Abstract
Interspecific hybridization plays a crucial role in plant genetics and breeding. The efficiency of interspecific crosses to a considerable extent depends on the genetic relatedness of genomes from parental species. Interspecific hybrids involving Brassica maurorum (2n = 16, MM) and two Brassica crop species, viz B. rapa (2n = 20, AA) and B. napus (2n = 38, AACC), were produced and analyzed for their meiotic chromosome pairings in pollen mother cells (PMCs) by using genomic in situ hybridization (GISH) with the labeled DNA of B. maurorum (MM) as probe. In hybrids B. maurorum × B. rapa (2n = 18, MA), all chromosomes remained unpaired in 28% PMCs, and the maximum of autosyndetic bivalents was two and one among the chromosomes of A and M genomes, with the average per cell being 0.27 and 0.12, respectively. Up to two allosyndetic bivalents between A and M genomes appeared, averagely 0.48 per cell. In hybrids B. maurorum × B. napus (2n = 27, MAC), the maximum of autosyndetic bivalents in M genome was two and the average was 0.11, while the maximum of allosyndetic bivalents between M and A/C genomes was two and the average was 0.78. The 2–7 bivalents formed by A/C-genome chromosomes showed their high homology. The results were compared and discussed with the chromosome pairings in the hybrids of B. maurorum with B. juncea and B. carinata with respect to the genome relationships and the potential for chromosome recombination.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In spite of recent advances in applications of genetic engineering and molecular biology techniques for plant breeding, interspecific hybridization still plays a crucial role in crop improvement, by generating new ecotypes or species, broadening the genetic basis of crops and introducing agronomically important genes from wild species into crop germplasm. Furthermore, by investigating the homeologous chromosome pairing during the meiotic divisions in the wide hybrids, the genomic affinities and phylogenetic relationships of the diverse genomes could be established, which serve as a guide in successfully utilizing the alien germplasm resources.
The family Brassicaceae consists of a large number of wild and weedy species which are an excellent reservoir of useful genes for resistance to biotic and abiotic stresses. In Brassica crops, for their economic improvement, many interspecific/intergeneric hybrid combinations have been obtained, and in several instances, introgression of nuclear genes conferring desirable traits have been accomplished (see review by Prakash et al. 2009).
In this communication, we report the development of hybrids of Brassica maurorum with B. rapa and B. napus and their meiotic behaviour using GISH. B. maurorum, a related wild species, is reported to carry resistance to fungal diseases viz. white rust caused by Albugo candida and alternaria blight caused by Alternaria spp. (Chrungu et al. 1999). Although, genomic affinities based on chromosome pairing between B. maurorum and B. rapa/B. napus have been studied in the past (Takahata and Hinata 1983; Chrungu et al. 1999; Garg et al. 2007), the precise degrees of allo- or auto-syndetic pairings could not be ascertained as the parental chromosomes were undistinguishable by conventional cytological methods.
GISH not only discriminates the parental chromosomes in interspecific and intergeneric hybrids, but can also detect the chromosome recombinations (Stevenson et al. 1998; Kamstra et al. 1999; Karlov et al. 1999; Ji et al. 2004). GISH is also effective for identification of Brassica interspecific and intergeneric hybrids (Fahleson et al. 1997; Snowdon et al. 2000; Benabdelmouna et al. 2003; Wang et al. 2005; Ge and Li 2007). In the present study, we applied the GISH technique to discriminate M genome from A/C genomes, and to visualize the extent of auto- and allosyndesis.
Materials and methods
Plant materials and crosses
The species used in the experiment were B. rapa L. (2n = 20, AA), B. napus L. (2n = 38, AACC) and B. maurorum Durieu (2n = 16, MM). B. maurorum, as the female parent, was pollinated by B. rapa (the combination designated as M.A) and B. napus (M.AC). The reciprocal cross B. napus × B. maurorum (AC.M) was also made, but the hybrids were not analyzed for chromosome pairings by GISH. About 3 weeks after pollination, the immature embryos were rescued on MS (Murashige and Skoog 1962) agar medium without hormones. The plantlets from embryo rescue were multiplied by continuously subculturing their axillary buds on MS medium with 1.5 mg/l 6-benzyl aminopurine (6-BA), 0.25 mg/l α-naphthalenacetic acid (NAA) to produce sufficient number of cloned plants, which were rooted and transferred to field for analyses.
