Abstract
Bruchid, Callosobruchus spp. (Coleoptera: Bruchidae), is a serious pest during storage of seeds of mungbean (Vigna radiata (L.) Wilczek) and other Vigna species. A source of resistance to this pest has been identified in Vigna sublobata (Roxb.) Bairig. accession TC1966. Two hundred recombinant inbred lines at the F12 generation have been developed for molecular mapping of bruchid resistance (Br) gene in TC1966. Through bulked segregant analysis (BSA), ten randomly amplified polymorphic DNA (RAPD) markers associated with the bruchid resistance gene were successfully identified. A total of four closely linked RAPDs were cloned and transformed into sequence characterized amplified region (SCAR) and cleaved amplified polymorphism (CAP) markers. Seven CAPs developed from the identified RAPD markers showed tighter linkage with the Br gene than the original RAPD. Through transformation of RAPDs into CAPs, codominant markers for bruchid resistance were successfully obtained. Homozygous genotypes of these PCR-based markers were estimated to contribute 85% of the variance for seed damage when the insect assay was performed under favorable growth conditions for bruchid.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bruchid beetle or bean weevil, Callosobruchus spp. (Coleoptera: Bruchidae), causes serious damage to several leguminous crops including cultivated Vigna species, such as mungbean (V. radiata [L.] Wilczek), azuki bean (V. angularis [Willd.] Ohwi & Ohasi) and cowpea (V. unguiculata [L.] Walp.) during storage (Fernandez and Talekar 1990). Control of this pest by fumigants or other insecticides is not practical because of the primitive nature of seed-storage facilities in most developing countries (where most of munbean and cowpea are produced), the small volume of the seeds and the frequent utilization of the seeds for human consumption (Talekar 1996). Complete resistance against bruchid has been reported in V. sublobata (Roxb.) Bairig. (Fujii and Miyazaki 1987; AVRDC 1991), V. umbellata (Thunb.) Ohwi & Ohashi, and V. mungo (L.) Hepper (Talekar 1996; Tomooka et al. 2000). Resistance could be mediated through hairiness of the pod or antibiosis (Talekar 1996). TC1966, a strain of V. sublobata, showed complete resistance to four species of bruchids, C. chinensis (L.), C. maculatus (Fabricius), C. phaseoli (Gyllenhal) and Zabrotes subfasciatus (Boheman) (Fuji et al. 1989). As no cross incompatibility exhibits between mungbean and V. sublobata, the resistance factor in TC1966 was regarded as a valuable source for mungbean resistance breeding (Fujii and Miyazaki 1987; Talekar 1988; Tomooka et al. 1992) and might provide useful clues for legume pest management in the future. Inheritance of resistance against azuki bean weevil (C. chinensis ) in TC1966 was suggested to be under the control of a single, dominant major gene with some modifying factors (Kitamura et al. 1988). Nevertheless, the exact genetic control and mechanism for resistance remained to be explored.
Mapping of the bruchid resistance (Br) gene in TC1966 has been conducted by restriction fragment length polymorphism (RFLP) analysis (Young et al. 1992). Two markers that mapped approximately 3.6 and 6.7 centimorgan (cM) from the gene were identified. Unfortunately, these markers were not satisfactory when applied at the advanced mungbean generation (Chen et al. unpublished data). Testing bean bug (Riptortus clavatus) within a segregating BC20F2 population, Kaga and Ishimoto (1998) identified eight random amplified polymorphic DNA (RAPD) linked with bean bug resistance. These fragments were subsequently used as RFLP probes and closely linked RFLP markers were reported. RFLP markers are regarded as codominant markers, which facilitate the distinction of homozygous and heterozygous individuals for breeders. Better reproducibility is also found in RFLPs than in RAPDs. However, the RFLP analysis is technically complicated and time consuming and not feasible for high throughput analysis (Powell et al. 1996).
