Abstract
The effects of soybean meal (SBM) in early diet of Sparus aurata larvae at two developmental windows were assessed. Prolonged (beyond 14 days post-hatch, dph) feeding with SBM decreased the activity of pancreatic enzymes of larvae. In the absence of SBM these larvae later resumed enzyme activities, but exhibited a significant delay in development. Larvae response to SBM involved up-regulation of extracellular matrix remodeling enzymes and pro-inflammatory cytokines, coupled with a drop in putative intestinal enzymes. Larvae receiving SBM at first feeding appear later to have lower expression of inflammation-related genes, especially those fed SBM until 14 dph. Multivariate analysis confirmed that the duration of the SBM early feeding period drives the physiology of larvae in different directions. Feeding larvae with SBM increased global histone H3 acetylation, whereas upon removal of SBM the process was reverted. A more in deep analysis revealed a dynamic interplay among several reversible histone modifications such as H3K14ac and H3K27m3. Finally, we showed that SBM feeding of larvae results in global hypomethylation that persist after SBM removal. This study is the first demonstrating an effect of diet on marine fish epigenetics. It is concluded that there are limitations for extending SBM feeding of S. aurata larvae beyond 14 dph even under co-feeding with live feed, affecting key physiological processes and normal growth. However, up to 14 dph, SBM does not affect normal development, and produces apparently lasting effects on some key enzymes, genes, and chromatin modifications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The use of plant proteins (PP) as alternatives to fish meal (FM) in aquafeeds have been thoroughly studied during the last decades but it remains a major issue in aquaculture research and fish physiology (Hardy 2010; Naylor et al. 2000). FM is expensive and it is expected to be limited available in the future, thus unable to sustain the expected increase in aquaculture production (Hardy 2010). Among several PP, soybean meal (SBM) is considered a suitable alternative to FM, due to its high availability, relatively low cost, and high nutritional value (Drew et al. 2007). However, there are species-specific limits for the inclusion of SBM in diets for different fish species. In salmonids, inclusion of SBM in diets produces alterations in the normal functioning of the digestive tract such as inflammation and failure to regulate mucosal integrity (Krogdahl et al. 2010), increased cell turnover and sloughing (Chikwati et al. 2013a), reduction in the activity of brush border enzymes and nutrient transport (Venold et al. 2013), increased activity of trypsin (Lilleeng et al. 2007) and other pancreatic proteases in the distal intestine (Chikwati et al. 2013b) and in feces (Krogdahl et al. 2003), and reduced cholesterol and bile salt reabsorption (Kortner et al. 2013). SBM contains different anti-nutritional factors (e.g. lectins, trypsin inhibitors, soy antigens, isoflavones), but soybean saponins have been suggested as the most likely contributing agent of these disorders (Krogdahl et al. 2010, 2015). Whereas salmonids are particularly sensitive to SBM and similar responses have been recently reported in turbot (Gu et al. 2016), higher SBM tolerance occurs in other fish species. For instance, S. aurata distal intestine lamina propria is moderately expanded due to an increase in mononuclear cells infiltration after feeding 30 % SBM diets, but this does not affect growth and feed conversion rates (Bonaldo et al. 2008). In this species, high level of FM substitution has been attained by using mixtures of vegetal protein sources (Dias et al. 2009) but SBM inclusion level has been regularly maintained well below 30 %. A higher inclusion level such as 40 % decreased feed conversion and plasma cholesterol (Venou et al. 2006). Advance to a higher use of SBM in feeds for this species would require an increased tolerance to SBM alone or in combination with others PPs.
Some studies in fish have provided evidence of nutritional programming (Geurden et al. 2014; Fang et al. 2014; Vagner et al. 2009; Koven et al. 2003; Geurden et al. 2013), including few recent reports on conditioning to SBM (Kemski et al. 2015; Perera and Yúfera 2016). The first step for conducting this kind of studies is the evaluation of the acceptance of the nutritional stimulus (or challenge) during early life without compromising further development. While SBM (∼5–10 % inclusion)-induced inflammatory response (i.e., enteropathy) in the distal intestine has been extensively documented in salmonids, now there are evidences that young salmon are relatively tolerant to SBM. SBM inclusion of 12 % induces cell proliferation in the distal intestine of salmon parr, but it produces no significant histological changes (Sanden et al. 2005). In line with these findings, a further study showed that a higher inclusion of SBM (i.e., 16.7 %) in diet did not produce intestinal inflammation, nor did it negatively affect growth, survival, enzyme activities, bile salt concentration or gene expression of various key functional markers in start feeding salmon fry (and up to three months following start-feeding) (Sahlmann et al. 2015). These studies showed that SBM can be used in diets for these life stages in salmon, but there is no information for other cultured marine species. The gilthead seabream, S. aurata, is one of the most important fish species cultured in Europe (Basurco et al. 2011) and it is known to be more tolerant to SBM (Bonaldo et al. 2008; Dias et al. 2009) than salmonids. While it is tempting to hypothesize that S. aurata larvae would also tolerate SBM in diet, this species has a prolonged larval period and the maturation of the digestive tract progress slower than in other fish (Moyano et al. 1996; Elbal et al. 2004).
Thus, this study accesses the possibility to include SBM in early diets of S. aurata. In addition, given that some components of SBM have both histone deacetylase and DNA methyl transferase inhibitory properties (Link et al. 2010), we hypothesized that SBM effects on fish larvae may include epigenetic modifications. Epigenetics refers to mechanisms for silencing or activating tissue specific gene expression independent of the DNA sequence, and mainly encompass changes in histone signatures and DNA methylation profiles that act in concert to induce transient and long-term changes in gene expression (Lillycrop and Burdge 2012). While several investigations on fish nutrition and digestion physiology have integrated ‘omic’ methodologies and there is now extensive information on the transcriptional responses of fish to different dietary components or nutritional conditions, knowledge on the molecular mechanisms regulating these responses is still limited. To our knowledge, there is no information on the effect of diet on marine fish epigenetics.
Results of this work revealed the limited capacity of S. aurata larvae to deal with SBM at first feeding. However, we determined a narrow window of time where SBM produces reversible and lasting effects on some key physiological variables without compromising further development. Also, this study shed some light on the putative role of different chromatin modifications on the regulation of the transcriptional response of larvae to SBM.
Material and methods
All experiments were conducted in compliance with Guidelines of the European Union (2010/63/UE) and the Spanish legislation (RD 1201/2005 and law 32/2007) for the use of laboratory animals. Experimental procedures used in this study received the approval of the CSIC Bioethics Subcommittee (ref. number 326,245/2013).
Experimental diets
Microdiets were prepared at the laboratory by the method of calcium internal gelation (Yúfera et al. 2005) and were formulated to have, on a dry weight basis, 50 % crude protein, 25 % lipid, 16 to 20 % carbohydrate (Table 1). One microdiet contained a combination of different animal protein sources including fish meal (FM) and was referred as the FM-diet. The other microdiet contained soybean meal (SBM) instead of FM, whereas keeping almost constant all other ingredients, and was referred as the SBM-diet.
