Abstract
Quantitative trait loci (QTL) analysis was conducted to identify QTL for seed yield and color retention following processing of a recombinant inbred line (RIL) black bean population. A population of 96 RILs were derived from the cross of black bean cultivars ‘Jaguar’ and 115M and evaluated in replicated trials at one location over 4 years (2004–2007) in Michigan. A 119-point genetic map constructed using simple sequence repeat (SSR), sequence related amplified polymorphism (SRAP), target region amplified polymorphism (TRAP) and phenotypic markers spanned fifteen linkage groups (LG) or 460 cM of the bean genome. Fourteen QTL for yield and color retention in four environments were identified by composite interval mapping on six linkage groups. A major QTL SY10.2J115 for seed yield was identified on LG B10 with additional QTL on B3, B5, and B11. Color retention following processing was associated with loci on B1, B3, B5, B8, and B11. 115M possessed positive alleles for yield, but negative alleles for color retention. Some QTL for yield and color retention co-localized with regions identified in previous studies while others, particularly for color retention, were unique. Additional QTL for agronomic and canning quality traits were detected and individual contributions to future black bean breeding are discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Bean breeding programs share a common goal to develop high yielding cultivars with desirable agronomic and quality traits (Beaver and Osorno 2009; Brick and Grafton 1999). However, genetic improvement of yield potential in common bean (Phaseolus vulgaris L.) has been lower than in other crops (Nienhuis and Singh 1988; Kelly et al. 1998) as breeding for the interrelated effects of growth habit, seed size, maturity, and gene pool further limit progress (Kornegay et al. 1992). Bean breeders have increasingly utilized wild germplasm as knowledge of the domestication syndrome (Koinange et al. 1996) has led to renewed procedures for capturing favorable alleles from the wild that enhance genetic potential for expression of seed yield (Acosta-Gallegos et al. 2007; Beaver and Osorno 2009). Breeding techniques such as the inbred-backcross line (IBL; Bliss 1993) method are especially well suited for introgressing exotic variability into cultivated backgrounds. Researchers at the Centro Internacional de Agricultura Tropical (CIAT, Cali, Colombia) have extensively utilized the IBL method to introgress diversity from wild bean accessions of Colombian origin into domesticated germplasm (Beebe et al. 2004; Blair et al. 2006a). One of these IBL populations derived from the Colombian wild bean G24423 and the black bean cultivar ‘Tacana’ (Lopez-Salinas et al. 1997) has been widely tested in Mexico (Acosta-Gallegos et al. 2007), and Michigan. One IBL designated 115M (aka ‘Black Rhino’) produced yields of 5790 kg ha−1 in Michigan, and has surpassed the yield of all local checks as well as the recurrent parent ‘Tacana’ by as much as 27% (Beaver et al. 2003; Kelly 2004). Black beans represent the largest commercial bean class in Michigan (46% acres harvested in 2008 and represents 57.6% of the U.S. production; NASS 2009) and growers need high-yielding cultivars to remain competitive (http://www.ers.usda.gov/Briefing/drybeans/PDFs/DBnOutlook.pdf.)
Consumers, bean processors and therefore plant breeders desire a canned bean product that retains strong black color and has intact beans that provide a desirable texture (Posa-Macalincag et al. 2002). The polyphenolic anthocyanin pigments in the black bean seed coat (Lin et al. 2008) are highly water soluble and are leached during thermal processing. Different processing methods change the antioxidant activities associated with anthocyanin pigments and seed coat color of black beans (Xu and Chang 2008, 2009). Maintaining these quality attributes in future cultivars is critical, since black beans that are especially prone to color loss during processing, produce a brown, washed-out, and generally unappealing canned bean product (Bushey and Hosfield 2007). Cultivars that retain more color and antioxidants following processing are valued by food processors. Balancing yield potential with seed quality while utilizing wild germplasm can be especially challenging since the wild bean may contribute alleles for superior yield but inferior quality.
To better understand the genetic architecture contributing to the yield potential of 115M across such diverse environments, a quantitative trait loci (QTL) study was initiated using a recombinant inbred line (RIL) population (‘Jaguar’/115M population). This population facilitated the simultaneous study of both yield and seed quality in black bean by determining the location of genetic loci associated with components of both traits. Previous QTL studies in bean have examined agronomic components of yield potential (Beattie et al. 2003; Tar’an et al. 2002; Blair et al. 2006a). Although these studies have identified some similar QTL, many of the results have been unique to specific populations or environments, suggesting that further studies are warranted. Similar results have been observed when examining QTL controlling canning quality. Although previous work has conclusively shown that components of canning quality are quantitatively inherited and exhibit a continuous range of phenotypes (Hosfield et al. 1984; Walters et al. 1997; Posa-Macalincag et al. 2002; Lu et al. 1996), little is known about the number or location of loci contributing to color retention following processing of black bean. Consequently, developing cultivars that possess a balance of positive agronomic attributes as well as acceptable canned bean color retention remains challenging. The current study was conducted to collect phenotypic data for QTL analysis of yield and canned bean quality in a black bean RIL population grown over multiple seasons in Michigan, USA.
Materials and methods
‘Jaguar’ (Kelly et al. 2001) and the breeding line 115M (CIAT) were used as parents to develop 96 F4:5 RILs via single seed descent. ‘Jaguar’ is a black bean cultivar with acceptable canning quality adapted to Michigan growing conditions. 115M is a high-yielding, poor canning quality black bean IBL developed at CIAT from the backcross of ‘Tacana’ (Lopez-Salinas et al. 1997) with G24423, a Colombian wild bean selected for its unique molecular marker pattern in a genetic diversity study (Tohme et al. 1996). The two black bean parents have similar agronomic traits but differ in yield potential, disease reactions and canning quality.