Insecticidal factors against bruchids have been reported and reviewed in common bean (Phaseolus vulgaris L.) (Fabre et al. 1998; Paes et al. 2000), cowpea (V. unguiculata (L.) Walp.) (Macedo et al. 1993) and other legumes (Grossi-de-Sá and Chrispeels 1997; Sales et al. 2000; Carlini and Grossi-de-Sá 2003). The two chemical factors, vignatic acid (Sugawara et al. 1996; Kaga and Ishimoto 1998) and VrCRP (cysteine-rich protein of the plant defensin family) (Chen et al. 2002), have also been isolated from TC1966 and its progenies. Both compounds were not confirmed as the antibiotic factors directly responsible for bruchid resistance in TC1966.
In addition to bruchid infestation, mungbean yellow mosaic virus (MYMV) is a serious disease in South Asia. To generate PCR-based molecular markers for bruchid and MYMV resistance and to pyramid both traits in mungbean breeding, 200 recombinant inbred lines (RILs) at the F12 generation were developed by crossing of MYMV-resistant cultivated mungbean NM92 (AVRDC 2002) with TC1966. Recombinant inbred populations are characteristically a group of segregating pure lines that can be reproduced indefinitely. Thus, this allows repeated phenotyping for different traits and accumulation of useful markers. Here we describe the development of PCR-based markers for bruchid resistance in mungbean by use of a recombinant inbred population. Polymorphism was first explored by the RAPD technique through bulked segregant analysis approach (Michelmore et al. 1991). The associated RAPDs were then cloned and sequenced for further SCAR (sequence characterized amplified region) and CAP (cleaved amplified polymeric region) analyses. In this study, we identified codominant CAP markers for bruchid resistance, which explained up to 85% of the variation for bruchid damage in the tested population.
Materials and methods
Plant materials and DNA extraction
A recombinant inbred (RI) population of 200 lines was generated from an inter-subspecific cross between bruchid susceptible (but MYMV resistant) mungbean variety NM92 and a bruchid resistant subspecies accession TC1966. The RI population was generated as follows: F1 were selfed for two generations to generate 200 F3 lines. Two hundred RILs were developed from these F3 plants by single seed descent for 9 generations to F12. The DNA of these RILs was extracted by a modified CTAB procedure described by Saghai Maroof et al. (1984).
Assay for bruchid resistance
A bruchid (C. chinensis) population was maintained on a susceptible mungbean variety VC1973A at the AVRDC insectary. Forty seeds of each RIL, parents, and resistant and susceptible checks were placed in individual 50 ml Erlenmeyer flasks. Twenty newly emerged bruchid adults were placed into each flask, for mating and laying eggs on the seeds. Seven days after infestation (DAI), all adults were removed and each seed was checked to make sure each had more than two eggs. The flasks were kept in a compartment room of the insectary. The samples were examined at 30 DAI to record the number of bruchid that had emerged and percentage of seed damage. The bruchid assay was repeated three times, in February 2002, May 2003 and July 2005. Mean maximum/minimum temperatures during the bruchid assays were 26.0°/20.5°C, 28.0°/24.5°C and 28.5°/25.0°C for February 2002, May 2003 and July 2005 respectively. In the July 2005 assay, adult emergence was recorded three times a week from 21 DAI to 40 DAI.
Bulked segregant approach and RAPD analysis
After the February 2002 assay, DNA of RILs that showed 0% and 80–100% bruchid infestation were pooled. Two resistant bulks (22 RILs each of 0% damage) and two susceptible bulks (20 RILs of 80–90% damage and 18 RILs of 90–100% damage, respectively) were formed. For the RAPD analysis, 10-mer sets of random primer were purchased from OPERON Inc. (CA, USA) and the University of British Columbia, Nucleic Acid-Protein Service Unit (Canada). Primers were first screened with parental DNA for successful PCR amplification and polymorphic bands. Primers that generated clear and multiple bands were further used to test the bulked samples. The PCR regime used was 94°C for 3 min, followed by 3 cycles of 94°C for 1 min, 40°C 1 min, 72°C 2 min linked to the second program with 40 cycles of 94°C for 30 s, 40°C 30 s, 72°C 1 min and a final extension at 72°C for 10 min. Random primers that showed polymorphism between resistant and susceptible bulks were further analyzed in the 200 RILs.
