Abstract
The rice cultivar ‘Chumroo’ is commonly cultivated in the mid- and high-altitude areas of Bhutan. This cultivar has shown durable blast resistance in that area, without evidence of breakdown, for over 20 years. Chumroo was inoculated with 22 blast isolates selected from the race differential standard set of Japan. The cultivar showed resistance to all the isolates. To identify the resistance gene(s), Chumroo was crossed with a susceptible rice cultivar, Koshihikari. The F1 plants of the cross showed resistance. Segregation analyses of 300 F3 family lines fitted the segregation ratio of 1:2:1 and indicated that a single dominant gene controls the resistance to a blast isolate Ao 92-06-2 (race 337.1). The Chumroo resistance locus (termed Pi46(t)) was mapped between two SSR markers, RM6748 and RM5473, on the terminal region of the long arm of chromosome 4, using linkage analysis with SSR markers. The nearest marker, RM5473, was linked to the putative resistance locus at a map distance of 3.2 cM. At the chromosomal region, no true resistance genes were identified, whereas two field resistance genes were present. Therefore, we designated Pi46(t) as a novel blast resistance locus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Magnaporthe oryzae (anamorph, Pyricularia oryzae) is the causal agent of blast disease in rice (Oryza sativa L.) (Couch and Kohn 2002). Blast resistance is the most effective means of controlling the disease without chemical applications. Race–cultivar specificity between the rice and M. oryzae complies with the gene-for-gene hypothesis (Flor 1956; Silué et al. 1992), which states that for every race-specific resistance gene in the host, a corresponding avirulence gene in the pathogen causes an incompatible response. More than 40 resistance genes, and several avirulence genes, conferring blast resistance have been identified in rice and the blast pathogen, respectively (Chen et al. 1999; Yasuda et al. 2004, 2005).
The extensive use of blast-resistant rice cultivars with an introduced resistance gene has engendered the development of new blast races that are virulent to the resistant cultivars, leading to subsequent breakdown of that resistance (Kiyosawa 1982; Bonman 1992). Based on the gene-for-gene hypothesis, the International Rice Research Institute (IRRI)-Japan collaborative research project is successfully constructing an international differential system of blast resistance genes by developing near-isogenic lines and monogenic lines with the genetic backgrounds of three blast susceptible varieties (Kobayashi et al. 2007; Fukuta et al. 2009; Hayashi and Fukuta 2009). This differential system will improve our understanding of the characteristics of blast disease and facilitate the development of blast-resistant rice breeding (Telebanco-Yanoria et al. 2008; Kobayashi et al. 2009).
The rice cultivar ‘Chumroo’ was selected from eight Bhutanese cultivars using the race differential standard set of Japan and identified as a potential donor of a novel resistance gene (Thinlay et al. 2000a). Chumroo is cultivated mostly in the mid- and high-altitude rice growing areas in Bhutan (Thinlay et al. 2000b). Chumroo can be grown under irrigated or upland conditions; the area under cultivation for this one cultivar is estimated to be about 20–30% of the high altitude rice area in Bhutan. Chumroo shows high resistance to Bhutanese blast fungus isolates; no evidence of breakdown has been found for over 20 years after its introduction from Nepal (Thinlay, pers. commun.). Chumroo was the only variety that was unaffected during the 1995/1996 rice blast epidemic in Bhutan (Thinlay, pers. commun.).
This study examines this remarkable blast resistance of Chumroo, by inoculating different Japanese isolates onto Chumroo and by linkage mapping of a locus that contributes to this resistance.
Materials and methods
Plant materials
The blast resistant rice cultivar from Bhutan, Chumroo, was used in this study. This cultivar is a Japonica-type with a bold grain and a red pericarp. A lowland rice cultivar, Koshihikari, harboring a blast resistance gene, Pik-s, was used as a susceptible cultivar. A cross between a female Koshihikari and pollen from Chumroo was conducted. For dominance analysis, 30 F1 individuals were used. An F1 plant was cultivated and self-fertilized, and 300 F2 plants were self-fertilized to develop 300 F3 family lines. This F3 line group was used for F2 segregation analysis and linkage analysis.
