Abstract
Synthetic hexaploid wheat (SHW) is known to be an excellent vehicle for transferring large genetic variations especially the many useful traits present in the D genome of Aegilops tauschii Coss. (2n = 2x = 14, DD) for improvement of cultivated wheat (Triticum aestivum L., 2n = 6x = 42, AABBDD). The objectives of the present study were to (1) evaluate genetic diversity among 32 selected SHW accessions with resistance to several fungal diseases using Amplified Fragment Length Polymorphism (AFLP) and Simple Sequence Repeat (SSR) markers and (2) identify diverse SHWs for pyramiding genes conferring resistance to different diseases. These SHWs containing different accessional sources of the D genome were identified from about 1000 SHW accessions developed by the Wheat Wide Crosses program at the International Maize and Wheat Improvement Center, Mexico. Of the 32 SHW accessions eight had resistance to Fusarium head blight (Fusarium graminearum Schw.), seven were resistant to leaf rust (Puccinia triticina Eriks.), eight resistant to Helminthosporium spot blotch [Cochliobolus sativus (Ito et Kurib.) Drechsler ex Dastur (syn.: Bipolaris sorokiniana (Sacc.) Shoem., Helminthosporium sativum Pammel, King et Bakke)], seven resistant to Septoria tritici blotch (Septoria tritici Roberge in Desmaz.), while two were resistant to both Fusarium head blight and leaf rust. Seventeen EcoRI/MseI AFLP primer combinations and 27 highly polymorphic SSR markers including 20 D genome specific markers were screened over all 32 SHW accessions. Among the 703 AFLP fragments scored, 225 were polymorphic across the 32 SHW accessions. Polymorphic information content (PIC) among the SHWs for AFLP ranged from 0.06 to 0.50 with an average PIC of 0.24. Major allelic frequency from SSR analysis ranged from 0.23 to 0.81 with an average of 0.45. Number of alleles per locus for the SSR markers ranged from 3 to 15 with an average allele number of 7.4. Average gene diversity and PIC for the SSR markers was 0.69 and 0.66, respectively, with the highest values being for the D genome specific markers. Cluster analysis showed distinct groups among the SHW accessions studied. Mantel statistics between the distance matrices from AFLP and SSR analyses showed a moderate but significant correlation (r = 0.52**). Our results indicate that the SHW accessions studied possess substantial genetic diversity and are valuable user-friendly pre-breeding materials for breeding improvement of wheat with resistance to Fusarium head blight, leaf rust, Helminthosporium spot blotch, and Septoria tritici blotch. The most diverse SHW accessions can be used for pyramiding resistance genes for different diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In a global context, fungal diseases are major constraints to wheat production. Therefore, breeding wheat with biotic stress resistance is a major goal in most wheat breeding programs that are global or location specific. Since the variation for traits such as disease resistance is limited in conventional wheat, introgression of genes from wild relatives into elite cultivars has been a major objective at International Maize and Wheat Improvement Center (CIMMYT) and in many other breeding programs (Mujeeb-Kazi et al. 2013; Ogbonnaya et al. 2013). Aegilops tauschii, the donor of the D genome of common wheat has many desirable genes for wheat improvement (Mujeeb-Kazi and Rajaram 2002). In fact, the first attempt to use SHW in a breeding program was inspired by the need to develop germplasm resistant to biotic stresses (Mujeeb-Kazi and Asiedu 1990) and specifically to Karnal bunt disease of wheat (Warburton et al. 2006). Between 1987 and 2003 about 1000 SHW accessions were developed at CIMMYT from a wide array of A. tauschii accessions (Mujeeb-Kazi et al. 1996, 2001b). These CIMMYT SHWs brought variations for morphological and agronomic traits and resistance to biotic and abiotic stresses (Villareal et al. 1994; Rajaram and van Ginkel 2001; Lage et al. 2004; Trethowan and Mujeeb-Kazi 2008; Mujeeb-Kazi et al. 2008; Ogbonnaya et al. 2013; Rasheed et al. 2014; Ali et al. 2015). High levels of resistance were obtained for Fusarium head blight, leaf rust, Septoria tritici blotch (Mujeeb-Kazi et al. 2000, 2001b), Karnal bunt (Tilletia indica Mitra) (Mujeeb-Kazi et al. 2001c), tan spot (Pyrenophora tritici-repentis (Died.) Drechs.), Helminthosporium spot blotch (Mujeeb-Kazi et al. 2001a) and Stagonospora nodorum blotch (Phaeosphaeria nodorum (Berk.) Castellani et Germano) (Cox 1998; Xu et al. 2004). In a study involving 282 synthetic backcross derived lines, higher grain yield and kernel weight in the SHWs as compared to their recurrent common wheat parents were reported by del Blanco et al. (2001). Cooper et al. (2013) reported use of CIMMYT derived SHWs in backcrosses with two winter wheat cultivars TAM111 and TAM112 in Texas. In this study SHWs producing progeny superior to TAM 111 or TAM 112 were identified. Recently, four wheat cultivars developed using CIMMYT SHWs have been released in China (Yang et al. 2009). CIMMYT SHW-derived wheat cultivars have also been released in other countries including Spain, Mexico and the USA.