Culture conditions, feeding protocol and sampling procedure
Two experiments were performed to evaluate the effects of early feeding of Fish- and SBM-diet on S. aurata larvae. Eggs were obtained from broodstock maintained at the Instituto de Ciencias Marinas de Andalucía, Cadiz, Spain, and incubated in 200 L incubators at 19 °C and 35 g L−1. Hatched larvae remained in incubators for 24 h with flow-through seawater and aeration, and then they were randomly sorted to two different experimental groups in triplicated 100 L cylindro-conical fiberglass tanks at an initial stocking density of 100 larvae L−1. Larvae were cultured at 19.2 ± 0.06 °C (first experiment) and 19.6 ± 0.08 °C (second experiment), 35 g Kg−1 salinity, pH 8.3, and 12 h light: 12 h darkness. Constant but slight aeration was provided and oxygen level was maintained around 6.0 mg L−1. Rotifers, Brachionus rotundiformis and B. plicatilis (5–10 individuals mL−1), and microalgae Nannochloropsis gaditana (0.5 x l06 to 1 x l06 cells mL−1) were supplied from day 4 post-hatch (dph) following the temporal sequence described by Polo et al. (1992). From 18 dph to 25 dph, the larvae were provided with Artemia sp. at 0.5 nauplii mL−1 in addition to rotifers. From 25 dph to 30 dph rotifers were substituted by 1 metanauplii mL−1, and from 30 dph to 36 dph live feeding only consisted on 10 metanauplii mL−1. The two experimental groups were fed two times a day (15:00 and 17:00 h), in addition to live feed as above, with the FM- and SBM-diet since first feeding until 18 dph (14 d of feeding) or 14 dph (10 d of feeding) in the first and second experiment, respectively. Then, all groups were fed the FM-diet up to 36 dph. Microdiet ingestion since mouth opening was checked by the observation of larvae under a microscope. Samples (entire larvae) were taken at 4, 14 or 18, and 36 dph, rinsed with distilled water, and immediately frozen at -80 °C for protein/enzyme/DNA assays or stored on RNAlater for gene expression analysis. Pools of 100 larvae (4 dph), 50 larvae (14 and 18 dph) and 20 larvae (36 dph) were collected in duplicate from each tank for protein/enzyme/DNA assays, whereas duplicated pools of 5 larvae per tank for expression analysis. Total length was determined each sampling day by measuring 10 larvae per tank (n = 30 per treatment) under a stereomicroscope.
Digestive enzyme activities
Trypsin, chymotrypsin, amylase, lipase, and leucine aminopeptidase activities were assayed in supernatants of homogenates of entire S. aurata larvae in ice-cold 200 mM Tris-HCl, 20 mM NaCl, 20 mM CaCl2, pH 7.5, after centrifugation at 10,000 x g, 4 °C. Brush border anchored enzymes such as alkaline phosphatase and matrix metallo-protease activities were measured after extraction with the same buffer containing 0.2 % Triton X-100.
Trypsin activity was measured using 1.25 mM N-benzoyl-DL-arginine-p-nitroanilide (BApNA) and chymotrypsin activity with 0.1 mM Suc-Ala-Ala-Pro-Phe-p-nitroanilide (SApNA), both in 200 mM Tris-HCl, 20 mM NaCl, 20 mM CaCl2, pH 7.5 (referred henceforth as the assay buffer). Esterase activity was assessed by the hydrolysis of 0.3 mM p-nitrophenyl butyrate (pNPB) in the assay buffer. Lipase activity was measured using 1 mM 4-nitrophenyl palmitate (4NPP) in assay buffer containing 0.3 % Triton X-100. Leucine aminopeptidase activity was assessed by the hydrolysis of 0.5 mM L-Leucine-p-nitroanilide (LeupNA) in assay buffer. Substrate stock solutions were prepared in DMSO (BApNA, SApNA), acetone (pNPB), 1:4 v:v acetonitrile/isopropanol (4NPP), and methanol (LeupNA). The liberation of p-nitroaniline and p-nitrophenol were kinetically followed at 405 nm and extinction coefficients of 6800 M−1·cm−1 and 18,000 M−1·cm−1 were used for p-nitroaniline and p-nitrophenol respectively. Amylase activity was determined using the Amylase Activity Assay Kit (Sigma-Aldrich, St. Louis, MO, USA), and alkaline phosphatase and matrix metallo-protease activities were measured using SensoLyte pNPP Alkaline Phosphatase Assay Kit and SensoLyte Generic MMP Assay Kit, respectively, from AnaSpec, Inc. (Fremont, CA, USA). Commercial kits were used according to manufacturer instructions. All assays were performed in a microplate reader BioTek ELx808 (BioTek Instrument, Inc). One unit of enzyme activity (U) was defined as the amount of enzyme that produces one μmol of product per minute (but 1 nmol min−1 for amylase) at 37 °C. All enzyme activities were expressed as specific activity (U μg−1 of protein for all enzymes, but per mg of protein for MMP).
Expression rate of target genes
Expression was assessed by real-time qPCR for genes involved in inflammation/immunity (proinflammatory cytokines interleukin 1β [il1b], tumor necrosis factor α [tnfa], anti-inflammatory cytokine interleukin 10 [il10]), the breakdown of extracellular matrix and tissue repair/remodeling (matrix metalloproteases 9 and 13 [mmp9, mmp13]), luminal digestion (trypsin [try]), and intestinal nutrient absorption/trafficking (intestinal alkaline phosphatase [alpi], solute carrier family 15, oligopeptide transporter, member 1b [slc15a1b] - here referred as proton/oligopeptides transporter PepT1, fatty acid binding protein 2, intestinal [fabp2], and fatty acid binding protein 6, ileal [fabp6]).
Total RNA was isolated using an Ultra-Turrax T25 (IKA-Werke) and the NucleoSpin RNA II kit (Macherey-Nagel, Düren, Germany) according to the manufacturer’s protocol, including an on-column RNase-free DNase digestion step. Concentration of total RNA was measured with the Qubit 2.0 Fluorometer and the corresponding RNA quantification assay (Invitrogen, Life Technologies), and its quality was determined in an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA) using the Agilent RNA 6000 Nano Kit. Only samples with a RNA Integrity Number (RIN) higher than 8.5 were used. Total RNA (500 ng) was reverse-transcribed in a 20 μL reaction using the qScript DNA synthesis kit (Quanta BioSciences) (5 min at 22 °C, 30 min at 42 °C and 5 min at 85 °C). Primers used (Online Resource 1) were synthesized by IDT (Integrated DNA Technologies, Leuven, Belgium). Various amounts of cDNA were used in triplicate (6 points of serial 1/5 dilutions from 10 ng to 3.2 pg per reaction) as templates to check the assay linearity (R2) and the amplification efficiency (E) of primers. Each qPCR reaction mixture (10 μL) contained 0.5 μL at 400 nM of each specific forward and reverse primers, 5 μL of PerfeCTa SYBR Green FastMix (Quanta Biosciences) and 4 μL of template (10 ng cDNA) in white wells twin.tec real-time PCR plates 96 (Eppendorf). Control reactions with DEPC water and RNA instead of cDNA were included to ensure the absence of contamination or genomic DNA. qPCR was performed with the Eppendorf Mastercycler eprealplex 2 S (Eppendorf, Hamburg, Germany). Cycling conditions were 95 °C, 10 min; [95 °C, 20 s; 60 °C, 35 s] X 40 cycles; melting curve [60 °C to 95 °C, 20 min], 95 °C, 15 s). The melting curve was used to ensure that a single product was amplified and confirm the absence of primer-dimer artifacts. Relative gene quantification was performed using the 2∆∆CT method (Livak and Schmittgen 2001). Data were normalized to both elongation factor 1α (ef1a) and ribosomal 18S, which were previously confirmed to express stably among all samples.
