Abstract
In present investigation, we studied how mitochondrial–nuclear interactions may give rise to the third abnormal whorl floral organ in cytoplasmic male sterility (CMS) stem mustard, which exhibited complete conversion of homeotic transformations of stamens into other floral organ structures. In this system, expressions of AP3-like, PI-like and AG-like genes, the counterpart of the Arabidopsis nuclear ABC model genes related to floral organ development, were specifically altered in floral organs of CMS plants. When mitochondrial-specific inhibitor was applied to wild type fertile stem mustard plants, expressions of AP3-like, PI-like and AG-like genes were repressed. As a consequence, abnormal floral organs were observed in its maintainer line treated with mitochondrial-specific inhibitor. Thereby, we suggest that nuclear MADS-box transcription factors subject to mitochondrial retrograde regulation were associated with cytoplasmic homeosis in CMS stem mustard and could be mimicked by specifically inhibiting mtETC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cytoplasmic male sterility (CMS), the maternally inherited trait of failure to produce viable pollen, exists in many plant species and is widely applicable for hybrid production. It was generally believed that CMS was associated with rearrangement of mitochondrial genomes, which, in many cases, was attributed to the generation of novel orfs (Schnable and Wise 1998; Budar and Pelletier 2001; Hanson and Bentolila 2004; Linke and Börner 2005; Yang and Zhang 2007). According to the abortive stage of pollen, they were defined different types of CMS and exhibited different CMS-related abnormalities in flower morphology (Linke and Börner 2005). Striking alterations in flower morphology were observed in CMS tobacco (Kofer et al. 1991; Zubko et al. 2001), CMS wheat (Murai and Tsunewaki 1993; Murai et al. 2002), CMS carrot (Linke et al. 1999, 2003) and CMS stem mustard (in supplementary data Fig. 1). In CMS stem mustard, we previously reported that CMS plants developed homeotic floral organs, in which stamens were replaced by other floral organ structures, such as, petaloidy, pinnate, carpelloidy or silk-like floral structure (Yang et al. 2008a).
In higher plants, the development of floral organs controlled by the homeotic genes has been well studied on dicotyledonary plants, especially on Arabidopsis and Antirrhinum (Theissen and Saedler 1999; Theissen 2001). Usually, an intact flower from dicotyledonary plant has four whorl structures, that is, sepals for the first whorl, petals for the second whorl, stamens for the third whorl and carpels inside of flower. ABC model in floral organ developmental could explain and predict flower organ families based on three classes of nuclear homeotic genes, called A, B, and C (Davies and Schwarz-Sommer 1994; Ma 1994; Weigel and Meyerowitz 1994). This model has been extended by another two class D and E genes (Theissen 2001). This model hypothesizes that all five classes of genes are endowed with their genetic functions through concerted interactions. The class-A together with class-E genes specify sepals development. The expressions of class-A, B and E genes determine petals development and the expressions of class-B, C, and E determine stamens development, respectively. Furthermore, the combinative actions of class-C and E genes are attributed to carpels formation and class-D genes are answered for ovule constructions. This is termed as nuclear homeosis. The detailed descriptions of functional evolution and diversification for ABC model genes were documented by several research groups (Kramer et al. 1998, 2004; Krizek and Fletcher 2005; Hernández-Hernández et al. 2007). Any mutations or alterations in the transcription of these nuclear homeotic genes can lead to variations in the four whorl structures of a flower. A certain type of flower organ is replaced by others (Coen and Meyerowitz 1991; Mandel et al. 1992). In recent research on mitochondrial mutant and CMS system, the nuclear MADS-box genes, such as AP3-like, GLOBOSA-like and DEFICIENS-like, were found to be transcriptionally down-regulated and involved in this kind of conversion (Zubko et al. 2001; Murai et al. 2002; Linke et al. 2003; Hama et al. 2004; Teixeira et al. 2005). This is termed cytoplasmic homeosis. Description of cytoplasmic homeosis was caused by the rearrangement or replacement of the mitochondrial genome and subsequently to alter the expressions of nuclear homeotic genes or genes related to homeotic function (Zubko 2004). Most studies of cytoplasmic homeosis were focused on the CMS systems.
