Abstract
The amino acid/auxin permease (AAAP) gene family plays an important role in the long-distance amino acid transport pathway and takes part in various stages of plant growth and development. However, little is known about the AAAP gene family in Medicago truncatula. Here, we identified 86 putative MtAAAP family members using genome sequence information. Based on phylogenetic analysis, these MtAAAP genes were categorized into eight distinct subfamilies. The MtAAAP genes were mapped on 8 chromosomes and duplication events appeared widely, with 19 and 21 pairs of MtAAAP genes showing segment and tandem duplication events, respectively. Ratio of Ka/Ks indicated that duplicated genes underwent purifying selection. Analysis of RNA-seq data showed that MtAAAP genes exhibited specific expression patterns among different tissues and abiotic stress, indicating that MtAAAP members were involved in plant developmental regulation and stress responses. Expression patterns of 16 MtAAAP genes under abiotic stress were verified by qRT-PCR. The present study provides a foundation for the functional analysis of MtAAAPs in developmental regulation and stress responses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amino acid/auxin permease (AAAP) protein is a type of amino acid transporter known to contribute to long-distance amino acid transport, such as in the transport of amino acids across the cellular membrane (Hirner et al. 1998; Wipf et al. 2002). In the majority of plants, organic nitrogen is mainly transported in the form of amino acids, which play an extremely important role in plant growth and development processes, such as in the biosynthesis and metabolism of key compounds (Tegeder et al. 2000). For these processes, amino acids are transported between tissues and organs via a transporter. The amino acid transporter is involved in resource allocation processes and can bind to a specific solute and undergo a conformational change that transfers the solute to the other side of the membrane (Bröer and Palacín 2011; Christensen 1990; Widdows et al. 2015).
The AAAP gene family is one of the largest amino acid transporter families and includes members from almost across all eukaryotic organisms (Wipf et al. 2002). The AAAP family is further grouped into amino acid permease (AAP), lysine and histidine transporter (LHT), γ-aminobutyric acid transporter (GAT), auxin transporter (AUX), proline transporter (ProT), aromatic neutral amino acid transporter (ANT), and the amino acid transporter-like (comprising ATLa and ATLb) subfamilies (Fischer et al. 1998; Okumoto et al. 2002; Saier et al. 2009). Each of the AAAP genes has a specific domain, Aa_trans. Most AAAP gene family members are known to transport amino acids with varying specificities and characteristics, and some AAAP gene family members have been fully studied in many model plants (Fischer et al. 1995; Sheng et al. 2014; Wu et al. 2015; Zhao et al. 2012, 2017), such as 8 members in Arabidopsis and 19 members in rice (Fischer et al. 1995; Zhao et al. 2012). Expression of AtAAP1 was high in the cotyledons and the endosperm, and in mediated-uptake of amino acids by embryos (Sanders et al. 2009). Similarly, OsAAP6 may be involved in amino acid uptake into the endosperm to provide amino acids for developing embryos (Peng et al. 2014). AtAAP2 functions in regulating amino acid transfer from xylem to phloem (Zhang et al. 2010). OsAAP8 and OsAAP15 participate in the uptake and long-distance transport of amino acids (Zhao et al. 2012), and AtAAP5 takes part in amino acid uptake processes in the root (Elashry et al. 2013). AtAUX1, which functions as an auxin influx carrier that can regulate root gravitropism, is primarily expressed in roots and can promote lateral root formation by transporting indole-3-acetic acid (Bennett et al. 1996; Marchant et al. 2002). AtAAP6 takes part in the transport regulation of amino acids in sieve elements (Hunt et al. 2010), and NtAAP2-2 can participate in the transport of Asp, Asn, Glu, and Gln in tobacco plants (Zhao et al. 2017).
Previous studies have shown that AAAP genes that encode transporters also have multifarious functions and can be involved in diverse developmental and physiological processes in plants (Elashry et al. 2013; Ortizlopez et al. 2000). In Arabidopsis, AAP3 and AAP6 are involved in root-knot nematode parasitism (Marella et al. 2013; Okumoto et al. 2004). AtAAP1, AtAAP2, and AtAAP8 are known to significantly reduce the number of nematodes developing in Arabidopsis (Elashry et al. 2013). The AAAP gene family members show potential for prevented and controled by biological stress responses and are essential in pathogen and abiotic stress responses (Rentsch et al. 1996).
Medicago truncatula is a diploid plant that has been frequently used for studying legume genomics (Benedito et al. 2008; Stacey et al. 2006). Although, AAAP genes have been identified and characterized in several plant species (Cheng et al. 2016; Liu et al. 2017; Wu et al. 2015; Zhao et al. 2012), there is currently no genome-wide analysis of M. truncatula. In the present study, we aimed to identify and characterize AAAP genes in M. truncatula by analyzing their phylogenetic relationship and gene duplication events, and by structuring conserved domain architecture. Additionally, expression profiles of these genes were examined under diverse abiotic stress. This study provides valuable information for the functional analysis of AAAP genes in M. truncatula.
Materials and methods
Identification of Medicago truncatula AAAP genes
The genome sequences were obtained from M. truncatula genome database (http://www.medicagogenome.org/). The Hidden Markov Model (HMM) profiles of the AAAP domain (PF01490) were downloaded from the Pfam database (http://pfam.xfam.org/). To find putative AAAP family member, we first searched AAAP domains from the M. truncatula protein database with e-value cut-off at 1.0 by using HMMER v3.0 software (http://hmmer.janelia.org/). Then, we performed BLASTP analysis against the M. truncatula genome database using Arabidopsis AAAP gene sequences as queries. The integrity of the AAAP domain was verified by using the online program SMART (http://smart.embl-heidelberg.de/) with an e-value < 0.1. The basic information of each gene, including length, molecular weight, isoelectric point, gene structure, and protein product characteristics, were predicted by the online ExPasy program (http://www.expasy.org/tools/) and GSDS website (http://gsds1.cbi.pku.edu.cn/index.php).
