Abstract
Spatio-temporal expression of an insecticidal gene (Cry1Ac) in pre existing transgenic lines of transgenic cotton was studied. Seasonal decline in expression of Cry1Ac differed significantly among different cotton lines tested in the field conditions. The leaves of the Bt cotton plants were found to have the highest levels of toxin expression followed by squares, bolls, anthers and petals. Expression of the gene decreased consistently with the age of plants. Toxin expression in fruiting parts was not enough to confer full resistance against bollworms. The reduction in efficacy of transgenic cotton plants late in the season was attributed to reduction in promoter activity. For this purpose, ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) small subunit (rbcS) promoter was isolated from Gossypium arboreum that was further cloned upstream of an insecticidal gene (Cry1Ac) in expression vector pCAMBIA 1301. A local cotton cultivar NIAB-846 was transformed with Cry1Ac driven by rbcS promoter. The same cotton cultivar was also transformed with Cry1Ac gene driven by 35SCaMV promoter to compare the expression pattern of insecticidal gene under two different promoters. The results showed that rbcS is an efficient promoter to drive the expression of Cry1Ac gene consistent throughout the life of cotton plant as compared to 35S promoter. The use of tissue specific promoter is also useful for addressing the biosafety issues as the promoter activity is limited to green parts of plants, hence no gene expression in roots, cotton seed and other cotton products and by products.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cotton is an important economic and fibre crop worldwide and likewise it serves as a jugular vein in the economy of Pakistan because of its major share in GDP. Transgenic cotton expressing Bt (Bacillus thuringiensis) toxins is currently cultivated on a large commercial scale in many countries, but data has shown that it behaves variably in toxin efficacy against target insects under field and greenhouse conditions (Benedict et al. 1996; Finnegan et al. 1998; Chen et al. 2000; Guo et al. 2001; Mahon et al. 2002; Olsen et al. 2005; Adamczyk et al. 2009; Manjunatha et al. 2009; Bakhsh et al. 2010). Understanding of the temporal and spatial variation in efficacy and the resulting mechanisms is essential for cotton protection and production.
The mechanisms of variation in endotoxin protein content in plant tissues are rather complicated. The reduction in Bt protein contents in late-season cotton tissues could be attributed to the overexpression of Bt gene at earlier stages, which leads to gene regulation at post-transcription levels and consequently results in gene silencing at a later stage (Xia et al. 2005). Methylation of the promotor may also play a role in the declined expression of endotoxin proteins (Xia et al. 2005; Dong and Li 2007). Several other factors might also be responsible for variation in the gene expression level, e.g. change in the nucleotide sequence of the gene, type of promoter, and the insertion point of the insert/cassette in the DNA of the transgenic variety, transgene copy number, internal cell environment, as well as several external factors in the environment (Hobbs et al. 1993; Guo et al. 2001; Rao 2009).
Nearly all transgenic crops around the world utilize the CaMV 35S promoter (or similar promoters from closely-related viruses) to drive transgenes (Odell et al. 1985). It is only now becoming clear that this promoter is not as robust as laboratory and glasshouse studies have suggested. Its function is influenced by as yet undefined physiological and perhaps environmental factors (Sunilkumar et al. 2002; Bakhsh et al. 2010). Compared with the temporal or spatial specific expression of the toxin, constitutive expression of foreign proteins in transgenic plants may cause adverse effects, such as the metabolic burden imposed on plants for constant synthesis of foreign gene products, and these may increase the potential risk of resistance of the target insects to Bt. There is also concern about the food safety of genetically modified plants (Kuiper et al. 2001; Shelton et al. 2002; Conner et al. 2003).
Consistent and targeted expression has become particularly important for the future development of value-added crops because the public may be more likely to accept ‘less intrusive’ expression of the transgenes. As sophistication in biotechnology improves, the need for more developmentally or environmentally regulated promoters is becoming evident and considerable efforts should be exerted for the discovery of specific tissue or biotic, hormonal or abiotic stress responsive genes and promoters (Potenza et al. 2003).
The present study was conducted to evaluate the expression of insecticidal gene (Cry1Ac) with the age of plants as well in different plant parts because the sustainable expression of Bt genes is crucial for its effectiveness to control the targeted insect pests. A strategy was devised out to make expression of Cry1Ac consistent in cotton by isolating a ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) small subunit (rbcS) promoter from Gossypium arboreum (cv. FDH-786) and was further transformed in Gossypium hirsutum (cv. NIAB-846).
Materials and methods
Twelve pre-existing transgenic cotton lines (cv. CIM-482) transformed with an insecticidal gene (Cry1Ac) (Rashid et al. 2008) were planted at Centre of Excellence in Molecular Biology (CEMB) campus experimental field along with non transgenic control using randomized complete block design with three replications.
