Abstract
Genetic variation of four populations of Sargassum thunbergii (Mert.) O. Kuntze and one outgroup of S. fusiforme (Harv.) Setchell from Shandong peninsula of China was studied with random amplified polymorphic DNA (RAPD) and inter-simple sequence repeat (ISSR) markers. A total of 28 RAPD primers and 19 ISSR primers were amplified, showing 174 loci and 125 loci, respectively. Calculation of genetic diversity with different indicators (P%, percentage of polymorphic loci; H, the expected heterozygosity; I, Shannon’s information index) revealed low or moderate levels of genetic variations within each S. thunbergii population. High genetic differentiations were determined with pairwise Nei’s unbiased genetic distance (D) and fixation index (F ST ) between the populations. The Mantel test showed that two types of matrices of D and F ST were highly correlated, whether from RAPD or ISSR data, r = 0.9310 (P = 0.008) and 0.9313 (P = 0.009) respectively. Analysis of molecular variance (AMOVA) was used to apportion the variations between and within the S. thunbergii populations. It indicated that the variations among populations were higher than those within populations, being 57.57% versus 42.43% by RAPD and 59.52% versus 40.08% by ISSR, respectively. Furthermore, the Mantel test suggested that the genetic differentiations between the four populations were related to the geographical distances (r > 0.5), i.e., they conformed to the IBD (isolation by distance) model, as expected from UPGMA (unweighted pair group method with arithmetic averages) cluster analysis. As a whole, the high genetic structuring between the four S. thunbergii populations along distant locations was clearly indicated in the RAPD and ISSR analyses (r > 0.8) in our study.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sargassum thunbergii is a common brown algal species inhabiting the coast of China, and widely distributed in warm and temperate water environments from northern Liaodong peninsula to southern Leizhou peninsula (Tseng 1983). S. thunbergii is important in maintaining the structure and function of coastal ecosystems because it is one of the dominant Sargassum species that forms seaweed beds in local intertidal zones. It can be used for alginate production and pharmacy development (Park et al. 2005), and is also an important feed source for holothurian or abalone aquaculture. Although its ecological and economic values are important, little is known about its population genetic structure. In perspectives of ecology and evolution, determining the patterns of genetic variation and population structure within one species would help in estimating the transmission of genes and such influential factors as dispersal, life cycle and habitat (Loveless and Hamrick 1984; Bohonak 1999), and in predicting the changes of long-term survival and the continued evolution of the populations and species (Sosa et al. 2002), as well as leading to a better understanding of the population dynamics and preservation of marine biodiversity (Butman and Carlton 1995).
Although allozyme analysis has been successfully applied to examine genetic structure for many organisms, it generally yielded few loci and a low level of polymorphism for seaweeds (Sosa et al. 1998; Sosa and Lindstrom 1999). In this context, a variety of molecular tools have been developed in several algal species for high-resolution analyses. Especially, the microsatellite marker, which is co-dominant in nature, has recently been used in algal population genetics studies on Fucus serratus (Coyer et al. 2003), Gracilaria gracilis (Engel et al. 2004) and G. chilensis (Guillemin et al. 2005), etc. Meanwhile, dominant markers, such as AFLP (the amplified fragment length polymorphism) and RAPD (random amplified polymorphic DNA), have also commonly been used for analyzing the genetic structure of macroalgal populations (Donaldson et al. 2000; Wright et al. 2000; Engelen et al. 2001; Bouza et al. 2006). Although dominate DNA markers cannot provide very high resolution results for population study (Hadrys et al. 1992; Van Oppen et al. 1996), they were considered advantageous in investigations of large-scale biogeographical patterns of populations hundreds to thousands of kilometers apart. ISSR (inter-simple sequence repeat) is another dominant DNA marker (Zietkiewicz et al. 1993) which has been successfully used in population genetic studies of many higher plants (Xue et al. 2004; Ge et al. 2005). However, few reports have been documented on seaweed population genetics (Hall and Vis 2002; Wang et al. 2006).
For Sargassum, most population studies have concentrated on ecology, such as seasonal pattern of population dynamics among or within species (Hwang et al. 2004; Rivera and Scrosati 2006). Engelen et al. (2001) examined the population genetic differentiation in S. polyceratium with RAPD marker, and detected significant differentiation between several bays ca. 25 km apart. Usually, geographical distance is regarded as an important factor influencing the genetic structure of populations via limiting the movements of gametes, propagules and individuals that are the media of spatial exchange of genes. Cases of genetic differentiation on various scales have been reported for many seaweed species (Sosa and Lindstrom 1999). For Sargassum, which is characterized by limited zygote dispersal capability (within 1–3 m off parental thalli) (Kendrick and Walker 1995; Stiger and Payri 1999), low genetic diversity within populations and strong differentiation among populations on a large geographical scale are expected.
