Abstract
Cytoplasmic male sterility (CMS) in higher plants is a maternally inherited trait and CMS-associated genes are known to be located in the mitochondrial genome. However, CMS-inducing genes in CMS-D2 and CMS-D8 of Upland cotton (Gossypium hirsutum L., AD1) are currently unknown. The objective of this study was to identify potential candidate DNA or gene sequences for CMS-D2 and CMS-D8 through restriction fragment length polymorphism (RFLP) analysis. Seven mtDNA gene probes and five restriction enzymes were first used to compare D2 (from G. harknessii Brandegee) and AD1 cytoplasms. With cox1, cox2, and atp1 as probes, RFLP polymorphisms were detected with one or more restriction enzyme digestions. The most notable difference was an additional fragment in the normal AD1 cytoplasm detected by cox2 in digests of three enzymes, and by cox1 and atp1 in digests with PstI. The RFLP analysis was then conducted among CMS-D2, CMS-D8 (from G. trilobum (DC.) Skovst.), and AD1 cytoplasms. Two probes from maize, atp1 and atp6, detected polymorphism among the different cytoplasmic lines. However, no difference in RFLP patterns was noted between male sterile (A) and restorer (R) lines with the D2 or D8 cytoplasm, indicating that the presence of the D2 or D8 restorer gene does not affect mtDNA organization in Upland cotton. The results demonstrate that RFLP using atp1 and atp6 as probes can distinguish the three cytoplasms. The atp1 and atp6 in CMS-D8 and these two genes together with cox1 and cox2 in CMS-D2 could be the candidates of CMS-associated genes in the mitochondrial genome, providing information for further molecular studies and developing PCR-based markers for the CMS cytoplasms in breeding. This research represents the first work using RFLP to analyze the genetic basis of CMS in cotton.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mitochondria are the cellular sites for key metabolic pathways such as the tricarboxylic acid cycle, respiratory electron transfer, and ATP synthesis (Chase 2006). To date, 14 mitochondrial genomes have been sequenced in plants including: Arabidopsis thaliana L. (Unseld et al. 1997), sugar beet (Beta vulgaris L. subsp. vulgaris, Kubo et al. 2000), canola (Brassica napus L., Handa 2003), Cycas taitungesis C.F. Shen, K.D. Hill, C.H. Tsou & C.J. Chen (Chaw et al. 2008), tobacco (Nicotiana tobacco L., Sugiyama et al. 2005), Oryza sativa L. subsp. indica (Tian et al. 2006), Oryza sativa L. subsp. japonica (Notsu et al. 2002), sorghum (Sorghum bicolor L., Allen et al. 2006), Tripsacum dactyloides L. (Allen et al. 2006), wheat (Tritium aestivum L., Ogihara et al. 2005), Zea luxurians L. (Allen et al. 2006), maize (Z. mays L. subsp. mays, Clifton et al. 2004), Z. mays L. subsp. parviglumis (Allen et al. 2006), and Z. parennis (Hitchc.) Reeves & Manglesdorf (Allen et al. 2006). The mitochondrial genome in plants encodes about 50–60 gene products, a small fraction of the gene products needed for mitochondrial functions. For example, the mitochondrial DNA of A. thaliana, has 366,924 nucleotides, 10% of which code for 57 genes, including those encoding for proteins that are components of the electron transport chain and the F1F0-ATPase, essential to produce ATP (Unseld et al. 1997).