The CIMMYT SHWs also possess great diversity at the molecular level (Zhang et al. 2005; Warburton et al. 2006). Genetic diversity of germplasm can be assessed through pedigree analysis (Cox et al. 1985; Martin et al. 1991) and DNA markers (Autrique et al. 1996; Barrett and Kidwell 1998; Soleimani et al. 2002). Lage et al. (2003) reported genetic diversity studies of SHWs using agronomic and AFLP data. They observed that the SHWs had a considerably higher level of AFLP diversity (39 %) than normally observed in conventional hexaploid wheats (12–21 %). Based on AFLP analysis, Das et al. (2007) observed substantial genetic diversity among several conventional wheat and CIMMYT derived SHWs with drought and salinity tolerance. Sharma et al. (2014) reported presence of genetic diversity for heat tolerance in CIMMYT derived SHWs using phenotypic and molecular marker data. The objective of the present study were to (1) assess genetic diversity among 32 selected SHW accessions from CIMMYT program based on AFLP and SSR molecular markers and (2) identify diverse SHWs for pyramiding genes conferring resistance to different diseases. This selected germplasm set contained different D genome accessional sources conferring resistance to Fusarium head blight, leaf rust, Helminthosporium spot blotch and Septoria tritici blotch.
Materials and methods
Plant materials
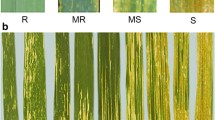
Thirty-two SHW accessions with resistance to several biotic stresses were used in this study (Table 1). These SHWs were identified from a main set of 1014 SHW accessions developed by the Wheat Wide Crosses program at CIMMYT, Mexico as resistant to main wheat diseases. Eight SHW accessions (1F to 8F, Table 1) had resistance to Fusarium head blight (type II) with mean scores ranging from 9 to 14 % infection across 5 years of data. Seven SHW accessions (1L to 7L, Table 1) had leaf rust resistance with rust severity ratings ranging from Trace to 5 MR-MS over several years of testing in CIMMYT, Mexico. Eight SHW accessions (1H to 8H, Table 1) were resistant to Helminthosporium spot blotch with severity scores ranging from 2-1 to 3-2 on a double-digit severity scale. Seven SHW accessions (1S to 7S, Table 1) were resistant to Septoria tritici blotch with disease severities ranging from 2-1 to 2-2 on a double-digit severity scale. Two SHW accessions (1FL and 2FL, Table 1) were resistant to both Fusarium head blight and leaf rust. Disease resistance levels of the SHW accessions presented above are from A. Mujeeb-Kazi (unpublished data).
DNA extraction and AFLP assays
Three leaves per entry were collected from wheat seedlings that were grown for about 10 days. Genomic DNA was extracted from these tissue samples using the CTAB procedure (Saghai-Maroof et al. 1984). AFLP analysis was done as described by Bai et al. (1999) and Das et al. (2007). Briefly, 300 ng genomic DNA from each of the wheat accessions was double digested with the restriction enzymes EcoRI and MseI. Adapters for both restriction enzymes were then ligated to the digested DNA fragments. For pre-amplification the ligated DNA was diluted tenfold. Forty micro-L of PCR reaction mixture contained 10 μl of the diluted DNA, 4 μl of 10X PCR buffer, 4 μl MgCl2 (25 mM), 1.6 μl dNTPs (5 mM), 0.75 μl EcoRI pre-amplification primer (100 ng μl−1), 0.75 μl MseI pre-amplification primer (100 ng μl−1), 0.15 μl Taq polymerase and 18.75 μl of deionized water. All PCR reactions were performed in an MJ PTC-100 thermocycler (MJ Research Inc., Waltham, MA, USA). The following thermal profile was used for pre-amplification: 94 °C for 1 min followed by 30 cycles at 94 °C for 30 s, 56 °C for 60 s and 72 °C for 60 s. Pre-amplification was confirmed by analyzing the PCR products on 1.5 % agarose gel.
