Abstract
Juniperus procera is economically highly important but threatened tree species. It is the only species among 67 taxa in the genus Juniperus that naturally grows in Africa and south of the equator extending up to 18°S in Zimbabwe. Ethiopia is assumed to host the largest J. procera populations, which are also believed to have high genetic variation owing to their wide ecological amplitude. This study assessed genetic variation at AFLPs of J. procera populations in the Ethiopian highlands. In the study six populations, namely Chilimo, Goba, Menagesha-Suba, Wef-Washa, Yabelo and Ziquala were included. A total of 20–24 trees from each population were investigated based on 128 AFLP band positions. AMOVA revealed that most of the variation (94%) resided within populations of J. procera suggesting extensive gene flow among populations which is attributable to the outcrossing mating system and effective gene transport mechanisms of the species. However, genetic differentiation among populations was still significant (P < 0.05), and the differentiation was significantly (P < 0.05) correlated with geographic distance. All population pairs were significantly (P < 0.05) differentiated except for Menagesha-Suba and Wef-Washa. These two populations also showed the highest gene diversity (H j = 0.301 and H j = 0.297, respectively). These results are in accordance with historical records that claim the establishment of the Menagesha-Suba juniper population as plantation of seedlings from Wef-Washa back in fifteenth century.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The history of the Ethiopian highland forests witnessed a huge impact of human activities and climate variability and changes (Gebru et al. 2009; Darbyshire et al. 2003; Teketay 1992). These activities and events have changed the forest ecosystems of the region; species in these ecosystems are believed to have undergone enormous changes in their genetic composition. According to recent findings, the Ethiopian highland forest cover has undergone four major phases in the last 3,000 years: the pre-history Podocarpus dominant podo-juniper phase, man-induced decline in podo-juniper forest, forest recovery of the juniper dominant phase and the last human induced decline in forest cover (Darbyshire et al. 2003). A consequence of these phases is that nowadays the highlands of Ethiopia are characterized by highly fragmented small patches of remnant forests in remote and protected areas. Forest fragmentation is a significant threat to the maintenance of genetic resources (Broadbent et al. 2008; Tabarelli and Peres 2002), since it is often accompanied by an erosion of genetic variation and increased interpopulation genetic divergence due to increased random genetic drift, elevated inbreeding and reduced gene flow (Young et al. 1996).
High genetic variations for Ethiopian highland forest species such as Hagenia abyssinica (Bruce) J.F. Gmel., Cordia africana Lam., Lobelia giberroa Hemsl., and Coffea arabica L. have been reported by Ayele et al. (2009), Derero et al. (2010), Kebede et al. (2007) and Aga et al. (2005), respectively. The authors suggest both human and non-human impacts on genetic variation patterns and propose possible conservation strategies. For example, Ayele et al. (2009) and Aga et al. (2005) discuss human mediated gene transport as an important factor for the current genetic structure of H. abyssinica and C. arabica.
Juniperus procera Hochst. ex Endl. (Cupressaceae), commercially known as the African Pencil Cedar, is an evergreen tree that grows to a height of over 40 m and a diameter above 3 m (Pohjonen and Pukkala 1992; Negash 1995). J. procera is the only species among 67 taxa in the genus Juniperus that naturally grows in Africa. Its habitat stretches from the Arabian Peninsula in Asia to Zimbabwe in Africa (Adams 2004). J. procera is assumed to have evolved from J. excelsa M. Bieb. or from a common ancestor, and expanded southwards from the western mountains of the Arabian Peninsula to Ethiopia across the Red Sea and then further to the present southernmost populations in Zimbabwe (Adams 2004; Farjon 2005). The species grows in small scattered populations in most areas. It is one of the economically most important tree species in Ethiopia having favourable wood qualities such as fine texture, straight grains, workability and high resistance to termite and decay. It is used for a variety of purposes like constructions, furniture and outdoor works (Negash 1995; Bekele-Tesema 2007). Consequently, it is one of the tree species that has been subjected to extensive logging (Negash 1995), leading its status to endangered in some provenances (Garzulia 2006; Borghesio et al. 2004) and threatened in its entire habitat (IUCN 2009).