Cytoplasmic male sterility (CMS) is defined as a maternally inherited trait that prevents the production of functional pollen or male gametes in plants (Schnable and Wise 1998). CMS has been widely used in plant breeding for the production of F1 hybrids, such as maize, sorghum, onions (Allium cepa L.), sugar beet, sunflower (Helianthus annuus L.), carrot (Daucus carota L.), canola, and rice (Budar and Pelletier 2001). CMS is associated with chimeric genes in the mitochondrial genome, which express novel open reading frames (ORFs) in most CMS plants (Schnable and Wise 1998). The mitochondrial genes associated with CMS are proposed to arise by aberrant recombination events or by intra-molecular rearrangement of the plant mitochondrial genome (Hanson and Bentolila 2004). The genes associated with CMS have been reported in numerous plant species: maize (Dewey et al. 1986), petunia (Petunia hybrida (Hook.) Vilm, Young and Hanson 1987), bean (Phaseolus vulgaris L., Johns et al. 1992), canola (Singh and Brown 1991; Bonhomme et al. 1991, 1992; Brown 1999; Grelon et al. 1994), radish (Raphanus sativus L., Makaroff et al. 1990), sunflower (Moneger et al. 1994), sorghum (Tang et al. 1996), rice (Akagi et al. 1995; Zhang et al. 2007), sugar beet (Yamamoto et al. 2005), and pepper (Capium annuum L., Kim et al. 2007). Most of these novel orfs are composed of fragments of genes for subunits of the ATP synthase complex, such as atp1 (or atpA), atp6, atp8, and atp9 (Hanson and Bentolila 2004; Chase 2006).
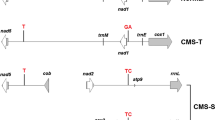
CMS systems have also been studied in cotton (Gossypium hirsutum L.). There are two internationally recognized cotton CMS systems, CMS-D2 and CMS-D8. CMS-D2 was developed by transferring the cytoplasm of wild G. harknessii Brandegee (D2) diploid cotton into tetraploid Upland cotton (G. hirsutum, AD1) (Meyer 1975). The alloplasmic CMS-D8 was derived from introducing G. trilobum (DC) Skovst (D8) cytoplasm into Upland cotton (Stewart 1992). Two different dominant genes, Rf 1 and Rf 2 , restore the fertility for CMS-D2 and CMS-D8, respectively. Rf 1 can also recover fertility of CMS-D8, whereas Rf 2 only restores fertility of CMS-D8 (Zhang and Stewart 2001a, b). Zhang et al. (2004) designed 36 primer pairs based on A. thaliana mitochondrial DNA gene sequences. These mtDNA universal primers were used to amplify three alloplasmic lines, i.e., fertile AD1, CMS-D2, and CMS-D8. The absence of sequence tagged site (STS) polymorphisms among the three cytoplasms proved that the mitochondrial DNA genes in cotton are highly conserved in sequence length as in other plant species. Zhang et al. (2008) further compared gene expression between CMS-D8 restored plants (Rf 2 rf 2 ) and normal non-restoring fertile plants (rf 2 rf 2 ) in cotton by mRNA differential display. The results identified four genes that were up-regulated and 22 genes, including starch synthase (SS), which were down-regulated. The down-regulated SS explained the lack of starch accumulation in sterile rf 2 pollen grains in the heterozygous restored plants.
The mitochondrial CMS loci in the alloplasmic CMS-D2 and CMS-D8 are currently unknown. The objective of this study was to identify potential candidate mtDNA regions or gene sequences uniquely found in CMS-D2 and CMS-D8 through restriction fragment length polymorphism (RFLP) analysis. These novel mtDNA regions, not found in fertile cytoplasms would be candidate CMS loci for the D2 and D8 systems in cotton.
Materials and methods
Plant materials and DNA extraction
In Experiment 1, three lines (Meyer 1975), i.e., CMS-D2 line, HAMS277 (A line, rf 1 rf 1 rf 2 rf 2 with D2 cytoplasm), its corresponding maintainer line, HAB277 (B line. rf 1 rf 1 rf 2 rf 2 with AD1 cytoplasm), and fertility restorer line (R line, Rf 1 Rf 1 rf 2 rf 2 with D2 cytoplasm) were used. In Experiment 2, five Upland cotton lines were used. (1) CMS-D8-8518 (Stewart, unpublished), a CMS line, carries CMS-D8 cytoplasm from American diploid wild species G. trilobum (D8 genome) and has the non-restoring rf 1 rf 1 rf 2 rf 2 nuclear background of 8518. The breeding line, 8518 (Bourland 1996), is a normal male fertile line with fertile AD1 cytoplasm and non-restoring rf 1 rf 1 rf 2 rf 2 nuclear background. (2) Stoneville 474 (ST474) is a normal male fertile commercial cultivar with fertile AD1 cytoplasm and non-restoring rf 1 rf 1 rf 2 rf 2 nuclear background. (3) D8R-8518 (Stewart, unpublished) is a restorer line with CMS-D8 cytoplasm and the restorer genotype rf 1 rf 1 Rf 2 Rf 2 in the 8518 background. (4) D8R-ST474 (Stewart, unpublished) is a restorer line with the CMS-D8 cytoplasm and the restoring genotype rf 1 rf 1 Rf 2 Rf 2 in ST474 background. (5) D2R-B418R (Cook and Namken 1995) is a restorer line with CMS-D2 cytoplasm from American diploid wild species G. harknessii (D2 genome) and the restorer genotype Rf 1 Rf 1 rf 2 rf 2 . Total DNA from the above genotypes were extracted using a cTAB method with (Zhang and Stewart 2000) or without modifications (Paterson et al. 1993).