The pre-amplified DNA was diluted tenfold and selective amplification was done using the following reaction mixture: 2 μl of the diluted DNA, 1 μl of 10X PCR buffer, 1 μl of 25 mM MgCl2, 0.4 μl of 5 mM dNTPs, 0.35 μl of MseI selective primer (50 ng μl−1), 0.4 μl fluorescence-labeled EcoRI selective primer (1 ρmol μl−1) from LI-COR (LI-COR Inc, Lincoln, NE, USA), 0.04 μl Taq polymerase, and 4.8 μl deionized water. Thermal cycles were as follows: 2 min at 94 °C followed by 13 cycles at 94 °C for 30 s, 65 °C for 30 s, 72 °C for 60 s with the annealing temperature lowered by 0.7 °C after each cycle; then followed by 23 cycles at 94 °C for 30 s, 56 °C for 30 s, 72 °C for 60 s. The PCR product was held at 72 °C for 5 min for final extension. Selective amplification was done with 17 AFLP selective primer combinations (Table 2). Using a 6.5 % denaturing gel (Li-Cor Inc., Lincoln, NE, USA) the PCR products from the selective amplification was run in 1X TBE buffer at 1500 V and 40 W for 3.5 h in a Li-Cor automated DNA sequencer (Li-Cor Inc., Lincoln, NE, USA) using DNA size standard from Li-Cor (Li-Cor Inc., Lincoln, NE).
SSR assays
Twenty-seven highly polymorphic SSR markers (Table 3) were analyzed for all 32 SHW accessions in a Li-Cor automated DNA sequencer (Li-Cor Inc., Lincoln, NE) by labeling one primer with an infrared fluorescence dye. Twenty of the markers were D genome specific. For PCR, each 10 μl of reaction contained 30 ng DNA, 1 X PCR buffer, 0.25 mM dNTP, 2.5 mM MgCl2, 0.5 pmol each of labeled and unlabeled SSR primers, and 1 unit of Taq polymerase. The following touchdown thermal profile was used for SSR amplification: 5 min of 95 °C followed by five cycles of 45 s at 95 °C, 5 min at 68 °C and 1 min at 72 °C, in which the annealing temperature was lowered by 2 °C in each cycle; five more cycles in which the annealing time was 2 min and the temperature was lowered by 2 °C in each following cycle; followed by 25 cycles in which the annealing temperature was held constant at 50 °C. Five minutes at 72 °C was used for final extension. Molecular size determination of the SSR fragments was done by comparison of amplified PCR fragments with the DNA size standard (Li-Cor Inc., Lincoln, NE).
Data analyses
A present (1) or absent (0) scoring of the AFLP bands ranging from 70 to 350 base pairs was used for the data analyses. Unresolvable bands were entered as 0.5 for missing data in the data matrix. For all possible pairs of accessions polymorphism rates were estimated by dividing the number of polymorphic bands by the total number of bands. Estimation of polymorphic information content (PIC) was done using the formula used by Anderson et al. (1993):
where pi is the frequency of the ith allele. An AFLP locus can have a maximum PIC value of 0.5. Genetic similarity between the SHW accessions was estimated using the similarity coefficients of Jaccard (1908). Similarity estimation was done for AFLP data set, SSR data set and also for combined AFLP and SSR data sets. Since 20 of the 27 SSR markers were D genome specific, similarity estimates were also done using the D genome specific SSR markers only. The resulting distance matrices were used for cluster analysis by the UPGMA (unweighted pair-group method with arithmetic averages, Sneath and Sokal 1973) method. The goodness of fit of the clustering to the data matrix was assessed by cophenetic correlation. NTSYS-pc software (Rohlf 2000) was used for cluster analyses and calculation of cophenetic correlation. Major allelic frequency, PIC and gene diversity for the SSR data were determined by the PowerMarker software (Liu and Muse 2005). Correlation between Jaccard’s similarity matrices from AFLP and SSR data set was calculated using the Mantel test (Mantel 1967) with 1000 random permutations.
