Abstract
Genotype by environment interaction remains a substantial issue in all breeding programs. Crop genotypes are generally developed in a central breeding location, but always require the evaluation of breeding products in different environments. This is particularly relevant in countries that have a wide range of climates. Eighteen cassava genotypes were evaluated in Cameroon in eight environments—varying in seasonal rainfall and temperature patterns and soil characteristics—over two cropping seasons. Soil nutrient content was analyzed and trials were established in a randomized complete block design in three replications. Response of genotypes to major cassava pests and diseases, yield and carotenoids content was evaluated. It was observed that four genotypes did not show cassava mosaic disease (CMD) symptoms irrespective of the environments. The local check had highest CMD incidence and severity across all environments. Average number of whitefly per plant across all environments was highest on TMS 96/0023. Average cassava green mite (CGM) infestation was low on all the genotypes. Fresh root yield of five genotypes ranged between 25 and 30 tons per ha for both years. Significant and positive correlation was found across locations between fresh root yield and soil K, P and Mg. AMMI analysis revealed highly significant differences among genotypes and environments and significant genotype × environment interaction for most of the estimated traits, indicating variability in genotypes performance with environment.
Similar content being viewed by others

Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Genotype by environment interaction remains a substantial issue in all breeding programs. Crop genotypes are generally developed in a central breeding location, but always require the evaluation of breeding products in different environments. This is particularly relevant in countries that have a wide range of climates. Cassava, a widely-grown crop throughout the tropics and sub-tropics, is continually being improved to respond to changing pest and disease pressure, utilization, changing climate, and improvement in yield. Multilocation trials are conducted to select the best genotypes for a set of environments for parameters such as crop yield and/or crop response to pests and diseases (Egesi et al. 2009; Akinwale et al. 2011). As such, G × E is essential for demonstrating the potential of a genotype in a given environment and for optimizing the production and utilization of that genotype (Nassir and Ariyo 2011). The net effect of G × E can be additive or multiplicative, depending on the significance of the G × E interaction. Significant interaction implies spatial variation, indicating that chosen sites belong to different environments and large magnitude of G × E is expected in contrasting environments for diverse genotypes (Ekanayake et al. 2000).
Several multilocation experiments focus mostly on agricultural biophysical properties dominated by climate but sometimes, social or cultural contexts are also included (Coe 2012). Many of these studies, however, do not specifically quantify environmental factors like soil nutrients that may account for underlying differences in genotype performance. They rely mainly on the physical and climatic description environments in which one or more genotype are stable or has performed best or worst (mega-environments) (Ngeve 1994; Otoo et al. 1994; Dixon and Nukenine 1998; Benesi et al. 2005; Ssemakula and Dixon 2007; Akinwale et al. 2011; Bradbury et al. 2011; Nassir and Ariyo 2011; Maroya et al. 2012; Callist Kundy 2015; Pariyo et al. 2015). Few authors have attempted to report on cassava performance in relation to soil physical properties and their nutrient contents in G × E experiments (Tan and Mak 1995; Mtunguja et al. 2016).
Cassava, Manihot esculenta (Crantz), is generally considered a widely-adapted crop; but there are numerous examples that show considerable variations in cassava genotype characteristics under varying environmental conditions (Mkumbira et al. 2003; Fu et al. 2014). In Africa and elsewhere in the world where cassava has largely been a subsistence crop, there has been considerable effort to enhance the cassava crop profile. To this end, several cassava genotypes have been developed by IITA cassava breeding unit in Nigeria with a focus in improving root yields, starch content, and resistance/tolerance to major pests and diseases, nutritional content (e.g., higher provitamin A content), and other characters that will satisfy the requirements of cassava based industry (Dixon and Ssemakula 2008). The deployment of these newly developed varieties into new production areas, requires basic understanding of their performance (root yield and stability, nutritional quality of yellow-root genotypes and response to pests and diseases) in relation to contrasting environments and identify some key elements, mainly soil nutrient content, that may account for the observed differences in their performance. Cameroon with five agroecologies ranging from humid forest with bimodal rainfall pattern in the south to the Sahelian zones in the north, is indicated for the evaluation of these varieties and results could be rapidly disseminated in neighboring countries and locations with similar soil characteristics.