Temporal and spatial expression of Cry1Ac endotoxin
As differential expression of Cry1Ac occurs among different plant structures (Greenplate 1999; Adamczyk et al. 2001), a single structure was selected for quantification of Cry1Ac. For each sample date and for all transgenic lines, a single main-stem terminal leaf was collected from five plants/transgenic line after every 15 days interval. For spatial expression, various plant parts i.e. leaf, anthers, pollens, square buds and bolls were selected at reproductive growth stage. The samples were transported to the laboratory and were processed the same day.
Expression of Cry1Ac in transgenic lines was quantified by enzyme linked immunosorbent assay (ELISA) using Envirologix Kit (Cat # AP051). Negative and positive controls were added to the wells along with test samples. ELISA was performed according to procedure given in the kit and quantification of Cry1Ac endotoxins was done by plotting absorbance values of Cry1Ac test samples on the standard curve generated with purified Cry1Ac standards on each of ELISA plates and expressed as nanogram of Cry1Ac per gram of fresh tissue weight. Protein expression data of Cry1Ac was further subjected to statistical analysis using SPSS software (version 11.0, SPSS Inc) to evaluate the differences among transgenic lines, sampling dates, interaction of transgenic lines and sampling dates at 5% level of significance.
Leaf biotoxicity assay
Based on spatio-temporal results, leaf biotoxicity assays were performed to counter check the variation in efficacy of endotoxin Cry1Ac against targeted insect pests after 30, 60 and 90 days of crop age. The five leaves from the upper, middle and the lower portion of each transgenic line along with the leaves from non transgenic control line in triplicates were detached on moist filter papers in petri plates and 2nd instar Heliothis larvae were fed to them. After 2–3 days, mortality rates of Heliothis larvae were recorded. The mortality rates were calculated as follows:
Isolation and sequencing of rbcS promoter
Total RNA extraction from leaves of cotton (G. arboreum cv. FDH-786) was followed using method of Jaakola et al. (2001) with some modifications. cDNA was synthesized by using Fermentas cDNA synthesis kit. The promoter region was amplified by polymerase chain reaction (PCR) using oligonucleotide primer including an KpnI restriction site in the forward and reverse primers (F: 5′-GCGGTACCCAAAAGAGTCTCACTGATCC-3′)and(R: 5′-GCGGTACCCTCTTTGTGGTCATAATGGT-3′). The promoter fragment was amplified by high fidelity PCR (1 reaction: 2.5 μl 10× buffer, 2.5 μl 2 mM dNTP, 1 μl each primer (10 μM), 1 μl Taq DNA polymerase, 0.5 μl DNA (100 ng/μl), 16.5 μl H2O) under the following conditions: initial denaturation 94°C for 5 min; followed by 40 cycles of denaturation at 94°C for 30 s, annealing at 60°C for 30 s, extension at 72°C for 1 min and a final extension at 72°C for 10 min. The rbcS promoter region (445 bp) was confirmed by sequencing.
Plasmid construction
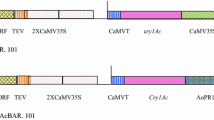
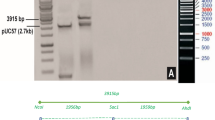
Plasmid construction was carried out in two steps. The Cry1Ac gene and NOS terminator fragment (2.11 kb) was amplified from its source plasmid PIA-2 (already available plasmid in laboratory having Cry1Ac under ubiquitin promoter) by PCR using BamHI and KpnI restriction sites and cloned in pre-digested plasmid pCAMBIA 1301. The binary vector pCAMBIA-1301 (11,837 bp) was used for cloning of Cry1Ac gene under rbcS promoter. The vector contains kanamycin gene for bacterial selection and hygromycin for plant selection resistance genes.
Furthermore, rbcS promoter was subcloned upstream of Cry1Ac + NOS fragment into the plasmid pCAMBIA 1301 pre-digested with KpnI. PCR, Restriction digestion and sequencing techniques were employed to confirm cloning of insert (promoter and insecticidal gene) in plant expression vector. The resulting plasmid was named Rb-Ac and was used to transform Agrobacterium tumefaciens strain LBA4404.