Southern blot analysis
Genomic DNA (10 or 20 μg) was digested separately with AvaI, BamHI, EcoRI, EcoRV, HindIII, PstI or XbaI; blots were prepared, hybridized and washed as described (Rodriguez-Uribe and O’Connell 2006). Probes for hybridization were prepared using cloned mtDNA genes: wheat for cox1, cotton for cox2 and cox3 (Wang 2008), maize for atp1 (atpA) (Braun and Levings 1985), atp6 (Dewey et al. 1986), cob and rrn26 (26S rRNA) (Stern et al. 1982), and petunia for atp9 (Young and Hanson 1987).
Results
Comparison between CMS-D2 and AD1 mtDNA
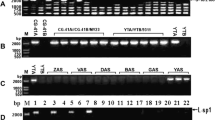
The CMS-D2 cytoplasm had the same mtDNA genome organization in the A or R nuclear background for all seven genes tested (Fig. 1). Four genes, atp6, atp9, cob, and rrn26 (Fig. 1d–g) showed no polymorphism in mtDNA from CMS-D2 line and the fertile Upland cotton (AD1) cytoplasm or the maintainer B line. With cox1 as a probe, a novel PstI fragment was observed in the AD1 cytoplasm (Fig. 1a). When cox2 was the probe, polymorphisms were detected in at least three digests from three of the five restriction enzymes (EcoRV, HindIII, and PstI). As compared with the CMS-D2 cytoplasm, the AD1 mtDNA had an extra fragment of various sizes (Fig. 1b). With the atp1 as the probe, the CMS-D2 cytoplasm had a larger EcoR1 fragment, while the AD1 cytoplasm had a smaller fragment. In the HindIII digest, the AD1 cytoplasm fragment was slightly larger (Fig 1c).
Comparison between CMS-D8 and AD1 mtDNA
To evaluate the possible effect of nuclear backgrounds on mtDNA organization RFLP analyses were performed on a B line, three alloplasmic lines with the CMS-D8 cytoplasm, and a D2 restorer line containing the D2 cytoplasm for a comparison. Probes for atp9, cox1, cox2, cox3, and rrn26 did not reveal polymorphic mtDNA RFLPs among the five lines containing three different cytoplasms; the results for cox3 are presented as an example in Fig 2. The maize atp1 probe revealed polymorphic RFLP among CMS-D8, CMS-D2, and AD1 cytoplasms (Fig. 2). Specifically, the atp1 probe detected RFLP polymorphisms with the EcoRI digestion. All the cytoplasms shared a common fragment of 3.0 kb. The 4.5 kb fragments were only present in CMS-D2 cytoplasm, while the 5.5 kb fragment was unique in CMS-D8 cytoplasm. The 6.0 kb fragment was present in AD1 cytoplasm but absent in both CMS-D8 and CMS-D2 cytoplasm.
Autoradiograph of EcoRI digested cotton genomic DNA, probed with mitochondrial genes. Lane 1. Stoneville 474 (ST474), a male fertile line with AD1 cytoplasm. Lane 2. D8R-ST474, a restorer line with CMS-D8 cytoplasm. Lane 3. D8R-8518, a restorer line with CMS-D8 cytoplasm. Lane 4. CMS-D8-8518, a CMS line, with CMS-D8 cytoplasm. Lane 5. D2R-B418R, a restorer line with CMS-D2-2 cytoplasm. Sizes of hybridizing fragments are indicated, polymorphic bands are marked with arrowheads. Fragment sizes of polymorphic fragments are indicated in kb
The maize atp6 probe also revealed RFLP polymorphisms among the three cytoplasms (Fig. 2). For EcoRI digested cotton genomic DNA, the atp6 probe detected polymorphism among the three cytoplasms. A 1.5 kb fragment was only present in CMS-D8 cytoplasm (Lane 2–4), but absent in AD1 cytoplasm. The fragment in CMS-D2 cytoplasm was larger (Lane 5). Both CMS-D8 and AD1 cytoplasm had two common fragments, i.e., 3.0 and 4.0 kb, while the respective fragments in CMS-D2 cytoplasm were larger.