CAP analysis
RAPD fragments highly correlated with bruchid resistance were then cloned into the pDrive cloning vector using QIAGEN PCR Cloning kit (QIAGEN Inc.). The inserts of each clone were sequenced using an ABI373 DNA Sequencer. Primer sets were designed based on both ends of the obtained sequences. PCR was carried out in a volume of 25 μl containing 50 ng of total genomic DNA, 5 pmoles of each primer, 0.6 U Taq DNA polymerase, 0.1 mM of each dNTP, 3 mM MgCl2, and 1× PCR buffer. PCR was processed with the following program: 1 cycle of 94°C for 3 min, 35 cycles of 94°C for 1 min, 55°C (depend on pretest results with gradient thermocycler) 1 min, 72°C 2 min and a final extension at 72°C for 10 min. The amplified products from the two parental DNAs were digested with different restriction enzymes to determine polymorphism within the insert. The identified CAPs were also tested for all 200 RILs.
Sequence analysis and marker linkage analysis
Sequence homologies of cloned RAPDs and SCARs were analyzed and compared at NCBI (http://www.ncbi.nlm.nih.gov/BLAST), using both blastn and blastx procedures. Associations between markers and traits were determined by QGENE program using single point regression analysis (Nelson 1997). Based on this analysis, coefficients of statistics (R 2, SE, genotype mean, etc.) were reported for all significant markers. Only marker-phenotype associations with a probability threshold of P < 0.001 were considered. Linkage analysis was performed using the software Mapmaker v. 3.0b (Lander et al. 1987) at a log likelihood 5.0 with a maximum Haldane distance of 50 cM. To verify the order of markers obtained by three-point analysis, the Ripple command was used at the window-size of 5 and log likelihood threshold of 6.0.
Results
Bruchid reaction among 200 RILs
Genetically, F12 RILs are deemed to be a set of highly homogeneous pure lines. However, the results of bruchid bioassays over 3 years were not consistent (Table 1). The results of bruchid assays in May 2003 and July 2005 were comparable, where as the February 2002 assay’s results deviated from the aforementioned two assays. Lower temperatures may not be favorable for growth of bruchid thus less damages was observed in the February 2002 assay. Interestingly, however, about one fourth of the total RILs consistently expressed complete resistance (0% damage) in all three assays (Table 1). More than half of the 200 RILs expressed complete susceptibility (100% damage) in May 2003 and July 2005, but only 18 lines showed similar susceptibility in the February 2002 assay. Damage differences between the 2002/2003 and 2005 assays could be attributed to temperature effect on infestation ability of bruchid. Low temperatures in the February 2002 assay could be a negative factor for larva emergence and growth hence resulting in a longer period for adult emergence (i.e., >30 DAI) and therefore less seed damage was recorded. Given the unstable factors frequently encountered for insect bioassays, marker-assisted selection, which is environment-independent, could circumvent this hurdle.
Bulked segregant and RAPD analyses
More than 600 primers were pre-tested for effective RAPD amplification in tested materials. In total, 360 primers were selected and RAPD tested with four bulked samples and two parents. Ten out of 360 primers produced polymorphic bands ranging from 700 to 1800 bp. Some primers (e.g., OPV02) produced unique bands in the two resistant bulks and resistant parent, whereas some primers (e.g., UBC223) produced a unique band in the two susceptible bulks and the susceptible parent, in addition, others (e.g., OPW02) produced polymorphisms between resistant and susceptible bulks but not between resistant and susceptible parents (Fig. 1).
Bulked segregant analysis for bruchid resistance using random decamers. OPV02 (a), UBC223 (b) and OPW02 (c). M: molecular weight marker; Lane 1: resistant bulk I; Lane 2: susceptible bulk I; Lane3: resistant bulk II; Lane 4: susceptible bulk II; Lane 5: resistant parent TC1966; Lane 6: susceptible parent NM92. Arrow indicates polymorphism between resistant and susceptible bulks
Ten polymorphic primers (i.e., UBC66, UBC168, UBC223, UBC313, UBC353, OPM04, OPU11, OPV02, OPW02 and OPW13) were further tested on a total of 200 RILs. Single marker regression analysis was processed using genotype and phenotype data to investigate their association using the QGene program. Four RAPDs with significant correlations were produced by OPW02, UBC223, OPU11 and OPV02 primers. The phenotypic correlation (R 2 value) explained by the markers were 0.72 for OPW02a4, 0.6 for OPU11a4, 0.55 for UBC223a7 and 0.5 for OPV02a3 when using data collected in 2005.