Materials and methods
Cassava genotypes
The present study consisted of advanced multilocation trials with three yellow-root and 15 white-root genotypes received from the cassava breeding unit of IITA-Ibadan and planted at IITA-Cameroon, and LMR, a farmer variety widely grown in Cameroon. Some varieties introduced in the 1980s (8034) and beginning of 1990s (92/0326, 92/0067 and 92/0057) were included for comparison. All the experimental material was collected on virus-free plants (i.e., CMD symptom-less plants) from the IITA cassava multiplication plots. Inoculation in the field was done naturally on this material by the whitefly Bemisia tabaci, the vector of the cassava mosaic disease. Subsequent CMD-infection proved to be current-season whitefly-borne infection as only the upper leaves displayed the symptoms (Okao-Okuja et al. 2004).
Environment, soil characterization and general procedure
The trials were conducted during two consecutive cropping seasons (2014/2015 and 2015/2016) at eight locations representing the four-contrasting cassava growing agroecologies in Cameroon. During Year 2, Ngaoundere was replaced by Meiganga (highland savannah) for logistical reasons. We used at each location a HOBO data logger (Onset Computing, USA) for continuous temperature and relative humidity. A rain gauge (Tru-chek®) was used to monitor daily rainfall (Table 1).
Cassava was grown for 12 months under natural conditions and without added fertilizer. Each trial consisted of a randomized complete block design with three replicate blocks, with 18 plots within each block. Each plot consisted of 42 plants in a 6 m × 7 m array, with 1 m-spacing within and between rows of plants. Weeds were controlled manually as necessary.
Data collection
A composite soil sample was taken from each plot at the onset of the experiment to determine soil physical and chemical properties (Soil pH, N, P, K, Ca, Mg, Organic C and Total N) relevant to cassava growth and development were determined in the soil laboratory of IITA-Cameroon. The incidence and severity of cassava mosaic disease (CMD), cassava bacterial blight (CBB) and cassava anthracnose (CAD) were scored at 3, 6, and 9 months after planting (MAP) following a 5-level scale where 1 = no symptoms and 5 = severe symptoms. Average incidence and severity score of CMD, CBB, and CAD were calculated based on data recorded at 3, 6, and 9 months after planting (MAP). Cassava green mite (CGM) active stages and whiteflies (both adults and nymphs) individuals were also counted at each evaluation using standard procedures. The presence of the predatory mite (Typhlodromalus aripo Bondar) and the parasitism of whitefly nymphs were also determined with standard sampling methods for both species (IITA 1990).
Cassava harvest yield was evaluated at approximately 12 MAP at all locations. We estimated fresh root yield (FYLD) per ha based on average fresh root weight of ten plants per plot, excluding border rows. Root dry matter content (DMC) was determined for each genotype from a random sample of three roots/plot. Roots were washed, sliced with peels and a sample of 500 g was dried for 48 h at 60 °C in an electric oven. Root dry matter content was calculated by dividing the dry weight by the fresh weight. Dry root yield (DYLD) was the product of FYLD and DMC: DYLD = FYLD × DMC.
For the determinations of total carotene concentration (TCC), five roots were collected from roots of the 10 harvested plants and processed following BioAnalyt protocol (Kulakow et al. 2015) and TCC was measured using iCheck™ Carotene device within 24 h after harvest.
Data analysis
Data of all parameters were statistically analyzed using a linear mixed model ANOVA that considered cassava genotype as fixed factor, environment and year as random factors. Replication was nested within environment. The model included genotype by environment (G × E) and genotype by year (G × Y) interactions. Tukey test was used as post hoc for the separation of means. Spearman correlation was used to evaluate relationship between soil nutrient content and CMD, and cassava agronomic yield. Analysis was done using SAS 9.2 package (SAS 2009).
There are several tools used to analyze G × E interaction data. We selected the Additive Main effects and Multiplicative Interaction (AMMI), widely used to analyze main effects and genotype by environment interactions in multilocation variety trials (Jalata 2011). This function generates biplots by plotting the first principal component (PC1) scores of the genotypes and the environments against their respective PC2 scores, resulting from the singular value decomposition of environment-centered or standardized G × E data (Crossa 1990; Yan et al. 2000; Yan and Kang 2003; Yan and Tinker 2006; Maroya et al. 2012; Pacheco et al. 2015). The generated plots graphically display environments in which genotypes performed best or worst.
Results
Soil nutrient analysis
Soil nutrient composition differed between environments and within environment for the 2 years (see Table 1 for description of environments). Greater variation was found in P and K content with Ekona having the highest levels. Soils were less acidic in Mbalmayo (Mid altitude forest with bimodal rainfall) and Foumbot. Soils in Foumbot and Bambui had higher Organic C while higher values of Mg were found in Ekona and Foumbot soils (Table 2).