In higher plants, mitochondria have genes essential for the synthesis of proteins involved in various mitochondrial functions: respiratory chain electron transport, ATP synthesis (cyanide-sensitive, cytochrome pathway), cyanide-insensitive, alternative pathway (uncoupled with ATP synthesis) and tricarboxylic acid (TCA) cycle producing carbon dioxide for cellular biosynthesis substrates. In most cases, mitochondrial specific inhibitors are used to depress mitochondrial functions allowing the study of the regulation of these two respiratory pathways (Siedow and Umbach 1995; Mackenzie and Mclntosh 1999). These mitochondrial inhibitors could act specifically on different sites of the oxidative phosphorylation (OXPHOS) complex system, such as antimycin A, NaN3 and myxothiazol acting on respiratory chain, oligomycin acting on the ATP synthesis and 2,4-dinitrophenol (2,4-DNP) acting as a proton ionophore that can shuttle protons across biological membranes as well. NaN3 acts on cytochrome oxidase complex restraining the transfer of electrons from upstream respiratory complex to O2 (Wagner and Wagner 1997; Finnegan et al. 1998; Mclntosh et al. 1998; Saisho et al. 2001).
In the present study, we used a novel CMS line of stem mustard, which exhibits complete conversion of homeotic transformations. We also make use of wild type stem mustard plants to which mitochondrion-specific inhibitors have been applied, allowing us to study mitochondria–nuclear interactions that could be involved in the induction of cytoplasmic homeosis. We suggest that nuclear MADS-box transcription factors subject to mitochondrial retrograde regulation was associated with cytoplasmic homeosis in CMS stem mustard and could be mimicked by specifically inhibiting mtETC.
Materials and methods
Plant materials
CMS stem mustard (Brassica juncea var. tumida Tsen et Lee) and its maintainer line were as described by colleagues (Chen et al. 1995; Yang et al. 2008a). The CMS stem mustard line was synthesized by distant hybridizations and subsequent back-crossings between Brassica campestris and Brassica juncea. The progenies of the advanced back-crossed BC13 generation and its corresponding maintainer lines (concomitantly self-crossed) were used as the sources of sterile and fertile cytoplasms, respectively. Genetically, both CMS and its maintainer lines could be treated as near-isogenic lines up to the 13th back-crossed generation, and were favorable for a comparative analysis of molecular aspects. It was a new unique alloplasmic CMS named after orf220 besides orf222 and orf224 sterile cytoplasm in Brassica family. Ten plants of CMS and its maintainer lines were prepared for the following experiments.
Nucleic acid isolation and reverse transcription
Samples were ground in liquid nitrogen, and total RNA was isolated from every whorl of floral organs collected from 50 flowers using Trizol reagent (Invitrogen, Carlsbad, Carlifornia, USA). The amount of total RNA was determined by UV spectrophotometry. Total RNA (3 μg) were exhaustively treated with RNase-free DNase (Sigma, St. Louis) for 15 min at ambient temperature before being used for RT-PCR to remove residual DNA contamination. Using reverse transcription kit (Toyobo, Osaka, Japan), aliquots of total RNA were reverse transcribed into cDNA with Oligo (T)18 primer.
Isolation of AP3-like, PI-like and AG-like genes
AP3-like, PI-like and AG-like genes of stem mustard were isolated using RT-PCR method from its maintainer line. PCR primers were designed according to their homological genes, AP3 from Arabidopsis thaliana, Brassica rapa and Brassica napus, AG from Arabidopsis thaliana and Brassica napus and PI from Arabidopsis thaliana. All primers were listed in Table 1. PCR reactions for all genes were processed at 94°C (60 s), 52°C (60 s) and 72°C (60 s). The cDNA amplifications of AP3-like, PI-like and AG-like genes were cloned using TA-cloning and sequenced at Invitrogen Comp.