Development for SCAR and CAP markers
OPW02a4, UBC223a7, OPU11a4 and OPV02a3, were successfully cloned and sequenced. Sets of primers were designed and tested for SCAR analysis. Both OPU11a4for/rev and OPV02a3for/rev primers produced dominant bands for the resistant parent. These two SCARs were successfully reproduced in segregating population. Both UBC223a7 and OPW02a4 for/rev primers produced monomorphic bands between the two parents.
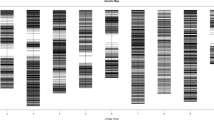
Further digestion with various restriction enzymes revealed that the UBC223a7 fragment generated a polymorphic band in the resistant parent when digested with HaeIII enzyme, however this polymorphism was irreproducible within the segregating population. In addition to the original 1.5 kb RAPD region, OPW02a4 for/rev primers also produced other bands located between 0.25 kb and 1.7 kb in size. This implied that multiple regions were tagged by these two primers. Although no clear polymorphism was observed in the SCAR profile, there was higher band intensity around the 1.5 kb region for the resistant parent than for susceptible parent. Further digestion of the OPW02a4 for/rev SCAR reaction with different restriction enzymes revealed many cleaved amplified polymorphisms (CAP) between resistant and susceptible parents, i.e., HaeIII (7 CAPs, W02s1-W02s7), MspI (3 CAPs, W02s8-W02s10) and HinfI (2 CAPs, W02s11-W02s12). With MspI digestion, two monomorphic bands were shown for both parents, and three polymorphic bands (i.e., 1.7, 0.6 and 0.48 kb) observed in the resistant parent. The 1.7 kb band was not related with the bruchid reaction as determined by genetic progeny testing. Among these CAPs, three of them revealed allelic polymorphism between R and S parents. Digestion with HaeIII enzyme produced two bands at about 600 and 900 bp for resistant parent and one bands at 1100 bp for susceptible parent. Testing among 200 RILs revealed that each RIL either produced two bands from resistant parent or one band from susceptible parent (Fig. 2). These bands can be used as codominant marker when making selection in the breeding program. According to the aforementioned results, it is suggested that OPW02a4 is the most informative RAPD for Br gene of TC1966. The complete sequence of OPW02a4 fragment and related RAPD and CAP primers are shown in Fig. 3.
CAP analysis for bruchid resistance gene using OPW02a4 for/rev primers followed by HaeIII enzyme digestion. Three bands (two from R parent and one from S parent) revealed allelic polymorphism between two parents when tested across 200 RILs. S: susceptible parent NM92; R: resistant parent TC1966; M: molecular weight marker. Arrows at left and right sides indicate polymorphic bands for susceptible and resistant parents respectively
Linkage and sequence analysis
Results of QGene analysis with the three bruchid assays revealed seven most closely linked CAPs and four RAPDs (Table 2). Among the four RAPDs, segregation of OPW02a4 (126:74) and OPV02a3 (116:84) skewed to the susceptible parent NM92. The segregation of UBC 223a7 (100:96) and OPU11a4 (104:94) did not deviate statistically from the expected ratio of 1:1 (P > 0.7). CAPs derived from OPW02a4 also skewed to the susceptible parent.
The R 2 value for each marker varied among three assays. The assay in the February 2002 assay had least association while the July 2005 assay showed the highest association for all listed markers. Nevertheless, these markers consistently showed high correlation with bruchid resistance comparing with other markers in the three assays. For OPW02a4 locus, mean bruchid damage was 90% for individuals that were homozygous for NM92 allele and 9% for individuals that were homozygous for TC1966 allele (Table 2). Six CAPs, such as W02s02, W02s03, W02s04, W02s09, W02s10 and W02s11, derived from RAPD OPW02a4 are more closely linked to Br gene (R 2 = 0.85) compared to the original RAPD (Table 2). Genetic mapping of RAPDs, SCARs and CAPs with MAPMAKER/EXP 3.0b also showed the same result (Fig. 4). The SCAR markers of OPU11a4, OPV02a3 and UBC223a7 were not able to generate more informative markers for this experiment. With low stringency PCR conditions, RAPD was not always reproducible to amplify all RILs. However, using a more stringent temperature regime for SCAR analysis, all samples were successfully amplified and the corresponding CAPs were identified.