Discussion
Since the CMS cytoplasms studied here are alloplasmic from two wild diploid D-genome cotton species, it is expected that both chloroplast and mitochondrial genomes in the CMS cytoplasms are different from one another and from that of tetraploid Upland cotton. Indeed, Wendel and Albert (1992) used chloroplast DNA RFLP analysis to construct a phylogenetic tree of 40 species in Gossypium. Both chloroplast and mitochondrial DNA RFLP analyses also confirmed that tetraploid cotton contains the A genome-like cytoplasm (Galau and Wilkins 1989; Wendel 1989; Small and Wendel 1999). Galau and Wilkins (1989) furthered demonstrated that the D2 chloroplast DNA was maintained in the alloplasmic CMS-D2 lines of Upland cotton. Our current mtDNA RFLP analysis indicates that both CMS-D2 and CMS-D8 have different mitochondrial genomes from that of Upland cotton. In all cases where polymorphic mtDNA RFLP was detected, the Upland cotton B lines had the same RFLPs, which were different from the D2 restorer line containing the D2 cytoplasm or the three lines (one A line and two R lines) containing the CMS-D8 cytoplasm which had the same RFLPs. Therefore, the present study provides evidence that the exotic mtDNAs have been maintained in the CMS-D2 and CMS-D8 cytoplasms. By including both CMS lines and restorer line(s) with the CMS-D2 or CMS-D8 cytoplasms, our study also clearly demonstrates that, as with most other CMS systems, the D2 or D8 restorer gene does not change the mitochondrial genome organization, since both A and R lines showed the same RFLP patterns when various probe and restriction enzyme combinations were used. It has been documented that tissue culture may induce structural mutations of mitochondrial genes in plants (Sadoch et al. 2000). The presence of the restorer gene (Fr) results in the permanent deletion of the mitochondrial CMS-inducing sequence (pvs) in common bean (Mackenzie and Chase 1990).
The current study represents the first attempt to employ mtDNA RFLP to analyze mitochondrial genomes of CMS cotton towards a better understanding of male sterility mechanism conditioned by exotic cytoplasms in cotton. Clear mtDNA RFLPs between CMS-D2 cytoplasm and AD1 cytoplasm were seen in cox1, cox2, and atp1, while no apparent polymorphism was observed in atp6 and atp9. Further analyses, included CMS D8, presented in Fig. 2 did confirm the results for atp1 and atp9; however, polymorphisms for cox1 and cox2 were not detected in CMS-D8, while polymorphisms for atp6 were seen. The discrepancies may be due to the use of different probes and enzymes in the two experiments. Taken together, CMS-D2 showed different mtDNA RFLPs from the AD1 cytoplasm in atp1, atp6, cox1, and cox2; while CMS-D8 showed mtDNA RFLPs in atp1 and atp6. The present study demonstrated that mtDNA RFLP is a reliable method to distinguish the two different alloplasmic CMS lines in cotton using atp1 and atp6 as probes. This provides useful information for developing more convenient and portable PCR-based markers to assay CMS cytoplasms for hybrid cotton breeding.
At present, it is still not understood as to how these mtDNA differences are related to the expression of the male sterility in CMS-D2 or CMS-D8. With the completion of sequencing of two chloroplast cotton genomes (Ibrahim et al. 2006; Lee et al. 2006) and a number of other plant mitochondrial genomes (http://www.ncbi.nlm.nih.gov/genomes/GenomesGroup.cgi?taxid=33090&opt=organelle), the 700-kb cotton mitochondrial genome (Hsu and Mullin 1989) should be sequenced in comparison with the CMS-D2 and CMS-D8 mtDNA genomes. This will allow a mitochondrial genome-wide comparative analysis, but tremendous sequence variations are expected between cytoplasms from different species. This will undoubtedly complicate the identification of CMS-causative factors, as exemplified in sugar beet and maize (Satoh et al. 2004, 2006; Allen et al. 2007). Other avenues will be needed to obtain a clearer picture regarding the mechanism of CMS and its restoration in cotton.
