Abstract
Systemic lupus erythematosus (SLE) is a multifactorial autoimmune disease with complex genetic inheritance that affecting different organs and systems. STAT4 has been newly identified as a susceptible gene in the development of SLE. According to recent studies, STAT4 has been associated with SLE in various populations. We investigated whether STAT4 single nucleotide polymorphisms (SNPs) were associated with susceptibility and clinical features of SLE in Iranian patients. The study group comprised 280 patients with SLE and 281 sex-, age-, and ethnicity-matched healthy controls of Iranian ancestry. Two SNPs (rs7574865 and rs7601754) were genotyped using the TaqMan MGB Allelic Discrimination method. Our results showed a significant association betweenrs7574865 T allele (odds ratio (OR) = 1.50, 95 % CI = 1.18–1.92, P = 0.002) and susceptibility to SLE. The rs7574865TT genotype (P = 0.02, OR = 1.94, 95 % CI = 1.74–3.19) and GT genotype (P = 0.008, OR = 1.71, 95 % CI = 1.19–2.45) showed a significant association with the risk of SLE in the Iranian population. We concluded that STAT4 rs7574865 is associated with SLE susceptibility in the Iranian population and this SNP might be a factor in the pathogenesis of SLE. However, further studies are required to investigate the mechanism by which polymorphisms in this gene lead to SLE.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Systemic lupus erythematosus (SLE) is a chronic inflammatory autoimmune disease that can affect different tissues including the skin, joints, blood, kidney, and nervous system [1]. The American College of Rheumatology established 11 clinical criteria such as arthritis, renal disorders, mucocutaneous involvement, photosensitivity, neurological disorders, production of variety of autoantibodies, and hematological disorders. SLE is characterized by the presence of any 4 of these 11 criteria [2, 3]. The SLE prevalence in menopause women is nine times more than men [4].
The etiology of SLE involves both genetic and environmental factors [5]. The heritability of SLE among siblings (>66 %) and the rates of concordance (>35 %) in monozygotic twins compared with (2–5 %) dizygotic twins document the importance of genetic factors in SLE [6, 7]. In various races, many genes can be implicated in SLE and these genes may have different effect size and correlate with subtypes of this disease. According to recent studies, SLE is characterized as a polygenic genetic model that may involve as many as 100 genes, and every gene has only a moderate effect size [8].
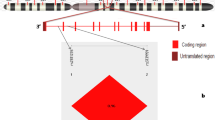
According to recent genome-wide association studies, STAT4 is a novel predisposing gene for rheumatoid arthritis and SLE [9]. The STAT4 gene is located on human chromosome 2q32.3 and includes of 24 exons spanning a 120-kb region. This gene encodes a transcription factor that can be activated by interleukin (IL)-12 and IL-23 and plays a central role in the signaling by interferon (IFN) type I receptor and Th1 differentiation [10–14]. STAT4 is also involved in IL-23 signaling and development of TH17 [15]. TH1 and TH17 are broadly implicated in several inflammatory and autoimmune diseases [16]. STAT4 seems to be a critical signaling molecule in both subsets of T cells. Therefore, STAT4 may also be an important factor in the pathogenesis of SLE [15]. Some SLE phenotypes have been associated with STAT4 polymorphisms, such as antidouble-stranded DNA (anti-dsDNA), autoantibodies and renal disorder [17]. rs7582694 shows complete linkage disequilibrium with rs7574865 in Asian and Caucasian populations presented in HapMap data (r 2 = 1) [18] (http://hapmap.ncbi.nlm.nih.gov/). These two SNPs have been identified as risk loci for SLE by several studies [18, 19].
In the present study, we analyzed the association between STAT4 polymorphisms (rs7574865 and rs7601754) and SLE disease in an Iranian population. We also investigated the association of these polymorphisms with different phenotypes of patients with SLE.