The differential rice varieties presented in Table 1 were supplied from the National Agricultural Research Center for Tohoku Region (Yamada et al. 1976; Kiyosawa 1984; Noda et al. 1999; Koizumi et al. 2007). Each variety possesses several resistance genes, chosen from Pik-s, Pish, Pia, Pii, Pik, Pik-m, Piz, Pita, Pita-2, Piz-t, Pik-p, Pib, Pit, and Pi19(t).
Fungal isolates, inoculation, and disease rating
Table 1 shows the races of the Magnaporthe oryzae isolates used for this study. All blast isolates were selected from the race differential standard set of Japan (Hayashi 2005). The race number of each fungus isolate was estimated according to the differential varieties (Yamada et al. 1976; Kiyosawa 1984; Noda et al. 1999). These fungus isolates are compatible with several resistance genes (Pik-s, Pish, Pia, Pii, Pik, Pik-m, Piz, Pita, Pita-2, Piz-t, Pik-p, Pib, Pit, and Pi19(t)).
For inoculation, fungal isolates were grown on oatmeal agar medium at 26°C for 11–12 days. Aerial mycelia of the agar culture were then washed off gently with a water-soaked paintbrush. They were placed at 21°C for 3–4 days under continuous illumination with fluorescent light to induce sporulation. To prepare a spore suspension, the mycelia were scraped and flooded with water containing 0.05% Tween 20. The suspension was filtered through a gauze mesh and adjusted to 1 × 105 spores/ml. Simultaneously, approximately 30 rice seeds were sown in a plastic tray (15 × 5 × 10 cm) and grown in a greenhouse at 20–30°C. Rice seedlings were inoculated at the 3.8-leaf to 5.0-leaf stages by spraying with 50 ml of the spore suspension described above. The inoculated seedlings were placed immediately in a dark chamber with a moisture-saturated atmosphere at 25°C for 20 h and then transferred to a greenhouse at 20–28°C.
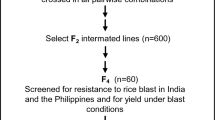
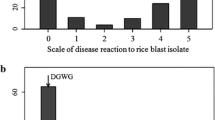
Approximately 7 days after inoculation, the plants were scored for resistance to the blast fungus isolates according to the classification of Hayashi et al. (2009). The classification comprises: 0 = no visible evidence of infection; 1 = uniform or scattered brown specks of smaller than 0.5 mm in diameter, no sporulation; 2 = small lesions with distinct tan centers surrounded by a darker brown margins approximately 1 mm in diameter, no sporulation; 3 = small eyespot lesions less than one and a half times the interval between thin veins or less than 1.5 mm in diameter surrounded by dark brown, lesions capable of sporulation; 4 = intermediate-size eyespot lesions less than twice the distance between thin veins or less than 2 mm diameter; and 5 = large eyespot lesions more than twice the interval between thin veins or more than 2 mm in diameter. Rice classified as type 0–2 is said to be resistant, and rice classified as type 3–5 is considered susceptible.
Segregation analysis and dominance analysis
A blast isolate, Ao 92-06-2 (race 337.1) was used for genetic analysis due to its stable virulence and high sporogenesis ability. In addition, compared to other isolates, it is compatible with many resistance genes shown in Table 1. The F3 plants were inoculated with Ao 92-06-2 to analyze the segregation of resistance in their parental F2 plants. The F3 family lines were classified into three phenotypes: resistant homozygote, segregating heterozygote, and susceptible homozygote, respectively. The reactions of 10 F1 plants were also confirmed using the same blast isolate.