Linkage mapping for RAPD and CAP markers with the Br gene. The linked markers included 3 RAPDs (UBC223a7, OPV02a3 and OPW02a4), 7 CAPs derived from OPW02a4 (W02s2, W02s3, W02s4, W02s6, W02s9, W02s10 and W02s11) and 6 Amplified fragment length polymorphism (AFLP) markers (m5pca382, m1pgg258, mg3pag431, mg3pag425, m1pgg256 and m5pca598)
As differences in SCAR intensity and CAPs were observed by OPW02a4for/rev primers, sequences of these SCAR products derived from resistant and susceptible lines might provide more inside information. To improve resolution, the SCAR products were electrophoresised on 1.3% agarose gel for longer period. The result indicated only one 1.45 kb band for the susceptible parent NM92. However, two bands (about 1.45 and 1.6 kb) were generated specifically for resistant parent TC1966 and the tested resistant lines. SCAR sequence of this band (1470 bp) of NM92 is the same as the sequence of the cloned RAPD (Fig. 3). Further comparison of these sequences suggested high homology of the 1470 bp band between resistant parent and resistant RIL (98%), the homology between resistant RIL and susceptible parent is 93% (data not shown). The CAPs were basically derived from this 1470 bp region. Sequence of this fragment did not show high homology with any reported gene on NCBI web site. However, comparison of the translated query with the protein database (blastx) suggested high correlation with RNA-directed DNA polymerase and gypsy/Ty-3 retroelement. Sequence alignment of the 1.6 kb fragment by blastn suggested high homology with chloroplast complete genome of soybean and other legumes. Alignment by blastx correlated this 1.6 kb fragment to NADH dehydrogenase subunit. Whether these regions play a role in plant defense to insects remains further study.
Discussion
Each RIL material used in this study, after selfing for 12 generation, were uniform in terms of agronomic phenotype in the field. However, their reactions to bruchid infestation obtained in three assays were temperature-dependent. In general, temperatures of 25–30°C are regarded as suitable for growth of bruchid pest (Dongre et al. 1996; Lambrides and Imrie 2000; Jackai and Asante 2003). During the February 2002 assay, mean max/min temperature (26.0°/20.5°C) fell below the range of favorable growth conditions for bruchid, consequently lowered seed damage. Correlation analysis confirmed that the same set of markers was linked to resistance for all three assays. However, these markers expressed their highest R 2 values and genetic effects in the July 2005 assays followed by the May 2003 assay then the February 2002 assay. In the July 2005 assay, mean bruchid damage for mungbean lines carrying homozygous susceptible and resistant alleles were 95% and 9%, respectively (Table 2), while those of the February 2002 assay were 62% and 18%, respectively. Even under environmental conditions unfavorable for bruchid growth and development, the results suggested that these markers were closely linked with the major gene responsible for bruchid resistance. Since 28–30°C is commonly used for bruchid assay, it is reasonable that the developed markers show the highest correlation with data collected in the July 2005 assay when the max/min temperature was 28.5°/25.0°C. The resistance of TC1966 was reported to be under the control of a single dominant allele together with modifier factors (Kitamura et al. 1988). Theoretically, one half of the materials are expected to carry the homozygous R allele, however, only about one fourth expressed stable resistance to the pest insect. The current assay with 200 RILs suggests that expression of bruchid resistance gene could be modified by some environmental factors. Sequences of the polymorphic markers were associated with putative plant disease resistance polyprotein and a retrotransposon protein by blastx. The occurrence of susceptible and resistant progenies from parents whose genotypes were estimated as homozygous R/R and homozygous r/r has been observed occasionally in our previous populations derived from different susceptible parents (VC3890A and VC1973A, our laboratory data) as well as reported by Kitamura et al. (1988). The presence of this retroelement-related gene may partially explain the reason behind high SNP frequency within the marker region and might be a potential reason for variations in bruchid resistance. It is not clear whether retro-element-related genes or NADH dehydrogenase subunit in the resistant line play a role on insect resistance or plant defense mechanism. This remains to be further explored.