MATERIALS AND METHODS
Patients and Controls
The study population included 280 unrelated patients recruited from the outpatient rheumatology clinic of Shariati Hospital from May 2011 to May 2012. All SLE patients fulfilled the revised 1982 American College of Rheumatology criteria for SLE [20]. The patient group had a mean age of 36.77 ± 11.72 years and included 244 females and 36 males. The mean age of 281 healthy controls was 38.04 ± 12.71 years (246 females and 35 males) with no clinical evidence or family history of any type of autoimmune disorders. Informed consent was obtained from all the study participants. The Ethics Committee of Tehran University of Medical Sciences approved this study. Healthy controls were gender-, ethnicity-, and age-matched with the patients. In order to investigate the association between two SNP variations and SLE phenotypes, the clinical manifestations of the patients were recorded.
Genotyping (DNA Preparation and Analysis)
Genomic DNA was extracted from peripheral blood leukocytes using the phenol–chloroform method [21]. The extracted DNA was stored at −20 °C until analyzed. Approximately 20 ng of the genomic DNA was used to genotype each sample. Genotyping of STAT4 (rs7574865 and rs7601754) was performed using the MGB TaqMan Allelic Discrimination method (Applied Biosystems, Foster City, CA, USA). Patient and control samples were genotyped for rs7574865 and rs7601754 using StepOnePlus Real-Time PCR System (Applied Biosystems). PCR was performed according to the manufacturer's protocols provided by Applied Biosystems.
Statistical Analysis
The distributions of rs7574865 and rs7601754 genotypes were tested for deviation from Hardy–Weinberg in controls. Genotypic and allelic distribution between patients and controls were assessed by chi-squares test. The odds ratio (OR) and 95 % confidence intervals were calculated. The association of two SNPs with clinical manifestations was determined by χ 2 test. To adjust for multiple comparisons, we used the Benjamini–Hochberg method to control for the false discovery rate (FDR) [22]. Analyses of data were carried out using the software SPSS for Windows (version 19.0, IBM SPSS Inc., USA).
RESULTS
We genotyped two SNPs (rs7574865 and rs7601754) in 280 SLE patients and 281 healthy controls. The distribution of rs7574865 and rs7601754 genotypes did not show any significant deviation from Hardy–Weinberg equilibrium between patients and healthy controls. The rs7574865 T allele in the patients was significantly higher than the control group (OR = 1.50, 95 % CI = 1.18–1.92, P = 0.002). The rs7574865 GT heterozygous genotype frequency was higher in patients than controls and amounted 46.8 and 38.1 %, respectively. According to Table 1, the TT genotype (OR = 1.94, 95 % CI = 1.74–3.19, P = 0.02) and GT genotype (OR = 1.71, 95 % CI = 1.19–2.45, P = 0.008) of the STAT4 rs7574865 were significantly associated with an increased risk of SLE when GG was taken as a reference.
We next investigated whether these two genotyped SNPs were correlated with specific clinical manifestations of SLE such as photosensitivity, malar rash, discoid rash, oral ulcer, arthritis, pleuritis, pericarditis, proteinuria, seizures, leucopenia, anti-ds DNA, and ANA. After FDR test, we did not detect any correlation between these SNPs and the mentioned clinical manifestations (Table 2).
DISCUSSION
Systemic lupus erythematosus is one of the most heterogeneous complex autoimmune diseases [23]. The severity and clinical manifestations of SLE differ by sex, race, and geography [17]. The prevalence of SLE in African-American, Hispanic, and Asian populations is more than other populations [8]. In current years, several specific genes have been reported to be associated with susceptibility and pathogenesis of SLE in some populations in the world [24–26]. The SNPs rs7574865 and rs7601754 located in intron 4 of STAT4 were shown to be strongly associated with SLE [26, 27]. This study produced results, which corroborate the findings of a great deal of the previous work in this field.