References
Akagi H, Nakamura A, Sawada R, Oka M, Fujimura T (1995) Genetic diagnosis of cytoplasmic male sterile cybrid plants of rice. Theor Appl Genet 90:948–951
Allen JO, Minx P, Fauron CM, Oddiraju S, Clifton SW, Newton KJ (2006) The complete mitochondrial genomes of five close relatives of maize (Unpublished http://www.ncbi.nlm.nih.gov/genomes/GenomesGroup.cgi?taxid=33090&opt=organelle)
Allen OJ, Fauron CM, Minx P, Roark L, Oddiraju S, Lin NG, Meyer L, Sun H, Kim K, Wang C, Du F, Xu D, Gibson M, Cifrese J, Clifon WS, Newton KJ (2007) Comparison among two fertile and three male sterile mitochondrial genomes of maize. Genetics 177:1173–1192
Bonhomme S, Budar F, Ferault M, Pelletier G (1991) A 2.5 Kb NocI fragment of Ogura radish mitochondrial DNA is correlated with cytoplasmic male sterility in Brassica cybrids. Curr Genet 19:21–127
Bonhomme S, Budar F, Lancelin D, Small I, Defrance MC, Pelletier G (1992) Sequence and transcript analysis of the Nco2.5 Ogura-specific fragment correlated with cytoplasmic male sterility in Brassica cybrids. Mol Gen Genet 235:340–348
Bourland FM (1996) Registration of ‘H1330’ cotton. Crop Sci 36:813
Braun JC, Levings SC (1985) Nucleotide sequence of the F1-ATPase alpha subunit gene from maize mitochondria. Plant Physiol 79:571–577
Brown GG (1999) Unique aspects of cytoplasmic male sterility and fertility restoration in Brassica napus. J Hered 90:351–356
Budar F, Pelletier G (2001) Male sterility in plants: occurrence, determinism, significance and usage. C R Acad Sci III 324:543–550
Chase CD (2006) Cytoplasmic male sterility: a window to the world of plant mitochondrial-nuclear interactions. Trends Genet 23:81–90
Chaw S, Shih AC, Wang D, Wu Y, Liu S, Chou T (2008) The mitochondrial genome of the gymnosperm Cycas taitungensis contains a novel family of short interspersed elements, Bpu sequences, and abundant RNA editing sites. Mol Biol Evol 25:603–615
Clifton SW, Minx P, Fauron CM, Gibson M, Allen JO, Sun H, Thompson M, Barbazuk WB, Kanuganti S, Tayloe C, Meyer L, Wilson RK, Newton KJ (2004) Sequence and comparative analysis of the maize NB mitochondrial genome. Plant Physiol 136:3486–3503
Cook CG, Namken LM (1995) Registration of B411R, B416R and B418R parental lines of cotton. Crop Sci 35:1518
Dewey RE, Levings CS III, Timothy DH (1986) Novel recombinations in the maize mitochondrial genome produce a unique transcriptional unit in the Texas male-sterile cytoplasm. Cell 44:439–449
Galau GA, Wilkins TA (1989) Alloplasmic male sterility in AD allotetraploid Gossypium hirsutum upon replacement of its resident A cytoplasm with that of D species G. harknessii. Theor Appl Genet 78:23–30
Grelon M, Budar F, Bonhomme S, Pelletier G (1994) Ogura cytoplasmic male-sterility (CMS)-associated orf138 is translated into a mitochondrial membrane polypeptide in male-sterile Brassica cybrids. Mol Gen Genet 243:540–547
Handa H (2003) The complete nucleotide sequence and RNA editing content of the mitochondrial genome of rape seed (Brassica napus L.): comparative analysis of the mitochondrial genomes of rape seed and Arabidopsis thaliana. Nucleic Acids Res 31:5907–5916
Hanson MR, Bentolila S (2004) Interactions of mitochondrial and nuclear genes that affect male gametophyte development. Plant Cell 16:S15–S169
Hsu CL, Mullin BC (1989) Physical characterization of mitochondrial DNA from cotton. Plant Mol Biol 13:467–468