In this study, bulked segregant analysis was successfully employed to identify molecular markers linked to a major gene for bruchid resistance. Several of the RAPDs, though not closely linked to Br, were allocated to the same linkage group with markers associated with the Br gene. This confirms the effectiveness of the bulked segregant approach to identify polymorphism associated with the trait of interested, especially when this is governed by the simple Mendelian inheritance. Conversion of RAPD markers into SCAR or CAP markers has been reported and applied to improve the low reproducibility of RAPD (Paran and Michelmore 1993). Our study also suggests the possibility of exploring codominant markers through transfer of RAPD into CAP. Besides C. chinensis, TC1966 is also known to be resistant to other bruchids, e.g., C. maculates, C. phaseolus, C. analis and Z. subfasciatus (Fuji et al. 1989). It would be interesting to investigate the correlation of the newly developed markers with the resistance to other Callosobruchus spp. using the obtained RILs. This will also help clarify whether single or more chemical compounds are responsible for resistance to different storage pests in TC1966.
References
AVRDC (1992) 1991 Progress Report. Asian Vegetable Research and Development Center. Shanhua, Tainan, Taiwan, pp 41–43
AVRDC (2002) AVRDC report 2001. Asian Vegetable Research and Development Center. Shanhua, Tainan, Taiwan, 25 pp
Carlini CR, Grossi-de-Sá MF (2003) Plant toxic proteins with insecticidal properties. A review on their potentialities as bioinsecticides. Toxicon 40:1515–1539
Chen KC, Lin CY, Kuan CC, Sung HY, Chen CS (2002) A novel defensin encoded by a mungbean cDNA exhibits insecticidal activity against bruchid. J Agr Food Chem 50:7258–7263
Dongre TK, Pawar SE, Thakare RG, Harwalkar MR (1996) Identification of resistant sources to cowpea weevil (Callosobruchus maculatus (F.)) in Vigna sp. and inheritance of their resistance in black gram (vigna mungo var. mungo). J Stored Prod Res 32:201–204
Fabre C, Causse H, Mourey L, Koninkw J, Rivière M, Hendriks H, Puzo G, Samama JP, Rougè P (1998) Characterization and sugar binding properties of arcelin-1, an insecticidal lectin-like protein isolated from kidney bean (Phaseolus vulgaris L. cv. RAZ-2) seeds. Biochem J 329:551–560
Fernandez GCJ, Talekar NS (1990) Genetics and breeding for bruchid resistance in Asiatic Vigna species. In: Fujii K, Gatehouse AMR, Johnson CD, Mitchell R, Yoshida T (eds) Bruchids and legumes: economics, ecology and coevolution. Kluwer Academic Pub., Dordrecht, Nethelands, pp 209–217
Fuji K, Ishimoto M, Kitamura K (1989) Patterns of resistance to bean weevils (Bruchidae) in Vigna radiata-mungo sublobata complex inform the breeding of new resistant varieties. Appl Ent Zool 24:126–132
Fujii I, Miyazaki S (1987) Infestation resistance of wild legumes (Vigna sublobata) to azuki bean weevil, Callosobruchus chinensis (L.) (Coleoptera: Bruchidae) and its relationship with cytogenetic classification. Appl Ent Zool 22:229–230
Grossi-de-Sá MF, Chrispeels MJ (1997) Molecular cloning of Bruchid (Zabrotes subfasciatus) α-amylase cDNA and interactions of the expressed enzyme with bean amylase inhibitors. Insect Biochem Mol Biol 27:271–281
Jackai LEN, Asante SK (2003) A case for the standardization of protocols used in screening cowpea, Vigna unguiculata for resistance to Callosobruchus maculatus (Fabricius) (Coleoptera: Bruchidae). J Stored Prod Res 39:251–263
Kaga A, Ishimoto M (1998) Genetic localization of a bruchid resistance gene and its relationship to insecticidal cyclopeptide alkaloids, the vignatic acids in mungbean (V. radiata L. Wilczek). Mol Gen Genet 258:378–384
