Abstract
The Emilia region (Northern Italy) is characterised by the occurrence of microclimates that permit olive growing. The presence of the species, albeit sporadic, in these territories for several centuries as a fruit crop is well documented, by both archaeological and written testimony, and by a large number of plants well over a century old, located in particular sites, favourable for growth and development of the tree.
Olive genetic diversity was studied using RAPD and SSR techniques, on plants growing in the Emilia territory (Reggio Emilia and Parma provinces). For genotype identification comparisons were made with 8 cultivars, some of which from Central Italy. Screening was obtained analysing patterns produced by 20 RAPD primers and 3 SSR primers, developed by other authors; the primers and we were able to discriminate olive cultivars with a sufficient degree of reliability. The dendrograms obtained from the analysis show the genetic relationship among accessions present in the Parma-Reggio Emilia district. Our results demonstrated the reliability of RAPDs and SSRs to identify all studied olive cultivars and to reveal the degree of their relatedness to each other. The analysis also reveals the presence of an interesting amount of genetic diversity among the studied individuals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Olive (Olea europaea L.) is, among cultivated fruit trees, the richest in genetic diversity. The longevity of this species and the low breeding pressure has contributed to the conservation of its variability, and the reduced extent of genetic erosion within its germplasm has allowed the persistence of olive diversity (Rallo et al. 2000). This is why the number of known cultivars is very high and has steadily increased since ancient times, especially since the more recent progresses in taxonomy and breeding. The described olive germplasm amounts to over 1,200 cultivars, not to mention a large number of synonyms and homonyms (Bartolini et al. 2005).
Italian germplasm alone includes about 48% of the known world germplasm (about 600 cultivars); in addition, numerous ancient trees are waiting to be identified and characterised (Bartolini et al. 2005).
The Italian olive industry is mostly spread in the Southern and Central regions of the Country, reaching Liguria and Tuscany at the far north edge of the traditional growing area; but olive is also present in restricted northern areas. Numerous historical sources date olive introduction in Northern Italy to the late empire times (Bignardi 1978), and olive industry has almost uninterruptedly existed along the piedmont belt of Emilia Romagna (Bianchedi 1880).
Emilia Romagna region is located in Northern Italy at about 44°–45°N and 11°–12°E. This region includes areas having microclimate similar to that of traditional olive growing areas of Central Italy.
The most visible evidence of the presence of olive as a cultivated fruit crop is the presence of a huge number of olive plants, stumps, bushy regrowths which were able to survive for centuries against the offences of time, environment and man. In these areas, mainly hilly environments, a number of olive varieties adapted and/or differentiated to constitute a rich and varied germplasm.
The study and characterisation of olive genetic diversity in Emilia is of great interest, because it might represent an interesting resource for breeding, with reference to biotic and abiotic factors of the olive environment, as well as an enrichment of the olive historical and cultural heritage.
Genetic variability can be evaluated in several ways (Ganino et al. 2006): morphology (Barranco et al. 2000), enzymatic studies (Loukas and Krimbas 1983; Truijllo et al.1995), RFLP markers (Gallitelli et al. 1991; Amane et al.1999; Besnard and Bervillè 2000), RAPD markers (Fabbri 1995; Belaj et al. 2004a; Ozkaya et al. 2006), AFLP markers (Angiolillo et al.1999; Sanz-Cortes et al. 2003; Owen et al. 2005), SSR markers (Rallo et al. 2000; Cipriani et al. 2002; Belaj et al. 2004b), SNPs (Diaz Bermudez 2005), chloroplast and mitochondrial DNA analysis (Amane et al. 1999; Besnard and Bervillè 2000; Besnard et al. 2002), Artificial Neural Network analysis (Mancuso and Nicese 1999), and NMR analysis of olive oils (Vlahov et al. 1999; Fragaki et al. 2005).