Phenotypic data
The RILs, along with ‘Jaguar’, ‘Tacana’, 115M, and the commercial black bean check cultivar ‘T-39’, were evaluated at the Saginaw Valley Bean and Beet Research Farm (Saginaw, MI) from 2004 to 2007. Four row plots 6.4 m in length with 0.5 m row spacing were organized in a 10 × 10 lattice with three replications. Standard agronomic practices were followed to ensure adequate crop growth and development. Agronomic data were collected for yield, seed size and moisture, days to flower, plant height, lodging, and days to maturity. Seed size was determined as the weight of 100 seeds and both yield and seed size were reported at 18% seed moisture. Lodging was recorded on scale of 1, erect to 5, prostrate. Following harvest, the population was canned according to the methods of (Hosfield et al. 1984) and evaluated for canned bean quality during the years 2005–2007. Since can sizes have changed from methods described in Hosfield et al. (1984), a 90 g seed sample (based on dry matter) of each individual in the population was used for canning. Canning samples were taken from the same replicate in the field. Color measurements on processed beans were recorded as a luminosity (L) value on the Hunter LAB scale using a LabScanXE (Hunter Laboratory, Reston, VA) where 1 = black and 100 = white. Texture measurements were made with a Kramer Shear Press (Food Technology Corp., Sterling, VA) for two 100 g samples taken from a single can according to the bean processing methodology posted on the Bean Improvement Cooperative website (http://www.css.msu.edu/bic/PDF/Bean_Processing.pdf, verified Feb. 2010). Texture data were recorded as the kg-force needed to compress a sample to the point of catastrophic bean failure. Visual appeal was subjectively rated on a 1 = undesirable to 7 = desirable scale by a group of panelists. This evaluation is based upon whole bean integrity (no splitting or clumping); uniformity of size (uniform water uptake); black color (limited leaching); and clear brine (no starch extrusion into canning liquid). Washed-drained weight (processor yield) was determined as the weight of the entire canned bean sample rinsed under cold water and allowed to drip dry for 2 min on a standard number 8 (2.36 mm) sieve. Segregation for resistance to race 73 of Colletotrichum lindemuthianum and to race 22:2 of Uromyces appendiculatus (Pastor-Corrales et al. 2010) were assayed in the greenhouse as phenotypic markers following the methods of Kelly et al. (1994) and Stavely (1983), respectively.
Genotypic data
The RIL population and parents were grown in the greenhouse and DNA was extracted from young trifoliate leaf tissue bulked from three to four individual plants per genotype using a modified CTAB method (Haley et al. 1994). DNA concentrations were determined with a fluorometer (Hoeffer DyNA Quant 200, San Francisco, CA) according to the manufacturer’s procedure and adjusted to 40 ng μl−1 for use in PCR. Molecular markers screened for polymorphisms between the parents 115M and ‘Jaguar’ included 444 simple sequence repeat (SSR) (Blair et al. 2003; Buso et al. 2006; Benchimol et al. 2007; Campos et al. 2007; Grisi et al. 2007; Hanai et al. 2007; http://www.css.msu.edu/bic/PDF/Bean_SSR_Primers_2007.pdf), 64 sequence related amplified polymorphism (SRAP) (Li and Quiros 2001), 220 target region amplified polymorphism (TRAP) (Hu and Vick 2003) and 7 sequence characterized amplified region (SCAR) markers. Those markers that were polymorphic between parents were used to genotype the population. Standard microsatellite amplification conditions were performed and products visualized according to Blair et al. (2003). For SRAP and TRAP markers, amplification and electrophoresis were performed as described by Terpstra et al. (2006).
Data analysis, linkage map construction, and QTL analysis
Analysis of variance for all traits by year was performed with Proc GLM in the Statistical Analysis System (SAS 2000) (SAS Institute Inc., Cary, NC). Linkage analysis was performed on genotypic data using JoinMap 3.0 (Van Ooijen and Voorrips 2001). The Kosambi mapping function was used with a minimum LOD threshold of 3.0 and recombination frequency ≤0.3 to divide 182 markers into linkage groups, determine marker order, and calculate relative map positions. Linkage groups were identified and named according to the core reference map (Freyre et al. 1998) based on microsatellite map locations previously assigned by Blair et al. (2003) and Grisi et al. (2007). Remaining linkage groups were anchored by mapping one or more markers in a subset of the BAT93/JaloEEP558 RIL population.
QTL analysis was performed for each trait and environment using Windows QTL cartographer v2.5 (Wang et al. 2007) and the mean value for each line in each year. The Composite Interval Mapping (CIM) function set to a window size of 10 cM, 5 background markers, 2 cM walk speed, and a forward and backward regression model was used to identify QTL. Significant QTL for individual traits were determined by the location of the peak LOD score at a genome wide empirical threshold of P = 0.05 after 1000 permutation tests (Churchill and Doerge 1994). Linkage maps and QTL were displayed using Mapchart v2.2 (Voorips 2002).