Sequence alignment and phylogenetic analysis
The deduced amino acid sequences of AP3-like, PI-like and AG-like genes from its maintainer line of stem mustard were performed using ORFfinder through NCBI online service. Full length amino acids alignments of AP3-like, PI-like and AG-like homologs from its maintainer line with 40 previously released AP3, PI and AG sequences from other species were initially complied using the computer program CLUSTAL W (Thompson et al. 1994). Clustal W multiple-aligment parameters were gap penalty 8 and gap extension penalty 2, using the PAM protein weight matrix for the amino acid alignment. A phylogenetic tree of the AP3, PI and AG genes was constructed by the neighbor-joining method provided by the above program. Amino acid sequences of the following genes were obtained from NCBI databases (Table 2).
Expressions analysis of MADS-box genes
Samples of every whorl floral organ were collected from CMS and its maintainer lines from 50 flowers for the measurement of AP3-like, PI-like and AG-like genes expressions using semi-quantitative RT-PCR. Meanwhile, the expressions of AP3-like, PI-like and AG-like genes from buds of maintainer line (termed CK) and CK treated with mitochondrial specific inhibitor were evaluated using semi-quantitative RT-PCR. Under the RT-PCR conditions developed in our laboratory, 27 cycles of amplification were performed to enable that quantification for all genes examined was within a linear range. PCR reactions for all genes were processed at 94°C (60 s), 52°C (60 s) and 72°C (60 s). RT-PCR was biologically repeated three times by using samples collected separately. In addition, stem mustard actin gene was used as internal control. A 20 μl from each PCR reaction was fractionated by 1% (w/v) agarose gel. The stained gels were digitally photographed with a JS380-A Auto Digital Imaging System (PEIQING Company, Shanghai, China).
NaN3 treatment
Wild type stem mustard (10 plants) were grown in a greenhouse and treated with different concentrations of 400 μmol/l NaN3 (mitochondrial cytochrome respiratory pathway specific inhibitor). Plants were treated at the stage of seedling and right before bolting (reproductive organs already fully differentiated). Wild-type stem mustard treated with distilled water was used as a control. A 400 μmol/l NaN3 (Sigma) solutions were sprayed every 3 days onto whole plant leaves before flowering.
Results
Flower phenotypes of stem mustard CMS line and its maintainer line
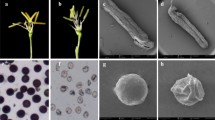
A dramatic variation in the morphology of floral organs was observed in CMS stem mustard. CMS had slightly smaller flowers in size as well as petals compared with its maintainer line. Severely curved pistils were observed in CMS stem mustard. CMS stem mustard exhibited complete homeotic transformations of stamens into pinnate, petaloidy, silk-like and carpelloidy structures (Supplementary data Fig. 1 or referred in Yang et al. 2008a). Single and mixed homeotic transformations occurred in one flower, in which a flower had one or two kind of transformed stamens structure and carpeloidy degenerative stamens with ovules were observed in this type of CMS.
Identification of AP3-like, PI-like and AG-like genes from stem mustard
The cDNAs of AP3-like (class-B, DQ060332), PI-like (class-B, DQ060333) and AG-like (class-C, DQ060334) genes were isolated from stem mustard, of which AP3-like, PI-like and AG-like genes were deduced to encode 224 amino acids, 208 amino acids and 252 amino acids, respectively. AP3-like, PI-like and AG-like genes belonged to MADS-box genes family due to their conserved MADS box domain in the N-terminal regions. One relative conserved K box domain was observed from AP3-like, PI-like and AG-like genes. Furthermore, PI-like gene was found to have the consensus PI motif (MPFxFRVQPxQPNLQE), which was identified in the C-terminal region of PI-type protein (Kramer et al. 1998) (Fig. 1).