For meiotic analysis, the young flower buds of F1 plants were collected and fixed in 3 ethanol : 1 acetic acid solution (Carnoy’s solution) overnight, transferred to the fresh solution and then stored in the refrigerator. For the chromosome counting in mitotic cells, the young ovaries were collected and then pre-treated in 2 mM 8-hydroxyquinoline for 3 h and finally fixed in Carnoy’s solution.
Probe labeling, chromosome preparation and GISH analyses
Total genomic DNA was extracted from young leaves by CTAB method. The DNA of B. maurorum was labeled with Bio-11-dUTP by nick translation and used as probe. The blocks were made by boiling the DNA of B. rapa and B. napus for 15 min.
Slide preparations of chromosomes for GISH were made mainly according to Zhong et al. (1996) and Ge and Li (2007) with some modifications. The enzyme mixture contained 0.6% cellulose “Onozuka” (Yakult Honsha, Japan), 0.4% pectinase (Merck, Germany) and 0.5% Snailase (Sabc, China).
In situ hybridization was carried out according to the protocol by Leitch et al. (1994). Hybridization signals of the B. maurorum probe were detected using Cy3-labeled streptavidin (Sigma, USA), and chromosomes were counterstained with 0.2% 4′-6-Diamidino-2-phenylindole (DAPI) solution (Roche, Basel, Switzerland), mounted in antifade solution (Vector) and examined under the fluorescent microscope (Nikon Eclipse 80i, Japan) equipped CCD camera. Images were processed by Adobe Photoshop (8.0).
Results
Hybridity in all the F1 hybrids of the three combinations obtained following embryo rescue was confirmed cytologically, viz. 2n = 18 for M.A and 2n = 27 for M.AC and AC.M. and by intermediate morphology between the parents. The hybrid plants flowered abundantly, but were totally male and female sterile, and set no seed after selfing and open pollination.
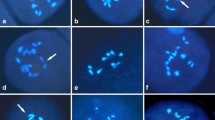
By applying GISH with the labeled DNA of B. maurorum as probe, the chromosomes of M genome could be unequivocally distinguished from those of A and C genomes in pollen mother cells (PMCs) of M.A and M.AC at diakinesis/metaphase I, which made it possible to investigate the inter- and intra-genomic pairings. In M.A, 28% PMCs had all the chromosomes unpaired (18I) with an average of 8.97 univalents in A genome and 7.28 in M genome (Table 1). Up to two bivalents were formed by A-genome chromosomes (Fig. 1 C2) with an average of 0.27 (Table 1), while a maximum of one bivalent with a mean of 0.12 was formed by M-genome chromosomes (Table 1, Fig. 1 B2). Allosyndetic pairing between A and M genome was noticed in 44% PMCs, with a maximum of two bivalents and an average of 0.48 bivalent (Table 1, Fig. 1 A2, C2).
GISH analyses of meiotic pairings in PMCs of hybrid B. maurorum × B. rapa. A1–C1, DAPI images (blue), M-genome chromosomes are marked by arrows. A2–C2 merged images, red signals are from B. maurorum probe. A2 Diakinesis with two IIAM (solid arrow). B2 Diakinesis with one IIAA (line arrow) and one IIMM (arrowhead). C2 Diakinesis with two IIAA (line arrow), and one IIAM (solid arrow). Bar 5 μm
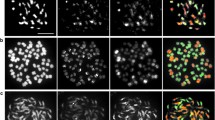
In M.AC, the averages of univalents and autosyndetic bivalents were 7.0 and 0.11 within M-genome chromosomes, respectively (Table 1). The maximum of autosyndetic bivalents was 2 (Table 1, Fig. 2 C2), higher than one in M.A. The mean of allosyndetic bivalents was 0.78 (Table 1), higher than 0.48 in M.A, while the maximal number (2) was the same as in M.A (Fig. 2 B2). All the PMCs observed showed at least two bivalents formed between A/C-genome chromosomes, even up to seven bivalents (Fig. 2 A2). In some PMCs at MI, the M-genome chromosomes were distributed around the periphery of the cells, while A/C chromosomes were at the center (Fig. 2 E). At anaphase I, individual M chromosomes divided precociously (Fig. 2 D2) with a low frequency, but A/C chromosomes did not show such divisions.