Results
Phenotypic data
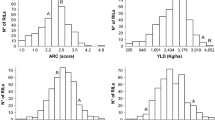
Genotypes and environments (years) differed significantly (P < 0.0001) for both agronomic and canned bean quality traits. Significant genotype by environment (GxE) interaction was also detected for all traits except days to flower. Transgressive segregation (Table 1) and nearly normal distributions (data not shown) were observed for all traits. Seed yield of individual RILs ranged from a low of 1214 kg ha−1 in 2004 to a high of 4261 kg ha−1 in 2006, a range of 3047 kg ha−1 (data not shown). These extremes were observed in the driest (195 mm) and wettest (328 mm) years, respectively. Nine lines yielded in the top 10% of the yield trial two or more years, and two lines appeared in this group all 4 years. As expected limited variation was observed among RILs for seed size, days to flower, plant height, lodging, and days to maturity and those data are summarized in Tables 1 and 2.
The population showed a range of variation for canned bean color, texture, visual appeal, and washed drained weight. Average values for each quality trait varied significantly (P < 0.05) among years (Table 3). Canned beans retained the most color in 2006 with a mean L-value (luminosity) of 14.2 and exhibited the greatest color loss in 2005 with a mean L-value of 17.6 (Table 3; Fig. 1). The two lines that consistently appeared in the top 10% of the trial based on yield retained less color than ‘Jaguar’ or the check ‘T-39’ The softest textures occurred in 2007 with a mean of 53.5 kg-force needed to compress a 100 g sample of processed beans to the point of catastrophic bean failure. Conversely, beans from 2005 had the firmest texture of the three seasons with a mean of 63.9 kg-force. Although lines were nearly normally distributed, there were several that more closely resembled the firmer texture of 115M. Visual appeal ranged from 1.6 to 3.2, indicating a generally undesirable appearance similar to 115M. Washed-drained-weight ranged from a low of 243 g in 2005 to a high of 254 g in 2007 (Table 3). These values correspond to values reported by Hosfield and Uebersax (1980) who had used higher dry weights (115 g) and larger can sizes in their canning process. Observed ratios of drained weight to dry matter (2.6) were also similar to those of Hosfield and Uebersax (1980).
Genotypic data
A total of 182 loci were included in the linkage analysis and 119 markers were placed on the linkage map divided among 15 linkage groups for a total map distance of 460 cM. The remaining 63 markers were not successfully linked. The number of markers per linkage group varied from 2 to 24. Markers clustered on regions of B5, B6, and B10, while their distribution was more uniform for the remaining regions of the linkage map. Three linkage groups (LG1–LG3) consisting of a total of seven markers were not successfully anchored to one of the 11 bean linkage groups (Freyre et al. 1998), and B9 was the only linkage group not represented in the current map. Fifty-six SSR, forty-two SRAP, and 81 TRAP loci were considered in the linkage analysis, with 117 of these loci placed on the map. Two phenotypic markers for disease resistance to race 73 of C. lindemuthianum, and race 22:2 of U. appendiculatus were also mapped.
QTL analysis
Composite interval mapping identified a total of 36 QTL for agronomic traits including yield and fourteen QTL for canned bean quality traits on 10 linkage groups (Tables 4, 5). Individual QTL explained 8–28% of the phenotypic variation for a given trait in a single year. The total phenotypic variation explained for a trait by all QTL detected in the same year varied from 10 to 50%. The number of QTL per trait ranged from 1 to 3. Individual environments varied in regard to total number of QTL and which traits for which those loci were detected (Tables 4, 5). Several of the QTL represent a similar genetic region associated with the same trait across multiple years, while others represent a genetic region that was only associated with a trait in one or 2 years.
Linkage groups B3, B5, B10, and B11 possessed significant QTL for yield in one or more environments from 2004 to 2006 (Table 4). The R2 values ranged from 0.08 to 0.28, and additive effects varied from 41 to 192 kg ha−1. The QTL on B10 consistently had the largest effect on yield, was detected in three environments, and accounted for up to 28% of variation in yield within a single year (Tables 4; Fig. 2). The only yield QTL associated with a ‘Jaguar’ allele was detected on linkage group B3 in the 2004 environment, and was located 20 cM from a QTL associated with a 115M allele in 2006. No yield QTL were identified in 2007, despite the detection of two or three QTL each year from 2004 to 2006 that accounted for 24–43% of the variation in yield each year.
Additional QTL for other agronomic traits were also identified (Table 4). Linkage groups B5, B6, B8, and B11 possessed QTL for seed size. B11 and the unanchored LG2 were associated with days to flower, and regions of B1, B3, B5, B7, and LG2 contributed QTL for maturity. B4b and B6 were implicated in lodging, and height was associated with regions of B3, B5, B6, and B11. Several of these loci were detected in only single environments, but a number of the QTL were consistent in two or more years (B6 and B11 for seed size, B11 and LG2 for days to flower, B5 and LG2 for maturity, B4b and B6 for lodging, and B6 for height).
Seven QTL influencing canned bean color retention resided on linkage groups B1, B3, B5, B8, and B11 and collectively accounted for as much as 47% of the variation for color within a single year (Table 5). QTL for color were detected in all three environments considered. B5 and B8 were associated with color in two of the 3 years, while the other regions were detected only in single environments. With the exception of one locus on B11, all color QTL were associated with 115M and an overall decrease in canned bean color retention. In 2007, a slight decrease in color was associated with the ‘Jaguar’ allele on B11. Regions of B6 and B11 were associated with texture, accounting for 11 and 12% of the variation in 2005 and 2006, respectively. Two QTL for visual appearance were detected. In 2005, a QTL on B5 accounted for 12% of the variation, while in 2006 a QTL that explained 19% of the variation in visual appeal was detected on B8. QTL for washed-drained weight were detected only in 2006 on B3 and two regions of B10. These QTL controlled 12, 11, and 10% of the variation in washed-drained weight.