Phylogenetic analysis and motif prediction
To investigate the phylogenetic relationship of the AAAP gene family in M. truncatula. Sequence alignments were performed using ClustalX 1.81 (Thompson et al. 2002). A neighbor-joining (NJ) phylogenetic tree was constructed using the MEGA 7.0 program, setting the bootstrap value at 1000 (Kumar et al. 2016). The online MEME analysis (http://meme-suite.org/) was used to identify the unknown conserved motifs using the following parameters, the maximum number of motifs was set to 20, the optimum motif width from 10 to 300, and use the 0 or 1 occurrence per sequence strategy.
Chromosomal localization and gene duplication analysis
The summary of gene localization information was downloaded from the phytozome database (https://phytozome.jgi.doe.gov/pz/portal.html). The genomic sequence for each AAAP gene was extracted from the whole-genomic sequence according to gene localization information using a programmed Perl script. A gene structure display server program (http://gsds.cbi.pku.edu.cn/index.php) was used to display the M. truncatula AAAP gene structures. Duplications between the MtAAAP genes were identified by using the PGDD database (http://chibba.agtec.uga.edu/duplication/). In addition, duplications were complemented using the MCScanX software (Wang et al. 2012). Duplicated gene pair were separated by four or fewer gene loci, will be identified as tandem duplications (TD). Others were identified as segmental duplications (SD). Ideograms were created by using Circos program. The Ka and Ks were used to assess selection history and divergence time (Li et al. 1981). The number of synonymous (Ks) and nonsynonymous (Ka) substitutions of duplicated AAAP genes were computed by using the MCScanX program (http://chibba.pgml.uga.edu/mcscan2/). The divergence time (T) was calculated using the formula T = Ks/(2 × 6.5 × 10−9) × 10−6 Millon years (Cannon et al. 2006; Lynch and Conery 2000).
Expression analysis of MtAAAP genes in different tissues and abiotic stress
Medicago truncatula transcriptome data from different development tissues and different abiotic stresses were downloaded from the Sequence Read Archive (SRA) of NCBI (http://www.ncbi.nlm.nih.gov/sra, Accession numbers SRX099057–SRX099062, SRX1056987–SRX1056992). The transcriptome data were derived from six tissues, including roots, nodules, blades, buds, seedpods, and flowers. The transcriptome data under stress were derived under six distinct factors, including cold, freezing, drought, salt, and high levels of ABA. Clean reads from six samples were mapped to the M. truncatula genome sequencs using Samtool (Li et al. 2009). RPKM was analyzed by Tophat and Cufflinks (Trapnell et al. 2014). The expression of AAAP genes were utilized for generating the heatmap and k-means clustering using R software.
Plant material and treatments
Seeds of M. truncatula (Jemalong) A17 were planted in a 3:1 (w/w) mixture of soil and sand, germinated, and irrigated with half-strength Hoagland solution once every 2 d. The seedlings were grown in the following environmental conditions: temperature of 18 °C (night) and 24 °C (day), relative humidity of 60–80% and a 14/10 h photoperiod (daytime, 06:00–20:00). The seedlings that germinated after 8 weeks were subjected to the following environmental conditions: temperatures of 4 (cold) or − 8 °C (freezing), treated with 300 mM mannitol (drought) or 200 mM NaCl solution (salt), and sprayed with 100 µM ABA solution (ABA). Control (untreated) and treated seedlings were harvested 3 h after treatment. All samples were frozen in liquid nitrogen and stored at − 80 °C until use.
RNA extraction and quantitative real-time PCR (qRT-PCR)
The transcriptome sequencing analysis was validated and quantified by qRT-PCR. Primers were designed according to AAAP CDS with Primer Express 3.0 software, the primer pairs are listed in Table S1. Total RNA were extracted with an RNA prep pure Plant Kit (Tiangen, Beijing, China), and cDNA was synthesized from total RNA using the PrimeScript RT reagent Kit (Toyobo, Shanghai, China), and qRT-PCR was performed by using SYBR Green and monitored on an ABI 7300 Real-Time PCR System (Applied Biosystems, CA, USA). The PCR conditions of were set as follows: 95 °C for 2 min, 40 cycles at 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 1 min, and a final dissociation at 95 °C for 15 s, followed by 1 cycle at 60 °C for 20 s and one cycle at 95 °C for 15 s. The reference gene was \(\beta\)-actin. \({2^{ - \Delta \Delta Ct}}\) method was used to analyze the relative expression level of each gene. All the samples were tested with three technical replicates and three independent biological replicates.
Results
Identification of AAAP family genes in Medicago truncatula
We identified 111 putative AAAP genes in the M. truncatula genome. SMART and Pfam were used to search gene sequences for the presence of the Aa_trans domain and genes whose characteristic domain was missing or incomplete were removed. A total of 86 non-redundant and complete AAAPs were obtained for further analysis and designated as MtAAAP1 to MtAAAP86 according to their chromosome locations. The encoding amino acid sequences of these genes extended from 204 to 830. Other detailed information, such as gene IDs, chromosome locations, and properties of encoding proteins is provided in Table S2.
Phylogenesis and multiple sequence alignment
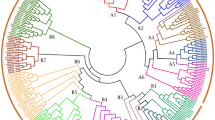
To explore the inherent laws of genetic evolutionary history of the MtAAAP gene family, we constructed a phylogenetic tree by using the Neighbor-Joining method (Fig. 1a), and a combined phylogenetic tree with AtAAAPs, OsAAAPs, and MtAAAPs using the method to identify orthologous genes between species (Fig. S1). Results revealed that AAAP proteins were separated into eight distinct clades based on the similarity of sequences with highly supported bootstrap values, and subclade distribution were consistent among the three species. We determined that the AAAP family could be divided into amino acid permease (AAP), lysine and histidine transporter (LHT), γ-aminobutyric acid transporter (GAT), auxin transporter (AUX), proline transporter (ProT), aromatic, neutral amino acid transporter (ANT), and amino acid transporter-like (including ATLa and ATLb) subfamilies.