Linkage mapping using SSR markers
Whole genomic DNA was extracted from leaves of each F2 plant according to the modified CTAB method (Murray and Thompson 1980). Polymerase chain reaction (PCR) was performed in a 20 μl reaction mixture containing 10 ng of DNA, 1 pmol of each primer, and 0.4 unit of Taq polymerase. Thermal cycling was conducted using a GeneAmp PCR System 9700 (PerkinElmer Inc., Boston, USA) programmed for 9 min at 95°C; 35 cycles of 1 min at 95°C, 1 min at 55°C and 1 min at 72°C; and 5 min at 72°C for the final extension. The PCR products were separated on 4% polyacrylamide denaturing gels and banding patterns were visualized using the silver staining method, as described by Panaud et al. (1996).
To identify the locus associated with resistance in Chumroo, bulk segregant analysis was conducted. Polymorphisms between Koshihikari and Chumroo were surveyed using 283 SSR markers covering all rice chromosomes (Temnykh et al. 2000; McCouch et al. 2002). Polymorphic markers were subjected to bulk segregant analysis in the F3 family lines. First, the map location of the resistant locus was roughly estimated in 94 randomly selected F3 lines. To detect the linkage with these SSR markers, we used an χ 2 test (Ise 1996) and the program MAPMAKER/EXP 3.0, based on the Kosambi function (Lander et al. 1987). For further detailed mapping, we selected 16 SSR markers on chromosome 4 and 143 F3 lines with a clear response to inoculation. The program MAPMAKER/EXP 3.0, based on the Kosambi function, was used to build the linkage map.
Results
Inoculation of Japanese fungus isolates of M. oryzae onto Chumroo
Table 1 shows the reactions of Chumroo and differential rice cultivars to the 22 fungal isolates. Chumroo was resistant to all the blast isolates. Figure 1 shows the symptoms of Koshihikari and Chumroo inoculated with a blast isolate Ao 92-06-2. Chumroo showed eyespot symptoms surrounded by brown necrotic margins, which is classified as a true resistant response according to the standard of Hayashi et al. (2009). Koshihikari showed typical susceptible symptoms with no brown lesions, which is classified as a susceptible response.
A cross between Koshihikari and Chumroo was conducted and the F1 individuals from the cross were inoculated with fungus isolate Ao 92-06-2. All the F1 individuals displayed brown necrotic lesions (Fig. 1). The symptoms of the F1 individuals were similar to those of Chumroo and were classified as a true resistant response.
Segregation analysis of F2 by F3 line group
Subsequently, 300 F3 family lines were developed from the cross between Koshihikari and Chumroo. Segregation analyses of the 300 F3 family lines showed that 86, 142, and 72 F2 lines were resistant, segregating, and susceptible, respectively, to blast isolate Ao 92-06-2 (Table 2). The segregation ratio corresponded to the expected segregation ratio governed by a single dominant locus, i.e. 1:2:1 (χ 2 = 2.16, P = 0.3396). This result suggests that at least one dominant locus contributes to the blast resistance of Chumroo.
Linkage mapping
The PCR screening and linkage analysis results are shown in Fig. 2. To analyze the locus containing the resistance gene, we surveyed polymorphisms between Chumroo and Koshihikari using 283 SSR markers covering all rice chromosomes. Polymorphisms were observed for 153 markers. Co-dominant polymorphisms were observed for 114 markers. For linkage analysis, we randomly selected 94 F3 lines and 39 co-dominant markers covering all rice chromosomes, where the map distance between each marker was at least 20 cM. Accordingly, linkage between the resistance locus of Chumroo and markers on chromosome 4 was detected (data not shown). Thus, for more detailed linkage mapping, we selected 16 co-dominant SSR markers on chromosome 4 and 143 F3 lines with a clear response to inoculation. As a result, the resistance locus was mapped between markers RM6748 and RM5473 on the terminal region of the long arm of chromosome 4 (Fig. 2). The putative map distances of RM6748 and RM5473 to the resistance locus were 7.4 and 3.2 cM, respectively.