Cassava mosaic disease incidence, severity and whitefly abundance
Four genotypes (TMS 01/0098, 95/0211, 98/0581 and 01/1086-55) did not show any visible CMD symptoms (Table 3). The local check LMR (G15) had the highest incidence (> 72% for both season), followed by 8034 (16%) and 92/0326 (15%) and 01/1814-9 (13%). The remaining genotypes had less than 9% infection across all the environments over the 2 years. Average CMD incidence across all environments was 13.0 ± 4.7 for year 1 and 12.6 ± 4.9 for year 2 while mean severity was 2.5 for both years (Table 3). Highest CMD incidence was recorded in Bertoua (26%), followed by Mbalmayo (22%), Ekona (18%), Bambui (17%), Meyomessala (16%) and Foumbot (16%), Meiganga (15%) and Ngaoundere (15%).
Average densities of whiteflies were higher on genotypes TMS 96/0023 and TMS 01/0098-36 (24/plant each) and lowest on TMS 95/0211 (8/plant). Whitefly density/plant across season and environment was higher in Bambui (47/plant) followed Meiganga (24/plant), Foumbot (18/plant), Bertoua (13/plant), Ekona (9/plant), and Mbalmayo, Meyomessala and Ngaoundere with less than 5/plant. Whitefly density on plants has no effect on CMD incidence and severity (Spearman correlation: r = − 0.01 and r = − 0.02 respectively).
All genotypes expressed mild CBB and CAD symptoms and CGM infestations were generally low. Less than 20% of the whitefly nymphs were parasitized. T. aripo presence were recorded on 12 genotypes, but its abundance was generally low probably due to low CGM infestations (see supplementary table for others pest and disease score).
The combined analysis of variance for CMD incidence and severity revealed that G contributed 84% and 19% respectively to the total sum of squares while E contributed 3% and 6% respectively. G × E contributed to 9% and 22% to the total sum of squares of CMD incidence and severity respectively (Table 4).
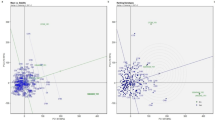
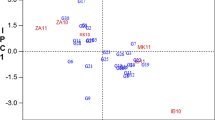
In the GGE biplot analysis of CMD incidence, PC1 and PC2 together explained 85% of the total variation and graphically summarized genotypes performance in relation to the eight environments. The genotypes located on the vertex of the polygon were TMS 92/0326 (G9), LMR (G15), and 8034 (G18) and were the most responsive with higher incidence. Two mega-environments could be defined: (1) Ekona and Meiganga where genotypes TMS 92/0326 (G9) and 8034 (G18) had higher CMD incidence; and (2) Bertoua, Bambui, and Meyomessala where genotype LMR (G15) had higher incidence. Mbalmayo with the shortest vector from biplot origin was less discriminating of the genotypes while Ngaoundere, Meiganga, Ekona, and Meyomessala were more discriminating (Fig. 1).
Fresh and dry root yield and correlation with soil properties
The highest mean for FYLD across locations was recorded from genotype TMS 01/0040-27 (28.4 t/ha) follow by TMS MM97/Jw2-2 (28.2 t/ha) in year 1 while in year 2 highest yield was obtained from genotype 01/0098 (28.8 t/ha) followed by TME 419 (26.7 t/ha). The highest mean for DYLD were recorded by TMS 01/0098-36 (12.5 t/ha) in year 1 and TMS 01/0098 (10.6 t/ha) in year 2. The highest mean for FYLD and DYLD was recorded in Ekona, followed by Foumbot and Bertoua for both years while Meiganga and Ngaoundere were the less favorable environments (Table 5).
The combined analysis of variance of FYLD and DYLD revealed that G contributed 5% and 7% respectively to the total sum of squares while E contributed 46.6% and 40.8% respectively. G × E contributed 12.8% and 12.7% to the total sum of squares (Table 6).
