prolonged periods (>2 h), significantly inhibited the activity and quantity of RuBisCo subunits (Lee et al. 2007).
Fructose bisphosphatase (spot 13), fructose-1,6-bisphosphate aldolase (spots 81 and 122), ferredoxin-NADP reductase (spots 60 and 61), photosystem II protein H (spot 14), and chloroplast-localized protein (spot 86) are the key enzymes of the Calvin cycle or participate in photosynthetic electron and energy transfer. Thus, the downregulated expression of these proteins further suggests that photosynthesis of P. haitanensis was inhibited by high-temperature stress. The speculation was validated by the measurement of optimum quantum yield of Z-61 blades during high-temperature stress (Fig. 1).
Energy metabolism-related proteins
Plants respond to stress conditions by triggering a network of events linked to energy metabolism (Hirayama and Shinozaki 2010; Cramer et al. 2011). As expected, 22.03 % of all proteins identified in the present study are related to energy metabolism. trans-2-Enoyl-CoA reductase (spot 34), beta-ketoacyl-acyl carrier protein synthase (spot 35), enoyl-(acyl-carrier-protein) reductase (NADH) (spots 43 and 47), and Rossmann-fold NAD(P)(+)-binding protein (spot 82) are involved in fatty acid biosynthesis. Aconitate hydratase (spot 30), glucose-6-phosphate 1-dehydrogenase (spot 38), transaldolase (spot 52), enolase (spot 54), cytosolic glyceraldehyde-3-phosphate dehydrogenase (spot 59), and pyruvate dehydrogenase E1 component beta subunit (spot 118) are involved in the tricarboxylic acid cycle, the pentose phosphate pathway, or the glycolytic pathway. These differentially expressed proteins, which were all upregulated at least once under high-temperature stress, would lead to the accumulation of acetyl-CoA and cause a subsequent increase in adenosine triphosphate (ATP) production. The result suggests that the blades of P. haitanensis require high levels of energy to cope with stress and to repair damage under high-temperature stress. This speculation is consistent with the upregulation of two proteins known to be involved in the energy production pathway: protein belonging to the H+/Na+-translocating f-type, v-type, and A-type ATPase superfamily (spot 24) and an inorganic pyrophosphatase (spot 73). ATPases represent a class of enzymes that catalyze the decomposition of ATP into adenosine diphosphate (ADP) and a free phosphate ion. Inorganic pyrophosphatase catalyzes the conversion of one molecule of pyrophosphate to two phosphate ions. These two proteins play essential roles in energy conservation and provide the energy for many biosynthetic pathways. ATPase and inorganic pyrophosphatase are also upregulated in many higher plants when exposed to heat stress (Neilson et al. 2010).
Other metabolic pathways-related proteins
In contrast to the upregulation of proteins related to energy metabolism, all of the differentially expressed proteins that function in other metabolism pathways were downregulated. Bifunctionals 3′-phosphoadenosine 5′-phosphosulfate synthetase 2 (spot 17) mediates two steps in the sulfate activation pathway. PAP2-like proteins (spot 25), alanine-glyoxylate transaminase (spot 65), d-3-phosphoglycerate dehydrogenase (spot 104), and the glycine cleavage system T protein (spot 80) participate in amino acid metabolism. Carbonyl reductase (spot 90) participates in arachidonic acid metabolism. Cinnamyl alcohol dehydrogenase (spot 102) is a key enzyme in the biosynthesis of lignin. The downregulation of these proteins suggests that the synthesis of lignin and other metabolites was inhibited under high-temperature stress. This suggests that the blades of P. haitanensis can avoid cell damage under stress by reducing unnecessary consumption of metabolites. The same phenomenon occurs in many plants under stress (Kosová et al. 2011).