Our research had the purpose to characterise the germplasm of Emilia, and particularly of the hilly areas of the provinces of Parma and Reggio Emilia, by using molecular methods. In this study 109 olive accessions were analysed to describe the genetic diversity in Parma and Reggio Emilia provinces, and to highlight any genetic relationship among the local accessions and some national cultivars. The final goal of the research was to investigate on the putative origin of Emilian genotypes.
Materials and methods
Plant material
A large number of ancient olive trees was retrieved along the piedmont zone of the emilian region, in the 1997–2004 period (Fabbri and Ganino 2003); although no specific age determination could be obtained, the trees selected for the study were all very old, or visibly deriving from very ancient stumps, indicating an age of the plant of more than one century; however, the real age of the sampled trees was in most cases in the 2–3 centuries range or more. Most trees were located at an altitude from 150 to 600 m with exposure south-east, south, or south-west. Fifty-eight accessions were found in the province of Parma and forty-three in the province of Reggio Emilia. These accessions were considered and compared with eight Italian cultivars (Table 1). Most of the accessions were abandoned, i.e., untended trees, located in woodlands, maquis, or at the edge of cultivated plots.
The accessions were tagged with an identification code and with the abbreviation of province (PR or RE).
The eight Italian genotypes were obtained from the World Olive Germplasm Bank of the Istituto Sperimentale per l’Olivicoltura of Rende (CS), Italy, and from the Collection created by Parma University in Salsomaggiore Terme (PR), with material originating from other research institutions.
The Italian cultivars were mainly from Tuscany and were chosen according to historical information indicating past imports of plants and oil from Tuscany (Bianchedi 1880). Four cultivars (Frantoio, Leccino, Cassanese, Carolea) were selected as inner controls in order to obtain a preliminary comparison with these genotypes, representatives of the Italian germplasm, and to get a varietal standard to insert in every analysis to check the correctness of PCR amplification and of sequencer analysis.
Moreover, during collection an additional control was added to assess the reliability of the RAPD and SSR markers by harvesting two leaf samples from the same tree (C1 and C2).
The collection of young leaves was carried out between April and May (intense vegetative growth). All plant material showing phytosanitary problems was discarded; care was taken not to collect shoots emerging from the tree stump, for fear of harvesting rootstock material.
DNA extraction
After collection the leaves were stored at −80°C; genomic DNA was extracted following Gounaris method (Gounaris et al. 2002), with minor modifications. DNA was quantified by spectrophotometric method (Spectrophotometer Uvikon 930, Kontron Instruments Inc., Boston, MA, USA).
RAPD analysis
The genomic DNA extracted was amplified by using 20 decamer primers (Table 2), previously used on international germplasm (Fabbri et al. 1995) and in another study on Emilian olive germplasm (Ganino and Fabbri 2005), showing a good discriminating ability.
The amplification reaction was obtained in a volume of 25 μl containing: 1× Reaction buffer (International PBI, Milan, IT), 1.9 mM MgCl2, 2.5 μM dNTPs (Amersham Biosciences, Amersham Place, UK), 0.4 μM of primer (Sigma-Genosys Ltd, Dorset, GB), 1 unit of taq DNA polymerase (International PBI, Milan, IT), and 40 ng DNA.
The amplification reaction was optimised in a thermal cycler MJ PCT 100 (MJ Research, Watertown, Mass.), programming a first step at 95°C for 5 min followed by 40 cycles of 40 s at 92°C, 40 s at 36°C, 2 min at 72°C, for denaturation, annealing, and primer extension; the last step included 10 min of incubation at 72°C. The reaction, for each oligonucleotide utilised, was performed three times.
The products of amplification were separated on agarose gel at 1.5% in TAE buffer and stained with ethidium bromide. The gels were photographed with a Kodak DC40 digital camera. The marker Gene Ruler 100 bp DNA Ladder plus (M—Medical, Milan, IT) was used to estimate the approximate molecular weight of the amplified products, with the aid of the Kodak digital science 1D Image Analysis Software (Eastman Kodak Company, Rochester, NY).