Cotton transformation
Cotton (G. hirsutum) cv. NIAB-846 was selected for transformation because it had high yield potential and desired fibre characteristics. The seeds were delinted and surface sterilized with Tween-20 for 3 min and further subjected to 0.1% HgCl2 and 0.1% Sodium dodecyl sulphate (SDS) solution mixture. The sterilized seeds were placed in dark at 30°C overnight for the germination. The shoot apices of germinating seedlings were used for Agrobacterium-mediated transformation according to the procedure described by Gould and Magallanes-Cedeno (1998) with some modifications adopted at CEMB as described by Rao (2009), Bakhsh (2010) and Khan et al. (2011). The inoculated explants were cultured on MS medium (Murashige and Skoog 1962) containing 1 mg/l Kinetin for 3 days at 28 ± 2°C. After co-cultivation, plantlets were sub-cultured on selection medium i.e. MS containing 75 μg/ml hygromycin, also supplemented with 0.5 mg/l benzylaminopurine (BAP) and 1 mg/l α-naphthaleneacetic acid (NAA). Cefataxime (250 mg/l) was also added to inhibit bacterial overgrowth. Sub culturing was done after every 10 days. After 2 months selection, these plantlets were shifted to hygromycin free MS medium until fully plants were developed. The putative transgenic plants were further shifted to pots containing soil of equal proportion of clay, sand and peat moss (1:1:1). Finally the plants were shifted to greenhouse and were subjected to various molecular analysis. Same cotton cultivar was also transformed with another plasmid pk2Ac (having Cry1Ac under 35S promoter) to study comparative spatio temporal expression of Cry1Ac in newly transformed cotton plants.
Molecular analysis of transgenic plants
Genomic DNA was isolated from newly transformed Rb-Ac plants (transformed with Cry1Ac under rbcS promoter) as well as pk2Ac plants (transformed with Cry1Ac under 35S promoter). PCR was run for the detection of integrated Cry1Ac to amplify internal fragments of 565 bp using gene specific primers by a modification of the method by Saiki et al. (1988). DNA extracted from untransformed plants was used as negative control and that of plasmid Rb-Ac as positive control. The PCR was performed at 94°C for 4 min, 94°C for 1 min, 52°C for 1 min and 72°C for 1 min followed by 35 cycles.
SDS-PAGE (polyacrylamide gel electrophoresis) of transgenic PCR positive Rb-Ac and pk2Ac plants was carried out by the method of Laemmli (1970) to confirm Cry1Ac expression. About 50 μg of isolated protein was mixed with loading dye denatured by heat shock in boiling water bath for 5 min and quick chilled on ice. Prepared 12% resolving, and 4% stacking gel in gel assembly. Stained the gel with commassie stain for 30 min and destained it with destaining solution.
Temporal and spatial expression of Cry1Ac in newly transformed plants
Newly transformed Rb-Ac and pk2Ac plants were further investigated to evaluate the temporal and spatial expression of Cry1Ac endotoxins. For each sample date, a single main-stem terminal leaf was collected from five plants/line at different intervals of plant growth stages. ELISA assay was performed using Envirologix Kit (Cat # AP051) and concentration of Cry1Ac gene in Rb-Ac and pk2Ac plants was quantified and expressed as nanogram per gram of fresh tissue weight. Different plants parts at reproductive stage were undertaken for spatial expression.
Results
Temporal and spatial expression of Cry1Ac endotoxin
Temporal expression of Cry1Ac gene was quantified in transgenic lines after 15 days interval. The result showed that the toxin level declined with the age of plant (Fig. 1a) being maximum at vegetative stage, 30 days crop. As the crop progressed towards the maturity, a decline in expression level of Cry1Ac was observed. Similar expression pattern was found in all these transgenic lines. Statistical analysis revealed that Cry1Ac differences among transgenic lines, sampling dates and their interaction were significant at 5% level of significance.
Spatio temporal expression of Cry1Ac in transgenic cotton lines. a Temporal expression of Cry1Ac was quantified after an interval of 15 days showing the maximum expression at vegetative growth stage while a decline in expression at the later stages of crop plants was observed. b Spatial expression of Cry1Ac quantified in different plant parts showing maximum in leaf followed by followed by square buds, bolls and anthers
To evaluate spatial expression, different plant parts i.e. leaves, square buds, bolls, anthers, petals and ovary were sampled from the field and subjected to ELISA assay. The result revealed that the expression of Cry1Ac was variable in different plant parts, being maximum in the leaves followed by square buds, bolls and anthers (Fig. 1b). Petals were showing less toxin expression as compared to other plant parts being only 8–10 ng/g of fresh tissues weight while Cry1Ac expression in ovary remained undetectable.
Leaf biotoxicity assay
Laboratory biotoxicity assays with 2nd Instar Heliothis larvae were conducted to confirm the variation in expression of the insecticidal gene at 30, 60 and 90 days of crop age. The results showed that transgenic lines showed a varying mortality rate of Heliothis larvae that ranged between 60 and 90% at different growth stages. The varying mortality rates indicated a variation in Cry1Ac protein levels. Transgenic lines that showed 100 mortality rate of Heliothis larvae at 30 days crop age, showed varying 60–80% mortality rate at 90 days of crop age (Fig. 2) while in case of non transformed control CIM-482, no any larval mortality was recorded.