Discussion
Phenotypic data
The evaluation of the recombinant inbred line population across 4 years provided the opportunity to observe these genotypes in the field under a range of environmental conditions. The four growing seasons represented a range of conditions from dry and hot environments that limited yields to years with more moderate temperatures and adequate precipitation that maximized yield potential. Breeding line B04431 produced the top mean yield over 4 years of 3654 kg ha−1, significantly exceeded the mean yield of the population, and checks ‘Jaguar’, and ‘T-39’, but failed to yield significantly more than the high yielding parent 115M. Eight additional lines also consistently produced yields above the test mean and in the top 10% of the population in two or more years. Conversely, eleven lines yielded significantly less than ‘Jaguar’, illustrating the difficulty of breeding for yield potential, even with two agronomically desirable parents. Although significant genotype by environment interactions were present in all years, those lines with stable yield potential across environments represent useful germplasm that will be utilized as parents to improve the yield potential of future breeding lines.
The variation observed for canned bean color among years agrees with previous studies that have shown seed quality traits vary widely based on environmental conditions among growing seasons (Fig. 1) (Hosfield et al. 1984; Posa-Macalincag et al. 2002; Lu et al. 1996; Walters et al. 1997). Differences in weather patterns such as air temperature or available moisture at critical times during the development of the bean crop have been implicated in the large range in bean quality traits from one season to the next (Hosfield et al. 1984). However, genetic differences still account for a significant portion of this variation (Shellie and Hosfield 1991; Posa-Macalincag et al. 2002).
‘Jaguar’ and 115M differed significantly in regard to seed quality, with 115M exhibiting particularly inferior color retention following processing. These contrasts led to a distribution of progeny suitable for detecting QTL. Unfortunately, some of the highest yielding lines had the worst color retention after processing. Visual appearance, including color, is important to bean processors and consumers, and these lines fail to meet minimum quality standards when compared with check cultivars. The prevalence of lines with undesirable canned bean color resembling that of 115M was unexpected since ‘Jaguar’ retains acceptable color levels when canned. The two parents also differed considerably in canned bean texture, and consequently the RILs ranged widely for this trait. Visual appeal was generally poor for this population, largely due to poor color retention. Despite the acceptable canning quality of ‘Jaguar’, the RILs tended to more closely resemble the unacceptable quality of 115M. Washed-drained weight was less variable than the other canning traits, but was within the acceptable range for black beans based on washed drained ratio of 2.6.Washed drained weight is an important parameter for food processors as it provides an estimate of yield of canned product from a specific quantity of dry beans. Low values are undesirable as fewer canned beans are being produced from the same quantity of dry product.
Linkage map and QTL
Common bean has 11 linkage groups, which correspond to the genome’s 11 chromosomes that are estimated to cover a total genetic distance of 1200 cM (Freyre et al. 1998; Pedrosa-Harand et al. 2008). The Jaguar x 115M map consisted of 15 linkage groups that spanned 460 cM representing 38% of the estimated genome size. Ten of the 11 linkage groups of the bean consensus map were anchored based on the placement of SSR markers (Blair et al. 2003), or mapping of SRAP and TRAP markers in the BAT93 x JaloEEP558 (BJ) core map population. Linkage group B9 was absent from the current map, although three small linkage groups remain unanchored and one may represent a small portion of B9. Linkage groups B1 and B4 were each represented by two un-joined linkage groups. The phenotypic marker(s) for resistance to anthracnose race 73 mapped to B1 (Vallejo and Kelly 2008) and resistance to rust race 22:2 mapped to B4 (Wright et al. 2008) in regions where disease resistance loci for both pathogens are located on the published bean map (Miklas et al. 2006a).
Relatively few of the markers screened across the two black bean cultivars were polymorphic in this study. Polymorphism varied by marker type: 12.6% for SSR, 25% for TRAP, and 33% for SRAP. However, SRAP and TRAP markers were overall much more polymorphic than SSR, which agrees with previous studies (V. Vallejo, personal comm.; Miklas et al. 2006b). The low number of polymorphic SSR markers available in this study also suggests that despite the polymorphism observed in genetic diversity studies (Blair et al. 2006b; Gomez et al. 2004), additional SSRs are needed to make the marker system widely applicable to elite populations or cultivar development. The number of publicly available common bean SSR markers more than doubled during this study (Buso et al. 2006; Benchimol et al. 2007; Campos et al. 2007; Grisi et al. 2007; Hanai et al. 2007), which enhanced the linkage map and subsequent QTL analysis (Tables 4, 5), but underscores the difficulty of finding adequate level of polymorphism between highly related parents used in breeding.
A total of fourteen QTL for yield and canned bean color were identified in this study. These data provide relevant information about location and relative effect of loci controlling these economically important traits even though not all QTL were detected each and every year. Given the genotype by environment interactions observed in this study, the detection of some QTL in only a single year was not unexpected. However, several QTL were detectable in two or three seasons (Tables 4, 5), which suggests these loci are repeatedly contributing a significant portion of the genetic variation for the trait. Even if their effect is not expressed in every environment, several of these regions appear to be reliably associated with a specific trait, and could be targeted for further study or for use in marker assisted breeding.