GISH analyses of meiotic pairings in hybrid M.AC. A1–D1, DAPI images (blue), M-genome chromosomes are marked by arrows. A2–D2, E merged images, red signals are from B. maurorum probe. A2 MI, seven bivalents formed between A/C-genome chromosomes, and M-genome chromosomes formed eight univalents. B2 Diakinesis with two allosyndetic bivalents between M- and A/C-genome chromosomes (solid arrow). C2 MI with two IIMM (arrowhead) and one M-genome chromosome associated with one A/C-genome chromosome (solid arrow). D2 AI with one M-genome chromosome lagged and another having sister chromatids separated (arrow). E MI with eight M-genome chromosomes (arrow) at the periphery. Bar 5 μm
Discussion
Because the ability to distinguish genomes from different species by GISH depends mainly on the degree of sequence homology (Ji et al. 2004), our results suggest that the sequence homology between M and A/C genomes is relatively low. This conclusion is consistent with earlier cytological and molecular studies which assigned B. maurorum to the nigra lineage while B. rapa to a distant rapa/oleracea lineage (Takahata and Hinata 1983; Warwick and Black 1991; Pradhan et al. 1992). Detectable GISH signals on the chromosomes of M genome are mainly restricted to pericentromeric heterochromatin blocks where repetitive DNA sequences are clustered, as in other Brassica genomes (Snowdon et al. 1997; Ge and Li 2007). Due to small size of Brassica chromosomes, recombinant segments are difficult to detect effectively by GISH (Snowdon et al. 1997, 2000).
Since it is now conclusively established that Brassica genomes evolved from a common ancestral genome of x = 3–7 (see Prakash et al. 2009), chromosome pairing due to both auto-and allosyndesis is expected in hybrids of Brassica species. However, the frequency of allosyndesis between two genomes would reduce due to reproductive isolation, genetic mutation and regulation of chromosome pairing after the divergence of two genomes. In hybrids between crop brassicas and B. maurorum reported previously (Takahata and Hinata 1983; Chrungu et al. 1999; Garg et al. 2007), the chromosome associations investigated by conventional method were speculated to result from both auto- and allo-syndesis, but did not reflect the precise genomic affinity. Herein, with the extent of intra- and intergenomic pairings of M genome with A/C genomes determined by the method of GISH (Fig. 1, 2), the frequency of allosyndesis (0.48 A–M bivalent) in hybrid M.A is higher than that of autosyndesis (0.27, 0.12 bivalents within A and M genomes), reflecting the higher intergenomic homology and more chances for intergenomic recombinations. The average of allosyndetic bivalents (0.48) is also comparable to that (0.43 A–M bivalent) in the hybrid M.AB from the cross between the same B. maurorum type and B. juncea (2n = 36, AABB) but the maximal number (2) is less than that (3), where the degrees of all the types of auto-/allosyndetic pairs for three genomes are evaluated by dual GISH (Yao et al. 2010). The average of autosyndetic bivalents (0.27) within A genome in M.A is lower than 0.39 in M.AB, but the maximal number is the same, two. For the autosyndesis within M genome, the maximal number of bivalents is one for M.A and M.BC/BC.M from the reciprocal crosses of the same B. maurorum type with B. carinata (Yao et al. 2010), and is two for M.AB and M.AC, while the average of bivalents is 0.26 for M.AB, higher that those in other four hybrids (0.11–0.14). One more autosyndetic bivalent in M.AC than M.A may be attributable to the preferential pairings of the chromosomes from A/C genome for their high homeology (up to seven bivalents formed), or the structural difference of A genome from B. rapa and B. napus.
Though A and C genome chromosomes distinction in B. napus has been recently realized by using DNA from B. oleracea as the probe and B. rapa DNA and the intergenic spacer of the B. oleracea 45S rDNA as the block (Howell et al. 2008), we failed to apply the procedure in the hybrid M.AC. Therefore, only the sum of allosyndetic pairings between A/C and M genomes are given, together with the autosyndetic pairing within M genome (Table 1). The average of 0.78 A/C–M bivalents in M.AC is nearly the same as the sums of C–M and B–M bivalents in M.BC/BC.M (0.70/0.68), but less the sum of A–M and B–M bivalents in M.AB (1.15), because the frequency for B–M (0.72) in M.AB was significantly higher than 0.39/0.37 in M.BC/BC.M. The average of A–B bivalent was higher than that of B–M and A–M in AB.M, and the average of B–C was higher than that for B–M or C–M in M.BC and BC.M. These results revealed a closer relationship between the basic genomes A and B or B and C than those between the three basic genomes and M. Furthermore, a maximum of three allosyndetic bivalents appeared between A–B, A–M, and B–M genomes in M.AB, respectively, but two were observed between B–C, B–M, and C–M genomes in M.BC and BC.M. The higher allosyndesis frequency of B–M than A–M or C–M in these hybrids also revealed their closer relationship (Takahata and Hinata 1983; Warwick and Black 1991; Pradhan et al. 1992).