Redox homeostasis-related proteins
Abiotic stresses can induce the production of reactive oxygen species (ROS). ROS can act as signaling molecules for stress responses; however, they can also cause damage to cellular components. To protect against oxidative damage, cells have developed a wide range of antioxidant systems that scavenge or reduce excessive ROS or ROS-induced toxic substances. In the present study, a group of antioxidant enzymes were identified, including pyridine nucleotide-disulfide oxidoreductase (spot 18), phosphomannomutase (spot 45), glutathione S-transferase (spot 85), ferritin 3 (spot 100), lactoylglutathione lyase (spot 101), cytosolic ascorbate peroxidase (spot 116), and cystathionine β-synthase (spot 36) domain-containing protein. Several reports have shown that these antioxidant enzymes play important roles in the pathway for scavenging ROS and enhance tolerance to environmental and biotic stresses (Dixon et al. 2002; Seppänen et al. 2000; Sweetlove et al. 2002; Moons 2003; Asada 1992; Mittler et al. 2004; Hegedus et al. 2008; Qian et al. 2007; Yadav et al. 2005; Mustafiz et al. 2010; Kumari et al. 2008; Kushwaha et al. 2009). The consistently upregulated expression of these identified antioxidant enzymes in this study suggests that they may detoxify ROS-induced lipid peroxidation products and contribute to the survival of the blades of P. haitanensis under high-temperature stress.
Proteins involved in responding to stimuli
High temperature leads to increased expression of several proteins that function as chaperones, particularly several members of the large family of heat shock proteins (HSPs), which are classified into five distinct subfamilies according to their molecular weights (HSP110, HSP90, HSP70, HSP60, and sHSPs) (Baniwal et al. 2004). In the transcriptome sequencing of gametophyte blades of Pyropia tenera, Choi et al. (2012) identified several high molecular weight HSPs whose expression was upregulated significantly under high-temperature stress for 1 h or 3 h, but no small HSPs (sHSPs) were identified. By contrast, the results of the present study show that high temperature increased the abundance of low molecular weight sHSPs (spot 19), but had little effect on the expression of high molecular weight HSPs. The difference may suggest that the changes in transcription do not often correspond with the changes of protein expression or that the expression of high molecular weight HSPs only increased significantly for a short time during high-temperature stress, but did not change as high-temperature stress continued for several days. Nevertheless, further studies should be conducted to clarify this point.
The chaperone activity of sHSPs, which function as a reservoir of intermediates of denatured proteins that prevent protein aggregation caused by stress, is usually undetectable in vegetative tissue under normal growth conditions. However, they can be induced by developmental stimuli or by environmental stresses (Sun et al. 2002). The upregulation of sHSPs in the present study suggests that they play an important role in counteracting high-temperature stress by reestablishing normal protein conformations and, thus, cellular homeostasis. Consistent with these findings, Lee et al. (2007) reported that several sHSPs are expressed in rice leaves during heat stress. Further, the overexpression of a tomato mitochondrial sHSP increased the thermotolerance of transgenic tobacco plants (Sanmiya et al. 2004).
Proteins involved in transcription and translation
Transcriptional and translational control of the expression of stress-responsive genes is a crucial part of the response of plants to abiotic stress. The elongation factor Tu (EF-Tu, spots 37 and 88) is a member of a highly conserved, nuclear-encoded multigene family (Momcilovic and Ristic 2007). During polypeptide synthesis, EF-Tu plays an important role in promoting the binding of aminoacyl-tRNA to the A site of the ribosome (Momcilovic and Ristic 2007). Moreover, EF-Tu acts as a molecular chaperone and protects chloroplast proteins from thermal aggregation and inactivation (Rao et al. 2004). In maize, Momcilovic and Ristic (2007) suggested that the expressions of EF-Tu genes correlate with the ability to tolerate stress. Elongation factor 1β (EF-1β, spot 87) is a member of the guanine nucleotide exchange factors family, which regulate the activity of G proteins and are critical for many cellular processes, including the stress response (Ejiri 2002). Transcription factor BTF3 (spot 113) is required for transcriptional initiation by RNA polymerase II. BTF3 is upregulated by drought in rice (Gorantla et al. 2005) and maize (Hu et al. 2011). In the present study, the upregulated expression of transcriptional and translational regulatory proteins suggested that they may play important roles in ensuring continued protein synthesis during high-temperature stress.