Seven loci on four linkage groups were associated with increased seed yield. These results support the hypothesis that a number of unique loci, each with relatively small effect, were associated with increased yield potential in this population. However, some regions accounted for a larger portion of the phenotypic variation, such as the QTL for yield on B10 that was associated with up to 28% of the variation in yield. This QTL was detected in three of the four environments, suggesting this region could be targeted by breeders for introgression into other elite germplasm to improve yield potential. The two markers (PV-BR181and PV-BR185) associated with this QTL were tested by Dr. Blair (CIAT) on the parents ‘Tacana’ and G24423 of the original population as we did not have access to DNA or seed of G24423. Dr. Blair determined that this particular genomic region on B10 originated in ‘Tacana’, the recurrent parent of 115M. Markers for the yield QTL on B10 were not detected in the wild bean parent, G24423, but it may be contributing to yield in regions not detected in the QTL analysis. ‘Tacana’ (aka DOR 390) is known to possess increased yield potential under ideal field conditions, but has less of a yield advantage when nutrients and water are limiting factors (Beebe et al. 2008). It is interesting that the B10 QTL for yield in Tacana was detected in a low rainfall year in 2004 but was not detected in 2007 when rainfall provided sufficient soil moisture throughout the growing season and overall growing conditions were generally favorable (Table 2).
Previous studies have identified similar QTL for yield on B3, B5 (Beattie et al. 2003) and B10 (Tar’an et al. 2002) in navy bean. The QTL identified on B10 in our study mapped adjacent to the yield QTL reported by Tar’an et al. (2002). This represents a second QTL for yield on B10 and we propose it be designated SY10.2J115 following the nomenclature proposed by Miklas and Porch (2010). Blair et al. (2006a) also identified two QTL for yield on B3 in an Andean bean population. These data support the complex genetic nature of yield potential described by previous studies and suggest that improvements in yield are possible only by accumulating favorable alleles at a series of unique loci. The SY10.2J115 QTL that demonstrated significant effects over multiple years represents a region that could be targeted for MAS to improve yield in black bean or introgressed into other bean seed types, following verification in other populations.
Seven QTL were identified for seed size on B5, B6, B8 and B11. All were associated with the 115M allele, accounted for no more than 15% of the variation, and increased seed size marginally (<0.5 g). These results were interesting as they suggest no negative effect of the small seeded wild bean, G24423, on seed size in 115M or the population. Similar QTL were identified in diverse bean seed types by Blair et al. (2006a) on B6, B8, and B11, Perez-Vega et al. (2008) on B6 and B8, Tar’an et al. (2002) on B11, and Park et al. (2000) on B5, B6, and B8.
QTL on B11 and the unanchored LG2 were associated with a slight increase in days to flower and accounted for up to 31% of the variation for this trait within a single year. The occurrence of QTL for both days to flower and maturity at the same location on LG2 supports the results of Tar’an et al. (2002) for navy bean and Blair et al. (2006a) for the wild bean G24404. Both of these studies showed co-localized QTL for days to flower and maturity in populations derived from similar genetic backgrounds as the ‘Jaguar’/115M RIL population. Together this information suggests LG2 could correspond to B9 where co-localized QTL for these traits were previously identified, but attempts to anchor this 15 cM linkage group to the core map were unsuccessful. Maturity was also associated with additional loci on B1, B3, B5, and B7, although these QTL tended to be environment specific with only B5 and LG2 QTL detected in multiple years. Blair et al. (2006a) also associated similar regions of B5 and B7 with maturity.
Two QTL on B4 and B6 increased lodging score slightly and were associated with the 115M allele. Beattie et al. (2003) associated a similar region of B4 with lodging in a navy bean population while the QTL on B6 has not been identified in previous studies. The location of the B4b QTL was interesting in that resistance to rust race 22:2 present in 115M mapped to the same region, and a higher than expected frequency of resistant lines was observed, suggesting segregation distortion. Closer examination of the genotypic data revealed all markers in this linkage group showed a distorted segregation pattern that favored the 115M allele, although the reason remains unknown.
The QTL residing on B3, B5, B6, and B11 that were associated with plant height were also environment specific with little consistency among years with the exception of QTL on B6 in 2004 and 2006. Similar QTL associated with increased plant height were reported by Checa and Blair (2008) on B3 and B11, Blair et al. (2006a) and Tar’an et al. (2002) on B6, and Tar’an et al. (2002) and Beattie et al. (2003) on B3. These studies used diverse cultivars, suggesting that plant height is controlled by similar genetic regions in a wide range of genetic backgrounds.
Seven QTL for color retention were identified, suggesting that the contrast between 115M and ‘Jaguar’ for seed quality characteristics resulted in a population suitable for dissecting the genetic control of canned bean color retention. QTL were detected in five regions of the genome, supporting the complex, quantitative, and environmentally specific nature of these traits as established in previous studies (Hosfield et al. 1984). Posa-Macalincag et al. (2002) previously identified a region of B3 that was associated with improved canning quality, however, direct comparison of other QTL locations with previous studies were not possible due to unanchored linkage groups of previous maps. Six of the QTL detected in the current study were associated with 115M and decreased color retention. The negative effect of these loci on canning quality could reflect the wild bean parentage of 115M, although markers associated with color QTL were not tested in the wild bean G24423. These results were not unexpected due to the poor canning quality of 115M. The remaining QTL on B11 slightly decreased color retention, but was associated with ‘Jaguar’, typically viewed as an acceptable cultivar in terms of color retention. Despite the negative effect of these QTL in the present study, knowledge of their genetic location could be helpful in targeting future studies of color retention that could identify more favorable alleles at these loci in other cultivars.