Global DNA methylation and histone (H3, H4) acetylation
Genomic DNA was extracted from whole larvae by the salting-out method (Martínez et al. 1998). DNA concentration was determined using the Invitrogen Qubit 2.0 Fluorometer (Life Technologies, ThermoFisher Scientific). Global methylation status was assessed using the colorimetric ELISA kit (EpiSeeker methylated DNA Quantification Kit, abcam) following manufacturer instructions, with 100 ng of DNA per well as the sample. Total histones were extracted and global acetylation of histones H3 and H4 measured with the EpiSeeker Histone H3 Acetylation Assay Kit (abcam) and the EpiSeeker Histone H4 Acetylation Assay Kit (abcam), respectively, following manufacturer instructions, with 2 μg of histone per well as the sample. Total histones were quantified by their OD230nm at a BioPhotometer Plus (Eppendorf). Methylated DNA and acetylated H3/H4 were quantified at 450 nm in an ELx808 BioTek (BioTek Instrument, Inc.) microplate reader. Methylation status (% of 5-methylcytosine, 5-mC%) was expressed relative to the positive control provided in the kit, whereas acetylation expressed as OD450nm.
Global histone (H3, H4) modifications
Acetylation at specific lysine residues of histones H3 and H4, as well as some other key post-translational modifications (i.e., mono-, di-, and tri-methylation at specific lysine residues) of these histones, were quantified at 450 nm in an ELx808 BioTek (BioTek Instrument, Inc.) microplate reader, using the Histone H3 Modification Multiplex Assay Kit (abcam) and the Histone H4 Modification Multiplex Assay Kit (abcam), following manufacturer instructions, with 150 ng of total histone (extracted and quantified as above) per well as the sample.
Statistical analysis
Results are presented as mean values plus standard error of means (SEM). Data were checked for normality and homogeneity of variance using Shapiro-Wilk and Levene’s tests, respectively. Data were analyzed by Student’s t-test. In few cases, due to heterogeneous variances, we used unequal variance t-test with the Welch correction. Differences with P values <0.05 were considered as significant. Also, we performed a forward stepwise discriminant analysis using the expression of studied genes to understand the combination of variables that can best explain the differences between old larvae with different early nutritional experiences. For the analysis, it was considered F to enter as 0.01, F to remove at 0.0, and tolerance of 0.01. After significant functions development, the relative importance of the original variables was gauged by standardized values. The software package Statistica 7.0 (StatSoft Inc.) was used for all tests performed.
Results
SBM feeding beyond 14 dph produces reversible effects on pancreatic enzymes activities and reduces larval growth
The specific activity of trypsin, chymotrypsin, and amylase (Fig. 1a, c, e) were significantly reduced in larvae fed with the SBM-diet for the first 14 d of feeding (at 18 dph), but size of larvae was not affected (Fig. 2a). The exception was lipase, for which no reduction in specific activity was found (Fig. 1g). When SBM was removed from the diet of these larvae, pancreatic enzymes attained at 36 dph similar levels of specific activity to those in fish fed the FM-diet (Fig. 1a, c, e), except for trypsin (Fig. 1a), and exhibited a significant delay in larval growth (Fig. 2a). Reduction of the SBM feeding to 10 d (up to 14 dph) lead to no differences in specific activity of pancreatic enzymes studied (Fig. 1b, d, h) but amylase (Fig. 1f), and following effects at 36 dph only for chymotrypsin (Fig. 1d). The size of these larvae did not differ between the experimental groups at 36 dph (Fig. 2b).
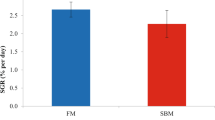
Effect of feeding gilthead seabream with fish meal (FM)- and soybean meal (SBM)-microdiet (black and gray columns, respectively) from mouth opening to 18 dph or 14 dph (equivalent to 14 and 10 d of feeding, respectively) under a co-feeding regimen with live feed, on specific activity of trypsin (A, B), chymotrypsin (Chy) (C, D), amylase (E, F), and lipase (G, H). After the initial feeding period (up to 18 or 14 dph), all fish were fed with the FM based diet and live feed up to 36 dph. One unit of enzyme activity (U) was defined as the amount of enzyme that catalyzed the release of 1 μmol of product per minute (1 nmol·min−1 for amylase), and activities were expressed as activity per μg of protein (U μg−1 of protein). Values are means + SEM, * P < 0.05
Growth of gilthead seabream larvae fed with fish meal (FM)- and soybean meal (SBM)-microdiet during two different periods. Gilthead seabream larvae were fed with a FM-microdiet (black columns) or a SBM-microdiet (gray columns) from mouth opening to 18 dph (A) or 14 dph (B) (equivalent to 14 and 10 d of feeding, respectively) under a co-feeding regimen with live feed, and then fed with the FM-diet and live feed up to 36 dph. Values are means + SEM, ** P < 0.01
Early SBM feeding produces both reversible and lasting effects on MMP, AP and LAP activities
The specific activity of matrix metalloproteinases (MMP) increased in larvae receiving the SBM-diet for 14 d (Fig. 3a), whereas this effect was not observed when the SBM feeding period was shorter (Fig. 3b). Change from the SBM-diet to live food resulted in a significantly decrease in MMPs activity at 36 dph irrespective of the early SBM feeding period (Fig. 3c). The specific activity of AP was significantly lower in S. aurata larvae feeding SBM for 14 d than in larvae fed the FM-diet (Fig. 3c). Later, feeding these larvae with live food up to 36 dph did not allow larvae to exhibit mean activities similar to those of larvae fed with the FM-diet since first feeding (Fig. 3c). Conversely, 10 d of feeding had no effect on AP activity either during or after SBM feeding (Fig. 3d). Moreover, we found that LAP specific activity significantly decreased after 14 d of SBM feeding, a change that was reverted after SBM removal from diet (Fig. 3e). The reduction of SBM period to 10 d leads to no differences in specific LAP activities or final size of larvae (Fig. 2b), but to an increase in activity in the absence of SBM at 36 dph (Fig. 3f).
Effect of feeding gilthead seabream with fish meal (FM)- and soybean meal (SBM)-microdiet (black and gray columns, respectively) from mouth opening to 18 dph (left side graphs) or 14 dph (right side graphs) (equivalent to 14 and 10 d of feeding, respectively) under a co-feeding regimen with live feed, on the specific activities of matrix metalloproteases (MMP) (A, B), alkaline phosphatase (AP) (C, D), and leucine aminopeptidase (LAP) (E, F). After the initial feeding period (up to 18 or 14 dph), all fish were fed with the FM-diet and live feed up to 36 dph. One unit of enzyme activity (U) was defined as the amount of enzyme that catalyzed the release of 1 μmol of product per minute, and activities were expressed as activity per μg of protein (U μg−1 of protein). Values are means + SEM, * P < 0.05, ** P < 0.01
Early SBM feeding produces both reversible and lasting effects on gene expression of different markers of inflammation and tissue remodeling
Feeding S. aurata with SBM for 14 d (up to 18 dph) up-regulated the inflammatory cytokine il1b (Fig. 4a) and the remodeling enzyme mmp13 (Fig. 4e) gene expression. We also observed a non significant trend for mmp9 expression to increase at this time (Fig. 4d). At 36 dph and in the absence of SBM from diet, these larvae exhibited lower expression of il1b (Fig. 4a) and both mmp9 (Fig. 4d) and mmp13 (Fig. 4e) than larvae fed with the FM-diet since first feeding. The analysis of expression data from larvae fed the SBM-diet only during 10 d (until 14 dph) allowed us corroborate the up-regulation of il1b (Fig. 5a), mmp9 (Fig. 5d), and mmp13 (Fig. 5e), but also to observe other changes that occurs earlier and become buried at 18 dph, such as the increased expression of tnfa (Fig. 5c) and alpi (Fig. 5g). After the removal of SBM from diet of these larvae, we observed lower expression of il1b (Fig. 5a), mmp9 (Fig. 5d), and fabp2 (Fig. 5i), and higher expression of the anti-inflammatory interleukin il10 (Fig. 5b) at 36 dph. Thus, both SBM feeding periods triggered changes in gene expression of some markers of inflammation and tissue remodeling, but these were reverted when SBM was removed from diet. Also, both feeding periods leaded to older larvae with higher expression of an anti-inflammatory cytokine.