Mortality % age of 2nd Heliothis larvae in different transgenic lines at 30, 60 and 90 days of crop plants. Lines showed 100% mortality of larvae at 30 days crop stage while this percentage varied at 90 days observation while no any larval mortality was recorded in non transgenic control cotton cultivar CIM-482
Cotton transformation
An optimized procedure by Gould and Magallanes-Cedeno (1998) was followed to transform cotton cultivar with some modifications (Fig. 3). A total of 4,000 mature embryos were used in all the transformation experiments. After 8 weeks of selection on 75 μg/ml hygromycin, 50 plants of Rb-Ac and 43 plants for pk2Ac plants were obtained. Among these 50 plants of Rb-Ac plants, 12 plants were shifted in the field while 10 pk2Ac plants established properly in soil. The overall transformation efficiency in both types of transgenic plants remained 0.55% (Table 1).
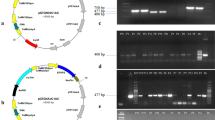
Confirmation of gene integration and expression in transgenic plants. a PCR assay showed the amplification of required Cry1Ac band. Lane 1 Lamba HindIII marker, lane 2–7 Transgenic plants of Rb-Ac, lane 8–12 transgenic plants of pk2Ac, lane 13 negative control (nontransformed NIAB-846), lane 14 positive control (plasmid DNA). b SDS-PAGE analysis of Rb-Ac and pk2Ac plants confirmed expression of 68 kDa band. Lane 1 high mol. weigh protein marker (Fermentas), lane 2–5 transgenic plants of Rb-Ac, lane 6–9 transgenic plants of pk2Ac, lane 10 negative control NIAB-846, lane 11 positive control protein isolated from HD-73 strain
Confirmation of gene integration and expression
Newly transformants Rb-Ac and pk2Ac were analyzed for the integration of Cry1Ac gene using gene specific primer by PCR assay. The amplified fragment of 565 bp revealed that Cry1Ac gene was integrated in Rb-Ac and pk2Ac plants successfully while no amplification was observed in non transformed plant sample (Fig. 4a). All the transgenic plants were phenotypically normal and grew well. Expression of Cry1Ac protein in transgenic plants of Rb-Ac and pk2Ac was confirmed by SDS-PAGE. A 68 kDa band of Cry1Ac protein was observed in all the transgenic plant samples including positive control while no such band was present in negative control (Fig. 4b); thereby confirming the expression of introduced gene (Cry1Ac) in transgenic plants.
Spatio-temporal expression of Cry1Ac in transgenic cotton plants of Rb-Ac and pk2Ac. a Temporal expression of Cry1Ac quantified showed higher and consistent expression of Cry1Ac in Rb-Ac plants while decline in expression was observed in pk2Ac plants. b Spatial expression of Cry1Ac quantified showed higher Cry1Ac levels in leaf, boll and squares in Rb-Ac plants as compared to pk2Ac plants
Temporal and spatial expression of Cry1Ac in newly transformed plants
The transgenic plants of Rb-Ac and pk2Ac were subjected to ELISA assay for the quantification of insecticidal protein Cry1Ac levels at every 15 days intervals. Expression of Cry1Ac in transgenic plants revealed that Rb-Ac plants were showing maximum expression of Cry1Ac protein and no decline in expression was observed in these plants. While in case of pk2Ac plants, Cry1Ac levels declined with the age of plants (Fig. 4a). Expression in Rb-Ac plants was up to 250 ng/g of fresh tissue weight even when the plant was 90 days old that conferred full protection against targeted insect pests.
Cry1Ac protein was also quantified in different plant tissues of Rb-Ac and pk2Ac plants. Plant tissue i.e. leaf, boll, square, anthers and petals were undertaken for this purpose. The results showed that leaves of both Rb-Ac and pk2Ac plants were having maximum Cry1Ac concentration followed by bolls and squares. Rb-Ac plants showed more expression of Cry1Ac in bolls and squares as compared to pk2Ac plants while no expression in anthers was recorded (Fig. 4b).
Discussions
This study was undertaken to determine the spatio temporal expression of insecticidal gene Cry1Ac in transgenic cotton lines. Spatio temporal studies revealed that expression level of Cry1Ac declined during the crop growth with toxin level falling to 15–20 ng/g of fresh tissue weight (Fig. 1a). These results were in agreement with the previous studies conducted by Fitt et al. (1998), Greenplate et al. (1998), Chen et al. (2000), Mahon et al. (2002), Xia et al. (2005), Olsen et al. (2005), Manjunatha et al. (2009) and Adamczyk et al. (2009), Bakhsh et al. (2010) and Bakhsh et al. (2011a, b) who have reported inconsistency in insecticidal gene expression over the crop growth period. Expression levels of insecticidal gene were also remained variable in different plant parts (Fig. 1b). The leaves of Bt cotton were having maximum expression as compared to certain fruiting parts (Greenplate 1999; Greenplate et al. 2000; Adamczyk et al. 2001; Gore et al. 2001). Furthermore, the results obtained from biotoxicity assays at different intervals also confirmed the variation in Cry1Ac expression (Fig. 2).