Ibrahim RIS, Azuma J, Sakamoto M (2006) Complete nucleotide sequence of the cotton (Gossypium barbadense L.) chloroplast genome with a comparative analysis of sequences among 9 dicot plants. Genes Genet Syst 81:311–321
Johns C, Lu M, Lyznik A, Mackenzie S (1992) A mitochondrial DNA sequence is associated with abnormal pollen development in cytoplasmic male sterile bean plants. Plant Cell 4:435–449
Kim DH, Kang JG, Kim BD (2007) Isolation and characterization of the cytoplasmic male sterility associated orf456 gene of chili pepper (Capsicum annuum L.). Plant Mol Biol 63:519–532
Kubo T, Nishizawa S, Sugawara A, Itchoda N, Estiati A, Mikami T (2000) The complete nucleotide sequence of the mitochondrial genome of sugar beet (Beta vulgaris L.) reveals a novel gene for tRNACys(GCA). Nucleic Acids Res 28:2571–2576
Lee SB, Kaittanis C, Jansen RK, Hostetler JB, Tallon LJ, Town CD, Daniell H (2006) The complete chloroplast genome sequence of Gossypium hirsutum: organization and phylogenetic relationships to other angiosperms. BMC Genomics 7:61
Mackenzie SA, Chase CD (1990) Fertility restoration is associated with loss of a portion of the mitochondrial genome in cytoplasmic male sterile common bean. Plant Cell 2:905–912
Makaroff CA, Apel IJ, Palmer JD (1990) Characterization of radish mitochondrial atpA: influence of nuclear background on transcription of atpA-associated sequences and relationship with male sterility. Plant Mol Biol 15:735–746
Meyer VG (1975) Male sterility from Gossypium harknessii. J Hered 66:23–27
Moneger F, Smart CJ, Leaver CJ (1994) Nuclear restoration of cytoplasmic male-sterility in sunflower is associated with the tissue specific regulation of a novel mitochondrial gene. EMBO J 13:8–17
Notsu Y, Masood S, Nishikawa T, Kubo N, Akiduki G, Nakazono M, Hirai A, Kadowaki K (2002) The complete sequence of the rice (Oryza sativa L.) mitochondrial genome: frequent DNA sequence acquisition and loss during the evolution of flowering plants. Mol Genet Genomics 268:434–445
Ogihara Y, Yamazaki Y, Murai K, Kanno A, Terachi T, Shiina T, Miyashita N, Nasuda S, Nakamura C, Mori N, Takumi S, Murata M, Futo S, Tsunewaki K (2005) Structural dynamics of cereal mitochondrial genomes as revealed by complete nucleotide sequencing of the wheat mitochondrial genome. Nucleic Acids Res 33:6235–6250
Paterson AH, Brubaker CL, Wendel JF (1993) A rapid method for extraction of cotton (Gossypium spp.) genomic DNA suitable for RFLP and PCR analysis. Plant Mol Biol Rep 11:122–127
Rodriguez-Uribe L, O’Connell MA (2006) A root-specific bZIP transcription factor is responsive to water deficit stress in tepary bean (Phaseolus acutifolius) and common bean (P. vulgaris). J Exp Bot 57:1391–1398
Sadoch Z, Majewska-Sawka A, Jazdzewska E, Niklas A (2000) Changes in sugar beet mitochondrial DNA induced during callus stage. Plant Breed 119:107–110
Satoh M, Kubo T, Nishizawa S, Estiati A, Itchoda N, Mikami T (2004) The cytoplasmic male-sterile type and normal type mitochondrial genomes of sugar beet share the same complement of genes of known function but differ in the content of expressed ORFs. Mol Genet Genomics 272:247–256
Satoh M, Kubo T, Mikami T (2006) The Owen mitochondrial genome in sugar beet (Beta vulgaris L.): possible mechanisms of extensive rearrangements and the origin of the mitotype-unique regions. Theor Appl Genet 113:477–484
Schnable PS, Wise RP (1998) The molecular basis of cytoplasmic male sterility and fertility restoration. Trends Plant Sci 3:175–180