In the SNP database (http://www.ncbi.nlm.nih.gov/snp/), the allele frequencies of G and T alleles of rs7574865 are 0.75 and 0.25, respectively (these data are based on 1,000 genomes project). According to our results, the frequencies of G and T alleles in the control group were 0.68 and 0.32, respectively, that is approximately similar to the SNP database. However, the frequencies of G and T alleles in our SLE patients were 0.59 and 0.41, respectively, which were significantly higher than that of the control group and the SNP database. A and G allele frequencies of rs7601754 in SNP database were 0.69 and 0.31, respectively, and these allele frequencies in our healthy control and SLE patient groups were approximately similar from the SNP database (Table 1).
The results of present study indicated that STAT4 rs7574865 GG and TT genotypes were significantly associated with the risk of SLE in the Iranian population. The present findings seem to be consistent with other studies, which have reported that rs7574865 is a risk factor for SLE. This SNP has been associated with SLE in Caucasians, southern Han Chinese (Hong Kong), Thai, and Japanese populations by multiple association and replication studies [15, 19, 23, 26–32]. However, these findings are not consistent with those of Zervou et al. [33] who found rs7574865 was not associated with SLE in the patients from Turkey [33]. In this study, we did not find any significant association between STAT4 (rs7601754) polymorphism and the SLE risk. Although this result differs from the study of Yang et al. in Chinese population in Hong Kong [27], it is inconsistent with those of Sigurdsson et al. and Kawasaki et al., who found that rs7601754 was not associated with SLE in the Swedish and Japanese patients, respectively [18, 28]. Differences between our results and results from previous studies may be due to different sample sizes and different race/ethnic groups. It also may be because our statistical analysis was different from others.
The present study showed that minor T allele in rs7574865 was not correlated with the specific SLE clinical features. According to previous studies, minor T allele in rs7574865 was associated with SLE patients with different clinical features such as, nephritis, anti-dsDNA antibody production, early age of onset, and oral ulcers [3, 18, 26, 28]. However, these findings were not consistent with those of Yin et al. who found no significant association between STAT4 rs7574865 and the clinical features of SLE [30]. According previous studies, different results have been reported regarding the association of STAT4 polymorphisms with clinical features. Possible explanations for this finding might be genetic heterogeneity, variable clinical features of SLE, different sizes of the studied groups, or patient interaction with different environmental factors [32].
It is well documented that the expression of IFN type 1 and IFN-inducible genes are increased in mononuclear cells in SLE patients [34–36]. IFN type 1 has an important role in the pathogenesis of SLE through maturation of myeloid dendritic cells [37, 38]. This myeloid dendritic cells differentiated by IFN can activate autoreactive T and B cells [37]. STAT4 is one of the IFN-inducible genes and also activated by IL-12 [39]. STAT4 produces IFN-γ and it maybe a component of IFN type 1 signaling. Furthermore, production of IFN-γ is induced by STAT4 in B cells. It seems that it mediates by transducing by IL-12 signaling [40].
Our data confirmed STAT4 as a susceptibility gene for SLE. However, further studies are needed to investigate the exact mechanism by which polymorphisms in this gene lead to SLE disorder. More in-depth research should also be done to identify all genetic risk factors or gene–gene interactions contributing to the risk of SLE. Therefore, it is important to collaborate in SLE genetic studies in the future.
References
Budarf, Goyette, Boucher, Lian, Graham, Claudio, Hudson, Gladman, Clarke, Pope, Peschken, Smith, Hanly, Rich, Boire, Barr, Zummer, Fortin, Wither, and Rioux. 2011. A targeted association study in systemic lupus erythematosus identifies multiple susceptibility alleles. Genes and Immunity 12: 51–58.
Hopkinson, Doherty, and Powell. 1994. Clinical features and race-specific incidence/prevalence rates of systemic lupus erythematosus in a geographically complete cohort of patients. Annals of the Rheumatic Diseases 53: 675–680.