RAPD data analysis
We used Jaccard’s similarity coefficient (Jaccard 1908) to discriminate between accessions and cluster analysis (UPGMA) with the Sin-Tax 2000 software (Podani, Department of Plant Taxonomy and Ecology, Budapest, Hungary).
SSR analysis
SSR assays were performed as described by Sefc et al. (2000), Cipriani et al. (2002), Marrazzo et al. (2002) and Carriero et al. (2002). In this study we have utilised 3 primers chosen among those tested by the above authors and which, according to the same authors, were best as polymorphism, absence of multiple bands, discrimination power. DNA amplification was carried out by using ssrOeUA-DCA3 (Sefc et al. 2000), UDO-043 (Cipriani et al. 2002; Marrazzo et al. 2002), GAPU101 (Carriero et al. 2002).
The amplification reaction was obtained in a volume of 25 μl containing: 10 mM Tris HCl, 50 mM KCl buffer, 1.5 mM MgCl2, (International PBI, Milan, IT), 0.2 mM dNTPs (Amersham Biosciences, Amersham Place, UK), 0.2 μM primers (MWG Biotech, Ebersberg, Germany), 20 ng genomic DNA, 0.4 U of Taq polymerase (International PBI, Milan, IT).
The amplification reaction was optimized in a thermal cycler MJ PCT 100 (MJ Research, Watertown, Mass.), programming a first step at 95°C for 5 min followed by 35 cycles of 45 s at 94°C, 45 s at 57°C, 45 s at 72°C, for denaturation, annealing, and primer extension; the last step included 8 min of incubation at 72°C. The reaction, for each oligonucleotide utilized, was performed three times.
The products of amplification were analysed by a DNA sequencer CEQ 2000 Genetic Analysis System on acrylamide CEQ separation gel LPA-1 (Beckman Coulter, Inc., Fullerton, CA, USA). The marker CEQ DNA Size Standard kit 400 (Beckman Coulter, Inc., Fullerton, CA, USA) was used to estimate the approximate molecular weight of the amplified products.
SSR data analysis
The fragments were sized using a conservative binning approach (Kirby 1990) using statistical R software base (R Development Core Team 2005), which takes into account the type of repeat and compensates for the limits of fragment resolution. A brief description of the approach is given in Matsuoka et al. (2002). Population biodiversity was evaluated by number of alleles, by their frequency and by observed and expected heterozygosity (H ○ and H e, respectively) (Nei 1973) using the software Identity 1.0 (Wagner and Sefc 1999) available at the URL http://www.boku.ac.at/zag/forsch/identity.htm. We calculated Dice’s similarity coefficient (Dice 1945) using software GenAlEx6 (Peakall and Smouse 2005) and we performed on distance matrix a cluster analysis using UPGMA method with the aid of Philip 3.65 software (Felsenstein 2005).
Results
RAPD polymorphism
Of the 20 RAPD oligonucleotides belonging to the AI, AH, OPA, OPC, OPX and OPK series, only 19 were selected which were able to successfully amplify, by producing polymorphic and reproducible amplification patterns (Table 3 and Fig. 1). Primer 544 only produced 5 monomorphic fragments, but no polymorphic fragments. The 19 selected oligonucleotides amplified a total of 365 RAPD fragments, only 20% of which resulted to be monomorphic. The analysis was carried out considering the most easily detectable and reproducible polymorphic amplification products; hence, the fragments with such characteristics were chosen as markers. The number of polymorphic fragments found per primer was between 7 (AH02 and AH17) and 23 (OPX01) (Table 3), and their size ranged from 150 to 2,900 bp.