Results
Analyses of 17 AFLP primer combinations produced 703 scorable AFLP fragments, 225 of which were polymorphic across the 32 SHW accessions with an average polymorphism of 32 % ranging from 16.7 to 56.7 %. The average polymorphic band per primer combination was 13.2 with the highest (34) and lowest (5) numbers of polymorphic bands produced by primer combinations e-AGT/m-CAGT and e-AAC/m-TGC, respectively (Table 2). Polymorphic information content for the AFLP markers ranged from 0.06 to 0.50 with an average PIC of 0.24.
Analyses of the 27 SSR markers produced 204 scorable fragments with 202 polymorphic fragments (99 % polymorphism) across the 32 SHW accessions. Major allelic frequency for the 27 SSR markers ranged from 0.23 for the markers WMC167 and Xgdm98 to 0.81 for the marker Xgwm558 with an average allelic frequency of 0.45 (Table 3). Number of alleles per locus ranged from three for the SSR markers Xgwm558, Xgwm334, Xgdm108, Xgdm40 and WMC326 to 15 for the marker WMC167 with an average allele number of 7.4. Gene diversity ranged from 0.33 for the marker Xgwm558 to 0.88 for the marker WMC167 with an average gene diversity of 0.69. Polymorphic information content ranged from 0.31 for the marker Xgwm558 to 0.86 for the marker WMC167 with an average PIC of 0.66.
Based on AFLP data Jaccard’s similarity coefficients ranged from 0.23 to 0.84 with an average coefficient of 0.57. The highest similarity (0.84) was between the SHW accessions 1S and 4S and the lowest similarity (0.23) was between the accessions 2FL and 6F. Jaccard’s similarity coefficients for the SSR data ranged from 0.018 to 0.97 with an average coefficient of 0.19. The highest similarity (0.97) was between the accessions 7H and 8H and the lowest similarity (0.018) was between the accessions 1FL and 2L. The 20 D genome specific SSR markers produced 167 polymorphic fragments. Jaccard’s similarity coefficients produced by the 20 D genome specific SSR markers ranged from 0.02 to 1.0 with an average coefficient of 0.17. The highest similarity (1.0) was between the accessions 1H and 2H while the lowest similarity (0.02) was between the accessions 1FL and 2L. Jaccard’s similarity coefficients based on the combined AFLP and SSR data ranged from 0.198 to 0.80 with an average similarity coefficient of 0.47. The highest similarity (0.80) was between accessions 7H and 8H and the lowest similarity coefficient (0.198) was between the accessions 6F and 2FL.
Cophenetic correlations for AFLP, SSR, D genome specific SSR and combined AFLP and SSR data were 0.84, 0.93, 0.95 and 0.88, respectively. Cophenetic correlation, r ≥ 0.9 indicates a very good fit; 0.8 ≤ r < 0.9 indicates a good fit; and 0.7 ≤ r < 0.8 indicates a poor fit (Rohlf 2000). Pejic et al. (1998) reported that 150 polymorphic fragments could provide a reliable estimate of genetic similarities among genotypes within a species. In our study, analyses on the 32 SHW accessions based on 225 (AFLP), 202 (SSR) and 167 (D genome SSR) polymorphic bands are expected to provide reliable estimates of the similarity values.
Cluster analyses based on UPGMA are given in Figs. 1, 2, 3, and 4. The dendrogram based on the 225 AFLP markers grouped the 32 SHW accessions into several clusters (Fig. 1). Several of the clusters joined together at similarity 0.58. The SHWs 1FL and 2FL clustered together and joined rest of the SHW accessions at similarity 0.40. The SHW 6F remained by itself and joined rest of the SHWs except 1FL and 2FL at similarity 0.43. The dendrogram based on the 27 SSR markers also grouped the 32 SHW accessions into several clusters each with a small number of entries (Fig. 2). In this grouping also SHWs 1FL and 2FL clustered together and joined the cluster with SHWs 4L and 5L at similarity 0.19. The dendrogram developed from the 20 D genome specific SSR markers gave a similar grouping of the 32 SHW accessions with few exceptions (Fig. 3). The dendrogram based on the combined data from 225 AFLP markers and 27 SSR markers also grouped the 32 SHW accessions into several small clusters (Fig. 4). In this dendrogram, like the dendrogram from the AFLP markers the SHWs 1FL and 2FL clustered together and the SHW 6F remained by itself. However, there are some differences in the groupings of the SHWs in the two dendrograms (Figs. 1, 4).