The Ethiopian highlands are described as the largest reservoir of the species (Bekele 1994; Yirdaw 2002). In Ethiopia, it is one of the dominant tree species in most remnant highland natural forests (Negash 1995). Historical evidences indicate that most of the northern Ethiopian highlands that are currently devoid of forests once were dominantly covered by J. procera (Logan 1946; White 1983; Friis 1992; Teketay 1992; Darbyshire et al. 2003). Today, remnants of the old stands have been reserved only around old churches and monasteries in these highlands (Bongers et al. 2006). These sanctuaries are presently the main source of information to reconstruct the historical forest status of the area (Wassie et al. 2005). In the Ethiopian highlands J. procera grows naturally in pure stands or mixed with Podocarpus falcatus (Thunb.) R. Br. ex Mirb. in lower altitudes (Negash 1995) and with Hagenia abyssinica at higher altitudes (Umer et al. 2007). J. procera is known to grow in Ethiopia as low as 1,700 m around Yabelo (Tigabu et al. 2007) and as high as over 3,300 m in the Bale Mountains (Umer et al. 2007).
Owing to its occurrence over a wide range of elevations, ecological heterogeneity is believed to have resulted in high genetic differentiation among populations of J. procera (Mamo et al. 2006; Demissie and Bjornstad 1997). On the other hand, obligatory outcrossing of this dioecious species (Negash 2002) and efficient gene flow via the wind-dispersed pollen and seed migration by frugivorous birds (Adams 2004) potentially decrease differentiation, but enhance high diversity within stands. Historical records indicate that Ethiopian highland forests in general and J. procera forests in particular have been subjected to over 3,000 years of anthropogenic influences (Gebru et al. 2009; Darbyshire et al. 2003; Teketay 1992). Anthropogenic gene transportation might have contributed remarkably to long distance seed dispersal and the establishment of Juniperus populations in the central highlands of Ethiopia in fifteenth century (Demissew 1988; Eshetu 2002).
Genetic variation of J. procera has rarely been studied. Mamo et al. (2006) and Tigabu et al. (2007) prove pronounced variation in seed morphology and germination percentage among Ethiopian populations of the species. However, these results do not allow to assess the variation of heritable traits of the species within or among populations.
Adams et al. (1993) included samples from Ethiopia (J. procera) and Greece (J. excelsa) to study past migration of Juniperus in Arabia and East Africa through RAPD fingerprints and terpenoids. However, diversity and differentiation within samples from Ethiopia was not analyzed. Mlangeni (2005) investigated genetic variation of J. procera from three countries including Ethiopia using AFLP markers. However, his study included only three populations from Ethiopia and low sample sizes, which makes it difficult to draw conclusions on the genetic structure of the species in Ethiopia.
The objectives of the present study are to assess genetic variation within and between populations of J. procera in Ethiopia, covering its wide geographic distribution and to explore the contribution of anthropogenic and natural causes for the current genetic structure of J. procera populations in Ethiopia.
Materials and methods
Study area
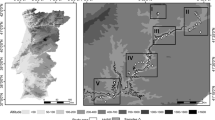
Samples were collected from six populations: Chilimo, Menagesha-Suba, Wef-Washa, Ziquala, Goba and Yabelo. Presence of natural barriers and historical information were considered for the selection of these populations (Table 1; Fig. 1). Chilimo, Menagesha-Suba and Wef-Washa represent populations in the central highland plateaus Northwest of the Rift Valley. On the other hand, Goba and Yabelo represent the Southeastern massif of the Southeastern and Southern highlands, respectively. Ziquala is an uplifted mountain within the Rift valley on the Northwestern side.
Sampling
From each population 20–24 trees were randomly sampled and needles (leaves) of each selected tree were collected for DNA analysis in separate plastic bags containing silica gel. In order to avoid the collection of related trees a minimum of 50 m distance was kept between trees within a population. Geographic coordinates of each population and distance from the preceding point were also measured using a GPS.