The two principal components (PC1 and PC2) of the biplot together explained 66.1% of the total variation of genotype performance in the eight environments. Most responsive genotypes were 01/1814-9 (G6), 92/0326 (G9), 95/0211 (G11), 98/0581 (G14), and TME 419 (G17) with the highest yield compared to other genotypes. The oldest genotype 8034 was less responsive in all environments except in Ekona for both years. Three mega-environments could be defined: (1) Foumbot, Meyomessala, Bambui, and Ngaoundere which were the winning niche of 98/0581 (G14); (2) Ekona for 92/0326 (G9) and TME 419 (G17); and (3) Bertoua and Mbalmayo for 95/0211 (G11). Ekona and Bertoua were more discriminating of the genotypes FYLD while Bambui and Meyomessala were less discriminating (Fig. 2).
Significant and positive correlation was found across locations between FYLD and soil K (r = 0.67; n = 140; p < 0.0001), soil P (r = 0.52; n = 140; p < 0.0001), and soil Mg (r = 0.60; n = 140; p < 0.0001), and significant negative correlation between FYLD and soil C/N (r = − 0.59; n = 140; p < 0.0001). A non-significant negative correlation was found between FYLD and CMD severity (r = − 0.14; n = 143; p = 0.09); however, when genotypes were pooled across locations, this correlation was significant (r = − 0.55; n = 18; p = 0.02).
Genotypes TMS 01/1086-55(G5), TMS 92/0057 (G7), TMS 95/0109 (G10), TMS 98/0581 (G14), and TME 419 (G17) were the most yielding and most stable across the environments for DYLD. Genotypes TMS 01/0040-27 (G1), TMS 01/0098 (G2), TMS 01/0098-36 (G4), and TMS MM97/JW2-2 (G16) also high yielding but their yield varied across the environments (Fig. 3).
Root total carotenoids content (TCC)
The mean TCC in fresh roots ranged from 3.62 µg/g for genotype TMS 01/0040-27 to 4.52 µg/g for TMS MM97/JW2-2 with an average of 4.1 ± 0.1 µg/g.
The combined analysis of variance of the three genotypes showed significant differences among G and Y but not among environments. The G × E and G × Y interaction were not significant. G, E, Y, and G × E contributed respectively 43.6%, 6.6%, 4.5% and 8.9% of the total sum of squares (Table 7).
In the GGE biplot analysis, PC1 and PC2 explained together 100% of the total variation and graphically summarized genotypes performance in relation to the eight environments. Three mega-environments could be defined: Bambui, Foumbot, and Mbalmayo which were the winning niche of TMSMM97/JW2-2 (G16); Meiganga, Ngaoundere, and Meyomessala for TMS 01/0040-27 (G1); and Ekona and Bertoua for TMS 01/1814-9 (G6). Foumbot, Bertoua, and Ngaoundere were more discriminating of the genotypes TTC; Bambui, Meiganga, and Mbalmayo were less discriminating (Fig. 4).
Discussion
Differential responses observed for the measured parameters to different environments justified the importance of testing genotypes in contrasting environments, to identify good performers for specific locations or those that are stable across locations. The large sum of square attributed to genotypes for CMD incidence showed that the susceptibility is more related to the genotype than on the environment (Thresh and Cooter 2005). The genotype however interacted significantly with the environment for the severity of the disease. This interaction accounted for 22% of the CMS severity sum of square and which could be attributed to the virus strain present in the environment. Bertoua and Meiganga are among environments where the virulent strain EACMV-UG virus was first recorded in Cameroon in 2009 (Akinbade et al. 2010). It has been demonstrated that virus has negative impact on genotypes by the severity of the infection varies greatly between genotypes, bringing evidence that the presence of a virus strain in the environment can impact the G × E interaction (van Mölken and Stuefer 2011). There was a negative correlation between CMD severity and fresh root yield. Some genotypes were heavily hit by the virus strain in those environments and the severity was often more than 3 (moderate mosaic pattern throughout the leaf, narrowing and distortion of the lower one-third of leaflets). Other varieties were not affected and these can be considered as good candidate for introduction in such areas. It is also an indication that they can be used in further cassava selection to improve resistance to CMD giving that finding a resistant or tolerant variety remain a major focus for CMD mitigation in cassava production areas (Chikoti et al. 2009).
Whitefly densities per plant were higher in environment with altitudes above 1000 m (Meiganga, Ngaoundere and Bambui). This finding contrasts with previous studies in western and eastern Kenya where whiteflies densities were higher in low midlands compared with highlands (Njoroge et al. 2016). Whitefly densities in savannah highlands (Ngaoundere and Meiganga) could have been higher without the impact of bacterial blight which defoliated the tip of many plants.