Dendrogram from the Jaccard’s similarity coefficients based on 225 polymorphic AFLP markers across 32 synthetic hexaploid wheat accessions. Accessions 1F to 8F, Fusarium head blight resistant; 1FL and 2FL, Fusarium head blight and leaf rust resistant; 1L to 7L, leaf rust resistant; 1H to 8H, Helminthosporium spot blotch resistant; 1S to 7S, Septoria tritici blotch resistant
Dendrogram from the Jaccard’s similarity coefficients based on 202 polymorphic SSR bands across 32 synthetic hexaploid wheat accessions. Accessions 1F to 8F, Fusarium head blight resistant; 1FL and 2FL, Fusarium head blight and leaf rust resistant; 1L to 7L, leaf rust resistant; 1H to 8H, Helminthosporium spot blotch resistant; 1S to 7S, Septoria tritici blotch resistant
Dendrogram from the Jaccard’s similarity coefficients based on 167 polymorphic SSR bands from the 20 D genome specific SSR markers screened across 32 synthetic hexaploid wheat accessions. Accessions 1F to 8F, Fusarium head blight resistant; 1FL and 2FL, Fusarium head blight and leaf rust resistant; 1L to 7L, leaf rust resistant; 1H to 8H, Helminthosporium spot blotch resistant; 1S to 7S, Septoria tritici blotch resistant
Dendrogram from the Jaccard’s similarity coefficients based on 225 AFLP markers and 202 SSR bands over 32 synthetic hexaploid wheat accessions. Accessions 1F to 8F, Fusarium head blight resistant; 1FL and 2FL, Fusarium head blight and leaf rust resistant; 1L to 7L, leaf rust resistant; 1H to 8H, Helminthosporium spot blotch resistant; 1S to 7S, Septoria tritici blotch resistant
Discussion
The importance and usefulness of more than 1000 SHW accessions developed at CIMMYT in the genetic improvement of wheat for biotic and abiotic stress tolerances has been repeatedly emphasized (Rajaram and van Ginkel 2001; Mujeeb-Kazi and Rajaram 2002; Warburton et al. 2006; Das et al. 2007; Trethowan and Mujeeb-Kazi 2008; Mujeeb-Kazi et al. 2008; Ogbonnaya et al. 2013; Sharma et al. 2014). In the present study, AFLP and SSR marker data showed substantial genetic diversity among the 32 SHW accessions. The average polymorphism (99 %) for SSR used was much higher than that of AFLP (32 %). Lage et al. (2003) used a group of 54 SHW accessions for investigating genetic diversity using AFLP. Based on polymorphism, they reported a considerably higher level of AFLP diversity (39 %) than normally observed in cultivated hexaploid wheats (12–21 %), suggesting usefulness of SHWs in introducing new genetic diversity into bread wheat gene pool. In the present study, average PIC for AFLP and SSR were 0.24 and 0.66, respectively. Using SSR markers and 60 A. tauschii and 60 T. aestivum genotypes, Lelley et al. (2000) reported an average PIC of 0.68 for A. tauschii and that of 0.40 for T. aestivum. This result suggests greater genetic diversity in the D genome of A. tauschii than that of cultivated wheat. In the present study, genetic similarity based on Jaccard’s similarity coefficients ranged from 0.23 to 0.84 with an average coefficient of 0.57 for AFLP. Using 78 wheat landraces from 22 countries and 5 AFLP primer combinations Strelchenko et al. (2004) reported genetic similarity ranging from 0.43 to 0.98 with an average similarity of 0.69. Similarity coefficients for the 27 SSR markers in our study ranged from 0.018 to 0.97 with an average coefficient of 0.19. For the 20 D genome specific SSR, the similarity coefficients ranged from 0.02 to 1.0 with an average coefficient of 0.17. Using 19 SSR primer pairs and the 78 landraces of wheat, Strelchenko et al. (2004) reported genetic similarity ranging from 0.00 to 0.95 with an average similarity of 0.19. Although the genetic diversity based on AFLP markers was slightly higher in the present study as compared to the study of Strelchenko et al. (2004), the genetic diversity among the SHW accessions in the present study was similar to the genetic diversity among the wheat landraces in the Strelchenko et al. (2004) study based on SSR markers. The low level of genetic similarity between the SHW accessions reported in the present study based on the D genome specific markers indicate that most of the Ae. tauschi entries (Table 1) used to create these SHWs are genetically diverse.