Alignment of deduced amino acid sequences of AP3-like, PI-like and AG-like genes isolated from its maintainer line of stem mustard (underlined) with other MADS-box amino acid sequences. The MADS-box and K-box domains are indicated by solid and dash frames, respectively. The PI motif (MPFxFRVQPNLQE) of PI-like genes was indicated in the C terminal of the PI family by double lines box
The phylogenetic tree was constructed based on the deduced amino acid sequences to inspect the genetic relationships among AP3-like, PI-like and AG-like homeotic genes from Brassica juncea and other members of the class-B and -C gene family (Fig. 2). Thereafter, AP3-like, PI-like and AG-like genes cloned from mustard were used as AP3, PI and AG ortholog for the following expression analysis in different whorls of floral organs.
Phylogenetic tree of class-B and -C MADS-box genes from some species. The deduced amino acid sequences were obtained from NCBI databases (Table 2). It was constructed from amino acids of each MADS-box genes
Expression of AP3-like, PI-like and AG-like genes
In this CMS stem mustard, we observed homeotic transformations of whorl three into other floral structures. Hence, we only study the expressions of class-B and class-C genes. The expression patterns of AP3-like, PI-like and AG-like genes were evaluated in stem mustard CMS line and its maintainer line. In its maintainer line, it was observed that AP3-like and PI-like genes were expressed at the second and third whorls of floral organs, and AG-like gene was expressed at the third and fourth whorl of floral organs. The expression patterns of these three genes were strictly controlled by the ABC model. However, the expression patterns of these three genes were dramatically altered in stem mustard CMS line. Expression of AP3-like gene was not only decreased at the third whorl floral organ, but also appeared at the fourth whorl floral organ in stem mustard CMS line. Transcripts of PI-like were also reduced at the third whorl floral organ of the CMS line. As to AG-like gene, it was unexpectedly expressed at the second whorl floral organs, reduced at the third whorl floral organ and enhanced at the fourth whorl floral organs (Fig. 3).
Expressions of AP3-like, PI-like and AG-like genes between stem mustard CMS line and its maintainer line. MF, its maintainer line; CMS, cytoplasmic male sterile line of stem mustard. Stamen* represents abnormal stamen in CMS stem mustard. The actin gene was used as a control showing that the same amount of cDNA was used in the PCR reaction of samples
Abnormal floral organ is induced by mitochondrion-specific inhibitor
Unfavorably, the restored line has not been discovered as yet for this type of CMS, and therefore, we fail to investigate whether the expressions of AP3-like, PI-like and AG-like genes could be recovered in its restored line to testify the involvement of these genes in the homeotic transformation. However, we may investigate whether the phenotypes and expressions of AP3-like, PI-like and AG-like genes were affected or not after artificially depressing the function of mitochondria using mitochondrial specific inhibitor in its maintainer line. As a consequence, expressions of AP3-like, PI-like and AG-like genes were observed to be reduced in the buds of its maintainer line treated with NaN3 (Fig. 4A). Importantly, we observed adhesive structure of petal and stamen when wild type stem mustard was treated with NaN3 (Fig. 4B). Two kinds of abnormal floral organs were shown in wild type stem mustard flowers treated with NaN3. Type 1 exhibited that both whorl two and whorl three organs were altered and sticking in flower (Fig. 4B2). Type 2 showed that whorl two and whorl three organs were sticking in flower and only whorl three was altered (Fig. 4B3). Apart from the abnormal floral organ, wild type stem mustard treated with could not NaN3 produce functional pollens (Yang et al. 2008c).
Expressions of AP3-like, PI-like and AG-like genes and floral organ structure in its maintainer line and its maintainer line after being treated by NaN3. A, Expressions of AP3-like, PI-like and AG-like genes, CK represented its maintainer line, Tr represented its maintainer line after being treated by NaN3; B, floral organ structure, B-1, maintainer line stamen, B-2 exhibits both whorl two and whorl three organs were altered and sticking, B-3 shows whorl two and whorl three organs were sticking and only whorl three was altered (Fig. 4-B3)
Discussion
In CMS system, changes in mitochondrial genes have been associated to male sterility occurrence. Together with the male sterility occurrence, homeotic transformations of floral organs happen (Linke and Börner 2005). However, mitochondrial factors do not directly control reproductive development. There must, therefore, be a signaling pathway from mitochondria to nucleus that is involved in the homeotic transformations of floral organs. How, and through which pathway, might mitochondrial factors determine homeotic transformations of floral organs?