In our study, genetic diversity and structure of the four natural S. thunbergii populations from Shandong peninsula were examined with RAPD and ISSR markers. Our principal objectives are to assess the level of genetic diversity and genetic structure of populations of S. thunbergii, and to explore the role of geographical distances on interpopulation genetic differentiation.
Materials and methods
Sampling
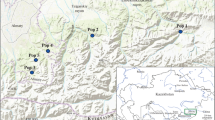
Algal materials were collected from May to July 2005 at four sites around the Shandong peninsula (Fig. 1; Table 1). At each site, over 20 different algal individuals were collected randomly, spaced at a minimum 1 m, and preserved shortly afterwards at −40°C for later DNA extraction.
DNA extraction
Three to ten leaf-like blades were plucked from a single frond of the frozen samples and washed thoroughly with distilled water. About 0.2 g of the blades was crushed and ground in liquid nitrogen. Total genomic DNA was extracted following the CTAB method using Plant Genomic DNA Extraction Kit (Tiangen Biotech, Beijing). The concentrations of the extracted DNA were tested by comparison with known quantities of λDNA/Hind III markers (Sangon, Shanghai, China) on a 0.8% agarose gel. DNA samples with 5 ∼ 20 ng μL-1 were prepared.
RAPD and ISSR amplification
A total of 200 RAPD primers and 50 ISSR primers were synthesized in the Sangon (Shanghai), of which 28 RAPD primers and 19 ISSR primers were finally selected based on polymorphism, quality and reproducibility of the amplifications (Table 2).
RAPD reactions were carried out in a 20 μL reaction mixture containing 1 μL of template DNA solution, 0.25 μM primers, 200 μM dATP, dGTP, dCTP, and dTTP, respectively (Promega, Shanghai), 2.0 mM Mg2+, 1× polymerase buffer (Promega, Shanghai) and 1.0 U Taq DNA polymerase (Promega, Shanghai). PCR amplification reaction was conducted with Master Thermal Cycler (Effendorf, German) in a procedure of initial denaturation for 5 min at 94°; 45 cycles of 1 min at 94°C, 1 min at 37°C and 2 min at 72°C; and final extension of 10 min at 72°C.
ISSR reactions were also performed in a 20-μl reaction mixture containing 1 μL of template DNA solution, 1.25 μM primers, 200 μM dNTP mix (Promega, Shanghai), 1.5 mM Mg2+, 1× polymerase buffer (Promega, Shanghai) and 1.0 U Taq DNA polymerase (Promega, Shanghai). PCR amplification reaction was conducted with Thermal Cycler (TaKaRa, Japan) in steps of initial denaturation of 94°C, 5 min and 40 cycles to denature at 94°C, 30 s; anneal at 52°C, 45 s; extend at 72°C, 2 min, then final extension of 72°C,10 min.
PCR products were separated on 1.5% agarose gels in a 1×TAE buffer system. DNA marker DL 2,000 (TaKaRa Biotech, Dalian) was used as a size marker. PCR products were recorded with a digital imager (Bio-rad, USA) after staining with ethidium bromide (0.5 μg mL-1).
Data analysis
The loci revealed by RAPD and ISSR analyses were regarded as phenotypes to estimate genotype information. The amplified DNA polymorphic fragments of each locus were scored as binary present (1) or absent (0), excluding smeared and weak bands, then the data matrices of the RAPD and ISSR were assembled for further analysis.
Software POPGENE version 1.31 (Yeh et al. 1997) was used for the calculations of (1) genetic variability from the percentage of polymorphic loci (P), at the 99% criterion; (2) the average expected heterozygosity (H), assuming a Hardy-Weinberg equilibrium; and (3) Shannon’s information index of genetic diversity (I).