Gene expression varies with the nucleotide sequence of the gene, promoter, and the insertion point of the gene in the DNA of the transgenic variety, transgene copy number, the internal cell environment, as well as several external factors in the environment (Hobbs et al. 1993; Guo et al. 2001; Rao 2009). A thorough research at molecular and environmental level was carried out to understand the differential expression of insecticidal protein in transgenic cotton lines and this variation in endotoxin efficacy was attributed to promoter activity (Bakhsh 2010). Results revealed that 35S CaMV promoter is cell-type-specific and is developmentally regulated promoter (Nilsson et al. 1992; Pauk et al. 1995; Wessel Vl Tom et al. 2001; Sunilkumar et al. 2002; Haddad et al. 2002; Yang and Christou 2005; Amarasinghe et al. 2006; Bakhsh et al. 2010). The study triggered research to find possible new promoter that would induce more consistent production of insecticidal gene throughout the life of the cotton plant. For this purpose, a rbcS promoter from desi cotton (G. arboreum) was isolated.
The small subunits promoters and their transit peptides are attractive candidates for expression of genes at high levels and have shown promising expression results in driving the expression of introduced genes (Wong et al. 1992; Fujimoto et al. 1993; Data et al. 1998; Song et al. 2000; Jang et al. 2002; Liu et al. 2005; Panguluri et al. 2005; Amarasinghe et al. 2006; Anisimov et al. 2007; Suzuki et al. 2009; Kim et al. 2009 and Bakhsh et al. 2011a, b). rbcS promoter was cloned in a plant expression vector pCAMBIA 1301. In the present study pCAMBIA 1301 vector was used to transform as it has been used by many researchers to transform the genes like cbnA in rice (Shimizu et al. 2002), acsA and acsB genes, USP-1 and HSP-26 genes in cotton (Li et al. 2004; Maqbool et al. 2010), HMGR gene (3-hydroxy-3-methylglutaryl-CoA reductase) in Taraxacum plants (Bae et al. 2005). A local cotton cultivar NIAB-846 was transformed with Cry1Ac driven by rbcS promoter. The same cotton cultivar was also transformed with Cry1Ac gene driven by 35SCaMV promoter to compare the expression pattern of insecticidal gene under two different promoters.
In the present study, agrobacterium-mediated transformation procedure established by Gould and Magallanes-Cedeno (1998) was followed with some modifications adopted at CEMB as described by Rao (2009), Bakhsh (2010) and Khan et al. (2011). The transformed plants were analyzed for DNA integration and expression through PCR and SDS-PAGE (Fig. 3).
The titer of Cry1Ac toxin in newly transformed Rb-Ac and pk2Ac plants quantified at different developmental stages indicated clearly that Rb-Ac plants were expressing Cry1Ac toxin at high concentration as compared to pk2Ac plants. The recorded data showed that rbcS is efficient promoter driving the expression of Cry1Ac consistent in cotton plant while a decline in Cry1Ac was observed in pk2Ac plants (Fig. 4a). Spatial expression of Cry1Ac gene in different plant parts further confirmed that promoter activity is limited to green parts of plants i.e., leaf, stems, and square buds. The high expression of Cry1Ac protein was quantified in Rb-Ac plants being maximum in leaf as compared to pk2Ac plants that were showing less concentration of Cry1Ac (Fig. 4b). Expression of insecticidal protein is most critical in leaves as majority of the newly hatched lepidopterans larvae initially feed by scraping chlorophyll in the tender leaves and, as they grow, moves over to the squares and bolls for further feeding and development (Jayaraj 1982; Gore et al. 2002).
The variability in expression of insecticidal gene was recorded in plants having Cry1Ac under rbcS promoter that induced consistent Cry1Ac expression at different growth stages of cotton plant. The use of this promoter is also useful for addressing the biosafety issues as this promoter activity is limited to green parts of plants, hence no gene expression in roots and cotton seed was observed (data not shown), ultimately no threat to soil organisms and safety of cotton products and by products can also be assured.