Phylogenetic analysis, conserved motifs and gene structure of the AAAP family in Medicago truncatula. a Phylogenetic tree analysis derived from neighbor-joining methods of MtAAAP proteins. Bootstrap values by using 1000 replicates are indicated at each node. The branches of eight subfamilies are marked by different colors. b Distribution of conserved motifs within AAAP family. Summary for the distribution of conserved motifs identified from 86 AAAP proteins by each group given separately. Each motif is represented by a number in colored box. c Exon–intron structure analysis of MtAAAP genes. Exon/intron organization are represented by yellow box and black lines, respectively. UTR are displayed using blue box, 3p_UTR and 5p_UTR are displayed using red box and green box, respectively
To further analyze phylogenetic relationships and explore potential functions, we analyzed conserved domains in the amino acid sequences of MtAAAPs using the MEME suite (Fig. 1b). A total of 20 distinct and highly conserved motifs were captured (Table S3). Most subfamilies were highly conserved in motif distribution pattern and several motifs were relatively pervasive among AAAP members, such as motif 5 and motif 19. In contrast, a large proportion of motifs displayed specificity to different subfamilies. For instance, motif 1 and motif 7 were specific to subfamily AAP, motif 2 was specific to LHT, and motif 9 and motif 16 were specific to ATLb. Similarly, most of the AAAP gene family members in the same subfamily had the same or similar gene structures and gene lengths (Fig. 1c). However, several members in the same group also showed differences in gene structures, such as MtAAAP26 which had a unique structure in the C terminal and MtAAAP75 which had a diverse structure in the intron compared with paralogs. Results indicated that the gene structure and conserved domain distribution were conserved in the same subfamily, which might indicate close evolutionary relationships.
Multiple sequence alignment of the MtAAAP genes showed that the distribution of most transmembrane regions (TM) in the same subfamily were highly conserved (Fig. S2). Furthermore, the length and amino acid composition of several TM regions in different members of the same subfamily were relatively similar. An example of the alignment of LHT subfamily members with high similarity is shown in Fig. S3. There were five conserved motifs in MtLHTs, namely motif 2, 11, 3, 4, and 6. The first three TM regions were located in motif 2, while TM4 and TM5 were located in motif 11. Motif 3 comprised the sixth and seventh TM regions. The remaining transmembrane regions, TM8 and TM10, were located in motif 4 and motif 6, respectively.
Gene chromosomal location, and gene duplication events of MtAAAP
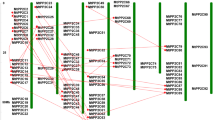
We mapped 84 of the 86 MtAAAPs on 8 chromosomes of M. truncatula (Fig. 2). It is noteworthy that MtAAAP1 and MtAAAP2 had a close relationship, yet unattributed scaffolds could not be conclusively mapped on any chromosome. The distribution of AAAP gene family members on each chromosome was not equal, ranging from 4 to 21 in chromosome 2 and chromosome 3, respectively. Most subfamilies were widely distributed among chromosomes. For example, AAPs were distributed among six chromosomes, excluding chromosome 06 and 07. Most chromosomes were comprised of more than five different subfamily members. Using gene duplication analysis, we determined that 52.3% (45 of 86) of MtAAAPs represent duplication events, including 21 pairs of genes with tandem duplications and 19 segmental duplication events. Duplication events mainly existed on chromosome 03, 05, 07, and 08, which might explain the AAAP gene distribution patterns on chromosomes, and also probably explain the abundance of AAAP genes in M. truncatula relative to Arabidoposis.
Chromosomal distribution and duplication events analysis of MtAAAP genes in Medicago truncatula. Duplicated genes in different subfamily were linked with colored lines in the diagrams using Circos. Red lines show duplications between members of the AAP subfamily, blue lines show duplication between members of the ATLa subfamily, yellow lines show duplications between members of the ATLb subfamily, black lines show duplication in GAT subfamily, purple lines show duplication between members of the AUX subfamily and green lines show duplications between members of the LHT subfamily
Additionally, we identified the synonymous substitution rate (Ks) and nonsynonymous substitution rate (Ka) of paralogs with duplication events, and relative Ks values were used as an expression of time (Table S4). As expected, the Ks value of tandem duplication pairs was smaller than the segmental duplication pairs, indicating that segmental duplication generally occurred in an ancient time period. The distribution of duplicate gene pair Ks values peaked at approximately 0.2 and 0.7, which suggested that AAAP genes in M. truncatula had undergone two rounds of large-scale duplication events approximately 15 and 52 million years ago, respectively. In general, Ka/Ks ratios of < 1, 1 and > 1 indicate negative or stabilizing selection, neutral selection, and positive selection, respectively (Cannon et al. 2004). The majority of Ka/Ks values for gene pairs were < 1, and 36 pairs of the MtAAAP duplicated genes were < 0.5, suggesting that most AAAP family homologous gene pairs had experienced strong purifying selection during evolution.
Expression of MtAAAP genes among tissues and response to abiotic stresses
To determine expression patterns of AAAP genes in different tissues of M. truncatula, a heatmap was produced by high-throughput sequencing data from NCBI (Fig. 3a). Genes could clearly be divided into six major groups from A to F, and most genes in AAAP family showed a specific expression pattern. For instance, genes in group A were highly expressed in the root, group B in the flower, group C in the bud, group D mainly in seedpod tissue, group E in the blade, and group F in the nodule. The MtAAP subfamily members were expressed among every single group. MtATL subfamilies were mainly distributed in groups E and F, and MtAUXs and MtLHTs were mainly distributed in groups B and D. However, there were also a number of genes that displayed similar expression profiles for distinct tissues, such as AAAP13 and AAAP57 in group A were also expressed in nodule and flower, and AAAP24, AAAP31, and AAAP46 in group E were expressed in nodule too. By analyzing genes expressed in different tissues, we found that all tissues contained at least five subfamilies, which indicated that AAAP family members might play important roles in many growth and development process of plants.