In the stem mustard CMS system here, complete homeotic conversions of stamens into other floral structures (Chen et al. 1995; Yang et al. 2008a). According to the ABC model of floral organ development, any mutation or over-expression of these genes would lead to homeotic transformations in flower architecture, with the replacement of certain floral organs by others, termed nuclear homeosis (Zubko 2004). Similarly, homeosis also has happened when mitochondrial genome were altered and subsequently affect expressions of nuclear homeotic genes or genes related to homeotic function, termed cytoplasmic homeosis (Zubko et al. 2001; Zubko 2004). In our investigations, we observed that AP3-like, PI-like and AG-like genes were strictly expressed in their whorl of floral organ according to ABC model in its maintainer line. However, it was observed that reduced transcription of AP3-like, PI-like and AG-like genes could possibly involve in the development of various types of aberrant stamen in CMS line. Although we did not determine which nuclear homeotic gene was responsible for each kind of respective type of abnormal floral organ.
AP3-like, PI-like and AG-like genes might therefore be target genes for mitochondrial retrograde regulation (MRR, Liao et al. 1991). In recent studies, variation in the morphology of flowers in CMS line was reported, of which nuclear MADS-box genes were subject to mitochondrial retrograde regulation, down-regulated expression and uncovered as targeted genes of interaction between nucleus and mitochondria (Zubko et al. 2001; Linke et al. 2003; Meguro et al. 2003; Hama et al. 2004; Teixeira et al. 2005). CMS systems offered a chance to explore the cytoplasmic homeosis. In higher plants, putative retrograde signals from mitochondria to the nucleus are thought to play a role in the regulation of many nuclear genes (Yu et al. 2001). Some eukaryotic transcription factor genes have been shown to be subject to redox regulation by mitochondria (Sun and Oberley 1996). Thus, changes in mitochondria of CMS stem mustard might account for the alterations on the expressions of AP3-like, PI-like and AG-like genes.
According to the ABC model, each type of nuclear homeotic gene is strictly expressed organ-specifically at the individual whorls of floral organs. However, ectopic expressions of AP3-like and AG-like genes were observed in stem mustard CMS line, of which AP3-like gene was expressed in pistil and AG-like gene in petal unexpectedly. Similar result was reported that AP3 gene was found to be ectopically expressed in CMS Brassica napus (Geddy et al. 2004). In stem mustard CMS line, we observed complete homeotic transformations of stamens into pinnate, petaloidy, silk-like and carpelloidy structures. Single and mixed homeotic transformations occurred in one flower, in which a flower had one or two kind of transformed stamens structure and carpeloidy degenerative stamens with ovules were observed in this type of CMS (Yang et al. 2008a). From the data obtained, it was not yet enough to answer whether alterations on ecotpic expressions of these genes were cause or consequence of developmental abnormality. However, it was an exceptional phenomenon observed in this CMS system. The ABC genetic model was not fully expected in this type of CMS and this finding had never been reported before, maybe the ABC model could be extended to more information. Therefore, we were trying to state the facts observed here to the readers as exceptions. Probably, ectopic expression of nuclear homeotic genes and cytoplasmic homeosis may offer a chance to learn more and extend the ABC model in higher plants.
In this type of CMS, we previously reported that genomic sequence and expression of mitochondrial nad2 gene, subunit of mitochondrial cytochrome oxidase respiratory pathway complex, were altered and may subsequently affect the mitochondrial cytochrome oxidase respiratory (Yang et al. 2008b). NaN3, as a mitochondrial inhibitor, restrains mitochondrial cytochrome oxidase respiratory electron transport chain. Therefore, NaN3 was employed to investigate whether depression of mitochondrial function could induce abnormal reproductive developments. Studies with mitochondrial inhibitors that differentially affect alternative respiratory pathways have been performed in Arabidopsis (Saisho et al. 2001), barley (Krömer and Heldt 1991) and Petunia (Wagner and Wagner 1997). However, whether a mitochondrial-specific inhibitor could induce extraordinary reproductive development or not has not been reported till now. We showed here that NaN3 could decrease the expressions of AP3-like, PI-like and AG-like genes at different levels (Fig. 4A). As a consequence, we observed the abnormal development of floral organs when we artificially inhibited the mitochondrial ETC. (Fig. 4B2/B3).