Effect of feeding gilthead seabream with fish meal (FM)- and soybean meal (SBM)-microdiet (black and gray columns, respectively) from mouth opening to 18 dph (equivalent to 14 d of feeding) under a co-feeding regimen with live feed, on relative expression of selected genes related to inflammation/immunity (interleukin 1β [A], interleukin 10 [B], tumor necrosis factor α [C]), breakdown of extracellular matrix and remodeling (matrix metalloprotease 9 [D], matrix metalloprotease 13 [E]), luminal digestion (trypsin [F]), and intestinal nutrient absorption/trafficking (intestinal alkaline phosphatase [G], solute carrier family 15, member 1 (proton/oligopeptide transporter) [H], fatty acid binding protein 2, intestinal [I], fatty acid binding protein 6, ileal (gastrotropin) [J]). After the initial feeding period (up to 18 dph), all fish were fed with the FM based diet and live feed up to 36 dph. Relative expression was assessed by real-time qPCR and normalized to both elongation factor 1 α (ef1a) and ribosomal 18S. Values are means + SEM. * P < 0.05, ** P < 0.01
Effect of feeding gilthead seabream with fish meal (FM)- and soybean meal (SBM)-microdiet (black and gray columns, respectively) from mouth opening to 14 dph (equivalent to 10 d of feeding) under a co-feeding regimen with live feed, on relative expression of selected genes related to inflammation/immunity (interleukin 1β [A], interleukin 10 [B], tumor necrosis factor α [C]), breakdown of extracellular matrix and remodeling (matrix metalloprotease 9 [D], matrix metalloprotease 13 [E]), luminal digestion (trypsin [F]), and intestinal nutrient absorption/trafficking (intestinal alkaline phosphatase [G], solute carrier family 15, member 1 (proton/oligopeptide transporter) [H], fatty acid binding protein 2, intestinal [I], fatty acid binding protein 6, ileal (gastrotropin) [J]). After the initial feeding period (up to 14 dph), all fish were fed with the FM based diet and live feed up to 36 dph. Relative expression was assessed by real-time qPCR and normalized to both elongation factor 1 α (ef1a) and ribosomal 18S. Values are means + SEM. * P < 0.05, ** P < 0.01
Duration of the SBM feeding period drives the physiology of larvae in different directions
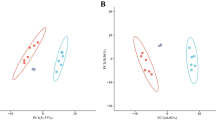
Two significant discriminant functions could be developed from the relative expression of studied genes in 36 dph larvae. These functions collectively accounted for 100 % of the total variance (Table 2). Expression of genes involved in inflammatory/anti-inflammatory processes, tissue remodeling, and nutrient transport were the variables with the highest relevance in the first discriminant function (Table 3), which accounted for 73 % of the total variance (Table 2). In this function, two groups of SBM feeding larvae showed opposed central values depending on early nutrition (Fig. 6, axis x). On the other hand, the second discriminant function better discriminated among SBM and FM diets irrespective of the SBM feeding period (Fig. 6, axis y).
Plots of the two axes from the forward stepwise discriminant function analysis of gene expression in gilthead seabream larvae. Larvae were fed, under a co-feeding regimen with live feed, from first feeding up to 14 dph with a FM-based diet or a SBM-based diet, or up to 18 dph with the same diets. Later, all larvae were fed with the FM-diet and live feed up to 36 dph. These older larvae were referred as FM 36 dph when fed on the FM-diet, SBM 14–36 dph if fed with the SBM-diet up to 14 dph and then with the FM-diet, and SBM 18–36 dph if fed with the SBM-diet up to 18 dph and then with the FM-diet
Early SBM feeding results in global hypomethylation
We observed no difference in global methylation level between S. aurata larvae fed with SBM up to 14 or 18 dph and larvae fed with the FM-diet (Fig. 7a,b). However, S. aurata larvae that faced SBM at first feeding showed a significant global DNA hypomethylation at 36 dph irrespective of the duration of the SBM feeding period (Fig. 7a,b).
Effect of feeding gilthead seabream with fish meal (FM)- and soybean meal (SBM)-microdiet (black and gray columns, respectively) from mouth opening to 18 dph or 14 dph (equivalent to 14 and 10 d of feeding, respectively) under a co-feeding regimen with live feed, on chromatin modifications. Global DNA methylation (A, B), histone H3 (C, D) and H4 (E, F) acetylation of entire larvae. After the initial feeding period (up to 18 or 14 dph), all fish were fed with the FM- based diet and live feed up to 36 dph. Values are means + SEM. * P < 0.05
H3K14ac and H3K27m3: the most dynamic, but not the only, epigenetic marks responding to SBM
Feeding S. aurata with SBM from mouth opening up to 14 dph (10 d of feeding) significantly increased global H3 acetylation (Fig. 7d). This SBM-induced H3 hyperacetylation was transient, as the level of acetylation was restored in later larvae in the presence (at 18dph, Fig. 7c) or absence of SBM (at 36 dph, Fig. 7d). We found a slight hypoacetylation of histone H4 at 18 dph in larvae fed with the SBM-diet (Fig. 7e), but no such effect if the feeding period was restricted up to 14 dph (Fig. 7f) or if SBM was removed from later diet (Fig. 7e,f).
To gain insight on the type of histone modifications that occurred as response to SBM in S. aurata larvae, we further assessed H3 and H4 modifications at 14 and 36 dph in fish fed with the SBM- and the FM-diet during the first 10 d of feeding. SBM feeding resulted in hyperacetylation of histone H3 at lysine 14 (H3K14ac) (Fig. 8a), whereas acetylation level of other H3 lysine residues was not modified or even decreased (H3K18ac) (Fig. 8a). Once SBM was removed from diet, we observed a significant hypoacetylation of the same lysine residue (H3K14ac) but also of H3K56ac (Fig. 8b). These reversible SBM-induced H3 acetylations occurred along with other histone modification such as the tri-methylation of the same histone at lysine residue 27 (H3K27m3). After 10 d of SBM feeding, we found a significant decrease in H3K27m3 (Fig. 8a), but upon removal of SBM from diet, H3K27m3 significantly increased over the level observed in fish fed the FM-diet (Fig. 8a). We found no difference between dietary treatments on acetylation level of H4 at lysine residues 5, 8, 12, and 16, or in other modifications such as mono- and tri-methylation of lysine 20 (Fig. 8c,d).
Effect of feeding gilthead seabream with fish meal (FM)- and soybean meal (SBM)-microdiet (black and gray columns, respectively) from mouth opening to 14 dph (equivalent to 10 d of feeding) under a co-feeding regimen with live feed, on global histone H3 modifications of entire larvae at 14 dph (A) and 36 dph (B), and on global histone H4 modifications of entire larvae at 14 dph (C) and 36 dph (D). After the initial feeding period (up to 14 dph), all fish were fed with the FM-based and live feed diet up to 36 dph. Values are means + SEM. * P < 0.05, ** P < 0.01, *** P < 0.001
Discussion
In this work we examined the effects of SBM in diet on S. aurata larvae at first feeding during two different developmental windows. Early SBM feeding of S. aurata larvae significantly affects the activity of most pancreatic enzymes in a time-of-exposure dependent form. More than 10 d of SBM feeding (i.e., beyond 14 dph) delayed the normal development of S. aurata larvae digestive capacities as the activities of all trypsin, chymotrypsin, and amylase were significantly reduced. This is opposed to the typical response of juvenile and adult fish to SBM. Protease inhibitors present in SBM can partially abrogate the activity of trypsin and chymotrypsin in the proximal intestine (Robaina et al. 1995), and juveniles of S. aurata (Santigosa et al. 2010) and other fish such as Atlantic salmon (Lilleeng et al. 2007; Chikwati et al. 2013b) exhibit a rapid compensatory increase in activities of these enzymes. The premature stage of pancreas development in early S. aurata larvae would lie behind their failure to respond to SBM with trypsin synthesis. Strengthening this hypothesis is the fact that we did not observe changes in trypsin expression in S. aurata larvae in response to SBM. This is also in line with the thought that regulatory mechanisms for the production of digestive enzymes in early fish larvae are not efficient (Zambonino-Infante and Cahu 2001). However, SBM-induced increase in trypsin activity in juvenile fish is more marked in the distal intestine and have been attributed to a reduced ability to reabsorb the pancreatic enzymes (Krogdahl et al. 2003) and the up-regulation of trypsin-like activity by immune cells (Lilleeng et al. 2007). Therefore, other plausible explanation for our observations is that these intestinal processes are not fully functional in early larvae.