Laboratory analyses
Total genomic DNA was extracted from a small amount of dried needles (about 20 mg) following the DNeasy 96 Plant Kit protocol of the manufacturer (Qiagen, Hilden, Germany). AFLP reactions were performed as described by Vos et al. (1995) with minor modifications. The genomic DNA was digested with the two restriction enzymes EcoRI/MseI. The corresponding double-stranded (ds) EcoRI-adapter and MseI-adapter were ligated to the sticky ends of the restriction fragments to generate template DNA for PCR amplification. Fragments were amplified with two primer pairs complementary to the adapters and restriction site sequences with one to three additional “selective” nucleotides at their 3′-ends. The EcoRI + 1 primer (E01) and MseI + 1 primer (M03) with selective nucleotides A and G, respectively, were used for a pre-selective amplification. The primer combination E35 and M69 (Keygene code), each +3 primers with selective nucleotides of ACA and GCG, respectively were used for selective amplification (nomenclature of AFLP primer enzyme combinations,). Primer E35 was labeled with the fluorescent dye 6-FAM. All PCR reactions were performed in the Peltier Thermal Cycler (PTC-200 version 4.0, MJ Research). PCR-amplification products were diluted in proportion of 1:20. Then, 2 μl diluted product of each probe was mixed with 12 μl Hi-DiTM formamide and the internal size standard GeneScan 500 ROX (2 μl for 96 samples) from Applied Biosystems, and run on an automatic sequencer (ABI PRISM 3100 Genetic Analyser, Applied Biosystems). The size of the AFLP fragments was assessed with the software packages GeneScan 3.7 and Genotyper 3.7 (Applied Biosystems).
Data analysis
The reproducibility of amplification products was tested by performing the AFLP procedure and analyses twice for five samples. Only 100% reproducible fragments were considered. Accordingly, a total of 128 AFLP band positions within a readable range from 50 to 475 bps were selected and scored for band presence (1) and absence (0) at a particular position. Each band was manually monitored and edited. The band positions were transformed into a binary matrix, ‘0’ for absence and ‘1’ for presence of a fragment. The matrices were saved in excel format for further analyses.
Within population genetic diversity
The percentage of polymorphic loci (PPL) at the 5% level and Nei’s (1973) gene diversity (H j) were computed using the software AFLP-Surv 1.0 (Vekemans et al. 2002) following a Bayessian approach with non-uniform prior distribution of allele frequencies (Zhivotovsky 1999). Computation was made after 10,000 permutations and 1,000 bootstraps. Shannon’s index was estimated using the software POPGENE version 1.32 (Yeh et al. 2000).
Among population differentiation
Analysis of molecular variance (AMOVA) (Excoffier et al. 1992) and pairwise population differentiation were computed using Arlequin 3.11 (Excoffier et al. 2006) at 10,000 permutations. For AMOVA, populations were grouped into three groups based on their geographic location with reference of the Great Rift Valley (Goba and Yabelo-Southeast, Ziquala within and the rest Northwest/Central highlands). In order to estimate the magnitude of variation among populations contributed by individual markers, standardized genetic differentiations (G′ST) at each band positions were calculated according to Hedrick (2005) from Nei’s (1973) coefficient of genetic differentiations (G ST) of each band computed in POPGENE version 1.32 (Yeh et al. 2000) and pattern of changes in G ST versus G′ST values are shown in Fig. 2. Frequency distributions of ten AFLPs with the highest G′ST values are shown in Fig. 3. Nei’s (1978) genetic distance was computed using AFLP-Surv 1.0 from which an UPGMA (Unweighted pair group method using arithmetic averages) dendrogram was generated using the method of SAHN clustering in NTSYSpc 2.0 (Rohlf 1998). In order to estimate the influence of geographic distance on genetic distance between population pairs, a Mantel test (Mantel 1967) was performed using GenAlEx6.3 (Smouse et al. 2008; Peakall and Smouse 2006).
Results
Within population genetic diversity
The lowest (H j = 0.23049) and the highest (H j = 0.31874) gene diversities, congruently Shannon indices were observed in Goba and Menagesha-Suba populations, respectively. The lowest PPL (49%) was computed for Goba. Generally, populations South East of the Rift valley (Goba and Yabelo) showed lower PPL and gene diversity than other populations (Table 2).