Discussion
We surveyed the reactions of Chumroo to blast fungus isolates selected from the differential standard set of Japan. Blast resistance is classified into two classes: true (vertical, complete) resistance comprising incompatible, hypersensitive symptoms; and field (horizontal, partial) resistance, which is a reduction in the degree of compatible symptoms (Ezuka 1972; Parlevliet 1979; Asaga 1981). Chumroo showed signs of true resistance to all fungal isolates presented in Table 1. F1 individuals from the cross of Koshihikari and Chumroo showed symptoms similar to those of Chumroo (Fig. 1). The estimated segregation ratio of the F2 generation of the cross between Koshihikari and Chumroo fitted well to the expected segregation ratio governed by a single dominant locus, suggesting that a dominant locus contributes to Chumroo’s resistance to Ao 92-06-2. Subsequently, we conducted linkage analysis to investigate relationships between this locus conferring resistance of Chumroo and known resistance genes. The locus of Chumroo was mapped on the terminal part of the long arm of chromosome 4. We tentatively designated this resistance locus in chromosome 4 of Chumroo as Pi46(t), under the international agreement of nomenclature (Kinoshita and Rothschild 1995). In future studies, one or several genes responsible for conferring resistance of Chumroo will be identified in this locus.
There is no report of a true resistance gene in this region, whereas the presence of two field resistance genes has been reported. Pi39(t) and Pikahei-1(t) are located on the terminal part of the long arm of chromosome 4 and these are tightly linked with RM5473 (Terashima et al. 2008; Xu et al. 2008). Rice variety Mineharuka, with Pi39(t), was susceptible to some blast races and developed the same type of blast lesions as susceptible cultivars (Terashima et al. 2008). Likewise, Pikahei-1(t) was described as a field resistance gene (Xu et al. 2008). Generally, blast resistance genes show isolate (race) specificity. There are numerous reports for multiple resistance genes, each detected from a separate resource and located in a single locus (Fukuta et al. 2009). Whereas, there is no report on the specificity of Pi46(t), Pi39(t) and Pikahei-1(t). To investigate this point, a near isogenic line (NIL), into which the resistance gene will be inserted in a susceptible variety’s genetic background, should be developed for accurate differentiation. The NIL will be challenged with various blast isolates, e.g. those shown in Table 1, to confirm that the incompatible response to these isolate are controlled by Pi46(t).
Variations in race-specificity have posed a problem in the breeding of blast-resistant cultivars. One strategy for developing a rice cultivar with durable blast resistance is the multiline concept. Jensen (1952) and Borlaug (1953) proposed multiline concepts for disease control in oat (Avena sativa L.) and wheat (Triticum aestivum L.), respectively. Thereafter, multiline rice cultivars were bred in Japan to suppress damage caused by rice blast disease. At present, three multiline cultivars—Sasanishiki BL (Abe 2004), Koshihikari Toyama BL (Kojima et al. 2003), and Koshihikari Niigata BL (Ishizaki 2007)—are used in practical cultivation by farmers. In Japan, 10 resistance genes (Pia, Pii, Piz, Piz-t, Pita, Pita-2, Pib, Pik, Pik-m, and Pik-p) were introduced into the multiline cultivars. However, the existence of blast isolates compatible with each gene or of multiple genes of these 10 genes, coupled with the breakdown of resistance of several genes, has led to fears of the appearance of a multi-compatible fungus isolate that infects all the multilines. Consequently, for sustainable use of multiline cultivars, it is important to introduce new resistance genes into the cultivar. Therefore, Pi46(t), when introduced into multiline cultivars, is potentially very useful.