Cytoskeleton-related proteins
Abiotic stress can also induce profound changes in the composition of the cytoskeleton (Abdrakhamanova et al. 2003). Here, the upregulation of the three cytoskeleton-related proteins (spot 16, 103, and 125) in response to high-temperature stress indicated that the dynamics of these proteins function in cellular homeostasis of P. haitanensis during high-temperature stress.
Proteins that mediate signal transduction
Under stress, plants usually begin responding by sensing and transferring stress signals through signal transduction networks. The response to stress is likely mediated through signaling pathways that regulate the expression levels of a range of proteins (Hirayama and Shinozaki 2010). One protein (spot 69) involved in signal transduction was identified here. The ADP-ribosylation factor (ARF) GTPase-activating proteins (GAPs) represent a family of proteins that induce hydrolysis of GTP bound to ARF and act as a modulator or transducer in various transmembrane signaling systems (Randazzo and Hirsch 2004). The upregulation of ARF-GAP in response to high-temperature stress reflects the role of this protein in cell signaling under stress.
In conclusion, the comparative proteomic analysis presented identified a significant number of stress-responsive proteins from P. haitanensis blades placed under high-temperature stress. Based on the dynamic changes in the levels of expression of these proteins, we conclude that the blades of Z-61 resist high-temperature stress by inhibiting photosynthesis and other unnecessary metabolic processes. Energy metabolism is increased, and upregulation of the expression of proteins related to redox homeostasis, as well as those that respond to stimuli, serves to maintain the physiological balance during stress. Concurrently, the expression levels of proteins related to transcription and translation, the cytoskeleton, and signal transduction are also upregulated to promote survival under high-temperature stress.
References
Abdrakhamanova A, Wang QY, Khokhlova L, Nick P (2003) Is microtubule disassembly a trigger for cold acclimation? Plant Cell Physiol 44:676–686
Asada K (1992) Ascorbate peroxidase—a hydrogen peroxide-scavenging enzyme in plants. Physiol Plant 85:235–241
Baniwal SK, Bharti K, Chan KY, Fauth M, Ganguli A, Kotak S, Mishra SK, Nover L, Port M, Scharf KD, Tripp J, Weber C, Zielinski L, Koskuli-Döring PV (2004) Heat stress response in plants: a complex game with chaperones and more than twenty heat stress transcription factors. J Biosci 29:471–487
Blouin NA, Brodie JA, Grossma AC, Xu P, Brawley SH (2010) Porphyra: a marine crop shaped by stress. Trends Plant Sci 16:29–37
Chen CS, Ji DH, Xie CT, Xu Y, Liang Y, Zhen YJ, Shi XZ, Wang FX, Zhao LM (2008) Preliminary study on selecting the high temperature resistance strains and economic traits of Porphyra haitanensis. Acta Oceanol Sin 30:100–106 (in Chinese with English abstract)
Choi S, Hwang MS, Im S, Kim N, Jeong WJ, Park EJ, Gong YG, Choi DW (2012) Transcriptome sequencing and comparative analysis of the gametophyte thalli of Pyropia tenera under normal and high temperature conditions. J Appl Phycol 24:79–87
Contreras L, Moenne A, Gaillard F, Potin P, Correa JA (2010) Proteomic analysis and identification of copper stress-regulated proteins in the marine alga Scytosiphon gracilis (Phaeophyceae). Aquat Toxicol 96:85–89
Cramer GR, Urano K, Delrot S, Pezzotti M, Shinozaki K (2011) Effects of abiotic stress on plants: a systems biology perspective. BMC Plant Biol 11:163