Among population variation
AMOVA
The AMOVA showed that only about 6% of the total variation was attributable to variation among populations (Table 3). A small, but significant amount of the genetic variation (3.16%) was distributed among geographic regions.
Pairwise population differentiation
Significant (P < 0.05) genetic differentiation was observed for all pairwise comparisons in the F ST analysis. Only populations Menagesha-Suba and Wef-Washa from the same geographic region showed no significant differentiation (P = 0.32031) (Table 4). Most of the population pairs showed highly significant differentiation (P < 0.01). The highest genetic differentiation (F ST = 0.12758) was observed between Yabelo from the Southeast and Chilimo from the Northwest.
Population clustering
The Mantel test revealed a significant correlation (P < 0.05) between geographic distances and pairwise genetic distances (Fig. 4). The UPGMA dendrogram based on Nei’s (1978) genetic distance also reflected the correlation between geographic and genetic distances (Fig. 5). However, the relatively distant northwestern populations (Fig. 1) Menagesha-Suba and Wef-Washa grouped together.
Markers contributing to high genetic differentiation
Out of the 128 markers ten had G′ST above 0.27. South eastern populations (Goba and Yabelo) tended to be unique in harboring 94 and 392 bps size markers at a higher frequency, a 51 bp marker at a relatively lower frequency and lacking the 276 bp marker. Yabelo further lacked the marker with the highest G′ST of 112 bp (Fig. 3).
Discussion
Within population variation and overall differentiation
The high within population variation and low population differentiation is most likely attributable to the reproductive biology and gene transportation mechanism of the species. J. procera is an obligatory outcrossing species (Negash 1995; Adams 2004; Farjon 2005) with efficient means of gene dispersal by pollen and seed; this promotes heterozygosity and the maintenance of high within population genetic diversity (White et al. 2007), but results in low differentiation among populations. The results obtained in this study are in accordance with many earlier studies that have reported high within population genetic variation as typical genetic structure for dioecious species (Billington 1991; Hamrick and Godt 1996; Weidema et al. 2000; Gerard et al. 2004). The production of large amounts of pollen by J. procera (Coetzee 1967) and the formation of pure stands in Ethiopia (Negash 1995) promote the efficiency of wind pollination.
Unlike most other conifers that produce stony cones, juniper seeds (except J. drupacea Labill.) are frequently dispersed by frugivorus birds (Adams et al. 1993; Adams 2004) which further contributes to gene exchange among distant populations. The relatively higher gene diversity of Ziquala, which is located on an uplifted isolated mountain in the rift valley, is possibly linked to the high movement of birds that have promoted reproductive contact of this stand with other populations (Adams 2004). Ziquala is known as haven of many bird species (BirdLife International 2009, available on http://www.birdlife.org).
In accordance with the present results, Mlangeni (2005) observed high within population genetic variation investigating samples of J. procera that originated from different countries including Ethiopia based on AMOVA of 264 AFLP markers which ranged from 83.32% for Kenyan populations to 98.65% for Malawian populations and where the Ethiopian population scored 85.34%.
Contribution of geographic distance and natural barriers for population genetic variability
Although the majority of the genetic variation resided within populations, the AMOVA revealed that there was significant (P < 0.05) variation among populations as well (Table 3). The highest differentiation (F ST = 0.1278, Table 4) was realized between Yabelo and Chilimo; which is in agreement with the long geographic distance between these two populations. Yabelo is the most divergent population as shown in the UPGMA dendrogram (Fig. 5). This population also appeared unique lacking an AFLP fragment of 112 bp that is most frequent at Chilimo and also present in all the other populations (Fig. 3). The strong differentiation of the Yabelo population from other populations is in line with the findings of Tigabu et al. (2007) and Mamo et al. (2006) who observed a high germination rate and unique morphology of Juniperus seeds from this population.
The significant (P < 0.05) variation among groups based on their location with reference to the Great Rift Valley suggests limited gene flow across this natural barrier. This view is supported by the strong differentiation of southeastern population (Goba and Yabelo) from the other populations on the northwestern side of the valley.