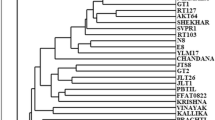
The dendrogram obtained from our data is shown in Fig. 2A. Within the 109 examined accessions the RAPD markers identified 26 different genotypes, and 83 synonyms. Of the 26 genotypes, 16 show a similarity index greater than zero (9PR, 16PR, 17PR, 44PR, 55PR, 32RE, 34RE, 36RE, 40RE, 42RE, 56PR, 58PR, A, C, D, LC).
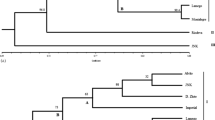
Dendrograms of 109 Olea europaea L. genotypes, generated with Jaccard similarity coefficient and UPGMA cluster analysis for the RAPD dendrogram (A) and by utilizing the Dice similarity coefficient for the SSR dendrogram (B). The small letters (a–k) indicate the genotype groups with high degree of similarity index
SSR polymorphism
The 3 oligonucleotides belonging to the series ssrOeUA-DCA, UDO and GAPU produced polymorphic and reproducible amplification fragments and permitted the discrimination of the 109 analysed accessions. They produced a sum total of 24 alleles with an average number of 8.0 alleles per locus (Table 4), ranging from a minimum of 6 for DCA3 to a maximum of 11 for GAPU101. The index of genetic variation (heterozygosity, H e) was high, ranging from a minimum of 0.684 for UDO43 to a maximum of 0.829 for GAPU101. Because of the negligible presence of null alleles, the heterozygosity reported here can be considered as a correct representation of the heterozygosity of the olive sample.
Following statistical analysis a dendrogram was generated and differences are evident among accessions within the studied population (Fig. 2B). Within the 109 examined accessions the SSR markers identified 26 different genotypes, and 83 synonyms (Table 5). Of the 26 genotypes, 16 show a similarity index smaller than one.
Discussion
From the analysis of molecular data by RAPD markers the presence of a genetic diversity could be detected. The oligonucleotides used as primers (Table 2) permitted the amplification of several DNA fragments, as shown in Fig. 1, useful for the study of olive germplasm. Of the 20 oligonucleotide decamers used only 544 appeared to be neutral for identification (it produced only amplified monomorphic fragments) (Table 3). Of the 19 oligonucleotides that resulted useful for analysis, 5 proved to be sufficient (AH09, AH18, AH19, AI11 e OPC16) to discriminate between accessions, producing a total of 104 fragments, 88 of which were polymorphic. Our results confirm the ability of the RAPD technique to discriminate olive germplasm in accordance with the study of Belaj et al. (2003), who demonstrated the high efficiency of RAPDs in detecting genetic similarities in olive. The RAPD results clearly show the marked genetic separation existing between certain local accessions. With this type of molecular markers, and with the number of primers utilised, it was possible to isolate 26 different genotypes within the analysed population (Fig. 2A). Another important feature is the high level of synonymy (76%) with known national cultivars.
The internal control (DNA samples extracted from the same tree, C1 and C2) gave a positive response: the analysed samples are genetically identical (data not shown), thus confirming the correct interpretation of data.
SSR analysis permitted to isolate, among local accessions, 26 different genotypes; 4 of them proved to be synonyms of 4 national cultivars present in the analysis (Ascolana Tenera, Frantoio, Maurino and Nostrana di Brisighella), while the other 22 belonged to16 genotypes represented by individual plants and to 6 groups of accessions which, with different names, are spread on the territory (Table 5). This result confirms the data obtained by RAPD analysis, although minor differences occurred (Fig. 2B): accessions 124 and 127 were considered as separate accessions by the SSRs, while they were included in the b group (Frantoio) by the RAPD markers; accessions 147 and 155 were classified by SSRs as belonging to the h group, the former, and to the d group, the latter, while RAPD markers considered them as individual accessions.
In the present research RAPD markers proved to be a reliable instrument to study population variability: through them it was possible to locate synonyms and single out some genotypes (Fig. 2).