Taylor, Remmers, Lee, Ortmann, Plenge, Tian, Chung, Nititham, Hom, Kao, Demirci, Kamboh, Petri, Manzi, Kastner, Seldin, Gregersen, Behrens, and Criswell. 2008. Specificity of the STAT4 genetic association for severe disease manifestations of systemic lupus erythematosus. PLoS Genetics 4: e1000084.
Sekigawa, Naito, Hira, Mitsuishi, Ogasawara, Hashimoto, and Ogawa. 2004. Possible mechanisms of gender bias in SLE: a new hypothesis involving a comparison of SLE with atopy. Lupus 13: 217–222.
Han, Zheng, Cui, Sun, Ye, Hu, Xu, Cai, Huang, Zhao, Xie, Fang, Lu, Li, Pan, Deng, Zeng, Ye, Zhang, Wang, Hao, Ma, Zuo, Zhou, Du, Cheng, Yang, Shen, Li, Sheng, Zuo, Zhu, Gao, Zhang, Guo, Li, Gao, Xiao, Quan, Zhang, Zhang, Zhu, Li, Hu, Lu, Huang, Liu, Li, Ren, Wang, Yang, Wang, Zhou, Lv, Zhang, Zhang, Lin, Low, Shen, Zhai, Wang, Zhang, Yang, Liu, and Zhang. 2009. Genome-wide association study in a Chinese Han population identifies nine new susceptibility loci for systemic lupus erythematosus. Nature Genetics 41: 1234–1237.
Moser, Kelly, Lessard, and Harley. 2009. Recent insights into the genetic basis of systemic lupus erythematosus. Genes and Immunity 10: 373–379.
Kaiser, and Criswell. 2010. Genetics research in systemic lupus erythematosus for clinicians: methodology, progress, and controversies. Current Opinion in Rheumatology 22: 119–125.
Yuan, Luo, and Shen. 2010. Current advances in lupus genetic and genomic studies in Asia. Lupus 19: 1374–1383.
Lee, Remmers, Le, Kastner, Bae, and Gregersen. 2007. Association of STAT4 with rheumatoid arthritis in the Korean population. Molecular Medicine 13: 455–460.
Farrar, Smith, Murphy, and Murphy. 2000. Recruitment of Stat4 to the human interferon-alpha/beta receptor requires activated Stat2. The Journal of Biological Chemistry 275: 2693–2697.
Morinobu, Gadina, Strober, Visconti, Fornace, Montagna, Feldman, Nishikomori, and O'Shea. 2002. STAT4 serine phosphorylation is critical for IL-12-induced IFN-gamma production but not for cell proliferation. Proceedings of the National Academy of Sciences of the United States of America 99: 12281–12286.
Lund, Chen, Scheinin, and Lahesmaa. 2004. Early target genes of IL-12 and STAT4 signaling in Th cells. Journal of Immunology 172: 6775–6782.
Kaplan. 2005. STAT4: a critical regulator of inflammation in vivo. Immunologic Research 31: 231–242.
O'Malley, Eri, Stritesky, Mathur, Chang, Hogenesch, Srinivasan, and Kaplan. 2008. STAT4 isoforms differentially regulate Th1 cytokine production and the severity of inflammatory bowel disease. Journal of Immunology 181: 5062–5070.
Kobayashi, Ikari, Kaneko, Kochi, Yamamoto, Shimane, Nakamura, Toyama, Mochizuki, Tsukahara, Kawaguchi, Terai, Hara, Tomatsu, Yamanaka, Horiuchi, Tao, Yasutomo, Hamada, Yasui, Inoue, Itakura, Okamoto, Kamatani, and Momohara. 2008. Association of STAT4 with susceptibility to rheumatoid arthritis and systemic lupus erythematosus in the Japanese population. Arthritis and Rheumatism 58: 1940–1946.