Major allelic frequency for the 27 SSR markers ranged from 0.23 to 0.81 with an average allelic frequency of 0.45. This numbers are greater than an average allele frequency of 0.07 reported by Strelchenko et al. (2004). Number of alleles per locus ranged from three to 15 with an average allele number of 7.4. Strelchenko et al. (2004) reported a range from 4 to 21 for number of alleles per locus in wheat landraces using SSR. Cluster analyses (Figs. 1, 2, 3, 4) using UPGMA and Jaccard’s similarity coefficient showed distinct groups among the wheat accessions studied for both AFLP and SSR data. This indicated presence of substantial genetic diversity among the SHW accessions studied. Dendrogram based on the 20 D genome specific SSR markers showed that most of the SHW accessions are distantly related and thus the A. tauschii accessions used for those distantly related accessions are genetically diverse (Fig. 3).
Results indicated that both AFLP and SSR markers used in this study were effective for the assessment of genetic variation among the 32 SHW accessions studied. Mantel statistics between the distance matrices from AFLP and SSR analyses was significant (r = 0.52**) indicating that both types of markers were similar in grouping the SHWs. In the present study, the Mantel test between Jaccard’s similarity matrices from 27 SSR marker data and from the 20 D genome specific SSR marker data was also significant and high (r = 0.96**). Our findings suggest that both AFLP and SSR markers indicated presence of substantial genetic diversity among the SHW accessions studied. This greater level of genetic diversity among the SHWs can be efficiently used in the improvement of cultivated wheat. In a recent review on the current advances in research and utilization of SHWs, Li et al. (2014) emphasized that the abundant genetic diversity in SHWs can effectively make breakthroughs in genetic improvements of wheat through the inclusion of genetic diversity. However, considering the fact that SHWs carry large number of unfavorable traits (Li et al. 2014), the desirable traits need to be transferred into cultivated bread wheat lines through backcross breeding. Two of the 32 SHW accessions used in our study possess resistances for leaf rust and Fusarium head blight. By crossing the most diverse accessions from the 32 SHW accessions, pyramiding of the resistance genes for leaf rust, Fusarium head blight, Helminthosporium spot blotch and Septoria tritici blotch will be possible. Use of molecular markers will certainly aid in this process.
Conclusion
Over the past two decades there has been a surge in the search by researchers to use alien genetic resources for exploiting unique allelic diversity that could be harnessed for wheat improvement. As a result, wide crosses between durum wheat and the D genome diploid accessions created SHWs for transferring useful genes from these species into bread wheat using their “homologous pairing” potential. This allows the exchanges to take place across all three (ABD) genomes when the SHWs are crossed onto recipient conventional bread wheats. We have used just one sub-set of 32 accessions and in this lot have identified only 2 entries that have dual resistances. The intrinsic potential of SHWs needs to be explored further and elaborate phenotyping is a need to detect SHWs with multiple resistances or tolerances. Our study has mainly shown that excellent resistance to different diseases and genetic diversity are present and molecular tools can be used to efficiently select “diverse” SHW parents for wheat breeding targets. We conclude by expressing that an output balance of productivity must recognize the applied needs and these targets should dictate breeding efforts that use novel diversity. Using resources that are not utilized or are so far under-utilized need re-visiting as we move forward and face global wheat productivity challenges of huge proportions.