Consumers and processers of black beans prefer beans that retain a dark color when cooked or canned. Anthocyanins are the water soluble compounds responsible for the seed color that accumulate in the seed coat. Variability in color retention may be attributed to overall amount of anthocyanins or amount of anthocyanin leaching (Benninger and Hosfield 2003; Bushey and Hosfield 2007). Since anthocyanins are known to have antioxidant capacity, beans that retain their color may have health benefits beyond those of beans that leach their color. Antioxidants combat the damaging effects of free radicals (Xu et al. 2007). Xu and Chang (2009) showed a positive correlation between anthocyanin amount and antioxidant activity in thermally processed black beans. A negative correlation between L-color value (seed coat whiteness) and antioxidant capacity has also been demonstrated in beans (Xu et al. 2007). Therefore improved color retention in canned or cooked black beans may also improve the nutritional value of black beans in addition to improving appearance.
Although several QTL were identified for canned bean color, significantly less information was revealed for other canning traits. Texture was associated with QTL on B6 and B11 but no QTL were detected in multiple years. However, the total variation explained within a year by all texture QTL in this population was greater than in any of three populations examined by Walters et al. (1997) or two populations studied by Posa-Macalincag et al. (2002).
Similarly, QTL for visual appearance were detected on B5 and B8 in 2005 and 2006, respectively. Washed-drained weight QTL were identified only in 2006 and were located on B3 and on two separate regions of B10. Together, these additional canning quality QTL were limited to single environments and accounted for only a small portion of the total variation. They provide additional preliminary information on the genetic control of these traits, but future studies on canning quality are needed to identify more robust QTL for use in breeding.
Conclusions
The QTL analysis for yield and canned bean color in four environments identified desirable loci from 115M that enhanced yield, and undesirable loci that decreased canned bean color retention. A single QTL on linkage group B10 accounted for up to 28% of the variation for yield within a year, while as much as 43% of the variation for yield was associated with all QTL detected in a single environment. The SY10.2J115 QTL on B10 enhanced yield in three of the four environments, suggesting that this locus should be utilized via marker assisted breeding to increase yield across a range of field conditions. Canned bean color was influenced by loci on at least five linkage groups with relatively equal effects, suggesting that maximizing color retention will require accumulating favorable alleles at all these loci. 115M was generally associated with an undesirable reduction in black seed coat color. However, alleles that enhanced yield could be separated from those that conferred color loss through future breeding, based on the independence of these loci. The utility of SRAP and TRAP markers for mapping and tagging QTL in bean was also validated, as many of the QTL were associated with one of these markers. Although this group of black bean lines reflected the yield potential of 115M, as well as its propensity for color loss when canned, several lines were identified that consistently exceeded the yield of 115M and represent parent material useful for enhancing the yield of other elite germplasm. The yield potential of 115M, and its progeny used in this study, supports the continued use of genetic diversity from wild bean to enhance the yield of cultivated common bean.
Abbreviations
- CIAT:
-
Centro Internacional de Agricultura Tropical
- CIM:
-
Composite interval mapping
- IBL:
-
Inbred backcross line
- MAS:
-
Marker assisted selection
- QTL:
-
Quantitative trait loci
- RIL:
-
Recombinant inbred line
- SCAR:
-
Sequence characterized amplified region
- SRAP:
-
Sequence related amplified polymorphism
- SSR:
-
Simple sequence repeat
- TRAP:
-
Target region amplified polymorphism
References
Acosta-Gallegos JA, Kelly JD, Gepts P. (2007). Pre-breeding and genetic diversity in common bean (Phaseolus vulgaris). Proceedings of international plant breeding symposium, Ciudad de México. Crop Sci 47(S3):S44–S59
Beattie AD, Larsen J, Michaels TE, Pauls KP (2003) Mapping quantitative trait loci for a common bean (Phaseolus vulgaris L.) ideotype. Genome 46:411–422
Beaver JS, Osorno JM (2009) Achievements and limitations of contemporary common bean breeding using conventional and molecular approaches. Euphytica 168:145–175
Beaver JS, Rosas JC, Myers J, Acosta J, Kelly JD, Nchimbi-Msolla S, Misangu R, Bokosi J, Temple S, Arnaud-Santana E, Coyne DP (2003) Contributions of the Bean/Cowpea CRSP to cultivar and germplasm development in common bean. Field Crops Res 82:87–102
Beebe S, Tohme J, Nienhuis J, Pedraza F, Rengifo J, Tovar E, Islam A (2004) Studies in Phaseolus germplasm diversity: a review of work at CIAT. Annu Rep Bean Improv Coop 47:33–34