Conversely, lipase activity was relatively insensitive to SBM in S. aurata larvae, as reported before in post-smolt Atlantic salmon (Chikwati et al. 2013b). The observed decrease in pancreatic proteases may be responsible for less lipase inactivation in the digestive tract explaining the stable lipase activity. Other, more likely explanation, is that regulation of lipase activity in S. aurata larvae and other pancreatic enzymes proceeds by distinct mechanisms. A recent study in S. aurata larvae supports this hypothesis (Mata-Sotres et al. 2016). The later removal of SBM from diet of S. aurata larvae that fed SBM during 14 d lead larva to resume the activity of most pancreatic enzymes but produced a significant reduction in size of older larvae suggesting that some physiological processes remain altered.
After 10 d of SBM feeding, S. aurata larvae did not modify LAP and AP activities. Conversely, a longer SBM feeding period resulted in decreased of both LAP and AP activities. Our observations on LAP activity in hole larvae resembles the decreased in LAP activity the distal intestine of salmon fed SBM (Chikwati et al. 2013b). Observations in salmonids lead to the suggestion that SBM-induced decrease in LAP activity results from increased sloughing of intestinal epithelia cells (Chikwati et al. 2013a) and epithelial proliferation with immature cell population expressing lower levels of brush border enzymes (Bakke-McKellep et al. 2007). This may be also a likely explanation for our result because LAP activity was recovered ones SBM was removed from diet. However, our analysis did not differentiate between intestinal LAP and other LAPs enzymes ubiquitously distributed so results must be taken with caution. The same is true for AP, an enzyme activity distributed all over the larvae though enriched at the intestine. IAP is involved in different pivotal process such as negative regulator of lipid absorption, pH regulation, lipopolysaccharide detoxification/tolerance to commensal microbiota, inflammation attenuation and intestine homeostasis (Lallès 2010) and its activity is known to decrease in fish feeding soybean anti-nutrients (Couto et al. 2015). Interestingly, after a restricted (10 d) exposure to SBM older larvae had more LAP activity in the absence of SBM, while such long-term effect was not observed for AP.
After 10 d of SBM feeding, S. aurata larvae increased MMPs gene expression but this did not result in higher activity. A longer SBM feeding period increased both gene expression and activity of MMPs in S. aurata larvae. Increased MMP activities (e.g., mmp9 and mmp13) have been well documented during inflammation (Pedersen et al. 2015) and tissue injury (Castillo-Briceño et al. 2009) in fish, but not studied before in S. aurata in response to SBM. As in other vertebrates, MMPs are induced in fish by the pro-inflammatory cytokines IL-1β and TNF-α (Johnson et al. 2004) and in accordance, we observed increased tnfa expression due to SBM feeding of S. aurata larvae. However, tnfa up-regulation was not sustained after 10 d of feeding, whereas a synchronized behavior for mmp9 and il1b expression was observed irrespective of the SBM time of exposure. These observations were not surprising, as tnfa from S. aurata is relatively irresponsive to some stimuli known to up-regulate il1b (García-Castillo et al. 2002; Castillo-Briceño et al. 2009). Other studies in fish (Chadzinska et al. 2008) and other vertebrates (Schӧnbeck et al. 1998) also pointed to a major functional interplay between fish MMP9 and IL-1β during both the initiation and the resolution of the inflammatory response. Larvae receiving SBM at first feeding appear later to have, in the absence of SBM, reduced il1b, mmp9, and 13 expressions, irrespective of the severity of the SBM short-term effects. The exception was il10, whose up-regulation was only observed in old larvae receiving SBM for the shortest period. It is not known if this is a result of different time-course responses of the different cytokines (Pérez-Cordón et al. 2014) or a true lasting effect of early nutrition.
The expression of PepT1, mostly in the proximal intestine, is known to be related with di- and tri-peptide transport efficiency and growth in fish (Romano et al. 2014). In S. aurata juveniles, expression of PepT1 was shown to be affected by some vegetal proteins and not by others (Terova et al. 2013). Here we showed that PepT1 expression in S. aurata larvae is not affected by SBM at the short-term, but that SBM feeding for either 10 d or 14 d lead to older larvae with slightly higher mean values of PepT1 expression, although not statistically different to those in larvae fed the FM-diet. However, this observation is noteworthy as we showed that the increased PepT1 expression of juvenile zebrafish in response to SBM was higher if they shortly faced SBM at first feeding (Perera and Yúfera 2016). On the other hand, we did not observe a significant impact of either 10 or 14 d of SBM feeding on the expression of fapp2 and fabp6. Conversely, fabp2, involved in fatty acid transport, decreased in Atlantic salmon at the transcript and protein levels during SBM-induced distal intestinal inflammation (Venold et al. 2013) as a result of a down regulation of regulatory proteins, rapid enterocyte turnover, and reduced lipid digestibility (Krogdahl et al. 2003). Also, fabp6, which is thought to be needed for the efficient intracellular transport of bile acids in enterocytes (Gong et al. 1994) did not vary its expression due to SBM or resulted altered later in larval development. Conversely, we found in zebrafish juveniles a decreased expression of fabp6 after SBM feeding of juveniles, whether or not they were provided with SBM at first feeding (Perera and Yúfera 2016).
Multivariate analysis performed confirmed that the duration of the SBM feeding period drives some features of the physiology of larvae in different directions, mostly evidenced by some genes related with inflammation, tissue remodeling, and nutrient transport. Further studies are necessary to determine if these effects persist in more advanced developmental stages (i.e., true nutritional programming).
We found that SBM feeding of S. aurata larvae globally increase H3K14ac, which is a well known activating mark of transcriptional initiation and enhancers expression (Karmodiya et al. 2012), elongation (Johnsson et al. 2009), and DNA repair (Duan and Smerdon 2014), although it can also mark inactive promoters to a lesser extent (Karmodiya et al. 2012). Concurrently, we observed that SBM decreased globally the repressive mark H3K27m3. H3K27me3 is known to occur along with histone activating marks such as H3K14ac at bivalent promoters (Karmodiya et al. 2012) and its decrease may also indicate the up-regulation of regulatory genes. Our results suggest that these chromatin modifications may be related with the transcriptional activation of SBM-affected genes during feeding. After SBM removal from diet, the level of the activating marks H3K14ac and H3K56ac decreased, restoring the global acetylation level. This behavior was expected as histone acetylation is a dynamic process and changes are reverted upon removal of activators or repressors (Katan-Khaykovich and Struhl 2002). Results suggest that after SBM removal from diet, larvae undergo overall gene down-regulation, which is also supported by the later increase of the repressive mark H3K27m3. It is interesting that H3K27m3, which is known to be up-regulated and persistent at advanced stages of inflammation, has been proposed to render affected genes such as cytokines less responsive to further inflammatory stimulation (i.e., reprogramming effect) (Angrisano et al. 2010). We also observed H3K56ac to decrease after SBM removal from diet. This mark is paramount for genome stability and its down-regulation promote DNA repair (Miller et al. 2010).