Five genotypes showed wider adaptability for root yield while others exhibited specific adaptation or performed well in specific environment (e.g. 8034). These difference in performance per environment justifies the large sum of square attributed to environments for roots yields (Kota et al. 2013). The pronounced effect of E on root yield was expected as many authors have indicated that root yield is a polygenic trait (Easwari and Sheela 1998; Cach et al. 2006). Aligning these observations to soil analysis results, Ekona and Foumbot had greater K content, while P content in Ekona was more than 20 folds that of Bambui. The nutrient element K is very important for cassava growth and low K level constitute a limiting factor to the uptake of other nutrients such as N and P (Boateng and Boadi 2010). Also, there is a linear increase in cassava fresh root yields with NPK uptake by plant (Byju et al. 2012; Temegne et al. 2015) and therefore, the high rate of K in Foumbot and Ekona could have played a major role in the uptake of other nutrients, resulting in greater root yields compared with other locations. Meiganga also had high P level but also low K content, negating therefore the effect of high P by reducing uptake (Boateng and Boadi 2010).
Cassava yield in Meiganga, Ngaoundere and Bambui was lower compared with other environments. Average minimal temperature in these locations ranged between 11.7 °C and 12.4 °C. Some authors have reported that cassava at higher altitude experience yield loss potential compared to lowland or mid-altitude environments due to low temperature and reduced solar radiation (Manrique 1992; Noerwijati and Budiono 2015).
Five genotypes—TMS 01/1086-55(G5), TMS 92/0057 (G7), TMS 95/0109 (G10), TMS 98/0581 (G14), and TME 419 (G17)—were stable for DYLD across the environments. Dry root yield is an important and favorable characteristic since cassava root dry matter has become an important character for acceptance by researchers and consumers, especially for processed cassava (Teye et al. 2011). These genotypes could therefore be promoted for industrial use.
The environment did not significantly affect the total carotenoid content (TCC) in biofortified genotypes and G × E interaction was not significant, but a small yet insignificant year effect was observed. The reverse was observed by Maroya et al. (2012) in Nigeria with other biofortified genotypes. TCC is known to be affected by the environment, especially in sweet potato (Manrique and Hermann 1999). In the present study, we used three genotypes with relatively low TTC content. The highest TTC recorded by the genotype MM97/JW2-2, was very low compared with more advanced clones such as TMS 07/0593 with TTC of 11 µg/g. The latter has already shown good performance and stability and is recommended for the food technologist and nutritionist (Maroya et al. 2012).
Conclusion
Newly developed cassava genotypes differed in their performance in tested environments. However, five genotypes (TMS 01/1086-55, TMS 92/0057, TMS 95/0109, TMS 98/0581, and TME 419) were high yielding and stable across the environments, and should be promoted in cassava development programs in regions sharing similar soil and climate characteristics as in our experiment. Four genotypes (TMS 01/0098, TMS 95/0211, TMS 98/0581 and TMS 01/1086-55) did not show CMD symptoms in any environment and constitute a good asset for cassava integrated pest management. Observed difference in genotypes performance could be predicted given the difference in soil nutrient content and virus strain found in each environment. The diffusion of CMD-resistant, high yielding and stable genotypes should promote the change of cassava from subsistence crop associated with low value and poor-quality products to an important commodity in a market-oriented agriculture, pending the evaluation of various utilization by consumers and industries.