Beebe SE, Rao IM, Cajiao C, Grajales M (2008) Selection for drought resistance in common bean also improves yield in phosphorous limited and favorable environments. Crop Sci 48:582–592
Benchimol LL, Campos T, Carbonell SAM, Colombo CA, Chioratto AF, Formighieri EF, Gouvea LRL, Souza AP (2007) Structure of genetic diversity among common bean (Phaseolus vulgaris L.) varieties of Mesoamerican and Andean origins using new developed microsatellite markers. Genet Resour Genet Resour Crop Evol 54:1747–1762
Benninger CW, Hosfield GL (2003) Antioxidant activity of extracts, condensed tannin fractions, and pure flavonoids from Phaseolus vulgaris L. seed coat color genotypes. J Agric Food Chem 51:7879–7883
Blair MW, Pedraza F, Buendia HF, Gaitan-Solis E, Beebe SE, Gepts P, Tohme J (2003) Development of a genome-wide anchored microsatellite map for common bean (Phaseolus vulgaris L.). Theor Appl Genet 107:1362–1374
Blair MW, Iriarte G, Beebe S (2006a) QTL analysis of yield traits in an advanced backcross population derived from a cultivated Andean x wild common bean (Phaseolus vulgaris L.) cross. Theor Appl Genet 112:1149–1163
Blair MW, Giraldo MC, Buendia HF, Tovar E, Duque MC, Beebe SE (2006b) Microsatellite marker diversity in common bean (Phaseolus vulgaris L.). Theor Appl Genet 113:100–109
Bliss FA (1993) Breeding common bean for improved nitrogen fixation. Plant Soil 152:71–79
Brick MA, Grafton KF (1999) Improvement of medium-seeded race Durango cultivars. In: Singh S (ed) Common bean improvement in the twenty-first century, 223–253. Kluwer, Dordrecht, the Netherlands
Bushey SM, Hosfield GL (2007) Absorbance of the soak water to predict canning quality. Annu Rep Bean Improv Coop 50:39–40
Buso GSC, Amaral ZPS, Brondani RPV, Ferreira ME (2006) Microsatellite markers for the common bean Phaseolus vulgaris. Mol Ecol Notes 6:252–254
Campos T, Benchimol LL, Carbonell SAM, Chioratto AF, Formighieri EF, Pereira de Souza A (2007) Microsatellites for genetic studies and breeding programs in common bean. Pesq Agropec Bras 42(4):589–592
Checa OE, Blair MW (2008) Mapping QTL for climbing ability and component traits in common bean (Phaseolus vulgaris L.). Mol Breed 22:201–215
Churchill GA, Doerge RW (1994) Empirical threshold values for quantitative trait mapping. Genetics 138:963–971
Freyre R, Skroch PW, Geffroy V, Adam-Blondon AF, Shirmohamadali A, Johnson WC, Llaca V, Nodari RO, Pereira PA, Tsai SM, Tohme J, Dron M, Nienhuis J, Vallejos CE, Gepts P (1998) Towards an integrated linkage map of common bean. 4. Development of a core linkage map and alignment of RFLP maps. Theor Appl Genet 97:847–856
Gomez OJ, Blair MW, Frankow-Lindberg BE, Gullberg U (2004) Molecular and phenotypic diversity of common bean landraces from Nicaragua. Crop Sci 44:1412–1418
Grisi MCM, Blair MW, Gepts P, Brondani C, Periera PAA, Brondani RPV (2007) Genetic mapping of a new set of microsatellite markers in a reference common bean (Phaseolus vulgaris) population BAT93 × Jalo EEP558. Genet Mol Res 6:691–706
Haley SD, Miklas PN, Afanador L, Kelly JD (1994) Random amplified polymorphic DNA (RAPD) marker variability between and within gene pools of common bean. J Am Soc Hort Sci 119:122–125
Hanai LR, Campos T, Camargo LEA, Benchimol LL, Souza AP, Melotto M, Carbonell SAM, Chioratto AF, Consoli L, Formighieri EF, Siqueira MVBM, Tsai SM, Vieira MLC (2007) Development, characterization, and comparative analysis of polymorphism at common bean SSR loci isolated from genic and genomic sources. Genome 50:266–277
Hosfield GL, Uebersax MA (1980) Variability in physico-chemical properties and nutritional components of tropical and domestic dry bean germplasm. J Am Soc Hort Sci 105:246–252
Hosfield GL, Uebersax MA, Isleib TG (1984) Seasonal and genotypic effects on yield and physico-chemical seed characteristics related to food quality in dry, edible beans. J Am Soc Hort Sci 109:182–189
Hu J, Vick BA (2003) Target region amplification polymorphism: a novel marker technique for plant genotyping. Plant Mol Bio Rep 21:289–294
Kelly JD (2004) Advances in common bean improvement: some case histories with broader applications. Acta Hort (ISHS) 637:99–122
Kelly JD, Afanador L, Cameron LS (1994) New races of Colletotrichum lindemuthianum in Michigan and implications in dry bean resistance breeding. Plant Dis 78:892–894
Kelly JD, Kolkman J, Schneider K (1998) Breeding for yield in dry bean (Phaseolus vulgaris L.). Euphytica 102:343–356
Kelly JD, Hosfield GL, Varner GV, Uebersax MA, Taylor J (2001) Registration of ‘Jaguar’ black bean. Crop Sci 41:1647
Koinange EMK, Singh SP, Gepts P (1996) Genetic control of the domestication syndrome in common bean. Crop Sci 36:1037–1045
Kornegay J, White JW, Ortiz de la Cruz O (1992) Growth habit and gene pool effects on inheritance of yield in common bean. Euphytica 62:171–180
Li G, Quiros CF (2001) Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: its application to mapping and gene tagging in Brassica. Theor Appl Genet 103:455–461
Lin L-Z, Harnly JM, Pastor-Corrales MS, Luthria DL (2008) The polyphenolic profiles of common bean (Phaseolus vulgaris L.). Food Chem 107:399–410