Analysis of molecular variance (AMOVA) was conducted to analyze the genetic diversity and genetic structure of Sargassum populations. AMOVA treats each RAPD and ISSR profile as a haplotype, from which a pairwise Euclidian distances matrix is used for analyses without pre-assumption of normality required by classical ANOVA (analysis of variance). The genetic variations within and among the four S. thunbergii populations and the fixation indices (F ST ) of total were analyzed using ARLEQUIN version 3.01(Excoffier et al. 2005). Through locus-by-locus AMOVA, F ST of each locus was calculated independently to discard possible problematic loci according to the level of significance testing. Permutation procedures (1,000 replicates) for the significance testing on variance components and F ST values were performed. The pairwise F ST between each pair of Sargassum populations was examined and displayed in a distance matrix, in which population 5 worked as outgroup.
Using TFPGA 1.3 software (Miller 1997), the Nei’s unbiased genetic distance (D) (Nei 1978) between each pair of populations was determined and a dendrogram based on it was constructed by the UPGMA (unweighted pair group method with arithmetic averages) grouping method. The population of S. fusiforme still acted as outgroup. During bootstrapping 1,000 permutations were performed to evaluate the robustness of the groupings.
The correlation between the two types of matrices of pairwise D and F ST was examined with the Mantel test (Mantel 1967) in TFPGA, as well as the two matrices obtained by RAPD and ISSR data in each type of pairwise analyses. To determine whether the genetic distance and genetic differentiation were correlated with geographical distance between population sampling sites, the Mantel tests were preformed between Nei’s unbiased genetic distance matrix, pairwise F ST matrix and geographical distance matrix respectively.
Results
RAPD amplification
A total of 174 loci ranging from 400 to 2,000 bp were scored, an average of 6.2 loci per primer. P, H and I were used to estimate genetic diversity within the S. thunbergii population (Table 3). Population 3 from Weihai exhibited the highest level of genetic diversity (P% = 54, H = 0.2153, I = 0.3138), whereas the lowest value was shown in Population 1 from Qingdao (P% = 27.6, H = 0.1094, I = 0.1602). Obviously, values of P, H and I showed a similar tendency in intrapopulation genetic diversities. Of 174 loci, 140 (80.5%) were polymorphic in all individuals of four populations. The mean genetic variations of total indicated by H and I, were 0.2729 and 0.4118 respectively.
ISSR amplification
The selected 19 ISSR primers consistently amplified a total of 125 scorable markers of which 97 (77.6%) were polymorphic for all individuals of S. thunbergii (Table 3). Each primer generated 5 to 11 loci (average 6.6). The percentage of polymorphic bands (P%) within each populations ranged from 32.0% to 57.6%, and I ranged from 0.1774 to 0.3470, correspondingly. The highest value of H was in Population 3, showing the same tendency with P and I, and consistent with that of RAPD analysis. The mean gene variations for the four populations were 0.2903 and 0.4282 by H and I, respectively (Table 3).
Cluster analysis
The Nei’s (1978) genetic distances (D) between pairs of Sargassum populations with RAPD analysis were from 0.1429 to 0.6419 (Table 4). The lowest value was between Populations 1 and 2, and the highest between Populations 1 and 5, while for ISSR analysis, pairwise D values were from 0.1589 (Populations 3 and 4) to 0.7680 (Populations 4 and 5). The highest value was between Populations 1 and 4 indicated by both RAPD and ISSR analysis when outgroup was excluded. In the Mantel test, high and significant correlation (r = 0.9715, P = 0.016) was shown between two matrices of D from RAPD and ISSR data.
The UPGMA dendrograms (Fig. 2a) based on pairwise D with RAPD data indicated that Populations 1 and 2 clustered together first, then with Populations 3 and 4 in turn, finally with the outgroup. With ISSR data (Fig. 2b), however, four S. thunbergii populations first clustered into two branches: Populations 3 and 4, and Populations 1 and 2, then clustered with outgroup. The bootstrap values for the UPGMA trees were moderate to high, from 71 to 100 (by RAPD) and from 64.9 to 100 (by ISSR). In addition, the Mantel correlation between genetic and geographical distances was moderate by RAPD (r = 0.5048, P = 0.2480) or high by ISSR (r = 0.9052, P = 0.0870).