References
Akinbade S, Hanna R, Nguenkam A et al (2010) First report of the East African cassava mosaic virus-Uganda (EACMV-UG) infecting cassava (Manihot esculenta) in Cameroon. New Dis Rep 21:22
Akinwale MG, Akinyele BO, Odiyi AC, Dixon AGO (2011) Genotype × environment interaction and yield performance of 43 improved cassava (Manihot esculenta Crantz) genotypes at three agro-climatic zones in Nigeria. Br Biotechnol J 1:68–84
Benesi I, Labuschagne M, Dixon A, Mahungu N (2005) Genotype × enviroment interaction effects on native cassava starch quality and potential for starch use in the commercial sector. Afr Crop Sci J 12:205–216. https://doi.org/10.4314/acsj.v12i3.27880
Boateng SA, Boadi S (2010) Cassava yield response to sources and rates of potassium in the forest–savanna transition zone of Ghana. Afr J Root Tuber Crop 8:1–5
Bradbury G, Potts B, Beadle C (2011) Genetic and environmental variation in wood properties of Acacia melanoxylon. Ann For Sci 68:1363–1373
Byju G, Nedunchezhiyan M, Ravindran CS et al (2012) Modeling the response of cassava to fertilizers: a site-specific nutrient management approach for greater tuberous root yield. Commun Soil Sci Plant Anal 43:1149–1162. https://doi.org/10.1080/00103624.2012.662563
Cach NT, Lenis JI, Perez JC et al (2006) Inheritance of useful traits in cassava grown in subhumid conditions. Plant Breed 125:177–182. https://doi.org/10.1111/j.1439-0523.2006.01192.x
Callist Kundy A (2015) Effect of G × E interaction on yield and yield components of cassava genotype by environment interaction. LAP LAMBERT Academic Publishing, Saarbrücken
Chikoti PC, Shanahan P, Melis R (2009) Evaluation of cassava genotypes for resistance to cassava mosaic disease and agronomic traits. Am J Plant Sci 7:1122–1128. https://doi.org/10.4236/ajps.2016.77107
Coe R (2012) Multi-environment trials: an overview. Stat Serv Centre, Univ Read UK World Agrofor Centre, Kenya 1–13
Crossa J (1990) Statistical analyses of multilocation trials. Adv Agron 44:55–85. https://doi.org/10.1016/S0065-2113(08)60818-4
Dixon AG, Nukenine EN (1998) Genotype × environment interaction and optimum resource allocation for yield and yield components of cassava. Afr Crop Sci J 8:1–10
Dixon A, Ssemakula G (2008) Prospects for cassava breeding in Sub-Saharan Africa in the next decade. J Food Agric Environ 6(4):256–262
Easwari ACS, Sheela MN (1998) Genetic analysis in a diallel cross of inbred lines of cassava. Madras Agric J 85:264–268
Egesi CN, Onyeka TJ, Asiedu R (2009) Environmental stability of resistance to anthracnose and virus diseases of water yam (Dioscorea alata). Afr J Agric Res 4:113–118
Ekanayake IJ, Ortiz R, of Tropical Agriculture II (2000) Genotype × environment interaction analysis of IITA mandate crops in Sub-Saharan Africa. International Institute of Tropical Agriculture, Ibadan
Fu J, Jiang D, Huang Y et al (2014) Evaluating the marginal land resources suitable for developing bioenergy in Asia. Adv Meteorol 2014:1–9. https://doi.org/10.1155/2014/238945
IITA (1990) Cassava in tropical Africa: a reference manual. IITA, Ibadan
Jalata Z (2011) GGE-biplot analysis of multi-environment yield trials of barley (Hordeum vulgare L.) genotypes in southeastern Ethiopia Highlands. Int J Plant Breed Genet 5:59–75. https://doi.org/10.3923/ijpbg.2011.59.75
Kota S, Singh S, Mohapatra T et al (2013) Genotype × environment interaction analysis for grain yield in new plant type (npt) wheat derivatives. SABRAO J Breed Genet 45:382–390
Kulakow PA, Parkes EY, Friedrich SK et al (2015) Linearity, reproducibility and comparison of iCheckTM carotene with spectrophotometer and HPLC for evaluation of total carotenoids in cassava roots. Eur J Nutr Food Saf 136:57881. https://doi.org/10.13140/2.1.4534.4007
Manrique LA (1992) Growth and yield performance of cassava grown at three elevations in Hawaii. Commun Soil Sci Plant Anal 23:129–141. https://doi.org/10.1080/00103629209368576
Manrique K, Hermann M (1999) CIP program report effect of G × E interaction on root yield and beta- carotene content of selected sweetpotato (Ipomoea batatas (L) Lam.) varieties and breeding clones. CIP Program Report 281–287
Maroya NG, Kulakow P, Dixon A, Maziya-Dixon B (2012) Genotype × environment interaction of mosaic disease, root yields and total carotene concentration of yellow-fleshed cassava in Nigeria. Int J Agron 2012:1–8. https://doi.org/10.1155/2012/434675