Dixon DP, Davis BG, Edwards R (2002) Functional divergence in the glutathione transferase superfamily in plants. Identification of two classes with putative functions in redox homeostasis in Arabidopsis thaliana. J Biol Chem 277:30859–30869
Ejiri S (2002) Moonlighting functions of polypeptide elongation factor 1, from actin bundling to zinc finger protein R1-associated nuclear localization. Biosci Biotechnol Biochem 66:1–21
Ferreira S, Hjerno K, Larsen M, Wingsle G, Larsen P, Fey S, Roepsrorff P, Pais MS (2006) Proteome profiling of Populus euphratica Oliv. upon heat stress. Ann Bot (Lond) 98:361–377
Ghazalpour A, Bennett B, Petyuk VA, Orozco L, Hagopian R, Mungrue IN, Farber CR, Sinsheimer J, Kang HM, Furlotte N, Park CC, Wen PZ, Brewer H, Weitz K, Camp DG, Pan C, Yordanova R, Neuhaus I, Tilford C, Siemers N, Gargalovic P, Eskin E, Kirchgessner T, Smith DJ, Smith RD, Lusis AJ (2011) Comparative analysis of proteome and transcriptome variation in mouse. PLoS Genet 7:e1001393
Gorantla M, Babu PR, Lachagari VBR, Feltus FA, Paterson AH, Reddy AR (2005) Functional genomics of drought-stress response in rice: transcript mapping of annotated unigenes of an indica rice (Oryza sativa L. cv. Nagina 22). Curr Sci 289:496–514
Gygi SP, Rochon Y, Franza BR, Aebersold R (1999) Correlation between protein and mRNA abundance in yeast. Mol Cell Biol 19:1720–1730
Hack CJ (2004) Integrated transcriptome and proteome data: the challenges ahead. Brief Funct Genomic Proteomic 3:212–219
Hang N, Xie CT, Chen CS, Ji DH, Xu Y (2011) Comparison of protein preparation methods of Porphyra haitanensis thalli for two-dimensional electrophoresis. J Fish China 35:1362–1368 (in Chinese with English abstract)
Hegedus A, Janda T, Horvath GV, Dudits D (2008) Accumulation of overproduced ferritin in the chloroplast provides protection against photoinhibition induced by low temperature in tobacco plants. J Plant Physiol 165:1647–1651
Hirayama T, Shinozaki K (2010) Research on plant abiotic stress responses in the post-genome era: past, present and future. Plant J 61:1041–1052
Hu XL, Lu MH, Li CH, Liu TX, Wang W, Wu JY, Tai FJ, Li X, Zhang J (2011) Differential expression of proteins in maize roots in response to abscisic acid and drought. Acta Physiol Plant 33:2437–2446
Jordan DB, Ogren WL (1984) The CO2/O2 specificity of ribulose 1,5-bisphosphate carboxylase/oxygenase. Dependence on ribulose bisphosphate concentration, pH and temperature. Planta 161:308–313
Kim EY, Kim DG, Kim YR, Hwang HJ, Nam TJ, Kong IS (2011) An improved method of protein isolation and proteome analysis with Saccharina japonica (Laminariales) incubated under different pH conditions. J Appl Phycol 23:123–130
Kosová K, Vítámvás P, Prášil IT, Renaut J (2011) Plant proteome changes under abiotic stress-contribution of proteomics studies to understanding plant stress response. J Proteomics 74:1301–1322
Kumari S, Sabharwal VP, Kushwaha HR, Sopory SK, Singla-Pareek SL, Pareek A (2008) Transcriptome map for seedling stage specific salinity stress response indicates a specific set of genes as candidate for saline tolerance in Oryza sativa L. Func Integr Genomics 9:109–123
Kushwaha HR, Singh AK, Sopory SK, Singla-Pareek SL, Pareek A (2009) Genome wide expression analysis of CBS domain containing proteins in Arabidopsis thaliana (L.) Heynh and Oryza sativa L. reveals their developmental and stress regulation. BMC Genomics 10:200
Lee DG, Ahsan N, Lee SH, Kang KY, Bahk JD, Lee IJ, Lee BH (2007) A proteomic approach in analyzing heat-responsive proteins in rice leaves. Proteomics 7:3369–3383
Lin SK, Chang MC, Tsai YG, Lur HS (2005) Proteomic analysis of the expression of proteins related to rice quality during caryopsis development and the effect of high temperature on expression. Proteomics 5:2140–2156