Differential expression analysis of MtAAAP genes a involved in tissue development and b in response to different abiotic stresses. Heatmap showing hierarchical clustering analyses of MtAAAP genes across different tissues including roots, nodules, blades, buds, seedpods and flowers, and hierarchical clustering analyses of MtAAAP genes response to abiotic stress including control, cold, freezing, drought, salt and ABA. Genes highly or weakly expressed are colored red and black, respectively. The expression values were log2 transformed using the R soft
To explore potential functions of MtAAAPs, we investigated expression patterns during various abiotic stresses, including cold, freezing, drought, salt, and ABA stress, using RNA-seq data from NCBI (Fig. 3b). We divided the MtAAAPs into six groups from A to F according to the expression profile. Group A and group C were highly expressed under ABA stress, and group A are also showed high expression under control. Group B showed high expression under drought and salt stress. Group D mainly showed high expression under cold stress. Similarly, group E was significantly expressed under cold and freezing libraries. Genes in group F was mainly expressed under drought and control conditions. When compared with the control, a total of 43 MtAAAPs were differentially expressed in abiotic stress conditions. Among them, 14 genes showed differential accumulation under cold stress, whereas 22 genes were differentially expressed in freezing stress. A total of 15 genes were differentially expressed in drought stress, 12 genes in salt stress, and 21 genes in ABA stress. Interestingly, LHT subfamily members were mainly distributed in groups D and E, indicating that they might mainly take part in the freezing response process and cold tolerance. AUX subfamily members were mainly distributed in group F, indicating that they might take part in the drought response process. It is noteworthy that three MtAAAP genes (MtAAAP10, MtAAAP36, and MtAAAP39) were significantly up-regulated under most stress conditions, suggesting that they might play an important role in the resistance of M. truncatula to abiotic stress.
To further verify the reliability of the transcriptome data of MtAAAPs under abiotic stress, we randomly selected 16 differentially expressed MtAAAP genes to verify their expression profiles under five abiotic stress conditions by qRT-PCR (Fig. 4). The expression profiles derived from qRT-PCR were consistent with those based on the number of reads from RNA-seq, indicating that the RNA-seq data was highly reliable.
Discussion
Members of the AAAP gene family play critical roles in the process of plant growth and development and have attracted much attention in recent years (Fischer et al. 1998). Genetic evolution patterns and characteristics of AAAP gene family members have been studied in some plants. For example, AAP subfamily (Fischer et al. 1995; Okumoto et al. 2004; Sanders et al. 2009) and AUX subfamily members (Okumoto et al. 2002; Swarup et al. 2004) have been reported in Arabidopsis, rice, and ginseng (Lu et al. 2012, 2018; Zhang et al. 2013). However, the numbers and characteristics of AAAP gene family members in M. truncatula have not been systematically analyzed. Here, we identified and analyzed a total of 86 members of the AAAP gene family in M. truncatula. Based on the sequence similarity of amino acids, MtAAAPs were divided into eight subfamilies. The ML-phylogenetic tree showed that the classification of subfamilies within Arabidopsis was highly consistent, which suggested that AAAP genes had been formed before Dilleniidae and Rosidae diverged. The number of AAAP genes in M. truncatula was higher than that of other plants, such as Arabidopsis (46), maize (71), bamboo (55), poplar (71), and rice (58), but much less than that of soybean (153) (Cheng et al. 2016; Liu et al. 2017; Sheng et al. 2014; Wu et al. 2015; Zhao et al. 2012). Additional members indicating the expansion of the AAAP genes in M. truncatula, and may existent as amino acids transporter evolved in nitrogen fixation (Kim et al. 2013; Rolletschek et al. 2005).
In general, gene functions are closely related to conserved domains in amino acid sequences. We combined motif analysis, gene structure analysis and phylogenetic analysis to investigate potential functions of MtAAAPs (Fig. 1). Results indicate that each cluster had its own individual motif distribution pattern that was highly conserved, and shared similar gene length and exon/intron structure patterns in every subcluster. This might imply that the functional differentiation of AAAP family was very diverse, whereas members in the same subfamily had similar functions (Hu et al. 2013). We also detected functions of each motif by searching the Pfam and SMART database. Results showed that among 20 motifs (Table S3), only 7 motifs (11, 12, 14, 15, and 18–20) did not encode any domain, whereas the other 13 all encoded the characteristic domain Aa_trans, which is consist with previously studies (Wu et al. 2015). Interestingly, motif 5 was widespread in almost the whole gene family members’ C terminal, and two transmembrane regions were detected in motif 5 by using the TMHMM tool. Similar distribution pattern were observed in other plants, such as motif 3 in bamboo, motif 3 in poplar and motif 2 in rice, and the amino acid sequences are species specificity (Liu et al. 2017; Wu et al. 2015; Zhao et al. 2012). This indicated that motif 5 might play an important role in long-distance transmembrane transport in M. truncatula, which is the major function of the AAAP gene family (Fischer et al. 1998).