Mitochondrial effects on the nuclear genes expressions (termed retrograde regulation, Liao et al. 1991) have been well documented mostly in yeast and mammals (Butow and Avadhani 2004; Liu and Butow 2006). In the case of plants, a group of genes related to programmed cell death (PCD) have been reported to be potential nuclear targets in CMS involving tapetum degeneration (Balk and Leaver 2001). In this case, PCD initiates in tapetal cells and subsequently extends to other anther tissues in association with partial release of cytochrome c from the mitochondria.
In conclusion, for the stem mustard CMS system exhibiting complete transformations of stamen into other floral structure, we hypothesize that AP3-like, PI-like and AG-like, nuclear genes encoding MADS-box transcription factor, are subject to mitochondrial regulation associated with the cytoplasmic homeosis. This might occur through and MADS-box gene direct or indirect response to a signal emanating from mitochondria. Nuclear transcription factor genes (AP3-like, PI-like and AG-like genes) may be considered as nuclear targeted genes on the signaling pathway involved in mitochondria-to-nucleus communication. Mitochondrial specific inhibitor offered us an efficient method to study the effects of mitochondrial dysfunction in higher plants. These possibilities remain to be elucidated.
Abbreviations
- mtETC:
-
Mitochondrial electron transport chain
- CMS:
-
Cytoplasmic male sterility
- TF:
-
Transcription factor
- AP3 :
-
APETALA3
- PI :
-
PISTILLATA
- AG :
-
AGAMOUS
- NZZ :
-
NOZZLE
- orf :
-
Open reading frame
References
Budar F, Pelletier G (2001) Male sterility in plant: occurrence, determinism, significance and use. Life Sci 324:543–550
Butow RA, Avadhani NG (2004) Mitochondrial signaling: the retrograde response. Mol Cell 14:1–15. doi:10.1016/S1097-2765(04)00179-0
Chen ZJ, Zhang MF, Wang BL, Dong WM, Huang SQ (1995) A study on fertility and agronomic characters of CMS lines for tuber mustard. Acta Hortic Sin 22(1):40–46
Coen ES, Meyerowitz EM (1991) The war of the whorls: genetic interactions controlling flower development. Nature 353:31–37. doi:10.1038/353031a0
Davies B, Schwarz-Sommer Z (1994) Control of floral organ identity by homeotic MADS-box transcription factors. In: Nover L (ed) Plant promoters and transcription factors. Springer, Berlin, Heidelberg, New York, pp 235–258
Finnegan PM, Djajanegara IN, Schuller LJ, Smith MK, Whelan J, Day DA (1998) Analysis o the antimycin A-inducible promoter region of soybean GmAOX1. In: MØller IM, Gardeström P, Glimelius K, Glaer E (eds) Plant mitochondria: from gene to function. Backhuys Publishers, Leiden, The Netherlands, pp 449–453
Geddy R, Mahe L, Brown GG (2004) Cell-specific regulation of a Brassica napus CMS-associated gene by a nuclear restorer with related effects on a floral homeotic gene promoter. Plant J 41:333–345
Hama E, Takumi S, Ogihara Y, Murai K (2004) Pistillody is caused by alteration to the class-B MADS-box gene expression pattern in alloplasmic wheat. Planta 218:712–720. doi:10.1007/s00425-003-1157-6
Hanson MR, Bentolila S (2004) Interaction of mitochondrial and nuclear genes that affect male gametophyte development. Plant Cell 16:S154–S169. doi:10.1105/tpc.015966
Hernández-Hernández T, Martínez-Castilla LP, Alvarez-Buylla ER (2007) Functional diversification of B MADs-box homeotic regulation of flower development: adaptive evolution in protein-protein interaction domain after major gene duplication events. Mol Biol Evol 24:465–481. doi:10.1093/molbev/msl182