References
Abe S (2004) Breeding of a blast resistant multiline variety of rice, Sasanishiki BL. JARQ 38:149–154
Asaga K (1981) A procedure for evaluating field resistance to blast in rice varieties. J Cent Agric Exp Stn 35:51–138 (in Japanese with English summary)
Bonman JM (1992) Durable resistance to rice blast disease—environmental influences. Euphytica 63:115–123
Borlaug NE (1953) New approach to the breeding of wheat varieties resistant to Puccinia graminis tritici. Phytopathology 43:467
Chen DH, dela Vina M, Inukai T, Mackill DJ, Ronald PC, Nelson RJ (1999) Molecular mapping of the blast resistance gene, Pi44(t), in a line derived from a durably resistant rice cultivar. Theor Appl Genet 98:1046–1053
Couch BC, Kohn LM (2002) A multilocus gene genealogy concordant with host preference indicates segregation of a new species, Magnaporthe oryzae, from M. grisea. Mycologia 94:683–693
Ezuka A (1972) Field resistance of rice varieties to blast disease. Rev Plant Prot Res 5:1–21
Flor HH (1956) The complementary genetic systems in flax and flax rust. Adv Genet 8:29–54
Fukuta Y, Xu D, Kobayashi N, Telebanco-Yanoria MJ, Hairmansis A, Hayashi N (2009) Genetic characterization of universal differential varieties for blast resistance developed under the IRRI-Japan Collaborative Research Project using DNA markers in rice (Oryza sativa L.). JIRCAS Working Report No 63, pp 35–68
Hayashi N (2005) MAFF microorganism genetic resources manual 18: rice blast fungus. National Institute of Agrobiological Sciences, Tsukuba
Hayashi N, Fukuta Y (2009) Proposal for a new international system of differentiating races of blast (Pyricularia oryzae Cavara) by using LTH monogenic lines in rice (Oryza sativa L.). JIRCAS Working Report No 63, pp 11–16
Hayashi N, Kobayashi N, Vera Cruz CM, Fukuta Y (2009) Protocols for the sampling of diseased specimens and evaluation of blast disease in rice. JIRCAS Working Report No 63, pp 17–33
Ise K (1996) Statistical analysis. In: Yamamoto R, Horisue N, Ikeda R (eds) Rice breeding manual. Yokendo, Tokyo, pp 244–251 (in Japanese)
Ishizaki K (2007) Studies on practical application of multiline cultivar, Koshihikari, with blast resistance genes in Niigata Prefecture. J Niigata Agric Res Inst 8:1–37 (in Japanese with English summary)
Jensen NF (1952) Intra-varietal diversification in oat breeding. Agron J 44:30–34
Kinoshita T, Rothschild G (1995) A report of the coordinating committee of rice genetics cooperative. Rice Genet Newsl 12:5–6
Kiyosawa S (1982) Genetics and epidemiological modeling of breakdown of plant disease resistance. Ann Rev Phytopathol 20:93–117
Kiyosawa S (1984) Establishment of differential varieties for pathogenicity test of rice blast. Rice Genet Newsl 1:95–97
Kobayashi N, Terebanco-Yanoria MJ, Tsunematsu H, Kato H, Imbe T, Fukuta Y (2007) Development of new sets of international standard differential varieties for blast resistance in rice (Oryza sativa L.). JARQ 41:31–37
Kobayashi N, Ebron LA, Fujita D, Fukuta Y (2009) Identification of blast resistance genes in IRRI-bred rice varieties by segregation analysis based on a differential system. JIRCAS Working Report No 63, pp 69–86
Koizumi S, Iwano M, Zenbayashi K, dela Peña FA, Sonoda R, Nakajima T, Arai M, Nakajima T, Miyasaka A, Ashizawa T, Yasuda N, Noguchi M (2007) Distribution of pathogenic races of the rice blast fungus in Japan in 2001. Misc Publ Natl Agric Res Cent 7:1–63 (in Japanese)
Kojima Y, Ebitani T, Kaneda H, Doi M, Ishibashi T, Kidani Y, Mukaino N, Yamaguchi T, Omoteno M, Yamamoto Y (2003) Development and utilization of isogenic lines Koshihikari Toyama II. Development of isogenic lines Koshihikari Toyama BL. Bull Toyama Agric Res Cent 20:13–32 (in Japanese with English summary)