Gene duplication events are a major evolutionary mechanism for generating novel genes, and these events are an important way to help plants deal with environmental change during growth and development (Gu et al. 2003; Wagner 1994). Gene duplication events probably caused the larger expansion of AAAPs in M. truncatula than in Arabidopsis. For example, we detected 14 ATLb subfamily members in M. truncatula, whereas only 10 ATLb were detected in Arabidopsis. Gene duplication analysis showed that tandem duplication events occurred in the ATLb subfamily and were observed in MtAAAP83, MtAAAP84, MtAAAP85, and MtAAAP86 (Fig. 2). Based on conserved motif analysis, two kinds of motif patterns were observed in MtATLb family members (Fig. 1b). In one pattern, a longer motif 11 was inserted after motif 5 on the C-terminal, and the other pattern showed that motif 11 was replaced by a shorter motif combination (motif 15 and motif 13). In duplication events of the ATLb family, MtAAAP84 and MtAAAP85 showed different motif patterns when compared to MtAAAP83 and MtAAAP86, which might be caused by a series of mutations that occurred in duplication during evolution and might induce the production of new functions (Panchy et al. 2016).
Members of the AAAP gene family show various expression patterns in different organizations of plants and can play important roles during plant growth and development (Ortizlopez et al. 2000). Genes in the AAP subfamily have been studied and have varied functions in different plant tissues. In Arabidopsis, AtAAP1 is highly expressed in cotyledons and endosperm, plays an important role in embryo-mediated amino acid uptake, and is important for storage protein synthesis and seed yield (Frommer et al. 1993; Sanders et al. 2009). As previously reported, AtAAP1 will import specific amino acids into root cells when supplied at ecologically relevant concentrations (Lee et al. 2007; Perchlik et al. 2014). In castor, RcAAP1 and RcAAP2 are highly expressed in cotyledons and roots during seedlings development, but expression is lower in endosperm and hypocotyl (Bick et al. 1998). However, RcAAP3 has a different pattern and is only high expressed in source and sink tissues (Neelam et al. 1999). In the current study, we analyzed expression patterns of 72 MtAAAP family members that were expressed in at least 1 tissue and divided them into 6 major groups (Fig. 3a). Among them, 21 AAP subfamily members were expressed among every single group, which suggests that AAP subfamily members in M. truncatula might take part in various processes in different tissues. This finding is consistent with results from previous studies (Chen and Bush 1997; Kwart et al. 2010; Ma et al. 2016; Reinhardt et al. 2003). Furthermore, we found that MtAAAP11, an ortholog of AtAAP1, showed high expression levels in seedpods and was also expressed in roots, flowers, and buds, which suggests that it might participate in amino acid transport in various tissues like AtAAP1. Additionally, MtAAAP63 and MtAAAP71, which were orthologous to AtLHT1, were highly expressed in roots and nodules. This indicates that they might have similar functions to AtLHT1, play an important role in improving the Lys content of Lys-deficient grains, and take part in the transport of ACC (Chen and Bush 1997; Shin et al. 2015). MtAAAP53, the ortholog of AtAUX1 which regulates root gravitropic curvature, was also highly expressed in flower, seedpod and root tissues in this study (Timpte et al. 2010). Overall, these results confirm the potential functions of the MtAAAP family in the uptake and long-distance transport of amino acids in various processes. However, the specific role of MtAAAP family members in these processes needs to be determined.
Periods of environmental stress change the expression pattern of genes involved in defense responses (Bennett et al. 1996; Matters and Scandalios 1986). Proline accumulates widely in response to various environmental stresses, and evidence suggests that a positive correlation exists between proline accumulation and plant stress tolerance (Dar et al. 2016; Hayat et al. 2012; Zanella et al. 2016). As a subfamily of the AAAP family, several proline transporter subfamily members are strongly induced under abiotic stress (Popova et al. 2003; Rentsch et al. 1996; Ueda et al. 2001). However, little research exists on the expression pattern of the whole AAAP family in response to abiotic stress in M. truncatula, and their functions under various tolerances are still unknown. To further confirm the relationship between MtAAAPs and abiotic stress response, we analyzed the expression profiles of MtAAAP genes under five stress treatments using RNA-seq data (Fig. 3b). Results showed that 67 genes were expressed under at least 1 abiotic stressor and the expression of 66 genes were also detected in tissue analysis (Fig. 3a). Among them, 42 genes were differentially expressed under 5 stress conditions when compared to the control and most of them were upregulated under abiotic stress. Among them, ANT subfamily members, which are primarily detected as a distinct family of Na+-dependent transporters, are similar to glutamate transporters (Kanai 1997). It has been reported that the ANT family is associated with early-onset of chronic kidney diseases (Vivante and Hildebrandt 2016), and AtANT1 can transport arginine, aromatic, and neutral amino acids in Arabidopsis, especially in flowers and cauline leaves (Chen 2001). Here, MtAAAP79 were induced in response to abiotic stress, especially in the salt and drought condition (Fig. 4b), indicating that a potential role of the ANT subfamily members MtAAAP79 is the regulation of salt and drought stress response in M. truncatula. Response of the Prot subfamily to abiotic stress has been previously studied, we found that MtAAAP19 had a close relationship with AtPro2, and AtProt2 could be strongly induced under water or salt stress (Rentsch et al. 1996). MtAAAP19 was induced under drought, salt and ABA stress in M. truncatula, indicating that MtAAAP19 may be involved in responses to abiotic stress conditions. Interestingly, MtAAAP52, which is orthologous to AtProt2, was down-regulated under drought stressors, which may be attributed to functional changes induced by genetic mutations (Zhao et al. 2012). In moso bamboo, the AAP subfamily gene PeAAAP18 was up-regulated under PEG, cold, and salt stress (Liu et al. 2017). MtAAAP42, an orthologous gene of PeAAAP18, was highly induced under drought, cold, salt, and ABA stress, indicating that MtAAAP42 may take part in several responses to abiotic conditions. Our results suggest that several MtAAAP genes have potential roles in abiotic stress responses in different tissues.