Kofer W, Glimelius K, Bonnett HT (1991) Modification of mitochondrial DNA cause changes in floral development in homeotic-like mutants of tobacco. Plant Cell 3:759–769
Kramer EM, Dorit RL, Irish VF (1998) Molecular evolution of genes controlling petal and stamen development: duplication and divergence within the APETALA3 and PISTILATA MADS-box gene lineages. Genetics 149:765–783
Kramer EM, Jaramillo MA, Di Stilio VS (2004) Patterns of gene duplication and functional evolution during the diversification of the AGAMOUS subfamily of MADS box genes in angiosperms. Genetics 166:1011–1023. doi:10.1534/genetics.166.2.1011
Krizek BA, Fletcher CJ (2005) Molecular mechanisms of flower development: an armchair guide. Nat Rev Genet 6:688–698. doi:10.1038/nrg1675
Krömer S, Heldt HW (1991) On the role of mitochondrial oxidative phosphorylation in photosynthesis metabolism as studied by the effect of oligomycin on photosynthesis in protoplasts and leaves of barley. Plant Physiol 95:1270–1276
Liao XS, Small WC, Srere PA, Butow RA (1991) Intramitochondrial functions regulate nonmitochondrial citrate synthase (CIT2) expression in Saccharomyces cerevisiae. Mol Cell Biol 11:38–46
Linke B, Börner T (2005) Mitochondrial effects on flower and pollen development. Mitochondrion 5:389–402. doi:10.1016/j.mito.2005.10.001
Linke B, Nothnagel T, Börner T (1999) Morphological characterization of modified flower morphology of three novel alloplasmic male sterile carrot sources. Plant Breed 118:543–548. doi:10.1046/j.1439-0523.1999.00402.x
Linke B, Nothnagel T, Börner T (2003) Flower development in carrot CMS plant: mitochondria affect the expression of MADS box genes homologous to GLOBOSA and DEFICIENS. Plant J 34:27–37. doi:10.1046/j.1365-313X.2003.01703.x
Liu ZC, Butow RA (2006) Mitochondrial retrograde signaling. Annu Rev Genet 40:159–185. doi:10.1146/annurev.genet.40.110405.090613
Ma H (1994) The unfolding drama of flower development: recent results from genetic and molecular analyses. Genes Dev 8:745–756. doi:10.1101/gad.8.7.745
Mackenzie S, Mclntosh L (1999) Higher plant mitochondria. Plant Cell 11:571–585
Mandel MA, Bowman JL, Kempin SA, Ma H, Meyerowitz EM, Yanofsky MF (1992) Manipulation of flower structure in transgenic tobacco. Cell 71:133–143. doi:10.1016/0092-8674(92)90272-E
Mclntosh L, Eichler T, Gray G, Maxwell D, Nichels R, Wang Y (1998) Biochemical and genetic controls exerted by plant mitochondria. Biochim Biophys Acta 1365:278–284. doi:10.1016/S0005-2728(98)00080-2
Meguro A, Takumi S, Ogihara Y, Muria K (2003) WAG, a wheat AGAMOUS homolog, is associated with development pistil-like stamens in alloplasmic wheat. Sex Plant Reprod 40:167–177
Murai K, Tsunewaki K (1993) Photoperiod-sensitive cytoplasmic male sterility in wheat with Aegilops crassa cytoplasm. Euphytica 67:41–48. doi:10.1007/BF00022723
Murai K, Takumi S, Koga H, Ogihara Y (2002) Pistillody, homeotic transformation of stamens into pistil-like structures, caused by nuclear-cytoplasm interaction in wheat. Plant J 29:169–181. doi:10.1046/j.0960-7412.2001.01203.x
Saisho D, Nakazono M, Tsutsumi N, Hirai A (2001) ATP synthesis inhibitor as well as respiratory inhibitor increase steady-state level of alternative oxidase mRNA in Arabidopsis thaliana. J Plant Physiol 158:241–245. doi:10.1078/0176-1617-00193