Although the A and C chromosomes share very high homology (Truco et al. 1996) and pair preferentially in B. napus haploids (Nicolas et al. 2008) and triploid hybrids involving A/B/C genome (Ge and Li 2007), the chromosome homeology between M and A/C genome is high enough to lead to the formation of a maximum of two bivalents. Furthermore, the B. napus chromosomes in M.AC may have experienced high frequency of rearrangements, as detected in the B. napus haploid (Nicolas et al. 2007). So the structurally differentiated A/C chromosomes show more allosyndesis, but the maximum of two allosyndetic bivalents between A/C and M genomes is lower than the expected five, for there are three A–M bivalents in M.AB, and two C–M bivalents in M.BC/BC.M (Fig. 3). Similarly, the significant difference in autosyndesis of the same M-genome in these hybrids (Fig. 3) could be due to the different genome combination or/and the structural differentiation of Brassica genomes from different origins, and the different chromosome numbers of three Brassica genomes (n = 8, 9, 10) will contribute to a different competitive condition to some extent.
The summary of autosyndesis and allosyndesis in hybrids of B. maurorum with Brassica species (B. rapa, A; B. napus, B; B. juncea, C; B. carinata, D). The maximum of auto- and allosyndetic bivalents is given for each type, and the multivalents are not shown for very low number and frequency. The data from the hybrid M.BC are listed here, and those for the hybrid BC.M are nearly the same. The double lines represent bivalents
The extent of autosyndesis in A, B, C genomes from three cultivated Brassica diploids was detected earlier from pairings in respective haploids. A maximum of two bivalents and one trivalent occurred in the haploid B. campestris (syn. B. rapa) (Armstrong and Keller 1981). The haploid B. nigra showed a maximum of two bivalents (Prakash 1973), B. oleracea haploid showed a maximum of two bivalents (Thompson 1956), or a maximum of one bivalent and one trivalent (Armstrong and Keller 1982). Two bivalents within B genome in trigenomic triploids (2n = 27, ABC) from B. carinata × B. rapa, natural and synthetic B. napus × B. nigra crosses were observed by GISH analysis (Li et al. 2005; Ge and Li 2007). These and present studies reveal the similar extent of autosyndesis in each Brassica genome from the extant diploids or allotetraploids (Fig. 3), suggesting that the main structures of these genomes are largely maintained during the evolutionary process of these allotetraploids after their formation by natural hybridizations between diploids, though genomic rearrangements are frequently detected in allopolyploids.
Extensive and diverse genomic changes can arise immediately at the onset of genome merging or within a few generations (Rieseberg et al. 1995; Song et al. 1995; Baack et al. 2005; Lukens et al. 2006). Homeologous exchanges have been shown to increase the range of genetic variation observed for important ecological and agronomic traits like flowering time (Pires et al. 2004) or seed yield (Osborn et al. 2003) in newly synthesized B. napus. The evidence presented here for chromosome association and chiasmata formation between M and A/C genomes demonstrates the feasibility of gene transfer from B. maurorum to B. rapa and B. napus. Therefore, exchange of chromosome fragments between M and A/C genomes via allosyndetic pairing is expected. The amphiploids originating from the B. maurorum × B. rapa hybrid also showed resistance to Alternaria brassicae and Albugo candida (Garg et al. 2007), indicating the expression of resistant genes. To transfer the useful genes from M to A/C chromosomes, production of allohexaploids from the B. napus × B. maurorum hybrids and backcrossing progenies with B. napus is being undertaken.