In conclusion, we identified 86 MtAAAP genes from M. truncatula genome data and investigated the classification, evolution, structure, duplication, and expression pattern of these genes. Results indicated that MtAAAP genes take part in the regulation of plant development, and several members may be involved in responses to different abiotic stress conditions. This study provides important information for further identifying and characterizing specific functions of AAAP genes in M. truncatula and provides guidance for future transgenic applications.
References
Benedito VA et al (2008) A gene expression atlas of the model legume Medicago truncatula. Plant J 55:504–513. https://doi.org/10.1111/j.1365-313X.2008.03519.x
Bennett MJ et al (1996) Arabidopsis AUX1 gene: a permease-like regulator of root gravitropism. Science 273:948–950. https://doi.org/10.1126/science.273.5277.948
Bick JA, Neelam A, Hall JL, Williams LE (1998) Amino acid carriers of Ricinus communis expressed during seedling development: molecular cloning and expression analysis of two putative amino acid transporters, RcAAP1 and RcAAP2. Plant Mol Biol 36:377–385. https://doi.org/10.1023/A:1005903229458
Bröer S, Palacín M (2011) The role of amino acid transporters in inherited and acquired diseases. Biochem J 436:193–211. https://doi.org/10.1042/BJ20101912
Cannon SB, Mitra A, Baumgarten A, Young ND, May G (2004) The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 4:10. https://doi.org/10.1186/1471-2229-4-10
Cannon SB et al (2006) Legume genome evolution viewed through the Medicago truncatula and Lotus japonicus genomes. Proc Natl Acad Sci USA 103:14959–14964. https://doi.org/10.1073/pnas.0603228103
Chen L (2001) ANT1, an aromatic and neutral amino acid transporter in Arabidopsis. Plant Physiol 125:1813–1820. https://doi.org/10.1104/pp.125.4.1813
Chen L, Bush DR (1997) LHT1, a lysine- and histidine-specific amino acid transporter in arabidopsis. Plant Physiol 115:1127–1134. https://doi.org/10.1104/pp.115.3.1127
Cheng L et al (2016) Genome-wide identification, classification, and expression analysis of Amino Acid Transporter gene family in Glycine Max. Front Plant Sci 7:515. https://doi.org/10.3389/fpls.2016.00515
Christensen HN (1990) Role of amino acid transport and countertransport in nutrition and metabolism. Physiol Rev 70:43–77. https://doi.org/10.1152/physrev.1990.70.1.43
Dar MI, Naikoo MI, Rehman F, Naushin F, Khan FA (2016) Proline accumulation in plants: roles in stress tolerance and plant development. In: Iqbal N, Nazar R, Khan A N (eds) Osmolytes and plants acclimation to changing environment: emerging omics technologies. Springer India, New Delhi, pp 155–166. https://doi.org/10.1007/978-81-322-2616-1_9
Elashry A, Okumoto S, Siddique S, Koch W, Kreil DP, Bohlmann H (2013) The AAP gene family for amino acid permeases contributes todevelopment of the cyst nematode Heterodera schachtii in roots of Arabidopsis. Plant Physiol Biochem 70:379–386. https://doi.org/10.1016/j.plaphy.2013.05.016
Fischer WN, Kwart M, Hummel S, Frommer WB (1995) Substrate specificity and expression profile of amino acid transporters (AAPs) in Arabidopsis. J Biol Chem 270:16315–16320. https://doi.org/10.1074/jbc.270.27.16315
Fischer WN, André B, Rentsch D, Krolkiewicz S, Tegeder M, Breitkreuz K, Frommer WB (1998) Amino acid transport in plants. Trends Plantence 3:188–195. https://doi.org/10.1016/S1360-1385(98)01231-X
Frommer WB, Hummel S, Riesmeier JW (1993) Expression cloning in yeast of a cDNA encoding a broad specificity amino acid permease from Arabidopsis thaliana. Proc Natl Acad Sci USA 90:5944–5948. https://doi.org/10.1073/pnas.90.13.5944
Gu Z, Steinmetz LM, Gu X, Scharfe C, Davis RW, Li WH (2003) Role of duplicate genes in genetic robustness against null mutations. Nature 421:63–66. https://doi.org/10.1038/nature01198
Hayat S, Hayat Q, Alyemeni MN, Wani AS, Pichtel J, Ahmad A (2012) Role of proline under changing environments: a review. Plant Signal Behav 7:1456–1466. https://doi.org/10.4161/psb.21949
Hirner B, Fischer WN, Rentsch D, Kwart M, Frommer WB (1998) Developmental control of H+/amino acid permease gene expression during seed development of Arabidopsis. Plant J Cell Mol Biol 14:535. https://doi.org/10.1046/j.1365-313X.1998.00151.x
Hu L-Z, Yin W-B, Chen Y-H, Hu Z-M (2013) Functional divergence and evolutionary dynamics of the putative AAAP gene family in Brassica rapa. Plant Mol Biol Report 32:517–530. https://doi.org/10.1007/s11105-013-0671-3
Hunt E, Gattolin S, Newbury HJ, Bale JS, Tseng HM, Barrett DA, Pritchard J (2010) A mutation in amino acid permease AAP6 reduces the amino acid content of the Arabidopsis sieve elements but leaves aphid herbivores unaffected. J Exp Bot 61:55–64. https://doi.org/10.1093/jxb/erp274
Kanai Y (1997) Family of neutral and acidic amino acid transporters: molecular biology, physiology and medical implications. Curr Opin Cell Biol 9:565. https://doi.org/10.1016/S0955-0674(97)80035-X
Kim DH, Parupalli S, Azam S, Lee SH, Varshney RK (2013) Comparative sequence analysis of nitrogen fixation-related genes in six legumes. Front Plant Sci 4:300. https://doi.org/10.3389/fpls.2013.00300