Kitamura K, Ishimoto M, Sawa M (1988) Inheritance of resistance to infestation with azuki bean weevil in Vigna sublobata and successful incorporation to V. radiata. Japan J Breed 38:459–464
Lambrides CJ, Imrie BC (2000) Susceptibility of mungbean varieties to the Bruchid species Callosobruchus maculatus (F.), C. phaseoli (GyII.), C. chinensis (L.), and Acanthoscelides obtectus (Say.) (Coleoptera:Chrysomelidae). Aust J Agric Res 51:85–89
Lander ES, Green P, Abrahamson J, Barlow A, Daly JM, Lincoln SE, Newberg L (1987) Mapmaker: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Macedo MLR, Andrade LBS, Morates RA, Xavier-Filho J (1993) Vicilin variants and the resistance of cowpea (Vigna unguiculata) seeds to the cowpea weevil (Callosobruchus maculates). Comp Biochem Physiol 105C:89–94
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci USA 88:9828–9832
Nelson JC (1997) QGENE: software for marker-based genomic analysis and breeding. Mol Breed 3:239–245
Paes NS, Gerhardt IR, Coutinho MV, Yokoyama M, Santana E, Harris N, Chrispeels MJ, Grossi-de-Sá MF (2000) The effect of arcelin-1 on the structure of the midgut of Bruchid larvae and immunolocalozation of the arcelin protein. J Insect Phys 46:393–402
Paran I, Michelmore RW (1993) Development of reliable PCR-based markers linked to downy mildew resistance gene in lettuce. Theor Appl Genet 85:985–993
Powell W, Morgante M, Andre C, Hanafey M, Vogel J, Tingey S, Rafalski A (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238
Saghai Maroof MA, Soliman KM, Jorgenson R, Allard RW (1984) Ribosomal DNA spacer length polymorphism in barley: Mendelian inheritance, chromosomal locations and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Sales MP, Gerhardt IR, Grossi-de-Sá MF, Xavier-Filho J (2000) Do legume storage proteins play a role in defending seeds against bruchids? Plant Phys 124:515–522
Sugawara F, Ishimoto M, Le-Van N, Koshino H, Uzawa J, Yoshida S, Kitamura K (1996) Insecticidal peptide from mungbean: a resistant factor against infestation with azuki bean weevil. J Agr Food Chem 44:3360–3364
Talekar NS (1988) Biology, damage and control of bruchid pests of mungbean. In: Shanmugasundaram S, McLean BT (eds) Mungbean. Proc 2nd Intl Sym on Mungbean. AVRDC, Taiwan, pp 329–342
Talekar NS (1996) Sources of resistance to major insect pests of mungbean in Asia. In: Asthana AN, Kim DH (eds) Recent advances in mungbean research. Indian Soc Pulse Res. IIPR, Kanpur 208 024, India, pp 40–51
Tomooka N, Kashiwaba K, Vaughan DA, Ishimoto M, Egawa Y (2000) The effectiveness of evaluating wild species: searching for sources of resistance to bruchid beetles in the genus Vigna subgenus Ceratotropis. Euphytica 115:27–41
Tomooka N, Lairungreang C, Nakeeraks P, Egawa Y, Thavarasook C (1992) Development of bruchid resistant mungbean line using wild mungbean germplasm in Thailand. Plant Breed 109:60–66
Young ND, Kumar L, Menancio-Hautea D, Talekar NS, Shanmugasundarum S, Kim DH (1992) RFLP mapping of a major bruchid resistance gene in mungbean (Vigna radiata L. Wilczek). Theor Appl Genet 84:839–844
Acknowledgments
This work was partially supported by a grant from the Council of Agriculture Taiwan during 2003–2005. The authors thank Dr. N. S. Talekar for technical support on bruchid assays. We also thank Dr. Anne Frary and Dr. Sami Doganlar for critical reading and helpful suggestions of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chen, HM., Liu, CA., George Kuo, C. et al. Development of a molecular marker for a bruchid (Callosobruchus chinensis L.) resistance gene in mungbean. Euphytica 157, 113–122 (2007). https://doi.org/10.1007/s10681-007-9400-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-007-9400-z