Schnable PS, Wise RP (1998) The molecular basis of cytoplasmic male sterility and fertility restoration. Trends Plant Sci 3:175–180. doi:10.1016/S1360-1385(98)01235-7
Siedow JN, Umbach AL (1995) Plant mitochondrial electron transfer and molecular biology. Plant Cell 7:821–831. doi:10.1105/tpc.7.7.821
Sun Y, Oberley LW (1996) Redox regulation of transcriptional activators. Free Radic Biol Med 21:335–348. doi:10.1016/0891-5849(96)00109-8
Teixeira RT, Farbos I, Glimelius K (2005) Expression levels of meristem identity and homeotic genes are modified by nuclear–mitochondrial interactions in alloplasmic male-sterile lines of Brassica napus. Plant J 42:731–742. doi:10.1111/j.1365-313X.2005.02407.x
Theissen G (2001) Development of floral organ identity: stories from the MADS house. Curr Opin Plant Biol 4:75–85. doi:10.1016/S1369-5266(00)00139-4
Theissen G, Saedler H (1999) The golden decade of molecular floral development (1990–1999): a cheerful obituary. Dev Genet 25:181–193
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighing, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. doi:10.1093/nar/22.22.4673
Wagner AM, Wagner MJ (1997) Changes in mitochondrial respiratory chain components of Petunia cells during culture in the presence of antimycin A. Plant Physiol 115:617–622
Weigel D, Meyerowitz EM (1994) The ABCs of floral homeotic genes. Cell 78:203–209. doi:10.1016/0092-8674(94)90291-7
Yang JH, Zhang MF (2007) Molecular genetics of mitochondrial respiratory/ATP-related genes in relation to cytoplasmic male-sterility of higher plants. Gene Genomes Genomics 1(1):21–26
Yang JH, Huai Y, Zhang MF (2008a) Mitochondrial atpA gene is altered in a new orf220-type cytoplasmic male-sterile line of stem mustard (Brassica juncea). Mol Biol Rep. doi:10.1007/s11033-007-9176-1
Yang JH, Zhang MF, Yu JQ (2008b) Mitochondrial nad2 gene is co-transcripted with CMS-associated orfB gene in cytoplasmic male-sterile stem mustard (Brassica juncea). Mol Biol Rep. doi:10.1007/s11033-007-9185-0
Yang JH, Zhang MF, Yu JQ (2008c) Relationship between cytoplasmic male sterility and SPL-like gene expression in stem mustard. Physiol Plant 133:426–434
Yu JP, Nickels R, Mcintosh L (2001) A genome approach to mitochondrial–nuclear communication in Arabidopsis. Plant Physiol Biochem 39:345–353. doi:10.1016/S0981-9428(01)01254-2
Zubko MK (2004) Mitochondrial tuning fork in nuclear homeotic functions. Trends Plant Sci 9(2):61–63. doi:10.1016/j.tplants.2003.12.001
Zubko MK, Zubko E, Ryban AV, Adler K, Mock HP, Misera S et al (2001) Extensive development and metabolic alteration in cybrids Nicotiana tabacum (+Hyoscyamus niger) are caused by complex nucleo-cytoplasmic incompatatibility. Plant J 25:627–639. doi:10.1046/j.1365-313x.2001.00997.x
Acknowledgements
This work was supported by a grant from the National Natural Science Foundation of China (NSFC30571270 and NSFC30772378). We thank Dr. Huang Jirong and Mikio Nakazono for providing critical comments on the paper.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Yang, JH., Qi, XH., Zhang, MF. et al. MADS-box genes are associated with cytoplasmic homeosis in cytoplasmic male-sterile stem mustard as partially mimicked by specifically inhibiting mtETC. Plant Growth Regul 56, 191–201 (2008). https://doi.org/10.1007/s10725-008-9300-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-008-9300-9