References
Ali A, Arshad M, Naqvi SMS, Rasheed A, Sher H, Kazi AG, Mujeeb-Kazi A (2015) Comparative assessment of synthetic-derived and conventional bread wheat lines under osmotic stress and implications for molecular analysis. Plant Mol Biol Rep. doi:10.1007/s11105.015-0884-8
Anderson JA, Churchill GA, Autrique JE, Tanksley SD, Sorrells ME (1993) Optimising parental selection for genetic linkage maps. Genome 36:181–186
Autrique E, Nachit MM, Monneveux P, Tanksley SD, Sorrells ME (1996) Genetic diversity in durum wheat based on RFLPs, morphophysiological traits, and coefficient of parentage. Crop Sci 36:735–742
Bai G-H, Ayele M, Tefera H, Nguyen H (1999) Amplified fragment length polymorphism analysis of tef [Eragrostis tef (Zucc.) Trotter]. Crop Sci 39:819–824
Barrett BA, Kidwell KK (1998) AFLP-based genetic diversity assessment among wheat cultivars from the Pacific Northwest. Crop Sci 38:1261–1271
Cooper JK, Ibrahim AMH, Rudd J, Hays D, Malla S, Baker J (2013) Increasing hard winter wheat yield potential via synthetic wheat: II. Heritability and combining ability of yield and its components. Crop Sci 53:67–73
Cox TS (1998) Deepening the wheat gene pool. J Crop Prod 1:1–25
Cox TS, Kiang YT, Gorman MB, Rodgers DM (1985) Relationship between coefficient of parentage and genetic similarity indices in soybean. Crop Sci 25:529–532
Das MK, Bai G-H, Mujeeb-Kazi A (2007) Genetic diversity in conventional and synthetic wheats with drought and salinity tolerance based on AFLP. Can J Plant Sci 87:691–702
del Blanco IA, Rajaram S, Kronstad WE (2001) Agronomic potential of synthetic hexaploid derived populations. Crop Sci 41:670–676
Jaccard P (1908) Nouvelles rescherches sur la distribution florale. Bull Soc Vaud Sci Nat 44:223–270
Lage J, Warburton ML, Crossa J, Skovmand B, Anderson SB (2003) Assessment of genetic diversity in synthetic hexaploid wheats and their Triticum dicoccum and Aegilops tauschii parents using AFLP and agronomic traits. Euphytica 134:305–317
Lage J, Skovmand B, Anderson SB (2004) Field evaluation of emmer wheat derived synthetic hexaploid wheats for resistance to Russian wheat aphid (Homoptera: Aphididae). J Econ Entomol 97:1065–1070
Lelley T, Stachel M, Grausgruber H, Vollman J (2000) Analysis of relationships between Aegilops tauschii and the D genome of wheat utilizing microsatellites. Genome 43:661–668
Li J, Wan HS, Yang WY (2014) Synthetic hexaploid wheat enhances variation and adaptive evolution of bread wheat in breeding processes. J Syst Evol 52(6):735–742
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Martin JM, Blake TK, Hockett EA (1991) Diversity among North American spring barley cultivars based on coefficient of parentage. Crop Sci 31:1131–1137
Mujeeb-Kazi A, Asiedu R (1990) Wide hybridization potential of alien genetic transfers for Triticum aestivum improvement. In: Bajaj YPS (ed) Biotechnology in agriculture and forestry, vol 13, Wheat, Springer Verlag, Berlin, pp 111–127
Mujeeb-Kazi A, Rajaram S (2002) Transferring alien genes from related species and genera for wheat improvement. In: Curtis BC, Rajaram S, Gomez Macpherson H (eds) Bread wheat: improvement and production. FAO, Rome, pp 199–215
Mujeeb-Kazi A, Rosas V, Roldán S (1996) Conservation of the genetic variation of Triticum tauschii (Coss.) Schmalh. (Aegilops squarrosa auct. non L.) in synthetic hexaploid wheats (T. turgidum L. s. lat. × T. tauschii; 2n = 6x = 42, AABBDD) and its potential utilization for wheat improvement. Genet Resour Crop Evol 43:129–134
Mujeeb-Kazi A, Gilchrist LI, Villareal RL, Delagado R (2000) Registration of 10 wheat germplasm lines resistant to Septoria tritici leaf blotch. Crop Sci 40:590–591
Mujeeb-Kazi A, Cano S, Rosas V, Cortes A, Delgado R (2001a) Registration of five synthetic hexaploid wheat and seven bread wheat germplasm lines resistant to wheat spot blotch. Crop Sci 41:1653–1654
Mujeeb-Kazi A, Cortes A, Rosas V, Cano J, Sanchez J, Juarez L, Delgado R (2001b) Genetic diversity for improving scab resistance in wheat. In: Proceedings of Warren Kronstad symposium CIMMYT, Mexico, pp 126–129