Singh M, Brown GG (1991) Suppression of cytoplasmic male sterility by nuclear genes alters expression of a novel mitochondrial gene region. Plant Cell 3:1349–1362
Small RL, Wendel JF (1999) The mitochondrial genome of allotetraploid cotton (Gossypium L.). J Hered 90:251–253
Stern BD, Dyer TA, Lonsdale DM (1982) Organization of the mitochondrial ribosomal RNA genes of maize. Nucleic Acids Res 10:3333–3340
Stewart JMcD (1992) A new cytoplasmic male sterility and restorer from cotton. Proceedings of Beltwide Cotton Conferences, National Cotton Council, Memphis, TN, USA, 610 pp
Sugiyama Y, Watase Y, Nagase M, Makita N, Yagura S, Hirai A, Sugiura M (2005) The complete nucleotide sequence and multipartite organization of the tobacco mitochondrial genome: comparative analysis of mitochondrial genomes in higher plants. Mol Genet Genomics 272:603–615
Tang HV, Pring DR, Shaw LC, Salazar RA, Muza FR, Yan B, Schertz KF (1996) Transcript processing internal to a mitochondrial open reading frame is correlated with fertility restoration in male-sterile sorghum. Plant J 10:123–133
Tian X, Zhang J, Hu S, Yu J (2006) The rice mitochondrial genomes and their variations. Plant Physiol 140:401–410
Unseld M, Marrienfeld JR, Brandt P, Brennicke A (1997) The mitochondrial genome of Arabidopsis thaliana contains 57 genes 366,924 nucleotides. Nat Genet 15:57–61
Wang F (2008) Molecular analysis of cytoplasmic male sterility (CMS) and fertility restoration in cotton. Ph.D. dissertation, New Mexico State University, Las Cruces, NM, USA
Wendel JF (1989) New World tetraploid cottons contain Old World cytoplasm. Proc Natl Acad Sci USA 86:4132–4136
Wendel JF, Albert VA (1992) Phylogenetics of the cotton genus (Gossypium): character-state weighted parsimony analysis of chloroplast-DNA restriction site data and its systematic and biogeographic implications. Syst Bot 17:115–143
Yamamoto MP, Kubo T, Mikami T (2005) The 5′-leader sequence of sugar beet mitochondrial atp6 encodes a novel polypeptide that is characteristic of Owen cytoplasmic male sterility. Mol Genet Genomics 273:342–349
Young EG, Hanson MR (1987) A fused mitochondrial gene associated with cytoplasmic male sterility is developmentally regulated. Cell 50:41–49
Zhang JF, Stewart JMcD (2000) Economic and rapid method for extracting cotton genomic DNA. J Cotton Sci 4:193–201
Zhang JF, Stewart JMcD (2001a) CMS-D8 restoration in cotton is conditioned by one dominant gene. Crop Sci 41:283–288
Zhang JF, Stewart JMcD (2001b) Inheritance and genetic relationships of the D8 and D2-2 restorer genes for cotton cytoplasmical male sterility. Crop Sci 41:289–294
Zhang JF, Mara-koosham G, Lu Y, Stewart JMcD (2004) Molecular analysis of mitochondrial genome in two cytoplasmic male sterile system of cotton. Proceedings of Beltwide Cotton Conferences, San Antonio, TX, USA, pp 1178–1182
Zhang H, Li S, Yi P, Wan C, Chen Z, Zhu Y (2007) A Honglian CMS line of rice displays aberrant F(0) of F(0)F(1)-ATPase. Plant Cell Rep 26:1065–1071
Zhang JF, Turley RB, Stewart JMcD (2008) Comparative analysis of gene expression between CMS-D8 restored plants and normal non-restoring fertile plants in cotton by differential display. Plant Cell Rep 27:553–561
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, F., Feng, C., O’Connell, M.A. et al. RFLP analysis of mitochondrial DNA in two cytoplasmic male sterility systems (CMS-D2 and CMS-D8) of cotton. Euphytica 172, 93–99 (2010). https://doi.org/10.1007/s10681-009-0055-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-009-0055-9