However the analysis of 109 samples implies reactions of amplification and electrophoretic runs, which, for an individual primer, require several sessions. The laboriousness of these operations emphasize low reproducibility of RAPD technique.
From the genetic analysis several synonymies emerged (Table 5 and Fig. 2) with cultivars of Central Italy.
From this brief description it appears evident that the emilian germplasm which has displayed synonymies has in common both the area of origin (or area in which a cultivar is presently most cultivated) and frost tolerance.
All cultivars which were synonyms of emilian germplasm had Central Italy as area of main diffusion; in particular, Frantoio and Maurino are cultivars that originated in Northern Tuscany (Cimato et al. 2001); Ascolana Tenera is widespread in Central Italy (Barranco et al. 2000); Nostrana di Brisighella, instead, seems to be present only in Romagna (Rotondi et al. 2004), which is however adjacent to Central Italy (Bartolini et al. 2005). It is therefore quite obvious to postulate that olive diffusion in Emilia has taken place from Northern Tuscany and Romagna, at least for the genotypes identified in the present research. It cannot be ruled out, however, that the still unknown accessions could have been imported from other regions, such as Liguria (Bianchedi 1880) (Fig. 3). The Tuscan origin can be justified by geographical and historical evidence; Northern Tuscany is closer to Parma and Reggio Emilia than the olive producing districts of Romagna, which had also been politically separated from Emilia for several centuries. Although separated by the Apennins, Emilia and the Tuscan provinces of Lucca and Massa Carrara had kept, in the same period, close commercial and political ties, which had sometimes produced unions under the same feudal ruler; in addition, the road network has always been well developed and properly maintained (Bianchedi 1880). Tuscany was the main oil supplier of Emilia, as proven by abundant historical evidence, and the passage of propagation material northwards is all but unlikely (Fabbri 2005).
As far as frost tolerance is concerned, all cultivars so far identified have tolerance to low temperatures as a common feature, with the exception of Frantoio, which is the main pollinator of the examined genotypes. Although the survived individuals were all in environmentally favoured niches, the occurrence of mutations favourable to frost resistance, and the consequent existence of more resistant clones of the said cultivars cannot be ruled out, and will be the object of future research.
References
Amane M, Lumeret R, Hany V, Ouazzani N, Debain C, Vivier G, Deguilloux MF (1999) Chloroplast-DNA variation in cultivated and wild olive (Olea europaea L.). Theor Appl Genet 99:133–139
Angiolillo A, Mencuccini M, Baldoni L (1999) Olive genetic diversity assessed using amplified fragment length polymorphism. Theor Appl Genet 98:411–421
Barranco D, Cimato A, Fiorino P, Rallo L, Touzani A, Castaneda C, Serafin F, Trujillo I (2000) World catalogue of olive varieties. International Olive Oil Concil, Madrid, 360 pp
Bartolini G, Prevost G, Messeri C, Carignani G (2005) Olive germplasm: cultivars and word-wide collections. Web site FAO: http://www.apps3.fao.org/wiews/olive/oliv.jsp
Belaj A, Satovic Z, Cipriani G, Baldoni L, Testolin R, Rallo L, Trujillo I (2003) Comparative study of the discriminating capacity of RAPD, AFLP and SSR markers and of their effectiveness in establishing genetic relationships in olive. Theor Appl Genet 107:736–744