Liu LN, Chen XL, Zhang YZ, Zhou BC (2005) Characterization, structure and function of linker polypeptides in phycobilisomes of cyanobacteria and red algae: an overview. Biochim Biophys Acta 1708:133–142
Majoul T, Bancel E, Triboi E, Ben Hamida J, Branlard G (2003) Proteomic analysis of the effect of heat stress on hexaploid wheat grain: characterization of heat-responsive proteins from total endosperm. Proteomics 3:175–183
Majoul T, Bancel E, Triboi E, Ben Hamida J, Branlard G (2004) Proteomic analysis of the effect of heat stress on hexaploid wheat grain: characterization of heat-responsive proteins from non-prolamins fraction. Proteomics 4:505–513
Mittler R, Vanderauwera S, Gollery M, Van Breusegem F (2004) Reactive oxygen gene network of plants. Trends Plant Sci 9:490–498
Momcilovic I, Ristic Z (2007) Expression of chloroplast protein synthesis elongation factor, EF-Tu, in two lines of maize with contrasting tolerance to heat stress during early stages of plant development. J Plant Physiol 164:90–99
Moons A (2003) Osgstu3 and osgtu4, encoding tau class glutathione S-transferases, are heavy metal- and hypoxic stress induced and differentially salt stress-responsive in rice roots. FEBS Lett 553:427–432
Mustafiz A, Sahoo KK, Singla-Pareek SL, Sopory SK (2010) Metabolic engineering of glyoxalase pathway for enhancing stress tolerance in plants. Methods Mol Biol 639:95–118
Nagai K, Yotsukura N, Ikegami H, Kimura H, Morimoto K (2008) Protein extraction for two-dimensional electrophoresis from the lamina of Ecklonia kurome (Laminariales), a recalcitrant tissue containing high levels of viscous polysaccharides. Electrophoresis 29:672–681
Neilson KA, Gammulla CG, Mirzaei M, Imin N, Haynes PA (2010) Proteomic analysis of temperature stress in plants. Proteomics 10:828–845
Palmblad M, Mills DJ, Bindschedler LV (2008) Heat-shock response in Arabidopsis thaliana explored by multiplexed quantitative proteomics using differential metabolic labeling. J Proteome Res 7:780–785
Qian W, Yu C, Qin H, Liu X, Zhang A, Johansen IE, Wang D (2007) Molecular and functional analysis of phosphomannomutase (PMM) from higher plants and genetic evidence for the involvement of PMM in ascorbic acid biosynthesis in Arabidopsis and Nicotiana benthamiana. Plant J 49:399–413
Randazzo PA, Hirsch DS (2004) Arf GAPs: multifunctional proteins that regulate membrane traffic and actin remodeling. Cell Signal 16:401–413
Rao D, Momcilovic I, Kobayashi S, Callegari E, Ristic Z (2004) Chaperone activity of recombinant maize chloroplast protein synthesis elongation factor, EF-Tu. Eur J Biochem 271:3684–3692
Sahoo D, Tang XR, Yarish C (2002) Porphyra—the economic seaweed as a new experimental system. Curr Sci India 83:1313–1316
Salvucci ME, Crafts-Brandner SJ (2004) Inhibition of photosynthesis by heat stress: the activation state of Rubisco as a limiting factor in photosynthesis. Physiol Plant 120:179–186
Samsonoff WA, MacColl R (2001) Biliproteins and phycobilisomes from cyanobacteria and red algae at the extremes of habitat. Arch Microbiol 176:400–405
Sanmiya K, Suzuki K, Egawa Y, Shono M (2004) Mitochondrial small heat-shock protein enhances thermotolerance in tobacco plants. FEBS Lett 557:265–268
Schreiber U, Bilger W, Neubauer C (1994) Chlorophyll fluorescence as a nonintrusive indicator for rapid assessment of in vivo photosynthesis. In: Schulze E, Caldwell M (eds) Ecophysiology of photosynthesis. Springer, Berlin, pp 49–70
Seppänen MM, Cardi T, Hyökki MB, Pehu E (2000) Characterization and expression of cold-induced glutathione S-transferase in freezing tolerant Solanum commersonii, sensitive S. tuberosum and their interspecific somatic hybrids. Plant Sci 153:125–133