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870. https://doi.org/10.1093/molbev/msw054
Kwart M, Hirner B, Hummel S, Frommer WB (2010) Differential expression of two related amino acid transporters with differing substrate specificity in Arabidopsis thaliana. Plant J 4:993–1002. https://doi.org/10.1046/j.1365-313X.1993.04060993.x
Lee YH, Foster JJ, Voll LM, Weber AP, Tegeder M (2007) AAP1 transports uncharged amino acids into roots of Arabidopsis. Plant J Cell Mol Biol 50:305–319. https://doi.org/10.1111/j.1365-313X.2007.03045.x
Li WH, Gojobori T, Nei M (1981) Pseudogenes as a paradigm of neutral evolution. Nature 292:237–239. https://doi.org/10.1038/292237a0
Li H et al (2009) Genome project data processing S: the sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Liu H, Min W, Zhu D, Feng P, Wang Y, Yue W, Yan X (2017) Genome-wide analysis of the AAAP gene family in moso bamboo (Phyllostachys edulis). BMC Plant Biol 17:29. https://doi.org/10.1186/s12870-017-0980-z
Lu Y, Song Z, Kai L, Lian X, Cai H (2012) Molecular characterization, expression and functional analysis of the amino acid transporter gene family (OsAATs) in rice. Acta Physiol Plant 34:1943–1962. https://doi.org/10.1007/s11738-012-0995-x
Lu K et al (2018) Blocking Amino acid transporter OsAAP3 improves grain yield by promoting outgrowth buds and increasing tiller number in rice. Plant Biotechnol J 16:1710–1722. https://doi.org/10.1111/pbi.12907
Lynch M, Conery JS (2000) The evolutionary fate and consequences of duplicate genes. Science 290:1151–1155. https://doi.org/10.1126/science.290.5494.1151
Ma H et al (2016) Genome-wide survey and expression analysis of the amino acid transporter superfamily in potato (Solanum tuberosum L.). Plant Physiol Biochem 107:164–177. https://doi.org/10.1016/j.plaphy.2016.06.007
Marchant A, Bhalerao R, Casimiro I, Eklöf J, Casero PJ, Bennett M, Sandberg G (2002) AUX1 promotes lateral root formation by facilitating indole-3-acetic acid distribution between sink and source tissues in the Arabidopsis seedling. Plant Cell 14:589–597. https://doi.org/10.1105/tpc.010354.2
Marella HH, Nielsen E, Schachtman DP, Taylor CG (2013) The amino acid permeases AAP3 and AAP6 are involved in root-knot nematode parasitism of Arabidopsis. Mol Plant Microbe Interact 26:44–54. https://doi.org/10.1094/MPMI-05-12-0123-FI
Matters GL, Scandalios JG (1986) Changes in plant gene expression during stress. Dev Genet 7:167. https://doi.org/10.1002/dvg.1020070402
Neelam A, Marvier AC, Hall JL, Williams LE (1999) Functional characterization and expression analysis of the amino acid permease RcAAP3 from castor bean. Plant Physiol 120:1049–1056. https://doi.org/10.1104/pp.120.4.1049
Okumoto S, Schmidt R, Tegeder M, Fischer W, Rentsch D, Frommer W, Koch W (2002) High affinity amino acid transporters specifically expressed in xylem parenchyma and developing seeds of Arabidopsis. J Biol Chem 277:45338–45346. https://doi.org/10.1074/jbc.M207730200
Okumoto S et al (2004) Root phloem-specific expression of the plasma membrane amino acid proton co-transporter AAP3. J Exp Bot 55:2155–2168. https://doi.org/10.1109/20.560044
Ortizlopez A, Chang HC, Bush DR (2000) Amino acid transporters in plants. Biochim Biophys Acta 1465:275–280. https://doi.org/10.1016/s0005-2736(00)00144-9
Panchy N, Lehti-Shiu M, Shiu SH (2016) Evolution of gene duplication in plants. Plant Physiol 171:2294–2316. https://doi.org/10.1104/pp.16.00523
Peng B et al (2014) OsAAP6 functions as an important regulator of grain protein content and nutritional quality in rice. Nat Commun 5:4847. https://doi.org/10.1038/ncomms5847
Perchlik M, Foster J, Tegeder M (2014) Different and overlapping functions of Arabidopsis LHT6 and AAP1 transporters in root amino acid uptake. J Exp Bot 65:5193–5204. https://doi.org/10.1093/jxb/eru278
Popova OV, Dietz KJ, Golldack D (2003) Salt-dependent expression of a nitrate transporter and two amino acid transporter genes in Mesembryanthemum crystallinum. Plant Mol Biol 52:569–578. https://doi.org/10.1023/A:1024802101057
Reinhardt D et al (2003) Regulation of phyllotaxis by polar auxin transport. Nature 426:255–260. https://doi.org/10.1038/nature02081
Rentsch D, Hirner B, Schmelzer E, Frommer WB (1996) Salt stress-induced proline transporters and salt stress-repressed broad specificity amino acid permeases identified by suppression of a yeast amino acid permease-targeting mutant. Plant Cell 8:1437–1446. https://doi.org/10.1105/tpc.8.8.1437
Rolletschek H et al (2005) Ectopic expression of an amino acid transporter (VfAAP1) in seeds of Vicia narbonensis and pea increases storage proteins. Plant Physiol 137:1236–1249. https://doi.org/10.1104/pp.104.056523
Saier MH, Reddy VS, Tsu BV, Ahmed MS, Li C, Moreno-Hagelsieb G (2009) The transporter classification database: recent advances. Nucleic Acids Res 37:D274–D278. https://doi.org/10.1093/nar/gkn862
Sanders A, Collier R, Trethewy A, Gould G, Sieker R, Tegeder M (2009) AAP1 regulates import of amino acids into developing Arabidopsis embryos. Plant J 59:540–552. https://doi.org/10.1111/j.1365-313X.2009.03890.x