Abbreviations
- CEMB:
-
Centre of Excellence in Molecular Biology
- rbcS :
-
Rubisco small subunit
- Rubisco:
-
Ribulose-1,5-bisphosphate carboxylase/oxygenase
- ELISA:
-
Enzyme linked immunosorbent assay
- PCR:
-
Polymerase chain reaction
- SDS-PAGE:
-
Sodium dodecyl sulphate-polyacrylamide gel electrophoresis
References
Adamczyk JJ, Hardee DD, Adams LC, Sumerford DV (2001) Correlating differences in larval survival and development of bollworms (Lepidoptera: Noctuidae) and fall armyworms (Lepidoptera: Noctuidae) to differential expression of Cry1Ac(c) δ-endotoxin in various plant parts among commercial cultivars of transgenic Bacillus thuringiensis cotton. J Econ Entomol 94:284–290
Adamczyk JJ, Perera JO, Meredith WR (2009) Production of mRNA from the Cry1Ac transgene differs among bollgard lines which correlate to the level of subsequent protein. Transgenic Res 18:143–149
Amarasinghe BHR, Nitschke EF, Wu Y, Udall JA, Dennis ES, Constable G, Llewellyn DJ (2006) Genomic approaches to the discovery of promoters for sustained expression in cotton (Gossypium hirsutum L.) under field conditions: expression analysis in transgenic cotton and Arabidopsis of a RuBisCo small subunit promoter identified using EST sequence analysis and cDNA microarrays. Plant Biotechnol 23:437–450
Anisimov A, Kimmo K, Anne K, Seppo K, Kari J, Viktor K (2007) Cloning of new RuBisCo promoters from Brassica rapa band determination of their activity in stably transformed Brassica napus and Nicotiana tabacum plants. Mol Breed 19:241–253
Bae TW, Park HR, Kwak YS, Lee HY, Ryu SB (2005) Agrobacterium tumefaciens-mediated transformation of a medicinal plant Taraxacum platycarpum. Plant Cell Tissue Organ Cult 80:51–57
Bakhsh A (2010) Expression of two insecticidal genes in cotton. PhD dissertation, University of the Punjab, Lahore, Pakistan
Bakhsh A, Rao AQ, Shahid AA, Husnain T, Riazuddin S (2010) CaMV 35S is a developmental promoter being temporal and spatial in expression pattern of insecticidal genes (cry1ac & cry2a) in cotton. Aust J Basic Appl Sci 4:37–44
Bakhsh A, Shahzad K, Husnain T (2011a) The variation in spatio temporal expression of insecticidal genes in transgenic Cotton. Czech J Genet Plant Breed 47:1–9
Bakhsh A, Rao AQ, Shamim Z, Husnain T (2011b) A minireview; RuBisCo small subunit as strong green tissue specific promoter. Arch Biol Sci Belgrade 63:299–307
Benedict JH, Sachs ES, Altman DW, Deaton WR, Kohel RJ, Berberich SA (1996) Field performance of cottons expressing transgenic Cry1A insecticidal proteins for resistance to Heliothis virescens and Helicoverpa zea (Lepidoptera: Noctuidae). J Econ Entomol 89:230–238
Chen S, Wu J, Zhou B, Huang J, Zhang R (2000) On the temporal and spatial expression of Bt toxin protein in Bt transgenic cotton. Acta Gossypii Sin 12:189–193
Conner AJ, Glare TR, Nap JP (2003) The release of genetically modified crops into the environment. Part II. Overview of ecological risk assessment. Plant J 33:19–46
Data K, Vasquez Z, Tu J, Torrizo L, Alam MF, Oliva N, Abrigo E, Khush GS, Datta SK (1998) Constitutive and tissue-specific differential expression of the cryI(A)b gene in transgenic rice plants conferring resistance to rice insect pests. Theor Appl Genet 97:20–30
Dong HZ, Li WJ (2007) Variability of endotoxin expression in Bt transgenic cotton. J Agron Crop Sci 193:21–29
Finnegan EJ, Lllewellyn DJ, Fitt GP (1998) What’s happing to the expression of the insect protection in field-grown Ingard cotton. In: Proceedings 9th Australian Cotton Conference, ACGRA, Wee Waa, pp 291–297
Fitt GP, Daly JC, Mares CL, Olsen K (1998) Changing efficacy of transgenic cotton plant patterns and consequences. In: Sixth Australian entomological research conference, Brisbane
Fujimoto H, Itoh K, Yamamoto M, Kyozuka J, Shimamoto K (1993) Insect resistant rice generated by introduction of a modified d-endotoxin gene of Bacillus thuringiensis. Biol Technol 11:1151–1155
Gore J, Leonard BR, Adamczyk JJ (2001) Bollworm (Lepidoptera: Noctuidae) survival on Bollgard® and Bollgard II® cotton flower buds (squares) and flowers. J Econ Entomol 94:1446–1451