UPGMA dendrograms using Nei’s (1978) unbiased genetic distance of Sargassum populations. a Tree built using RAPD data. b Tree built using ISSR data. Bootstrap values are indicated as percentages of 1,000 replicates
AMOVA analysis
The levels of interpopulation genetic differentiation were also examined with pairwise F ST besides pairwise D. The F ST values between each S. thunbergii population and outgroup were obviously higher than those between S. thunbergii populations (Table 5). When outgroup was excluded the pairewise F ST values were 0.4745–0.6955 by RAPD, and 0.4677–0.7004 by ISSR, and all with highly significant levels (P < 0.001). A similar tendency shown in genetic distance matrix was also found. Mantel test showed that the two types of matrices of D and F ST were highly and significantly correlated whether from RAPD or ISSR data, with r = 0.9310 (P = 0.008) and 0.9313 (P = 0.009), respectively, and two F ST matrices obtained by RAPD and ISSR were also highly correlated (r = 0.8737, P = 0.0520). In addition, for the four S. thunbergii populations, the pairwise F ST values were also positively related to the corresponding geographical distances, with r = 0.7217 (P = 0.1660) by RAPD and r = 0.7277 (P = 0.0920) by ISSR respectively.
AMOVA with the RAPD data indicated that the genetic variances between populations accounted for 57.57% of the total, while the rest 42.43% were within populations (Table 6). Similarly, with the ISSR data, 59.52% was ascribed to between populations, and 40.48% within populations. Significance tests of variance components among and within populations showed significant difference with P < 0.001 (Table 6).
Discussion
On the whole, P, H and I revealed a low or moderate level of genetic diversity within populations with RAPD and ISSR data (Table 3). It was not suitable enough to calculate allele frequencies due to the dominant nature of RAPD and ISSR markers that only allow estimation of genotypes. However, H, based on allele frequency, was still selected as a parameter to assess genetic variations within populations for higher plants or seaweeds, when the dominant markers were used for population genetic studies (Faugeron et al. 2001; Xue et al. 2004; Bouza et al. 2006). In this study, for S. thunbergii, we also used H to measure the genetic diversities within populations in conjunction with I, the Shannon’s information index, which is based on band phenotypes (i.e. band present/absent) without depending on assumption of Hardy-Weinberg equilibrium. It was clear that the similar level of intrapopulation genetic diversity was revealed with three different indicators P, H and I, even by different calculations (Table 3). The results were consistent with those reported previously for other species using RAPD or ISSR marker (Li and Xia 2005; Wei et al. 2005; Bouza et al. 2006).
The mean F ST values for the four S. thunbergii populations were 0.5757 by RAPD and 0.5952 by ISSR, respectively (Table 6), indicating that most of the total genetic variances (57.57% or 59.52%) were at the interpopulation level. According to Wright (1978), F ST exceeding 0.15 would indicate high genetic structuring. Here, among the four S. thunbergii populations, the F ST indicated a high level of genetic differentiation and suggested a low level of gene flow between the populations. The limited gene flow (genetic isolation) probably resulted from the large geographical distances that limited the long-distance dispersals of spores, gametes, or drifting reproductive fragments. In fucoids, zygote dispersal takes place within only a few meters even where there is relatively strong turbulence (Kendrick and Walker 1995; Stiger and Payri 1999), and the gamete is generally short-lived, only for a few days. Although long-distance dispersals of drifting fertile (zygote-bearing) thalli have been observed for some species of Sargassum (Deysher and Norton 1982), surveys on the distribution and composition of floating seaweeds showed that S. thunbergii was rare in offshore masses (Ohno 1984). Hence, long-distance dispersal of sperms, zygotes or drifting fertile thalli is quite rare for S. thunbergii, and the large spacing (>120 km) between two sample sites would be large enough to form a geographical barrier for gene flow of S. thunbergii across the four populations.
High genetic variations between pairs of Sargassum population were confirmed with Nei’s (1978) unbiased genetic distances matrix (Table 4) and fixation indices (F ST ) matrix (Table 5), although the two measures have different definitions and assumptions for analyzing the interpopulation differentiation. Nei’s unbiased genetic distances were based on allele frequency with an assumption of Hardy-Weinberg equilibrium. In contrast, F ST values were calculated with AMOVA that treated the dominant markers profiles as haplotypes under the assumption of linkage equilibrium. Significant and high correlations shown by the Mantel test between the two types of matrices of D and F ST from RAPD (r = 0.9310, P = 0.008) and ISSR (r = 0.9313, P= 0.009) data can be taken as the evidence for the consistency of the results. By examining the two types of matrices, it was found that the outgroup was separated from any other populations with the largest genetic distances, and this can easily be seen from the UPGMA dendrograms based on Nei’s unbiased genetic distances, in which the outgroup (Population 5) formed a clade of its own far from the other S. thunbergii populations with 100 bootstrap values (Fig. 2a,b). Therefore, it was valid to select S. fusiformis (Population 5) as the outgroup in our study.