On the other hand, we demonstrated that early SBM feeding of S. aurata larvae resulted in lasting global hypomethylation. Global methylation data are difficult to read as transcriptional activation has been associated with both hypomethylation (Reik and Dean 2001) and hypermethylation (Gavery and Roberts 2013). Hypomethylation has been associated with the re-programming of developmentally silenced genes during tissue regeneration (Thummel et al. 2006) and DNA repair (Reik 2007).
Nutrition is known to affects epigenetic phenomena by nutrients acting as a source of methyl groups or as co-enzymes for one-carbon metabolism (Jaenisch and Bird 2003), or by affecting the activity of histone modifier enzymes (Feil and Fraga 2012). The mechanisms by which SBM exerts the observed epigenetic effects on fish larvae are unknown. It may be related with some SBM component known to act as a histone deacetylase inhibitors such as genistein and other isoflavones (Link et al. 2010), or through the action of microbiota metabolites with histone deacetylase inhibitory properties such as butyrate (Chang et al. 2014), or with the capacity to increase the histone acetylation substrate acetyl-coA, such as acetate (Kendrick et al. 2010). Butyrate and acetate are known to modulate the expression of cytokine genes (Karmodiya et al. 2012; Fusunyan et al. 1999) and intestinal alkaline phosphatase (Hinnebusch et al. 2003) through histone acetylation. Genistein and other isoflavones may also be responsible for the DNA demethylation observed in this study through DNA methyl transferases inhibition (Link et al. 2010). These mechanisms deserved further studies in fish larvae.
Feeding trials with larvae are challenging, largely because of the limited ability to assess feed intake. Accordingly, there are some confounding factors in our experiments associated with possible differences in feed intake of microdiets (e.g. due to less palatability of the SBM-diet). These may lead to uncertainness on the role of SBM as origin of the differences detected. Considering these limitations, we ensure microdiet ingestion by undersupply live food and by providing microdiets at times of high ingestion rate in S. aurata larvae (Mata-Sotres et al. 2015). Ingestion of microdiets was confirmed in all cases by the observation of larvae under a microscope. This feeding regime did not affect per se larvae growth as compared to a traditional regime with only live feed. The size of larvae at 15 dph observed this work (4.5–5 mm) is similar to that observed in other studies for larvae of the same age (5–5.5 mm, Polo et al. 1992; 4.8–5 mm, Izquierdo et al. 2013). The same is true at 36 dph (7.5–8 mm, this work; 7–7.5 mm, Izquierdo et al. 2013) for larvae feed with the FM- or SBM-microdiets for the first 10 days. In addition, different results from SBM fed S. aurata larvae resemble those reported for juvenile fish of different species after SBM feeding as discussed above, confirming that larvae ingested enough microdiets for dietary treatments to exert the observed effects. It is also pertinent to remark that due to their small size, dissection of individual tissues/organs is usually impractical in fish larval studies. Thus, further studies are needed for sorting the observed results to different S. aurata organs, which will assist in advancing the knowledge on particular physiological processes affected by SBM inclusion in early diet.
In conclusion, the present study shows that i) there are limitations for expanding SBM feeding of S. aurata larvae beyond 10 d from mouth opening (i.e., 14 dph), affecting some key physiological processes and ultimately normal growth, and ii) it is worthy to explore the possibility to manage the future responses of S. aurata to SBM through early nutritional manipulation, even within a short developmental window of 10 d, given the absence of evident effects on normal development and growth, and the occurrence of apparently lasting effects on the activity of some key enzymes, genes, and chromatin modifications.
References
Angrisano T, Pero R, Peluso S, Keller S, Sacchetti S, Bruni CB, Chiariotti L, Lembo F (2010) LPS-induced IL-8 activation in human intestinal epithelial cells is accompanied by specific histone H3 acetylation and methylation changes. BMC Microbiol 10:172. doi:10.1186/1471-2180-10-172
Bakke-McKellep AM, Penn MH, Salas PM, Refstie S, Sperstad S, Landsverk T, Ringø E, Krogdahl Å (2007) Effects of dietary soyabean meal, inulin and oxytetracycline on intestinal microbiota and epithelial cell stress, apoptosis and proliferation in the teleost Atlantic salmon (Salmo salar L.). Br J Nutr 97:699–713. doi:10.1017/S0007114507381397
Basurco B, Lovatelli A, García B (2011) Current status of Sparidae aquaculture. In: Pavlidis MA, Mylonas CC (eds) Sparidae: Biology and Aquaculture of Gilthead Sea Bream and Other Species. Blackwell Publishing Ltd
Bonaldo A, Roem AJ, Fagioli P, Pecchini A, Cipollini I, Gatta PP (2008) Influence of dietary levels of soybean meal on the performance and gut histology of gilthead sea bream (Sparus aurata L.) and European sea bass (Dicentrarchus labrax L.). Aquac Res 39:970–978. doi:10.1111/j.1365-2109.2008.01958.x
Castillo-Briceño P, Sepulcre MP, Chaves-Pozo E, Meseguer J, García-Ayala A, Mulero V (2009) Collagen regulates the activation of professional phagocytes of the teleost fish gilthead seabream. Mol Immunol 46:1409–1415. doi:10.1016/j.molimm.2008.12.005
Chadzinska M, Baginski P, Kolaczkowska E, Savelkoul HFJ, Verburg-van Kemenade BML (2008) Expression profiles of matrix metalloproteinase 9 in teleost fish provide evidence for its active role in initiation and resolution of inflammation. Immunology 125:601–610. doi:10.1111/j.1365-2567.2008.02874.x
Chang PV, Hao L, Offermanns S, Medzhitov R (2014) The microbial metabolite butyrate regulates intestinal macrophage function via histone deacetylase inhibition. Proc Natl Acad Sci U S A 111:2247–2252. doi:10.1073/pnas.1322269111
Chikwati EM, Gu J, Penn MH, Bakke AM, Krogdahl Å (2013a) Intestinal epithelial cell proliferation and migration in Atlantic salmon Salmo salar L.: effects of temperature and inflammation. Cell Tissue Res 353:123–137. doi:10.1007/s00441-013-1631-9
Chikwati EM, Sahlmann C, Holm H, Penn MH, Krogdahl Å, Bakke AM (2013b) Alterations in digestive enzyme activities during the development of diet-induced enteritis in Atlantic salmon, Salmo salar L. Aquaculture 402-403:28–37. doi:10.1016/j.aquaculture.2013.03.023
Couto A, Kortner TM, Penn M, Bakke AM, Krogdahl Å, Oliva-Teles A (2015) Saponins and phytosterols in diets for European sea bass (Dicentrarchus labrax) juveniles: effects on growth, intestinal morphology and physiology. Aquacult Nut 21:180–193. doi:10.1111/anu.12146