References
Armstrong KC, Keller WA (1981) Chromosomes pairing in haploid of Brassica campestris. Theor Appl Genet 59:49–52. doi:10.1007/BF00275776
Armstrong KC, Keller WA (1982) Chromosomes pairing in haploid of Brassica oleracea. Can J Genet Cytol 24:735–739
Baack EJ, Whitney KD, Rieseberg LH (2005) Hybridization and genome size evolution: timing and magnitude of nuclear DNA content increases in Helianthus homoploid hybrid species. New Phytol 167:623–630. doi:10.1111/j.1469-8137.2005.01433.x
Benabdelmouna A, Guéritaine G, Abirached-Darmency M, Darmency H (2003) Genome discrimination in progeny of interspecific hybrids between Brassica napus and Raphanus raphanistrum. Genome 46:469–472. doi:10.1139/G03-020
Chrungu B, Verma N, Mohanty A, Pradhan A, Shivanna KR (1999) Production and characterization of interspecific hybrids between B. maurorum and crop Brassicas. Theor Appl Genet 98:608–613. doi:10.1007/s001220051111
Fahleson J, Lagercrantz U, Mouras A, Glimelius K (1997) Characterization of somatic hybrids between Brassica napus and Eruca sativa using species-specific repetitive sequences and genomic in situ hybridization. Plant Sci 123:133–142. doi:10.1016/s0168-9452(96)04575-x
Garg H, Banga S, Bansal P, Atri C, Banga SS (2007) Hybridizing Brassica rapa with wild crucifers Diplotaxis erucoides and Brassica maurorum. Euphytica 156:417–424. doi:10.1007/s10681-007-9391-9
Ge XH, Li ZY (2007) Intra- and intergenomic homology of B-genome chromosomes in trigenomic combinations of the cultivated Brassica species revealed by GISH analysis. Chromosome Res 15:849–861. doi:10.1007/s10577-007-1168-4
Howell EC, Kearsey MJ, Jones GH, King GJ, Armstrong SJ (2008) A and C genome distinction and chromosome identification in Brassica napus by sequential fluorescence in situ hybridization and genomic in situ hybridization. Genetics 180:1849–1857. doi:10.1534/Genetics,108.095893
Ji Y, Pertuze R, Chetelat RT (2004) Genome differentiation by GISH in interspecific and intergeneric hybrids of tomato and related nightshades. Chromosome Res 12:107–116. doi:10.1023/B:CHRO.0000013162.33200.61
Kamstra SA, Ramanna MS, De Jeu MJ, Kuipers GJ, Jacobsen E (1999) Homoeologous chromosome pairing in the distant hybrid Alstroemeria aurea × A. inodora and the genome composition of its backcross derivatives determined by fluorescent in situ hybridization with species-specific probes. Heredity 82:69–78. doi:10.1046/j.1365-2540.1999.00465.x
Karlov GI, Khrustaleva LI, Lim KB, Van Tuyl JM (1999) Homoeologous recombination in 2n-gamete producing interspecific hybrids of Lilium (Liliaceae) studied by genomic in situ hybridization (GISH). Genome 42:681–686. doi:10.1139/gen-42-4-681
Leitch AR, Schwarzacher T, Jackson D, Leitch IJ (1994) Microscopy handbook No.27. In situ hybridization: a practical guide. Bios Scientific, Oxford
Li MT, Li ZY, Zhang CY, Qian W, Meng JL (2005) Reproduction and cytogenetic characterization of interspecific hybrids derived from crosses between Brassica carinata and B. rapa. Theor Appl Genet 110:1284–1289. doi:10.1007/s00122-005-1965-0
Lukens LN, Pires JC, Leon E, Vogelzang R, Oslach L et al (2006) Patterns of sequence loss and cytosine methylation within a population of newly resynthesized Brassica napus allopolyploids. Plant Physiol 140:336–348. doi:10.1104/pp.105.066308
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15(3):473–497
Nicolas SD, Le Mignon G, Eber F, Coriton O, Monod H, Clouet V, Huteau V, Lostanlen A, Delourme R, Chalhoub B, Ryder CD, Chèvre AM, Jenczewski E (2007) Homeologous recombination plays a major role in chromosome rearrangements that occur during meiosis of Brassica napus haploids. Genetics 175:487–503. doi:10.1534/genetics.106.062968