Yabelo has the second lowest gene diversity possibly attributable to its geographical isolation from other J. procera populations. The Yabelo population is considered as ‘lowland’ juniper in Ethiopia (Tigabu et al. 2007) where the species is less frequent. Similarly, Chilimo harbored a comparatively low genetic diversity as compared to the other populations in the central highlands reflecting its marginal geographic position for species of dry evergreen forests such as J. procera (Negash 1995; Couralet et al. 2005).
Anthropogenic impact on genetic variability of J. procera
The Ethiopian highland forest cover has passed different phases of alterations due to human activities (Darbyshire et al. 2003). According to recent findings, gymnosperms (P. falcatus and J. procera) dominated forests in the Ethiopian highlands since 7,000 years ago where these species have been enormously subjected to human influences (Gebru et al. 2009). In the present study, the genetic similarity between Menagesh-Suba and Wef-Washa populations (Table 4) clearly illustrates anthropogenic effects on the genetic structure of J. procera in the Ethiopian highlands. Despite their isolation by larger geographic distance, they show a very low genetic differentiation (F ST = 0.00244). According to written and oral records, Menagesha-Suba was established as plantation of wildlings from Wef-Washa in the fifteenth century by an Abyssinian king Zere-Yacob (Gilbert 1970; Demissew 1988). Through carbon isotopes analyses, Eshetu and Högberg (2000) confirmed that J. procera at Menagesha-Suba was originally established as plantation over 550 years ago. However, their study gave no indication from where exactly the original planting material originated. Despite all efforts to trace the origin of this plantation, the aforementioned historical records could neither be supported nor rejected so far. Hence, our results appear to be the first to confirm century-years old records claiming Wef-Washa as seed source of Juniperus at Menagesha-Suba.
The high genetic diversity in the presumably planted population Menagesha-Suba (Table 2) is likely to be due to a high number and diversity of the founder plants during establishment (Tang et al. 2003; Parisod et al. 2005). The Menagesha-Suba population has been protected since the 1600s by Imperial decrees (Demissew 1988) until to date as State forest which enabled the occurrence of refuge trees as old as about 600 years (Eshetu and Högberg 2000; Eshetu 2002), young regeneration and hence the presence of all the age classes in this population. Less disturbed populations harbor higher genetic diversity than populations subjected to frequent disturbances (Lefevre et al. 2004; White et al. 2007). Anthropogenic gene transportation might also have contributed to the relatively high genetic diversity at Ziquala, a population under an old and big Monastery, which was reportedly established in the late twelfth century (Tamene 1998). It has very likely been visited by monks of other ancient monasteries that are sanctuaries of tree species in the Northern highlands (Bongers et al. 2006).
Even though the Goba population is in close distance to other populations of the Bale Mountains (Malcolm and Evangelista 2005; Umer et al. 2007) which might have favored higher gene diversity than in smaller and fragmented populations (Hamrick et al. 1992; Finkeldey and Hattemer 2007; White et al. 2007), it had the lowest gene diversity and a very low proportion of polymorphic loci (Table 2). On the other hand, the highest gene diversity was observed in this region for other species such as for Hagenia abyssinica (Ayele et al. 2008), Lobelia gimberroa (Kebede et al. 2007) and Coffea arabica (Aga et al. 2005). The Goba population has been highly disturbed by human activities as compared to the other population analyzed in this study. The proximity of this population to the Goba town might contribute for its sever disturbance. Thus, the Goba population possibly has experienced population bottlenecks (Nei et al. 1975) due to excessive disturbances.
Genetic structure of J. procera and its implication for conservation
Efficient mechanisms of gene dispersal via pollen and seed contributed to the maintenance of high genetic diversity within populations in the past, but significant genetic differentiation, a significant correlation between spatial and geographic distances, and evidence for natural barriers limiting gene dispersal was also observed. Accordingly, potentially unique genetic information might be harbored at least in some of the populations. This calls for conservation activities in multiple populations with special focus on populations characterized by high genetic differentiation. The result suggests to consider the geographic location and environmental barriers to set conservation priorities where genetic data are lacking.