Martinez, Varade, Marquez, Cenit, Espino, Perdigones, Santiago, Fernandez-Arquero, de la Calle, Arroyo, Mendoza, Fernandez-Gutierrez, de la Concha, and Urcelay. 2008. Association of the STAT4 gene with increased susceptibility for some immune-mediated diseases. Arthritis and Rheumatism 58: 2598–2602.
Harley, Kaufman, Langefeld, Harley, and Kelly. 2009. Genetic susceptibility to SLE: new insights from fine mapping and genome-wide association studies. Nature Reviews Genetics 10: 285–290.
Sigurdsson, Nordmark, Garnier, Grundberg, Kwan, Nilsson, Eloranta, Gunnarsson, Svenungsson, Sturfelt, Bengtsson, Jonsen, Truedsson, Rantapaa-Dahlqvist, Eriksson, Alm, Goring, Pastinen, Syvanen, and Ronnblom. 2008. A risk haplotype of STAT4 for systemic lupus erythematosus is over-expressed, correlates with anti-dsDNA and shows additive effects with two risk alleles of IRF5. Human Molecular Genetics 17: 2868–2876.
Hellquist, Sandling, Zucchelli, Koskenmies, Julkunen, D'Amato, Garnier, Syvanen, and Kere. 2010. Variation in STAT4 is associated with systemic lupus erythematosus in a Finnish family cohort. Annals of the Rheumatic Diseases 69: 883–886.
Gladman, Ginzler, Goldsmith, Fortin, Liang, Urowitz, Bacon, Bombardieri, Hanly, Hay, Isenberg, Jones, Kalunian, Maddison, Nived, Petri, Richter, Sanchez-Guerrero, Snaith, Sturfelt, Symmons, and Zoma. 1996. The development and initial validation of the Systemic Lupus International Collaborating Clinics/American College of Rheumatology damage index for systemic lupus erythematosus. Arthritis and Rheumatism 39: 363–369.
Roe, Crabtree and Khan. 1995. Methods for DNA isolation. Part III. Protocols for recombinant DNA isolation, cloning, and sequencing [Internet edition]. Norman, OK: University of Oklahoma: 2488–2498.
Benjamini, and Hochberg. 1995. Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society Series B 57: 289–300.
Yuan, Feng, Pan, Qiu, Li, Zhang, and Ye. 2010. A meta-analysis of the association of STAT4 polymorphism with systemic lupus erythematosus. Modern Rheumatology 20: 257–262.
Hochberg. 1997. Updating the American College of Rheumatology revised criteria for the classification of systemic lupus erythematosus. Arthritis and Rheumatism 40: 1725.
Remmers, Plenge, Lee, Graham, Hom, Behrens, de Bakker, Le, Lee, Batliwalla, Li, Masters, Booty, Carulli, Padyukov, Alfredsson, Klareskog, Chen, Amos, Criswell, Seldin, Kastner, and Gregersen. 2007. STAT4 and the risk of rheumatoid arthritis and systemic lupus erythematosus. The New England Journal of Medicine 357: 977–986.
Yang, Shen, Ye, Liu, Zhang, Qian, Hirankarn, Ying, Pan, Mok, Chan, Wong, Lee, Mok, Wong, Leung, Li, Avihingsanon, Wong, Lee, Ho, Lee, Chang, Li, Li, Zhang, Wong, Ng, Lau, Sham, and Lau. 2010. Genome-wide association study in Asian populations identifies variants in ETS1 and WDFY4 associated with systemic lupus erythematosus. PLoS Genetics 6: e1000841.
Yang, Ng, Zhao, Hirankarn, Lau, Mok, Chan, Wong, Lee, Mok, Wong, Avihingsanon, Lee, Ho, Lee, Wong, and Lau. 2009. Population differences in SLE susceptibility genes: STAT4 and BLK, but not PXK, are associated with systemic lupus erythematosus in Hong Kong Chinese. Genes and Immunity 10: 219–226.