The genetic differentiation by distance was assessed with the Mantel test between genetic distances matrices of D or F ST and graphical distances matrix, showing a moderate to high correlation. For example, with ISSR data, r = 0.9052, P = 0.0870 for the matrix of D and r = 0.7277, P = 0.0920 for the matrix of F ST , which indicated that genetic differentiation between populations agreed with the IBD (isolation by distance) model proposed by Wright (1946). Namely, in general, the greater the separation caused by distance or barrier between two populations, the higher would be the genetic differentiation between them. Our interpopulation grouping in the dendrograms supported this model (Fig. 2a,b), as the populations that are close in space clustered easily and vice versa. The high genetic differentiation between two geographically separated S.thunbergii populations indicated once again that geographical distance was an important factor for genetic divergence, and genetic drift might be the primary evolutionary force during the process.
In conclusion, with RAPD and ISSR markers, low levels of genetic diversity within populations and high levels of genetic differentiation between four populations of S. thunbergii have been revealed, which supports the traditional IBD model: the geological distances among populations would explain the limited gene flow and high genetic structuring among the detected populations of S. thunbergii from Shandong peninsula.
References
Bohonak AJ (1999) Dispersal, gene flow and population structure. Q Rev Biol 74:21–45
Bouza N, Caujapé-Castells J, González-Pérez MÁ, Sosa PA (2006) Genetic structure of natural populations in the red algae Gelidium canariense (Gelidiales, Rhodophyta) investigated by random amplified polymorphic DNA (RAPD) markers. J Phycol 42:304–311
Butman CA, Carlton JT (1995) Understanding marine biodiversity. National Academic Press, Washington, D.C.
Coyer JA, Peters AF, Stam WT, Olsen JL (2003) Post-ice age reclonization and differentiation of Fucus serratus L. (Phaeophyceae; Fucaceae) populations in Northern Europe. Mol Ecol 12:1817–1829
Deysher L, Norton TA (1982) Dispersal and colonization in Sargassum muticum (Yendo) Fensholt. J Exp Mar Biol Ecol 56:179–195
Donaldson SL, Chopin T, Saunders G (2000) An assement of the AFLP method for investigating population structure in the red alga Chondrus crispus Stackhouse (Gigartinales, Florideophyceae). J Appl Phycol 12:25–35
Engel CR, Destombe C, Valero M (2004) Mating system and gene flow in the red seaweed Gracilaria gracilis: effect of haploid-diploid life history and intertidal rocky shore landscape on fine-scale genetic structure. Heredity 92:289–298
Engelen AH, Olsen JL, Breeman AM, Stam WT (2001) Genetic differentiation in Sargassum polyceratium (Fucales: Phaeophyceae) around the island of Curacao (Netherlands Antilles). Mar Biol 139:267–277
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Faugeron S, Valero M, Destombe C, Martínez EA, Correa JA (2001) Hierarchical spatial structure and discriminant analysis of genetic diversity in the red alga Mazzaella laminarioides. J Phycol 37:705–716
Ge JP, Cai B, Ping WX, Song G, Ling H, Lin P (2005) Mating system and population genetic structure of Bruguiera gymnorrhiza (Rhizophoraceae), a viviparous mangrove species in China. J Exp Mar Biol Ecol 326:48–55
Guillemin ML, Destombe C, Faugeron S, Correa JA, Valero M (2005) Development of microsatellites DNA markers in the cultivated seaweed, Gracilaria chilensis (Gracilariales, Rhodophyta). Mol Ecol Notes 5:155–157
Hadrys H, Balick M, Schierwater B (1992) Applications of random amplified polymorphic DNA (RAPD) in molecular ecology. Mol Ecol 1:55–63
Hall MM, Vis ML (2002) Genetic variation in Batrachospermum helminthosum (Batrachospermales, Rhodophyta) among and within stream reaches using intersimple sequence repeat molecular markers. Phycol Res 50:155–162
Hwang RL, Tsai CC, Lee TM (2004) Assessment of temperature and nutrient limitation on seasonal dynamics among species of Sargassum from coral reef in southern Taiwan. J Phycol 40:463–473