Mujeeb-Kazi A, Fuentes-Davila G, Villareal RL, Cortes A, Rosas V, Delgado R (2001c) Registration of 10 synthetic hexaploid wheat and six bread wheat germplasms resistant to karnal bunt. Crop Sci 41:1652–1653
Mujeeb-Kazi A, Gul A, Farooq M, Rizwan S, Ahmad I (2008) Rebirth of synthetic hexaploids with global implications for wheat improvement. Aust J Agric Res 59:391–398
Mujeeb-Kazi A, Kazi AG, Dundas I, Rasheed A, Ogbonnaya F, Chen P, Kishi M, Bonnett D, Wang R-C, Xu S, Bux H, Mahmood T, Farrakh S (2013) Genetic diversity for wheat improvement as a conduit for food security. Adv Agron 122:179–259
Ogbonnaya FC, Abdalla O, Mujeeb-Kazi A, Kazi AG, Xu SX, Gosman N, Lagudah ES, Bonnett D, Sorrells ME, Tsujimoto H (2013) Synthetic hexaploids: harnessing species of the primary gene pool for wheat improvement. J Plant Breed Rev 37:35–122
Pejic I, Ajmone-Marson P, Morgante M, Kozumplick V, Castiglioni P, Taramino G, Motto M (1998) Comparative analysis of genetic similarity among maize inbred lines detected by RFLPs, RAPDs, SSRs and AFLPs. Theor Appl Genet 97:1248–1255
Rajaram S, van Ginkel M (2001) Mexico, 50 years of international wheat breeding. In: Bonjean AP, Angus WJ (eds) The world wheat book: a history of wheat breeding. Lavoisier Publishing, Paris, pp 579–608
Rasheed A, Xia XC, Ogbonnaya FC, Mahmood T, Zhang Z, Mujeeb-Kazi A, He Z (2014) Genome wide association for grain morphology in synthetic hexaploid wheats using digital imaging analysis. BMC Plant Biol. doi:10.1186/1471-2229-14-128
Rohlf FJ (2000) NTSYS-pc. Numerical taxonomy and multivariate analysis system. ver. 2.1, Exeter software, Setauket, New York
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Sharma P, Sindhu S, Saini M, Verma A, Tyagi BS, Sharma I (2014) Assessing genetic variation for heat tolerance in synthetic wheat lines using phenotypic data and molecular markers. Aust J Crop Sci 8(4):515–522
Sneath PHA, Sokal RR (1973) Numerical taxonomy: the principles and practice of numerical classification. W H Freeman, San Francisco
Soleimani VD, Baum BR, Johnson DA (2002) AFLP and pedigree-based genetic diversity estimates in modern cultivars of durum wheat [Triticum turgidum L. subsp. durum (Desf.) Husn.]. Theor Appl Genet 104:350–357
Strelchenko P, Street K, Mitrofanova O, Mackay M, Balfourier F (2004) Genetic diversity among hexaploid wheat landraces with different geographical origins revealed by microsatellites: comparison with AFLP and RAPD data. In: Proceedings of the 4th international crop sci congress, Brisbane, Australia
Trethowan R, Mujeeb-Kazi A (2008) The use of novel germplasm resources to improve the environmental stress tolerance of hexaploid wheat. Crop Sci 48:1255–1265
Villareal RL, Mujeeb-Kazi A, Del Toro E, Crossa J, Rajaram S (1994) Agronomic variability in selected Triticum turgidum x T. tauschii synthetic hexaploid wheat. J Agron Crop Sci 173:307–317
Warburton ML, Crossa J, Franco J, Kazi M, Trethowan R, Rajaram S, Pfeiffer W, Zhang P, Dreisigacker S, van Ginkel M (2006) Bringing wild relatives back into the family: recovering genetic diversity in CIMMYT improved wheat germplasm. Euphytica 149(3):289–301
Xu SS, Friesen TL, Mujeeb-Kazi A (2004) Seedling resistance to tan spot and Stagonospora nodorum blotch in synthetic hexaploid wheat. Crop Sci 44:2238–2245
Yang W, Liu D, Li J, Zhang L, Wei H, Hu X, Zheng Y, He Z, Zou Y (2009) Synthetic hexaploid wheat and its utilization for wheat genetic improvement in China. J Genet Genom 36:539–546
Zhang PS, Dreisigacker AE, Melchinger M, van Ginkel M, Hoisington D, Warburton ML (2005) Quantifying novel sequence variation in CIMMYT synthetic hexaploid wheats and their backcross-derived lines using SSR markers. Mol Breed 12:1–10
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors do not have any conflicts of interest. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture.
Rights and permissions
About this article
Cite this article
Das, M.K., Bai, G., Mujeeb-Kazi, A. et al. Genetic diversity among synthetic hexaploid wheat accessions (Triticum aestivum) with resistance to several fungal diseases. Genet Resour Crop Evol 63, 1285–1296 (2016). https://doi.org/10.1007/s10722-015-0312-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-015-0312-9