Kawasaki, Ito, Hikami, Ohashi, Hayashi, Goto, Matsumoto, Ito, Tsutsumi, Koga, Arinami, Graham, Hom, Takasaki, Hashimoto, Behrens, Sumida, and Tsuchiya. 2008. Role of STAT4 polymorphisms in systemic lupus erythematosus in a Japanese population: a case–control association study of the STAT1-STAT4 region. Arthritis Research & Therapy 10: R113.
Kelley, Hughes, Malik, Danila, Edberg, Alarcon, Conn, Jonas, Callahan, Smith, Brasington, Edberg, Kimberly, Moreland, and Bridges. 2010. Genetic variants of STAT4 associated with rheumatoid arthritis in persons of Asian and European ancestry do not replicate in African Americans. Annals of the Rheumatic Diseases 69: 625–626.
Su, Zhao, Liu, Guo, Jiang, Liu, Zhang, Zheng, Li, Song, Huang, Huang, Wang, Pan, Li, Zhu, Zhang, and Li. 2010. Variation in STAT4 is associated with systemic lupus erythematosus in Chinese Northern Han population. Chinese Medical Journal 123: 3173–3177.
Li, Cao, Luan, Li, Hu, Zhang, Zeng, Zhang, Zeng, and Li. 2011. Association of genetic variations in the STAT4 and IRF7/KIAA1542 regions with systemic lupus erythematosus in a Northern Han Chinese population. Human Immunology 72: 249–255.
Piotrowski, Lianeri, Wudarski, Olesinska, and Jagodzinski. 2012. Contribution of STAT4 gene single-nucleotide polymorphism to systemic lupus erythematosus in the Polish population. Molecular Biology Reports 39: 8861–8866.
Zervou, Vazgiourakis, Yilmaz, Kontaki, Trouw, Toes, Bicakcigil, Boumpas, Yavuz, and Goulielmos. 2011. TRAF1/C5, eNOS, C1q, but not STAT4 and PTPN22 gene polymorphisms are associated with genetic susceptibility to systemic lupus erythematosus in Turkey. Human Immunology 72: 1210–1213.
Baechler, Batliwalla, Karypis, Gaffney, Ortmann, Espe, Shark, Grande, Hughes, Kapur, Gregersen, and Behrens. 2003. Interferon-inducible gene expression signature in peripheral blood cells of patients with severe lupus. Proceedings of the National Academy of Sciences of the United States of America 100: 2610–2615.
Crow. 2003. Interferon-alpha: a new target for therapy in systemic lupus erythematosus? Arthritis and Rheumatism 48: 2396–2401.
Crow. 2010. Type I interferon in organ-targeted autoimmune and inflammatory diseases. Arthritis Research & Therapy 12(Suppl 1): S5.
Banchereau, and Pascual. 2006. Type I interferon in systemic lupus erythematosus and other autoimmune diseases. Immunity 25: 383–392.
Kyogoku, and Tsuchiya. 2007. A compass that points to lupus: genetic studies on type I interferon pathway. Genes and Immunity 8: 445–455.
Watford, Hissong, Bream, Kanno, Muul, and O'Shea. 2004. Signaling by IL-12 and IL-23 and the immunoregulatory roles of STAT4. Immunology Reviews 202: 139–156.
Durali, de Goer de Herve, Giron-Michel, Azzarone, Delfraissy, and Taoufik. 2003. In human B cells, IL-12 triggers a cascade of molecular events similar to Th1 commitment. Blood 102: 4084–4089.
Acknowledgments
The authors would like to declare their gratitude to all contributors who have made the completion of this study. This work was supported by a research grant [grant number: 90-04-41-14909] from the Deputy of Research, Tehran University of Medical Sciences.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mirkazemi, S., Akbarian, M., Jamshidi, A.R. et al. Association of STAT4 rs7574865 with Susceptibility to Systemic Lupus Erythematosus in Iranian Population. Inflammation 36, 1548–1552 (2013). https://doi.org/10.1007/s10753-013-9698-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10753-013-9698-8