The low diversity observed in the Goba population suggests that anthropogenic activities leading to heavy population disturbances can affect the genetic composition of the species considerably. Anthropogenic activities therefore appear to be potential threats for the loss of genetic information particularly in spatially isolated small populations where genetic drift is possible. Gene flow from larger populations possibly enhances diversity in disturbed neighboring populations. The occurrence of large juniper forests close to the Goba population suggests the possibility to restore its genetic diversity by natural processes (gene flow of pollen and seed migration) if conservation measures allowing natural regeneration are implemented.
References
Adams RP (2004) Juniperus of the world: The genus Juniperus. Trafford Publishing, Victoria, pp 275
Adams RP, Demeke T, Abulfatih HA (1993) RAPD DNA fingerprints and terpenoids: clues to past migrations of Juniperus in Arabia and east Africa. Theor Appl Genet 87:22–26
Aga E, Bekele E, Bryngelsson T (2005) Inter-Simple Sequence Repeat (ISSR) variation in forest coffee trees (Coffea arabica L.) populations from Ethiopia. Genetica 124:213–221
Ayele TB, Gailing O, Umer M, Finkeldey R (2009) Chloroplast DNA haplotype diversity and postglacial recolonization of Hagenia abyssinica (Bruce) J.F.Gmel. in Ethiopia. Plant Syst Evol 280:175–185
Bekele T (1994) Phytosociology and ecology of a humid afromontane forest on the central plateau of Ethiopia. J Veg Sci 5:87–98
Bekele-Tesema A (2007) Useful trees and shrubs for Ethiopia: identification, propagation and management for agricultural and Pastoral Community. World Agroforestry Center, Nairobi, pp 559
Billington HL (1991) Effect of population size on genetic variation in a dioecious conifer. Conserv Biol 5:115–119
BirdLife International (2009) Important bird area factsheet: Mount Zuquala, Ethiopia. Downloaded from the Data Zone at http://www.birdlife.org on 30 Dec 2009
Bongers F, Wassie A, Sterck FJ, Bekele T, Teketay D (2006) Ecological restoration and church forests in northern Ethiopia. J Drylands 1:35–44
Borghesio L, Giannetti F, Ndang’ang’a K, Shimelis A (2004) The present conservation status of Juniperus forests in the South Ethiopian Endemic bird area. Afr J Ecol 42(2):137–143
Broadbent EN, Asner GP, Keller M, Knapp DE, Oliviera PJC, Silva JN (2008) Forest fragmentation and edge effects from deforestation and selective logging in the Brazilian Amazon. Biol Conserv 141:1745–1757
Coetzee JA (1967) Pollen analytical studies in East and Southern Africa. Palaeoecol Afr 3:1–146
Couralet C, Sass-Klaassen U, Sterck F, Bekele T, Zuidema PA (2005) Combining dendrochronology and matrix modeling in demographic studies: an evaluation for Juniperus procera in Ethiopia. For Ecol Manag 216:317–330
Darbyshire I, Lamb H, Umer M (2003) Forest clearance and re-growth in northern Ethiopia during the last 3000 years. Holocene 13:537–546
Demissew S (1988) The floristic composition of the Menagesha state forest and the need to conserve such forests in Ethiopia. Mt Res Dev 8:243–247
Demissie A, Bjornstad A (1997) Geographical, altitude and agro-ecological differentiation of isozyme and hordein genotypes of landrace barleys from Ethiopia: implications to germplasm conservation. Genet Resour Crop Evol 44:43–55
Derero A, Gailing O, Finkeldey R (2010) Maintenance of genetic diversity in Cordia africana Lam., a declining forest species in Ethiopia. Tree Genet Genomes. doi:10.1007/s11295-010-0310-1
Eshetu Z (2002) Historical C3–C4 vegetation pattern on forested mountain slopes: its implication for ecological rehabilitation of degraded highlands of Ethiopia by afforestation. J Trop Ecol 18:743–758
Eshetu Z, Högberg P (2000) Reconstruction of forest site history in Ethiopian highlands based on 13C natural abundance of soils. Ambio 29:87–89