Belaj A, Satovic Z, Rallo J, Trujillo I (2004a) Optimal use of RAPD markers for identifying varieties in olive (Olea europaea L.) germplasm collection. J Am Soc Hortic Sci 129(2):266–270
Belaj A, Cipriani G, Testolin R, Rallo L, Trujillo I (2004b) Characterization and identification of the main Spanish and Italian olive cultivars by simple-sequence-repeat markers. HortScience 39(7):1557–1561
Besnard G, Khadari B, Baradat P, Bervillé A (2002) Olea europaea (Oleaceae) phylogeography based on chloroplast DNA polymorphism. Theor Appl Genet 104:1353–1361
Besnard G, Bervillé A (2000) Multiple origins for Mediterranean olive (Olea europaea L. ssp. europaea) based upon mitochondria DNA polymorphisms. Life Science 323:173–181. CR Acad Sci Paris
Bianchedi C (1880) L’Olivo sulle colline parmensi. Ferrari e figli, Parma Italy, 60 pp
Bignardi A (1978) Le campagne emiliane nel rinascimento e nell’età barocca. Forni, Bologna Italy, 401 pp
Carriero F, Fontanazza G, Cellini F, Giorio G (2002) Identification of simple sequence repeats (SSRs) in olive (Olea europaea L.). Theor Appl Genet 104:301–307
Cimato A, Cantini C, Sani G (2001) L’olivo in Toscana: il germoplasma autoctono. ARSIA Toscana-IPSL, CNR-Regione Toscana, Florence, Italy
Cipriani G, Marrazzo MT, Marconi R, Cimato A, Testolin R (2002) Microsatellite markers isolated in olive (Olea europaea L.) are suitable for individual fingerprinting and reveal polymorphism within ancient cultivars. Theor Appl Genet 104:223–228
Diaz Bermudez A (2005) Desarrollo y caracterización de nuevos microsatélites y SNPs y aplicación en la mejora genética del olivo (Olea europaea L.), Ph.D. tesis, Departamento de Genética, Universidad de Córdoba—ETSIAM Córdoba, Spain, 180 pp
Dice LR (1945) Measures of the amount of ecological association between species. Ecology 26:297–302
Fabbri A (2005) Olivicoltura emiliana? Perchè no! Olivo e Olio 4:12–17
Fabbri A, Ganino T (2003) Il germoplasma olivicolo in Emilia. Atti Convegno Germoplasma olivicolo e tipicità dell’olio Perugia, Italy, pp 48–52
Fabbri A, Hormaza JI, Polito VS (1995) Random amplified polymorphic DNA analysis of olive (Olea europaea L.) cultivars. J Am Soc Hort Sci 120:538–542
Felsenstein J (2005) PHYLIP (Phylogeny Inference Package) version 3.6. Distributed by the author. Department of Genome Sciences, University of Washington, Seattle
Fragaki G, Spyros A, Siragakis G, Salivaras E, Dais P (2005) Detection of extra vergin olive oil adulteration with lampante olive oil and refined olive oil using nuclear magnetic resonance spectroscopy and multivariate statistical analysis. J Agric Food Chem 53:2810–2816
Gallitelli M, Semeraro L, Antonelli NM (1991) RFLP analysis in the olive (Olea europaea L.). EMBO Course, Cologne
Ganino T, Bartolini A, Fabbri A (2006) The classification of olive germplasm—a review. J Hortic Sci Biotech 81(3):319–334
Ganino T, Fabbri A (2005) Genetic characterization of Olea europaea L. germplasm in northern Italy. 5th international symposium on olive growing, 27 September–2 October 2004, Izmir (Turkey), 127 pp
Gounaris Y, Skoula M, Fournaraki C, Drakakaki G, Makris A (2002) Comparison of essential oils and genetic relationship of Origanum × intercedens to its parental taxa in the island of Crete. Biochem Syst Ecol 30:249–258
Jaccard P (1908) Nouvelle recherches sur la distribution florale. B Soc Vaud Sci Nat 44:223–270
Kirby LT (1990) DNA fingerprinting: an introduction, Stockton Press, New York, 365 pp
Loukas M, Krimbas CB (1983) History of olive cultivars based on their genetic distances. J Hortic Sci 58:121–127
Mancuso S, Nicese FP (1999) Identifying olive (Olea europaea L.) cultivars using Artificial Neural Networks. J Am Soc Hortic Sci 124(5):527–531