Nicolas SD, Leflon M, Liu Z, Eber F, Chelysheva L, Coriton O, Chèvre AM, Jenczewski E (2008) Chromosome ‘speed dating’ during meiosis of polyploid Brassica hybrids and haploids. Cytogenet Genome Res 120:331–338. doi:10.1159/000121082
Osborn TC, Butrulle DV, Sharpe AG, Pickering KJ, Parkin IAP et al (2003) Detection and effects of a homeologous reciprocal transposition in Brassica napus. Genetics 165:1569–1577
Pires JC, Zhao JW, Schranz ME, Leon EJ, Quijada PA et al (2004) Flowering time divergence and genomic rearrangements in resynthesized Brassica polyploids (Brassicaceae). Biol J Linn Soc 82:675–688. doi:10.1111/j.1095-8312.2004.00350.x
Pradhan AK, Prakash S, Mukhopadhyay A, Pental D (1992) Phylogeny of Brassica and allied genera based on variation in chloroplast and mitochondrial DNA patterns: molecular and taxonomical classifications are incongruous. Theor Appl Genet 85:331–340. doi:10.1007/BF00222878
Prakash S (1973) Haploidy in Brassica nigra Koch. Euphytica 22:613–614. doi:10.1007/BF00036663
Prakash S, Bhat SR, Quiros CF, Kirti PB, Chopra VL (2009) Brassica and its close allies: cytogenetics and evolution. Plant Breed Rev 31:21–187
Rieseberg LH, Vanfossen C, Desrochers AM (1995) Hybrid speciation accompanied by genomic reorganization in wild sunflowers. Nature 375:313–316. doi:10.1038/375313a0
Snowdon RJ, Kohler W, Friedt W, Kohler A (1997) Genomic in situ hybridization in Brassica amphidiploids and interspecific hybrids. Theor Appl Genet 95:1320–1324. doi:10.1007/s001220050699
Snowdon RJ, Winter H, Diestal A, Sacristan MD (2000) Development and characterization of Brassica napus-Sinapis arvensis addition line exhibiting resistance to Leptosphaeria maculans. Theor Appl Genet 101:1008–1014. doi:10.1007/s001220051574
Song KM, Lu P, Tang KL, Osborn TC (1995) Rapid genome change in synthetic polyploids of Brassica and its implications for polyploid evolution. Proc Natl Acad Sci USA 92:7719–7723
Stevenson M, Armstrong SJ, Ford-Lloyd BV, Jones GH (1998) Comparative analysis of crossover exchanges and chiasmata in Allium cepa × fistulosum after genomic in situ hybridization (GISH). Chromosome Res 6:567–574. doi:10.1023/A:1009296826942
Takahata Y, Hinata K (1983) Studies on cytodemes in the subtribe Brassicinae. Tohoku J Agric Res 33:111–124
Thompson KF (1956) Production of haploid plants of narrow stem kale. Nature 178:748. doi:10.1038/178748a0
Truco MJ, Hu J, Sadowski J, Quiros CF (1996) Inter- and intra-genomic homology of the Brassica genomes: implications for their origin and evolution. Theor Appl Genet 93:1225–1233. doi:10.1007/BF00223454
Wang YP, Zhao XX, Sonntag K, Wehling P, Snowdon RJ (2005) Behaviour of Sinapis alba addition chromosomes in a Brassica napus background revealed by genomic in situ hybridization. Chromosome Res 13:819–826. doi:10.1007/s10577-005-1017-2
Warwick SI, Black LD (1991) Molecular systematics of Brassica and allied genera (subtribe Brassicinae, Brassiceae)—chloroplast genome and cytodeme congruence. Theor Appl Genet 82:81–92. doi:10.1007/BF00231281
Yao XC, Du XZ, Ge XH, Chen JP, Li ZY (2010) Intra- and intergenomic chromosome pairings revealed by dual-color GISH in Brassica trigenomic hybrids of Brasscia juncea and B. carinata with B. maurorum. Genome 53:14–22. doi:10.1139/G09-082
Zhong XB, Hans JJ, Zabel P (1996) Preparation of tomato meiotic pachytene and mitotic metaphase chromosomes suitable for fluorescence in situ hybridization (FISH). Chromosome Res 4:24–28. doi:10.1007/BF02254940
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yao, XC., Ge, XH., Chen, JP. et al. Intra- and intergenomic relationships in interspecific hybrids between Brassica (B. rapa, B. napus) and a wild species B. maurorum as revealed by genomic in situ hybridization (GISH). Euphytica 173, 113–120 (2010). https://doi.org/10.1007/s10681-010-0131-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-010-0131-1