Dias J, Conceição LEC, Ribeiro AR, Borges P, Valente LMP, Dinis MT (2009) Practical diet with low fish-derived protein is able to sustain growth performance in gilthead seabream (Sparus aurata) during the grow-out phase. Aquaculture 293:255–262. doi:10.1016/j.aquaculture.2009.04.042
Drew MD, Borgeson TL, Thiessen DL (2007) A review of processing of feed ingredients to enhance diet digestibility in finfish. Anim Feed Sci Tech 138:118–136. doi:10.1016/j.anifeedsci.2007.06.019
Duan MR, Smerdon MJ (2014) Histone H3 lysine 14 (H3K14) acetylation facilitates DNA repair in a positioned nucleosome by stabilizing the binding of the chromatin remodeler RSC (remodels structure of chromatin). J Biol Chem 289:8353–8363. doi:10.1074/jbc.M113.540732
Elbal MT, García-Hernández MP, Lozano MT, Agulleiro B (2004) Development of the digestive tract of gilthead sea bream (Sparus aurata L.). Light and electron microscopic studies. Aquaculture 234:215–238. doi:10.1016/j.aquaculture.2003.11.028
Fang L, Liang XF, Zhou Y, Guo XZ, He Y, Yi TL, Liu LW, Yuan XC, Tao YX (2014) Programming effects of high-carbohydrate feeding of larvae on adult glucose metabolism in zebrafish, Danio rerio. Br J Nut 111:808–818. doi:10.1017/S0007114513003243
Feil R, Fraga MF (2012) Epigenetics and the environment: emerging patterns and implications. Nat Rev Genet 13:97–109. doi:10.1038/nrg3142
Fusunyan RD, Quinn JJ, Fujimoto M, MacDermott RP, Sanderson IR (1999) Butyrate switches the pattern of chemokine secretion by intestinal epithelial cells through histone acetylation. Mol Med 5:631–640
García-Castillo J, Pelegrín P, Mulero V, Meseguer J (2002) Molecular cloning and expression analysis of tumor necrosis factor α from a marine fish reveal its constitutive expression and ubiquitous nature. Immunogenetics 54:200–207
Gavery MR, Roberts SB (2013) Predominant intragenic methylation is associated with gene expression characteristics in a bivalve mollusc. PeerJ 1:e215. doi:10.7717/peerj.215
Geurden I, Mennigen J, Plagnes-Juan E, Veron V, Cerezo T, Mazurais D, Zambonino-Infante J, Gatesoupe J, Skiba-Cassy S, Panserat S (2014) High or low dietary carbohydrate:protein ratios during first feeding affect glucose metabolism and intestinal microbiota in juvenile rainbow trout. J Exp Biol 217:3396–3406. doi:10.1242/jeb.106062
Geurden I, Borchert P, Balasubramanian MN, Schrama JW, Dupont-Nivet M, Quillet E, Kaushik SJ, Panserat S, Médale F (2013) The positive impact of the early-feeding of a plant-based diet on its future acceptance and utilization in rainbow trout. PLoS One 8:e83162. doi:10.1371/journal.pone.0083162
Gong Y-Z, Everett ET, Schwartz DA, Norris JS, Wilson FA (1994) Molecular cloning, tissue distribution, and expression of a 14 kDa bile acid-binding protein from rat ileal cytosol. Proc Natl Acad Sci U S A 91:4741–4745
Gu M, Bai N, Zhang Y, Krogdahl Å (2016) Soybean meal induces enteritis in turbot Scophthalmus maximus at high supplementation levels. Aquaculture 464:286–295. doi:10.1016/j.aquaculture.2016.06.035
Hardy RW (2010) Utilization of plant proteins in fish diets: effects of global demand and supplies of fishmeal. Aquac Res 41:770–776. doi:10.1111/j.1365-2109.2009.02349.x
Hinnebusch BF, Henderson JW, Siddique A, Malo MS, Zhang W, Abedrapo MA, Hodin RA (2003) Transcriptional activation of the enterocyte differentiation marker intestinal alkaline phosphatase is associated with changes in the acetylation state of histone H3 at a specific site within its promoter region in vitro. J Gastrointest Surg 7:237–245
Izquierdo MS, Scolamacchia M, Betancor M, Roo J, Caballero MJ, Terova G (2013) Witten PE (2013) effects of dietary DHA and a-tocopherol on bone development, early mineralisation and oxidative stress in Sparus aurata (Linnaeus, 1758) larvae. Br J Nutr 109:1796–1805. doi:10.1017/S0007114512003935
Jaenisch R, Bird A (2003) Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat Genet 33(Suppl):245–254
Johnson MC, Sangrador-Vegas A, Smith TJ, Cairns MT (2004) Molecular cloning and expression analysis of rainbow trout (Oncorhynchus mykiss) matrix metalloproteinase-9. Fish Shellfish Immunol 17:499–503. doi:10.1016/j.fsi.2004.05.005
Johnsson A, Durand-Dubief M, Xue-Franzén Y, Rönnerblad M, Ekwall K, Wright A (2009) HAT-HDAC interplay modulates global histone H3K14 acetylation in gene-coding regions during stress. EMBO Rep 10:1009–1014. doi:10.1038/embor.2009.127
Karmodiya K, Krebs AR, Oulad-Abdelghani M, Kimura H, Tora L (2012) H3K9 and H3K14 acetylation co-occur at many gene regulatory elements, while H3K14ac marks a subset of inactive inducible promoters in mouse embryonic stem cells. BMC Genomics 13:424. doi:10.1186/1471-2164-13-424
Katan-Khaykovich Y, Struhl K (2002) Dynamics of global histone acetylation and deacetylation in vivo: rapid restoration of normal histone acetylation status upon removal of activators and represors. Gene Dev 16:743–752
Kemski MM, Kwasek K, Wojno M, Wick M, Dabrowski K (2015) Soybean-dependent nutritional programming: effect of soybean diet intake during early life stages on adult growth in yellow perch Perca flavescens. Aquaculture America 2015-New Orleans, Louisiana-Meeting Abstract. https://www.was.org/meetings/mobile/MG_Paper.aspx?i=34860
Kendrick SFW, O’Boyle G, Mann J, Zeybel M, Palmer J, Jones DEJ, Day CP (2010) Acetate, the key modulator of inflammatory responses in acute alcoholic hepatitis. Hepatology 51:1988–1997. doi:10.1002/hep.23572
Kortner TM, Gu J, Krogdahl Å, Bakke AM (2013) Transcriptional regulation of cholesterol and bile acid metabolism after dietary soyabean meal treatment in Atlantic salmon (Salmo salar L.). Br J Nutr 109:593–604. doi:10.1017/S0007114512002024
Koven WM, van Anholt R, Lutzky S, Ben AI, Nixon O, Ron B, Tandler A (2003) The effect of dietary arachidonic acid on growth, survival, and cortisol levels in different-age gilthead sea bream larvae (Sparus aurata) exposed to handling or daily salinity change. Aquaculture 228:307–320. doi:10.1016/S0044-8486(03)00317-X
Krogdahl Å, Bakke-McKellep AM, Baeverfjord G (2003) Effects of graded levels of standard soybean meal on intestinal structure, mucosal enzyme activities, and pancreatic response in Atlantic salmon (Salmo salar L). Aquac Nutr 9:361–371. doi:10.1046/j.1365-2095.2003.00264.x
Krogdahl Å, Penn MH, Thorsen J, Refstie S, Bakke AM (2010) Important antinutrients in plant feedstuffs for aquaculture: an update on recent findings regarding responses in salmonids. Aquac Res 41:333–344. doi:10.1111/j.1365-2109.2009.02426.x
Krogdahl Å, Gajardo K, Kortner TM, Penn M, Gu M, Berge GM, Bakke AM (2015) Soya saponins induce enteritis in Atlantic salmon (Salmo salar L.). J Agric Food Chem 63:3887–3902. doi:10.1021/jf506242t
Lallès JP (2010) Intestinal alkaline phosphatase: multiple biological roles in maintenance of intestinal homeostasis and modulation by diet. Nutr Rev 68:323–332. doi:10.1111/j.1753-4887.2010.00292.x