Gore J, Leonard BR, Church CE, Cook DR (2002) Behavior of bollworm (Lepidoptera: Noctuidae) larvae on genetically engineered cotton. J Econ Entomol 95:763–769
Gould J, Magallanes-Cedeno M (1998) Adaptation of cotton shoot apex culture to agrobacterium-mediated transformation. Plant Mol Biol Rep 16:1–10
Greenplate JT (1999) Quantification of Bacillus thuringiensis insect control protein Cry1Ac over time in Bollgard cotton fruit and terminals. J Econ Entomol 92:1377–1383
Greenplate JT, Head GP, Penn SR, Kabuye VT (1998) Factors potentially influencing the survival of Helicoverpa zea on Bollgard cotton. In: Dugger P, Richter DA (eds) Proceedings, 1998 beltwide cotton conference. National Cotton Council of America, Memphis, TN, pp 1030–1033
Greenplate JT, Penn SR, Mullins JW, Oppenhuizen M (2000) Seasonal Cry1Ac levels in DP50B: the Bollgard basis for Bollgard II. In: Dugger P, Richter R (eds) Proceedings, beltwide cotton conference, San Antonio, TX, 4–8 Jan 2000. National Cotton Council of America, Memphis, TN, pp 1039–1041
Guo WZ, Sun J, Guo YF, Zhang TZ (2001) Investigation of different dosage of inserted Bt genes and their insect-resistance in transgenic Bt cotton. Acta Genet Sin 28:668–676
Haddad R, Morris K, Buchanan-Wollaston V (2002) Regeneration and transformation of oilseed (Brassica napus) using CaMV 35S promoter-β-glucuronidase gene. J Agric Sci Technol 4:151–160
Hobbs SLA, Warkentin TD, Delong CMO (1993) Transgene copy number can be positively or negatively associated with transgene expression. Plant Mol Biol 2:17–26
Jaakola L, Pirttila AM, Halonen M, Hohtola A (2001) Isolation of high quality RNA from Bilberry (Vaccinium myrtillus L.) fruit. Mol Biol 19:201–203
Jang IC, Lee KH, Nahm BH, Kim JK (2002) Chloroplast targeting signal of a rice rbcS gene enhances transgene expression. Mol Breed 9:81–91
Jayaraj S (1982) Biological and ecological studies of Heliothis. In: Reed W, Kumble V (eds) Proceedings of the international workshop on Heliothis management. ICRISAT Center, Patancheru, AP, India, pp 17–28
Khan GA, Bakhsh A, Riazuddin S, Husnain T (2011) Introduction of cry1Ab gene into cotton (Gossypium hirsutum) enhances resistance against Lepidopteran pest (Helicoverpa armigera). Span J Agr Res 9:296–300
Kim EH, Suh SC, Park BS, Shin KS, Kweon SJ, Han EJ, Park SH, Kim YS, Kim JK (2009) Chloroplast- targeted expression of synthetic Cry1Ac in transgenic rice as an alternative strategy for increased pest protection. Planta 230:397–405
Kuiper HA, Kleter GA, Noteborn HP, Kok EJ (2001) Assessment of the food safety issues related to genetically modified foods. Plant J 27:503–528
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Li X, Wang XD, Zhao X, Dutt Y (2004) Improvement of cotton fiber quality by transforming the acsA and acsB genes into Gossypium hirsutum L. by means of vacuum infiltration. Plant Cell Rep 22:691–697
Liu QQ, Yu HX, Zhang WJ, Wang H, Gu MH (2005) Zhi Wu Sheng Li Yu Fen Zi Sheng Wu Xue Xue Bao. J Plant Physiol Mol Biol 31:247–253
Mahon R, Finnergan J, Olsen K, Lawrence L (2002) Environmental stress and the efficacy of Bt cotton. Austrians Cotton Grow 22:18–21
Manjunatha R, Pradeep S, Sridhar S, Manjunatha M, Naik MI, Shivanna BK, Hosamani V (2009) Quantification of Cry1Ac protein over time in different tissues of Bt cotton hybrids. Karnataka J Agric Sci 22:609–610
Maqbool A, Abbas W, Rao AQ, Irfan M, Zahur M, Bakhsh A, Riazuddin S, Husnain T (2010) Gossypium arboreum GHSP26 enhances drought tolerance in Gossypium hirsutum. Biotechnol Prog 26:21–25
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Nilsson O, Aldén T, Sitbon F, Anthony CH, Chalupa V, Sandbergand G, Olsson O (1992) Spatial pattern of cauliflower mosaic virus 35S promoter-luciferase expression in transgenic hybrid aspen trees monitored by enzymatic assay and non-destructive imaging. Transgenic Res 1:209–220
Odell JT, Nagy F, Chua NH (1985) Identification of DNA sequences required for activity of the cauliflower mosaic virus 35S promoter. Nature 313:810–812