Kendrick GA, Walker DI (1995) Dispersl of propagules of Sargassum spp. (Sargassaceae, Phaeophyta): observation of local patterns of dispersal and consequences for recruitment and population structure. J Exp Mar Biol Ecol 192:273–288
Loveless MD, Hamrick JL (1984) Ecological determinants of genetic structure in plant populations. Ann Rev Ecolog Syst 15:65–95
Li F, Xia N (2005) Population structure and genetic diversity of an endangered species, Glytostrobus pensilis (Cupressaceae). Bot Bull Acad Sin 46:155–162
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Miller MP (1997) Tools for population genetic analyses (TFPGA) 1.3: a Windows program for the analysis of allozyme and molecular population genetic data. Computer software distributed by author
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Ohno M (1984) Observation on the floating seaweeds of near-shore waters of southern Japan. Hydrobiologia 116/117:408–412
Park PJ, Heo SJ, Park EJ et al (2005) Reactive oxygen scavenging effect of enzymatic extracts from Sargassum thunbergii. J Agric Food Chem 53:6666–6672
Rivera M, Scrosati R (2006) Population dynamics of Sargassum lapazeanum (Fucales, Phaeophyta) from the Gulf of California, Mexico. Phycologia 45:178–189
Sosa PA, Bouza N, Cabrera-Pérez MA, Luque A (1998). Genetic variation in Phoenix canariensis and P. dactylifera (Arecaceae) populations of Gran Canaria using isoezyme electroforesis. Bol Mus Mun Funchal 5: 443
Sosa PA, Lindstrom SC (1999) Isozymes in microalgae (seaweeds): genetic differentiation, genetic variability and applications in systematics. Eur J Phycol 34:427–442
Sosa PA, González-Pérez MA, Batista F, Bouza N (2002) Conservación genética de especies vegetales amenazadas. In: Bañares-Baudet A (ed) Biología de la conservación de plantas amenazadas. Organismo Autónomo de Parques Nacionales, Madrid, Spain, pp 133–160
Stiger V, Payri CE (1999) Spatial and temporal patterns of settlement of the brown macroalga Turbinaria ornate and Sargassum mangarevense in a coral reef on Tahiti. Mar Ecol Prog Ser 19:91–100
Tseng CK (1983) Common seaseeds of China. Science Press, Beijing
Van Oppen JH, Klerk H, De Graaf M, Stam WT, Olsen JL (1996) Assessing the limits of Random Amplified Polymorphic DNAs (RAPDs) in seaweed biogeography. J Phycol 32:433–444
Wang XL, Wang D, Li DP, Duan DL (2006) Genetic analysis of the gametophytes of Undaria pinnatifida (Phaeophyceae) with ISSR method. Aquaculture 258:250–256
Wei X, Wei JQ, Cao HL, Li F, Ye WH (2005) Genetic diversity and differentiation of Camellia euphlebia (Theaceae) in Guangxi, China. Ann Bot Fenn 42:365–370
Wright S (1946) Isolation by distance under diverse systems of mating. Genetics 31:39–59
Wright S (1978) Evolution and genetics of populations. University of Chicago Press, Chicago
Wright JT, Zuccarello GC, Steinberg PD (2000) Genetic structure of the subtidal red alga Delisea pulchra. Mar Biol 136:439–448
Xue DW, Ge XJ, Hao G, Zhang CQ (2004) High genetic diversity in a rare, narrowly endemic primrose species: Primula interjacens by ISSR analysis. Acta Bot Sin 46:1163–1169
Yeh FC, Yang RC, Boyle T, Ye ZH, Mao JX (1997) POPGENE, the user-friendly shareware for population genetic analysis. Molecular Biology and Biotechnology Center, University of Alberta, Edmonton, Alberta, Canada
Zietkiewicz E, Rafalski A, Labuda D (1993) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20:176–183
Acknowledgements
The authors gratefully acknowledge NSFC (40618001) and the Shandong Agriculture Seedstock Breeding Project for financial assistance and anonymous reviewers for comments. Thanks are also kindly given to Mr. Roger Yu for English revision, to Mr. J. T. Yao and Z. M. Hu for the assistance in sample collection, and to Miss Y. X. Liu for her help in figure drawing.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhao, F., Wang, X., Liu, J. et al. Population genetic structure of Sargassum thunbergii (Fucales, Phaeophyta) detected by RAPD and ISSR markers. J Appl Phycol 19, 409–416 (2007). https://doi.org/10.1007/s10811-006-9147-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-006-9147-2