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA aplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Excoffier L, Laval G, Schneider S (2006) An integrated software package for population genetics data analysis; Arlequin Ver. 3.1 user manual. University of Berne, Switzerland. http://cmpg.unibe.ch/software/arlequin3
Farjon A (2005) A monograph of Cupressaceae and Sciadopitys. Royal Botanic Gardens, Kew, pp 648
Finkeldey R, Hattemer HH (2007) Tropical forest genetics. Springer, Berlin, pp 315
Friis I (1992) Forests and forest trees of Northeast Tropical Africa: their natural habitats and distribution patterns in Ethiopia, Djibouti and Somalia. Kew Bulletin, London, pp 396
Garzulia M (2006) Threatened, endangered and vulnerable tree species: a comparison between FRA 2005 and the IUCN red list. FAO, Forestry Department, working paper 108/E, Rome, Italy
Gebru T, Eshetu Z, Huang Y, Woldemariam T, Strong N, Umer M, DiBlasi M, Terwilliger VJ (2009) Holocene palaeovegetation of the Tigray Plateau in northern Ethiopia from charcoal and stable organic carbon isotopic analyses of gully sediments. Palaeogeogr Palaeoclimatol Palaeoecol 282:67–80
Gerard J, Oostermeijer B, Knegt B (2004) Genetic population structure of the wind-pollinated, dioecious shrub Juniperus communis in fragmented Dutch heathlands. Plant Species Biol 19:175–184
Gilbert EF (1970) Mount Wachacha: a botanical commentary. Walia 2:3–12
Hamrick JL, Godt MJW (1996) Effects of life history traits on genetic diversity in plant species. Trans R Soc Lond B 351:1291–1298
Hamrick JL, Godt MJW, Sherman-Broyles SL (1992) Factors influencing levels of genetic diversity in woody plant species. New For 6:95–124
Hedrick PW (2005) A standardized genetic differentiation measure. Evolution 59:1633–1638
IUCN (2009) Juniperus procera: red list of threatened species. Conifer Specialist Group Report, 1998
Kebede M, Ehrich D, Taberlet P, Nemomissa S, Brochmann C (2007) Phylogeography and conservation genetics of giant lobelia (Lobelia giberroa) in Ethiopia and Tropical East African Mountains. Mol Ecol 16:1233–1243
Lefevre F, Fady B, Rubio DF, Ghosn D, Bariteau M (2004) Impact of founder population, drift and selection on the genetic diversity of a recently translocated tree population. Heredity 93:542–550
Logan WEM (1946) An introduction to the forests of Central and Southern Ethiopia. Imperial Forestry Institute of Oxford Paper, vol 24, pp 58
Malcolm J, Evangelista PH (2005) The range and status of Mountain Nyala: case study in Bale Mountains, Ethiopia, pp 42 Also be accessed www.ethiopianwolf.org/publications
Mamo N, Mihretu M, Fekadu M, Tigabu M, Teketay D (2006) Variation in seed and germination characteristics among Juniperus procera populations in Ethiopia. For Ecol Manag 225:320–327
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Mlangeni ET (2005) Genetic diversity and population structure in the East African Pencil Cedar, Juniperus procera (Cupressaceae). MSc thesis, Natural History Muesuem, Botanical Garden, University of Oslo
Negash L (1995) Indigenous trees of Ethiopia: biology, uses and propagation techniques. SLU Rprocentralen, Umea, pp 285
Negash L (2002) Successful vegetative propagation techniques for threatened African Pencil Cedar (Juniperus procera Hochst.ex Endl.). For Ecol Manag 161:53–64
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci 70:3321–3323
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Nei M, Maruyama T, Chakraborty R (1975) The bottleneck effect and genetic variability in populations. Evolution 29:1–10
Parisod C, Trippi C, Galland N (2005) Genetic variability and founder effect in the Pitcher Plant Sarracenia purpurea (Sarraceniaceae) in populations introduced into Switzerland: from inbreeding to invasion. Ann Bot 95:277–286