Sheng L et al (2014) A genome-wide analysis of the AAAP gene family in maize. J Proteom Bioinform 07:023–033. https://doi.org/10.4172/jpb.1000299
Shin K et al (2015) Genetic identification of ACC-RESISTANT2 reveals involvement of LYSINE HISTIDINE TRANSPORTER1 in the uptake of 1-aminocyclopropane carboxylic acid in Arabidopsis thaliana. Plant Cell Physiol 56:572–582. https://doi.org/10.1093/pcp/pcu201
Stacey G, Libault M, Brechenmacher L, Wan J, May GD (2006) Genetics and functional genomics of legume nodulation. Curr Opin Plant Biol 9:110–121. https://doi.org/10.1016/j.pbi.2006.01.005
Swarup R et al (2004) Structure–function analysis of the presumptive Arabidopsis auxin permease AUX1. Plant Cell 16:3069–3083. https://doi.org/10.1105/tpc.104.024737
Tegeder M, Offler CE, Frommer WB, Patrick JW (2000) Amino acid transporters are localized to transfer cells of developing pea seeds. Plant Physiol 122:319–325. https://doi.org/10.1104/pp.122.2.319
Thompson JD, Gibson TJ, Higgins DG (2002) Multiple sequence alignment using ClustalW and ClustalX. Wiley, New York. https://doi.org/10.1002/0471250953.bi0203s00
Timpte C, Lincoln C, Pickett FB, Turner J, Estelle M (2010) The AXR1 and AUX1 genes of Arabidopsis function in separate auxin-response pathways. Plant J Cell Mol Biol 8:561–569. https://doi.org/10.1046/j.1365-313X.1995.8040561.x
Trapnell C et al (2014) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7:562. https://doi.org/10.1038/nprot.2012.016
Ueda A, Shi W, Sanmiya K, Shono M, Takabe T (2001) Functional analysis of salt-inducible proline transporter of barley roots. Plant Cell Physiol 42:1282–1289. https://doi.org/10.1093/pcp/pce166
Vivante A, Hildebrandt F (2016) Exploring the genetic basis of early-onset chronic kidney disease. Nat Rev Nephrol 12:133. https://doi.org/10.1038/nrneph.2015.205
Wagner A (1994) Evolution of gene networks by gene duplications: a mathematical model and its implications on genome organization. Proc Natl Acad Sci USA 91:4387–4391. https://doi.org/10.1073/pnas.91.10.4387
Wang Y et al (2012) MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res 40:e49. https://doi.org/10.1093/nar/gkr1293
Widdows KL et al (2015) Integration of computational modeling with membrane transport studies reveals new insights into amino acid exchange transport mechanisms. Faseb J 29:2583. https://doi.org/10.1096/fj.14-267773
Wipf D, Ludewig U, Tegeder M, Rentsch D, Koch W, Frommer WB (2002) Conservation of amino acid transporters in fungi, plants and animals. Trends Biochem Sci 27:139. https://doi.org/10.1016/S0968-0004(01)02054-0
Wu M, Wu S, Chen Z, Dong Q, Yan H, Xiang Y (2015) Genome-wide survey and expression analysis of the amino acid transporter gene family in poplar. Tree Genet Genomes 11:83–103. https://doi.org/10.1007/s11295-015-0908-4
Zanella M et al (2016) β-Amylase 1 (BAM1) degrades transitory starch to sustain proline biosynthesis during drought stress. J Exp Bot 67:1819–1826. https://doi.org/10.1093/jxb/erv572
Zhang L, Tan Q, Lee R, Trethewy A, Lee YH, Tegeder M (2010) Altered xylem-phloem transfer of amino acids affects metabolism and leads to increased seed yield and oil content in Arabidopsis. Plant Cell 22:3603–3620. https://doi.org/10.1105/tpc.110.073833
Zhang R, Zhu J, Cao HZ, Xie XL, Huang JJ, Chen XH, Luo ZY (2013) Isolation and characterization of LHT-type plant amino acid transporter gene from Panax ginseng Meyer. J Ginseng Res 37:361–370. https://doi.org/10.5142/jgr.2013.37.361
Zhao H, Ma H, Yu L, Wang X, Zhao J (2012) Genome-wide survey and expression analysis of amino acid transporter gene family in rice (Oryza sativa L.). PloS One 7:e49210. https://doi.org/10.1371/journal.pone.0049210
Zhao Y et al (2017) Genome-wide identification and characterization of an amino acid permease gene family in Nicotiana tabacum. Rsc Adv 7:38081–38090. https://doi.org/10.1039/C7RA05610A
Acknowledgements
This work was supported by the grant from the National Nature Science Foundation of China (nos. 31470571; 31770575). The National Major Project for Cultivation of Transgenic Crops (#2016ZX08004-002-003), and The National Key Research and Development Program of China (2017YFD0101303).
Author information
Authors and Affiliations
Contributions
All authors contribute equally.
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10709_2019_62_MOESM1_ESM.tif
Fig. S1 Phylogenetic tree analysis of the AAAP family in Medicago truncatula, Arabidopsis thaliana and Oryza sativa L (TIF 1648 KB)
10709_2019_62_MOESM3_ESM.tif
Fig. S3 Multiple sequence alignment and transmembrane region of LHT subfamily in Medicago truncatula. Identical (100%), conservative (75-99%), and block (50-74%) of similar amino acid residues are shaded in deep, lilac, and Cambridge blue, respectively. The transmembrane regions are marked by gray lines. The conserved motifs are marked by yellow lines (TIF 4888 KB)
Rights and permissions
About this article
Cite this article
Qu, Y., Ling, L., Wang, D. et al. Genome-wide identification and expression analysis of the AAAP family in Medicago truncatula. Genetica 147, 185–196 (2019). https://doi.org/10.1007/s10709-019-00062-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-019-00062-6