Lopez-Salinas E, Acosta-Gallegos JA, Becerra-Leor EN, Frayre-Vazquez G, Orozco SH, Beebe SE (1997) Registration of Negro Tacana common bean. Crop Sci 37:1022
Lu W, Chang KC, Grafton KF, Schwarz PB (1996) Correlations between physical properties and canning quality attributes of navy bean (Phaseolus vulgaris L.). Cereal Chem 73:788–790
Miklas PN, Porch T (2010) Guidelines for common bean QTL nomenclature. Annu Rep Bean Improv Coop 53:202–205
Miklas PN, Kelly JD, Beebe SE, Blair MW (2006a) Common bean breeding for resistance against biotic and abiotic stresses: from classical to MAS breeding. Euphytica 147:105–131
Miklas PN, Hu J, Grunwald NJ, Larsen KM (2006b) Potential application of TRAP (targeted region amplified polymorphism) markers for mapping and tagging disease resistance traits in common bean. Crop Sci 46:910–916
NASS (2009) USDA crop production 2008 summary. http://www.ers.usda.gov/Briefing/drybeans/PDFs/DBnOutlook.pdf
Nienhuis J, Singh SP (1988) Genetics of seed yield and its components in common bean (Phaseolus vulgaris L.) of Middle-American origin. II: genetic variance, heritability and expected response from selection. Plant Breed 101:155–163
Park SO, Coyne DP, Jung G, Skroch PW, Arnaud-Santana E, Steadman JR, Ariyaranthe HM, Nienhuis J (2000) Mapping of QTL for seed size and shape traits in common bean. J Am Soc Hort Sci 125:466–475
Pastor-Corrales MA, Wright EM, Markel SG, Awale HE, Kelly JD, Jordahl JG, Lamppa RS, Mathew FM, Osorno JM, Goswami RS (2010) Comparing the virulence of new races of the common bean rust pathogen from Michigan and North Dakota. Annu Rep Bean Improv Coop 53:128–129
Pedrosa-Harand A, Porch T, Gepts P (2008) Standard nomenclature for common bean chromosomes and linkage groups. Annu Rep Bean Improv Coop 51:106–107
Perez-Vega E, Campa A, Giraldez R, Ferreira JJ (2008) Mapping of QTL involved in the genetic control of seed traits in common bean. Annu Rep Bean Improv Coop 51:116–117
Posa-Macalincag MCT, Hosfield GL, Grafton KF, Uebersax MA, Kelly JD (2002) Quantitative trait loci (QTL) analysis of canning quality traits in kidney bean (Phaseolus vulgaris L.). J Am Soc Hort Sci 127:608–615
SAS Institute, Inc. (2000) SAS version 8. SAS Institute Inc, Cary, NC
Shellie KC, Hosfield GL (1991) Genotype × environmental effects on food quality of common bean: resource-efficient testing procedures. J Am Soc Hort Sci 116:732–736
Stavely JR (1983) A rapid technique for Inoculation of Phaseolus vulgaris with multiple pathotypes of Uromyces phaseoli. Phytopathology 73:676–679
Tar’an B, Michaels TE, Pauls KP (2002) Genetic mapping of agronomic traits in common bean. Crop Sci 42:544–556
Terpstra KA, Wright EM, Kelly JD (2006) Protocol for visualizing sequence related amplified polymorphism (SRAP) and target region amplified polymorphism (TRAP) markers on agarose gels. Annu Rep Bean Improv Coop 49:191–192
Tohme J, Gonzalez DO, Beebe S, Duque MC (1996) AFLP analysis of gene pools of a wild bean core collection. Crop Sci 36:1375–1384
Vallejo VA, Kelly JD (2008) Molecular tagging and genetic characterization of alleles at the Co-1 anthracnose resistance locus in common bean. ICFAI Univ J Genet Evol 1:7–20
Van Ooijen JW, Voorrips RE (2001) JoinMap version 3.0: software for the calculation of genetic linkage maps. CPRODLO Plant Research International, Wageningen, The Netherlands
Voorips RE (2002) Mapchart version 2.2: windows software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Walters KJ, Hosfield GL, Uebersax MA, Kelly JD (1997) Navy bean canning quality: correlations, heritability estimates and randomly amplified polymorphic DNA markers associated with component traits. J Am Soc Hort Sci 122:338–343
Wang S, Basten CJ, Zeng ZB (2007) Windows QTL cartographer 2.5. Department of Statistics, North Carolina State University, Raleigh, NC
Wright EW, Awale HE, Kelly JD (2008) Use of TRAP markers to map resistance to a new race of common bean rust in Michigan. Annu Rep Bean Improv Coop 51:210–211
Xu BJ, Chang SKC (2008) Total phenolic content and antioxidant properties of Eclipse black beans (Phaseolus vulgaris L.) as affected by processing methods. J Food Sci 73:H19–H27
Xu BJ, Chang SKC (2009) Total phenolic, phenolic acid, anthocyanin, flavan-3-ol, and flavonol profiles and antioxidant properties of pinto and black beans (Phaseolus vulgaris L.) as affected by thermal processing. J Agric Food Chem 57:4754–4764
Xu BJ, Yuan SH, Chang SKC (2007) Comparative analyses of phenolic composition, antioxidant capacity, and color of cool season legumes and other selected food legumes. J Food Sci 72:S167–S177
Acknowledgments
The authors wish to thank Matthew Blair for screening parental lines with SSR markers linked to QTL on B10 at CIAT.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wright, E.M., Kelly, J.D. Mapping QTL for seed yield and canning quality following processing of black bean (Phaseolus vulgaris L.). Euphytica 179, 471–484 (2011). https://doi.org/10.1007/s10681-011-0369-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-011-0369-2