Marrazzo T, Cipriani G, Marconi R, Cimato A, Testolin R (2002) Isolation and characterisation of microsatellite DNA in Olive (Olea europaea L.). Acta Hort 586:61–64
Matsuoka Y, Mitchell SE, Kresovich S, Goodman M, Doebley J (2002) Microsatellites in Zea—variability, patterns of mutations, and use for evolutionary studies. Theor Appl Genet 104:436–450
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Owen CA, Bita EC, Banilas G, Hajjar SE, Sellianakis V, Aksoy U, Hepaksoy S, Chamoun R, Talhook SN, Metzidakis I, Hatzopoulos P, Kalaitzis P (2005) AFLP reveals structural details of genetic diversity within cultivated olive germplasm from the Eastern Mediterranean. Theor Appl Genet 110:1169–1176
Ozkaya MT, Cakir E, Gokbayrak Z, Ercan H, Taskin N (2006) Morphological and molecular characterization of Derik Halhali olive (Olea europaea L.) accession grown in Derik-Mardin province of Turkey. Sci Hortic 108:205–209
Peakall R, Smouse PE (2005) GenAlEx 6: Genetic Analysis in Excel. Population genetic software for teaching and research. The Australian National University, Canberra, Australia. http://www.anu.edu.au/BoZo/GenAlEx/
Podani J (2001) Sin-tax 2000 Computer program for data analysis in ecology and systematics: user’s manual. Budapest, Hungary
R Development Core Team (2005) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria, ISBN 3-900051-07-0, URL http://www.R-project.org
Rallo P, Dorado G, Martin A (2000) Development of simple sequence repeats (SSRs) in olive tree (Olea europaea L.). Theor Appl Genet 101:984–989
Rotondi A, Mari M, Babini AR, Govoni M, Cristoferi G (2004) L’attitudine alla propagazione e la certificazione genetica e sanitaria dell’olivo in Emilia-Romagna. Regione Emilia-Romagna, CNR-IBIMET, Bologna, Italy, 117 pp
Sanz-Cortés F, Parfitt DE, Romero C, Struss D, Liácer G, Badenes M (2003) Intraspecific olive diversity assessed with AFLP. Plant Breed 122:173–177
Sefc KM, Lopes MS, Mendonça D, Rodrigues Dos Santos M, Laimer Da Câmara Machado M, Da Câmara Machado A (2000) Identification of microsatellite loci in olive (Olea europaea) and their characterization in Italian and Iberian olive trees. Mol Ecol 9:1171–1173
Trujillo I, Rallo L, Arus P (1995). Identifying olive cultivars by isozyme analysis. J Am Soc Hortic Sci 120:318–324
Wagner HW, Sefc KM (1999) Identity 1.0: freeware program for the analysis of microsatellite data. Vienna, http://www.boku.ac.at/zag/forsch/identity.htm
Vlahov G, Shaw AD, Kell DB (1999) Use of 13 C nuclear magnetic resonance distortionless enhancement by polarization transfer pulse sequence and multivariate analysis to discriminate olive oil cultivars. J Am Oil Chem Soc 76(10):1223–1231
Acknowledgements
The Authors wish to thank Dr. Enzo Perri, Director of Istituto Sperimentale per l’Olivicoltura of Rende (Cosenza, Italy), for the germplasm made available for this research, and Dr. Paolo Piovani, for reviewing the manuscript. The research was supported by the Project “Studio sullo sviluppo dell’olivicoltura da olio in Emilia-Romagna, nelle province di Modena, Reggio Emilia, Parma e Piacenza”, made possible by a joint grant of the said Provinces and of the Region Emilia-Romagna.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ganino, T., Beghè, D., Valenti, S. et al. RAPD and SSR markers for characterization and identification of ancient cultivars of Olea europaea L. in the Emilia region, Northern Italy . Genet Resour Crop Evol 54, 1531–1540 (2007). https://doi.org/10.1007/s10722-006-9145-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-006-9145-x