Skylas DJ, Cordwell SJ, Hains PG, Larsen MR, Basseala DJ, Walsha BJ, Blumenthalb C, Rathmellb W, Copelandc L, Wrigley CW (2002) Heat shock of wheat during grain filling: proteins associated with heat tolerance. J Cereal Sci 35:175–188
Sule A, Vanrobaeys F, Hajos G, Van Beeumen J, Devreese B (2004) Proteomic analysis of small heat shock protein isoforms in barley shoots. Phytochemistry 65:1853–1863
Sun W, Van MM, Verbruggen N (2002) Small heat shock proteins and stress tolerance in plants. Biochim Biophys Acta 1577:1–9
Sweetlove LJ, Heazlewood JL, Herald V, Holtzapffel R, Day DA, Leaver CJ, Millar AH (2002) The impact of oxidative stress on Arabidopsis mitochondria. Plant J 32:891–904
Wahid A, Gelani S, Ashraf M, Foolad MR (2007) Heat tolerance in plants: an overview. Environ Exp Bot 61:199–223
Wong PF, Tan LJ, Nawi H, AbuBaker S (2006) Proteomics of the red alga, Gracilaria changii (Gracilariales, Rhodophyta). J Phycol 42:113–120
Xie CT, Chen CS, Xu Y, Ji DH (2010) Construction of a genetic linkage map for Porphyra haitanensis (Bangiales, Rhodophyta) based on sequence-related amplified polymorphism and simple sequence repeat markers. J Phycol 46:780–787
Yadav SK, Singla-Pareek SL, Reddy MK, Sopory SK (2005) Transgenic tobacco plants overexpressing glyoxalase enzymes resist an increase in methylglyoxal and maintain higher reduced glutathione levels under salinity stress. FEBS Lett 579:6265–6271
Yan JX, Wait R, Berkelman T, Harry RA, Westbrook JA, Wheeler CH, Dunn MJ (2000) A modified silver staining protocol for visualization of proteins compatible with matrix-assisted laser desorption/ionization and electrospray ionization-mass spectrometry. Electrophoresis 21:3666–3672
Yan XH, Lv F, Liu CJ, Zheng YF (2010) Selection and characterization of a high-temperature tolerant strain of Porphyra haitanensis Chang et Zheng (Bangiales, Rhodophyta). J Appl Phycol 22:511–516
Yotsukura N, Nagai K, Kimura H, Morimoto K (2010) Seasonal changes in proteomic profiles of Japanese kelp: Saccharina japonica (Laminariales, Phaeophyceae). J Appl Phycol 22:443–451
Zhang XC, Qin S, Ma JH, Xu P (2005) The genetics of marine algae. China Agriculture, Beijing
Zhang BY, Yang F, Wang GC, Peng G (2010a) Cloning and quantitative analysis of the carbonic anhydrase gene from Porphyra yezoensis. J Phycol 46:290–296
Zhang MH, Li GW, Huang W, Bi T, Chen GY, Tang ZC, Su W, Sun WN (2010b) Proteomic study of Carissa spinarum in response to combined heat and drought stress. Proteomics 10:3117–3129
Zhang BL, Yan XH, Huang LB (2011a) Evaluation of an improved strain of Porphyra yezoensis Ueda (Bangiales, Rhodophyta) with high-temperature tolerance. J Appl Phycol 23:841–847
Zhang Y, Xie CT, Chen CS, Ji DH, Zhou WW (2011b) Preliminary study on physiological response of gametophytic blades of Porphyra haitanensis to high temperature stress. J Fish China 35:379–385 (in Chinese with English abstract)
Acknowledgments
This research was supported in part by the National Natural Science Foundation of China (grant no. 41176151, 41276177), the National High Technology Research & Development Program of China (grant no. 2012AA100811), and the Fund for Distinguished Young Scientists of Fujian Province of China (grant no. 2010J06016).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xu, Y., Chen, C., Ji, D. et al. Proteomic profile analysis of Pyropia haitanensis in response to high-temperature stress. J Appl Phycol 26, 607–618 (2014). https://doi.org/10.1007/s10811-013-0066-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-013-0066-8