Peakall R, Smouse PE (2006) GenAlEx 6: genetic analysis in excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pohjonen V, Pukkala T (1992) Juniperus procera Hochst. ex Endl. in Ethiopian forestry. For Ecol Manag 49:75–85
Rohlf FJ (1998) NTSYSpc. Numerical taxonomy and multivariate analysis system. Version 2.0. User guide: exeter software. Setauket, New York, pp 31
Smouse PE, Peakall R, Gonzales E (2008) A heterogeneity test for fine-scale genetic structure. Mol Ecol 17:3389–3400
Tabarelli M, Peres CA (2002) Abiotic and vertebrate seed dispersal in the Brazilian Atlantic forest: implication for forest regeneration. Biol Conserv 106:165–176
Tamene G (1998) Features of the Ethiopian Orthodox Church and the clergy. Asian Afr Stud 7:87–104
Tang T, Zhong Y, Jian S, Shi S (2003) Genetic diversity of Hibiscus tiliaceus (Malvaceae) in China assessed with AFLP markers. Ann Bot 92:409–414
Teketay D (1992) Human impact on a natural montane forest in Southeastern Ethiopia. Mt Res Dev 12:393–400
Tigabu M, Fjellström J, Odén PC, Teketay D (2007) Germination of Juniperus procera seeds in response to stratification and smoke treatments, and detection of insect-damaged seeds with VIS + NIR spectroscopy. New For 33:155–169
Umer M, Lamb HF, Bonnefill R, Le′zin AM, Tierceline JJ, Gibert E, Cazet JP, Watrin J (2007) Late Pleistocene and Holocene vegetation history of the Bale Mountains, Ethiopia. Quat Sci Rev 26:2229–2246
Vekemans X, Beauwens T, Lemaire M, Roldan-Ruiz I (2002) Data from amplified fragment length polymorphism (AFLP) markers show indication of size homoplasy and of a relationship between degree of homoplasy and fragment size. Mol Ecol 11:139–151
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acid Res 23(21):4407–4414
Wassie A, Teketay D, Powell N (2005) Church forests in North Gonder Administrative Zone, Northern Ethiopia. For Trees Livelihoods 15:349–374
Weidema IR, Magnussen LS, Philipp M (2000) Gene flow and mode of pollination in a dry-grassland species, Filipendula vulgaris (Rosaceae). Heredity 84:311–320
White F (1983) The vegetation of Africa, a descriptive memoir to accompany the UNESCO/AETFAT/UNSO vegetation map of Africa (3 Plates, Northwestern Africa, Northeastern Africa, and Southern Africa, 1:5,000,000). UNESCO, Paris
White TL, Adams WT, Neale DB (2007) Forest genetics. Oxford University Press, Oxford, pp 500
Yeh F, Yang RC, Boyle T (2000) POPGENE (Version 1.32), Microsoft windows-based freeware for population genetic analysis. Retrieved from http://www.ualberta.ca/~fyeh/pr01.htm
Yirdaw E (2002) Restoration of the native woody-species diversity, using plantation species as foster trees, in the degraded highlands of Ethiopia: PhD dissertation. Faculty of Agriculture and Forestry of the University of Helsinki
Young A, Boyle T, Brown T (1996) The population consequences of habitat fragmentation for plants. Trends Ecol Evol 11:413–418
Acknowledgments
The German Academic Exchange Service (DAAD) is highly appreciated for financial support to the first author. We thank Olga Artes, Oleksandra Dolynska and Gerold Dinkel for technical support during the laboratory work. We also thank Mulualem Tigabu for his valuable comments and Demeke Nigussie for his assistance in plotting the populations on Ethiopian map. The inestimable suggestions by the anonymous reviewers are gratefully acknowledged.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Sertse, D., Gailing, O., Eliades, NG. et al. Anthropogenic and natural causes influencing population genetic structure of Juniperus procera Hochst. ex Endl. in the Ethiopian highlands. Genet Resour Crop Evol 58, 849–859 (2011). https://doi.org/10.1007/s10722-010-9623-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-010-9623-z