Mkumbira J, Mahungu NM, Gullberg U (2003) Grouping locations for efficient cassava evaluation in Malawi. Exp Agric 39:167–179. https://doi.org/10.1017/S0014479702001199
Mtunguja MK, Laswai HS, Kanju E et al (2016) Effect of genotype and genotype by environment interaction on total cyanide content, fresh root, and starch yield in farmer-preferred cassava landraces in Tanzania. Food Sci Nutr 4:791–801. https://doi.org/10.1002/fsn3.345
Nassir AL, Ariyo OJ (2011) Genotype × environment interaction and yield-stability analyses of rice grown in tropical inland swamp. Not Bot Hort Agrobot Cluj 39:220–225
Ngeve JM (1994) Yield stability parameters for comparing cassava varieties. Acta Hortic. https://doi.org/10.17660/ActaHortic.1994.380.21
Njoroge MK, Kilalo DC, Miano DW et al (2016) Whiteflies species distribution and abundance on cassava crop in different agro-ecological zones of Kenya. J Entomol Zool Stud 258:258–262
Noerwijati K, Budiono R (2015) Yield and yield components evaluation of cassava (manihot esculenta crantz) clones in different altitudes. Energy Procedia 65:155–161. https://doi.org/10.1016/j.egypro.2015.01.050
Okao-Okuja G, Legg JP, Traore L, Alexandra Jorge M (2004) Viruses associated with cassava mosaic disease in Senegal and Guinea Conakry. J Phytopathol 152:69–76. https://doi.org/10.1046/j.1439-0434.2003.00797.x
Otoo JA, Dixon AGO, Asiedu R et al (1994) Genotype × environment interaction studies with cassava. Acta Hortic. https://doi.org/10.17660/ActaHortic.1994.380.22
Pacheco Á, Vargas M, Alvarado G et al (2015) GEA-R (Genotype × environment analysis with R for Windows) Version 3.0—CIMMYT Research Software Dataverse—CIMMYT Dataverse Network
Pariyo A, Baguma Y, Alicai T et al (2015) Stability of resistance to cassava brown streak disease in major agro-ecologies of Uganda. J Plant Breed Crop Sci 7:67–78. https://doi.org/10.5897/JPBCS2013.0490
SAS (2009) JMP® Version 8.0.2. SAS Institute, Cary, NC
Ssemakula G, Dixon A (2007) Genotype × environment interaction, stability and agronomic performance of carotenoid-rich cassava clones. Sci Res Essay 2:390–399
Tan SL, Mak C (1995) Genotype × environment influence on cassava performance. Field Crop Res. https://doi.org/10.1016/0378-4290(95)00016-J
Temegne NC, Ajebesone FN, Fotso KA (2015) Influence de la composition chimique du sol sur la teneur en éléments nutritifs et le rendement du manioc (Manihot esculenta Crantz, Euphorbiaceae) dans deux zones agro-écologiques du Cameroun. Int J Biol Chem Sci 9:2776–2788. https://doi.org/10.4314/ijbcs.v9i6.21
Teye E, Asare AP, Amoah RS, Tetteh JP (2011) Determination of the dry matter content of cassava (Manihot esculenta Crantz) tubers using specific gravity method. ARPN J Agric Biol Sci 6:23–28
Thresh JM, Cooter RJ (2005) Strategies for controlling cassava mosaic virus disease in Africa. Plant Pathol 54:587–614. https://doi.org/10.1111/j.1365-3059.2005.01282.x
van Mölken T, Stuefer JF (2011) The potential of plant viruses to promote genotypic diversity via genotype × environment interactions. Ann Bot 107:1391–1397. https://doi.org/10.1093/aob/mcr078
Yan W, Kang MS (2003) GGE biplot analysis: a graphical tool for breeders, geneticists, and agronomists. CRC Press, Boca Raton. https://doi.org/10.1201/9781420040371
Yan W, Tinker NA (2006) Biplot analysis of multi-environment trial data: principles and applications. Can J Plant Sci 86:623–645. https://doi.org/10.4141/P05-169
Yan W, Hunt LA, Sheng Q, Szlavnics Z (2000) Cultivar evaluation and mega-environment investigation based on the GGE Biplot. Crop Sci 40:597. https://doi.org/10.2135/cropsci2000.403597x
Acknowledgements
This work was supported by the Agricultural Investment and Market Development Project (AIMDP) jointly funded by the Cameroonian government and the World Bank, and CGIAR Research Program on Roots, Tubers and Bananas (RTB). The administrative and logistic support from IRAD Office is acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fotso, A.K., Hanna, R., Kulakow, P. et al. AMMI analysis of cassava response to contrasting environments: case study of genotype by environment effect on pests and diseases, root yield, and carotenoids content in Cameroon. Euphytica 214, 155 (2018). https://doi.org/10.1007/s10681-018-2234-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-018-2234-z