Lilleeng E, Froystad MK, Ostby GC, Valen EC, Krogdahl Å (2007) Effects of diets containing soybean meal on trypsin mRNA expression and activity in Atlantic salmon (Salmo salar L). Comp Biochem Physiol 147A:25–36
Lillycrop KA, Burdge GC (2012) Epigenetic mechanisms linking early nutrition to long term health. Best Pract Res Clin Endocrinol Metab 26:667–676. doi:10.1016/j.beem.2012.03.009
Link A, Balaguer F, Goel A (2010) Cancer chemoprevention by dietary polyphenols: promising role for epigenetics. Biochem Pharmacol 80:1771–1792. doi:10.1016/j.bcp.2010.06.036
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C (T)) method. Methods 25:402–408
Mata-Sotres JA, Martínez-Rodríguez G, Pérez-Sánchez J, Sánchez-Vázquez FJ, Yúfera M (2015) Daily rhythms of clock gene expression and feeding behavior during the larval development in gilthead seabream, Sparus aurata. Chronobiol Int 32:1061–1074. doi:10.3109/07420528.2015.1058271
Mata-Sotres JA, Moyano FJ, Martínez-Rodríguez G, Yúfera M (2016) Daily rhythms of digestive enzyme activity and gene expression in gilthead seabream (Sparus aurata) during ontogeny. Comp Biochem Physiol 197A:43–51. doi:10.1016/j.cbpa.2016.03.010
Martínez G, Shaw EM, Carrillo M, Zanuy S (1998) Protein salting-out method applied to genomic DNA isolation from fish whole blood. BioTechniques 24:238–239
Miller KM, Tjeertes JV, Coates J, Legube G, Polo SE, Britton S, Jackson SP (2010) Human HDAC1 and HDAC2 function in the DNA-damage response to pro-mote DNA nonhomologous end-joining. Nat Struct Mol Biol 17:1144–1151. doi:10.1038/nsmb.1899
Moyano FJ, Diaz M, Alarcón FJ, Sarasquete MC (1996) Characterization of digestive enzyme activity during larval development of gilthead seabream (Sparus aurata). Fish Physiol Biochem 15:121–130
Naylor RL, Goldburg RJ, Primavera JH, Kautsky N, Beveridge MCM, Clay J, Folke C, Lubchenco J, Mooney H, Troell M (2000) Effect of aquaculture on world fish supplies. Nature 405:1017–1024
Pedersen ME, Vuong TT, Rønning SB, Kolset SO (2015) Matrix metalloproteinases in fish biology and matrix turnover. Matrix Biol 44-46:86–93. doi:10.1016/j.matbio.2015.01.009
Perera E, Yúfera M (2016) Soybean meal and soy protein concentrate in early diet elicit different nutritional programming effects on juvenile zebrafish. Zebrafish 13:61–69. doi:10.1089/zeb.2015.1131
Pérez-Cordón G, Estensoro I, Benedito-Palos L, Calduch-Giner JA, Sitjà-Bobadilla A, Pérez-Sánchez J (2014) Interleukin gene expression is strongly modulated at the local level in a fish-parasite model. Fish Shellfish Immunol 37:201–208. doi:10.1016/j.fsi.2014.01.022
Polo A, Yúfera M, Pascual E (1992) Feeding and growth of gilthead seabream (Sparus aurata L.) larvae in relation to the size of the rotifer strain used as food. Aquaculture 103:45–54
Reik W (2007) Stability and flexibility of epigenetic gene regulation in mammalian development. Nature 447:425–432
Reik W, Dean W (2001) DNA methylation and mammalian epigenetics. Electrophoresis 22:2838–2843
Robaina L, Izquierdo MS, Moyano FJ, Socorro J, Vergara JM, Montero D, Fernández-Palacios H (1995) Soybean and lupin seed meals as protein-sources in diets for gilthead seabream (Sparus aurata): nutritional and histological implications. Aquaculture 130:219–233. doi:10.1016/0044-8486(94)00225-D
Romano A, Barca A, Storelli C, Verri T (2014) Teleost fish models in membrane transport research: the PEPT1(SLC15A1) H+-oligopeptide transporter as a case study. J Physiol 592:881–897. doi:10.1113/jphysiol.2013.259622
Sahlmann C, Gu J, Kortner TM, Lein I, Krogdahl Å, Bakke AM (2015) Ontogeny of the digestive system of Atlantic salmon (Salmo salar L.) and effects of soybean meal from start-feeding. PLoS One 10:e0124179. doi:10.1371/journal.pone.0124179
Sanden M, Berntssen MHG, Krogdahl Å, Hemre G-I, Bakke-McKellep A-M (2005) An examination of the intestinal tract of Atlantic salmon, Salmo salar L., parr fed different varieties of soy and maize. J Fish Dis 28:317–330
Santigosa E, Sáenz de Rodrigáñez MA, Rodiles A, Barroso FG, Alarcón FJ (2010) Effect of diets containing a purified soybean trypsin inhibitor on growth performance, digestive proteases and intestinal histology in juvenile sea bream (Sparus aurata L.). Aquac Res 41:e187–e198. doi:10.1111/j.1365-2109.2010.02500.x
Schӧnbeck U, Mach F, Libby P (1998) Generation of biologically active IL-1β by matrix metalloproteinases: a novel caspase-1-independent pathway of IL-1β processing. J Immunol 161:3340–3346
Terova G, Robaina L, Izquierdo M, Cattaneo A, Molinari S, Bernardini G, Saroglia M (2013) PepT1 mRNA expression levels in sea bream (Sparus aurata) fed different plant protein sources. Springerplus 2:17. doi:10.1186/2193-1801-2-17
Thummel R, Burket CT, Hyde DR (2006) Two different transgenes to study gene silencing and re-expression during zebrafish caudal fin and retinal regeneration. ScientificWorldJournal 6(Suppl 1):65–81. doi:10.1100/tsw.2006.328
Vagner M, Robin JH, Zambonino-Infante JL, Tocher DR, Person-Le Ruyet J (2009) Ontogenic effects of early feeding of sea bass (Dicentrarchus labrax) larvae with a range of dietary n-3 highly unsaturated fatty acid levels on the functioning of polyunsaturated fatty acid desaturation pathways. Br J Nutr 101:1452–1462. doi:10.1017/S0007114508088053
Venou B, Alexis MN, Fountoulaki E, Haralabous J (2006) Effects of extrusion and inclusion level of soybean meal on diet digestibility, performance and nutrient utilization of gilthead sea bream (Sparus aurata). Aquaculture 261:343–356. doi:10.1016/j.aquaculture.2006.07.030
Venold FF, Penn MH, Thorsen J, Gu J, Kortner TM, Krogdahl Å, Bakke AM (2013) Intestinal fatty acid binding protein (fabp2) in Atlantic salmon (Salmo salar): localization and alteration of expression during development of diet induced enteritis. Comp Biochem Physiol 164A:229–240. doi:10.1016/j.cbpa.2012.09.009
Yúfera M, Fernandez-Díaz C, Pascual E (2005) Food microparticles for larval fish prepared by internal gelation. Aquaculture 248:253–262. doi:10.1016/j.aquaculture.2005.04.026
Zambonino-Infante JL, Cahu CL (2001) Ontogeny of the gastrointestinal tract of marine fish larvae. Comp Biochem Physiol 130C:477–487
Acknowledgments
This research was funded by the European Union through the grant FISHPROG (326245). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the article. The views expressed in this work are the sole responsibility of the authors. Special thank to Yannis Kotzamanis who kindly donated the SBM used in this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(PDF 75 kb)
Rights and permissions
About this article
Cite this article
Perera, E., Yúfera, M. Effects of soybean meal on digestive enzymes activity, expression of inflammation-related genes, and chromatin modifications in marine fish (Sparus aurata L.) larvae. Fish Physiol Biochem 43, 563–578 (2017). https://doi.org/10.1007/s10695-016-0310-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10695-016-0310-7