Olsen KM, Daly JC, Holt HE, Finnegan EJ (2005) Season-long variation in expression of Cry1Ac gene and efficacy of Bacillus thuringiensis toxin in transgenic cotton against Helicoverpa armigera (Lepidoptera: Noctuidae). J Econ Entomol 98:1007–1017
Panguluri SK, Sridhar J, Jagadish B, Sharma PC, Kumar PA (2005) Isolation and characterization of a green tissue-specific promoter from pigeon pea (Cajanus cajan L. Millsp.). Indian J Exp Biol 43:369–372
Pauk J, Stefanov I, Fekete S, Bögre L, Karsai I, Fehér A, Dudits D (1995) A study of different (CaMV 35S and mas) promoter activities and risk assessment of field use in transgenic rapeseed plants. Euphytica 85:411–416
Potenza C, Aleman L, Sengupta-Gopalan C (2003) Targeting transgene expression in research, agricultural, and environmental applications: promoters used in plant transformation. In Vitro Cell Dev Biol Plant 40:1–22
Rao CK (2009) Transgenic Bt technology: 4. Expression of Transgenes (4.2 Variation in expression of Bt genes) Available online http://plantbiotechnology.org.in/issue38.html
Rao AQ (2009) Expression of phytochrome B gene in cotton. PhD dissertation, The University of Punjab, Lahore
Rashid B, Saleem Z, Husnain T, Riazuddin S (2008) Transformation and inheritance of Bt genes in Gossypium hirsutum. J Plant Biol 51:248–254
Saiki R, Chang CA, Levenson CH, Warren TC, Boehm CD, Kazazian HHJ, Erlich HA (1988) Diagnosis of sickle cell anemia and beta-thalassemia with enzymatically amplified DNA and nonradioactive allele-specific oligonucleotide probes. N Engl J Med 319:537–541
Shelton AM, Zhao JZ, Zhao RT (2002) Economic, ecological, food safety and social consequences of the development of Bt transgenic plants. Annu Rev Entomol 47:845–881
Shimizu M, Kimura T, Koyama T, Suzuki T, Ogawa N, Miyashita K, Sakka K, Ohmiya K (2002) pCambia vector molecular breeding of transgenic rice plants expressing a bacterial chlorocatechol dioxygenase gene. Appl Environ Microbiol 68:4061–4066
Song P, Heinen JL, Burns TH, Allen RD (2000) Expression of two tissue-specific promoters in transgenic cotton plants. J Cotton Sci 4:217–223
Sunilkumar G, Mohr L, Lopata-Finch E, Emani C, Rathore KS (2002) Developmental and tissue-specific expression of CaMV 35S promoter in cotton as revealed by GFP. Plant Mol Biol 50:463–474
Suzuki Y, Kaori N, Ryuichi Y, Tadahiko M, Amane M (2009) Differences in expression of the RBCS multigene family and RuBisCo protein content in various rice plant tissues at different growth stages. Plant Cell Physiol 50:1851–1855
Wessel Vl Tom R, Antoinette B, Linus VP, Alexander VK (2001) Characterization of position-induced spatial and temporal regulation of transgene promoter activity in plants. J Exp Bot 52:949–959
Wong EY, Hironaka CM, Fischhoff DA (1992) Arabidopsis thaliana small subunit leader and transit peptide enhance the expression of Bacillus thuringiensis proteins in transgenic plants. Plant Mol Biol 20:81–93
Xia L, Xu Q, Guo S (2005) Bt insecticidal gene and its temporal expression in transgenic cotton plants. Acta Agron Sin 31:197–202
Yang NS, Christou P (2005) Cell type specific expression of a CaMV 35S-GUS gene in transgenic soybean plants. Dev Genet 11:289–293
Acknowledgments
The author (Dr. Allah Bakhsh) is highly thankful to Higher Education Commission (HEC), Pakistan for providing funds to CEMB, University of the Punjab for his research activities during PhD studies. Many thanks to Dr. Ghazanfar Ali Khan, Incharge Cotton Research Station Vehari for providing seed of cotton cultivar FDH-786 and Dr. Muhammad Aslam, Principal Scientist NIAB, Faisalabad for providing seed of cotton cultivar NIAB-846. The services rendered by CEMB insect rearing laboratory (Muhammad Illyas, Abdul kareem and Raza Ali Zaidi) are also acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bakhsh, A., Siddique, S. & Husnain, T. A molecular approach to combat spatio-temporal variation in insecticidal gene (Cry1Ac) expression in cotton. Euphytica 183, 65–74 (2012). https://doi.org/10.1007/s10681-011-0497-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-011-0497-8