Lander ES, Green O, Abrahanson J (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
McCouch SR, Teytelman L, Xu Y, Lobos KB, Clare K, Walton M, Fu B, Maghirang R, Li Z, Xing Y, Zhang Q, Kono I, Yano M, Fjellstrom R, DeClerck G, Schneider D, Cartinhour S, Ware D, Stein L (2002) Development and mapping of 2240 new SSR markers for rice (Oryza sativa L.). DNA Res 9:199–207
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Noda T, Hayashi N, Du PV, Dinh HD, E LV (1999) Distribution of pathogenic races of rice blast fungus in Vietnam. Ann Phytopathol Soc Jpn 65:526–530
Panaud O, Chen X, McCouch SR (1996) Development of microsatellite markers and characterization of simple sequence length polymorphism (SSLP) in rice (Oryza sativa L.). Mol Gen Genet 252:597–607
Parlevliet JE (1979) Components of resistance that reduce the rate of epidemic development. Ann Rev Phytopathol 17:203–222
Silué D, Notteghem JL, Tharreau D (1992) Evidence of a gene-for-gene relationship in the Oryza sativa–Magnaporthe grisea pathosystem. Phytopathology 82:577–580
Telebanco-Yanoria MJ, Imbe T, Kato H, Tsunematsu H, Ebron LA, Vera Cruz CM, Kobayashi N, Fukuta Y (2008) A set of standard differential blast isolates (Magnaporthe grisea (Herbert) Barr.) from the philippines for rice (Oryza sativa L.) resistance. JARQ 42:23–34
Temnykh S, Park WD, Ayres N, Cartinhour S, Hauck N, Lipovich L, Cho TG, Ishii T, McCouch SR (2000) Mapping and genome organization of microsatellite sequences in rice (Oryza sativa L.). Theor Appl Genet 100:697–712
Terashima T, Fukuoka S, Saka N, Kudo S (2008) Mapping of a blast field resistance gene Pi39(t) of elite rice strain Chubu 111. Plant Breed 127:485–489
Thinlay, Koizumi S, Ashizawa T, Zenbayashi K (2000a) Characterization of resistance to blast in Bhutanese rice cultivars. Jpn J Phytopathol 66:267
Thinlay, Zeigler RS, Finckh MR (2000b) Pathogenic variability of Pyricularia grisea from the high- and mid-elevation zone of Bhutan. Phytopathology 90:621–628
Xu X, Chen H, Fujimura T, Kawasaki S (2008) Fine mapping of a strong QTL of field resistance against rice blast, Pikahei-1(t), from upland rice Kahei, utilizing a novel resistance evaluation system in the greenhouse. Theor Appl Genet 117:997–1008
Yamada M, Kiyosawa S, Yamaguchi T, Hirano T, Kobayashi T, Kushibuchi K, Watanabe S (1976) Proposal of a new method for differentiating races of Pyricularia oryzae Cavara in Japan. Ann Phytopathol Soc Jpn 42:216–219
Yasuda N, Fujita Y, Noguchi M (2004) Identification of avirulence genes in the rice blast fungus corresponding to three resistance genes in Japanese differentials. J Gen Plant Pathol 70:202–206
Yasuda N, Noguchi MT, Fujita Y (2005) Identification of an Avirulence gene in Magnaporthe grisea corresponding to a resistance gene at Pik locus. Phytopathology 95:768–772
Acknowledgments
We thank Dr. T. Nagamine and Dr. T. Ishii, National Agricultural Research Center for Western Region (WeNARC), Dr. M. Kawase, National Institute of Agrobiological Sciences, and Dr. K. Matsuya, Japan International Cooperation Agency for their technical advice. We also thank Dr. E. Araki, WeNARC, for providing the SSR markers. We also thank the field technicians of the Technical Support Section, WeNARC, who provided the plant materials. We also thank Ms. N. Kobayashi, Ms. I. Ikeda, Ms. K. Hanada, Ms. Y. Tashima, and Ms. M. Oda for their technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Matsushita, K., Yasuda, N., Thinlay et al. A novel blast resistance locus in a rice (Oryza sativa L.) cultivar, Chumroo, of Bhutan. Euphytica 180, 273–280 (2011). https://doi.org/10.1007/s10681-011-0405-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-011-0405-2