Abstract
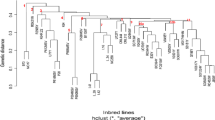
Establishment of the best combination among heterotic groups, heterotic patterns, is crucial to the development of successful maize (Zea Mays L.) hybrids. The use of molecular markers in maize-breeding programs might or might not increase the efficiency of heterosis prediction by classifying diverse inbred lines into heterotic groups. The objectives of present research were to classify elite North Dakota (ND) maize inbred lines into heterotic groups and evaluate the consistency between simple sequence repeat (SSR) grouping and testcross data. Thirteen ND inbred lines representing diverse genetic background were crossed in a diallel mating design in 2000. The crosses and 12 checks were evaluated across four ND environments in 2001 and 2002. In addition, these lines were crossed to commercial inbred testers representing known heterotic groups in 2002. Hybrids between public and private lines were evaluated across three ND environments in 2003. Inbred lines representing Lancaster Sure Crop, Iowa Stiff Stalk Synthetic (BSSS), Minnesota #13, Northwestern Dent, Golden Glow pedigrees and ND inbred lines were screened with 49 SSR markers. Inbred lines ND246, ND278, ND280, ND281, ND282 and ND284 were clustered within the BSSS heterotic group. Inbreds ND277, ND285, ND286, ND290, and ND291 grouped closer to the Lancaster Sure Crop heterotic group. Inbred lines ND257 and ND288 grouped within Minnesota #13. Data from ND278 and ND290 testcrosses showed good combining ability with testers representing more than one heterotic group. Our research shows that groups of genetically similar germplasm could not be identified accurately and reliably with molecular markers even when the available germplasm was diverse contrary what has been suggested. Therefore, extensive field evaluation is recommended to classify unrelated inbred lines of maize.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bernardo R (2002) Breeding for quantitative traits in plants. Stemma Press, Woodbury, MN
Bernardo RJ, Romero-Severson J, Ziegle J, Hauser J, Joe L, Hookstra G, Doerge RW (2000) Parental contribution and coefficient of co-ancestry among maize inbreds: pedigree, RFLP, and SSR data. Theor Appl Genet 100:552–556
Carena MJ (2005) Maize commercial hybrids compared to improved population hybrids for grain yield and agronomic performance. Euphytica 141:201–208
Carena MJ, Hallauer AR (2001) Expression of heterosis in Leaming and Midland Corn Belt dent populations. Jour Iowa Acad Sci 108:73–78
Carena MJ, Wanner DW (2003) Registration of ND2000 inbred line of maize. Crop Sci 43:1568–1569
Comstock RE, Robinson HF (1948) The components of genetic variance in populations of bi-parental progenies and their use in estimating the average degree of dominance. Biometrics 4:254–266
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–302
Falconer DS, Mackay TFC (1996) Introduction to quantitative genetics. 4th ed. Longman Group, Ltd., Essex, England
Fulton TM, Chunwongse J, Tanksley SD (1995) Micropep protocol for extraction of DNA from tomato and other herbaceous plants. Plant Mol Bio Rep 13:207–209
Gethi JG, Labate JA, Lamkey KR, Smith ME, Kresovich S (2002) SSR variation in important U.S. maize inbred lines. Crop Sci 42:951–957
Godshalk EB, Lee M, Lamkey KR (1990) Relationship of restriction fragment length polymorphisms to single cross hybrid performance of maize. Theor Appl Genet 80:273–280
Griffing B (1956) Concept of general and specific combing ability in relation to diallel crossing systems. Aust J Biol Sci 9:463–493
Hallauer AR (1999) Temperate maize and heterosis. In: Coors JG, Pandey S (ed.) The Genetics and Exploitation of Heterosis in Crops. ASA, CSSA & SSSA, Madison, WI, pp 353–361
Hallauer AR, Russel WA, Lamkey KR (1988) Corn breeding. In: Sprague GF, Dudley JW (ed.) Corn and corn improvement, 3rd ed. Agron. Monogr. 18. ASA, CSSA & SSSA, Madison, WI
Hallauer AR, Miranda JB (1988) Quantitative genetics in maize breeding (2nd ed.) Iowa State University Press, Ames, IA
Hoisington D, Khairallah M, Reeves T, Ribaut JM, Skovmand B, Taba S, Warburton M (1999) Plant genetic resources: What can they contribute toward increase in crop productivity? Proc. Natl Acad Sci USA 96:5937–5943
Jaccard P (1908) Nouvelles researches sur la distribuition florale. Bull Soc Vaudoise Sci Natl 44:223–270
Labate JA, Lamkey KR, Mitchell SE, Kresovich S, Sullivan H, Smith JSC (2003) Molecular and historical aspects of corn belt dent diversity. Crop Sci 43:80–91
Liu K, Goodman MM, Muse S, Smith JSC, Buckler E, Doebley J (2003) Genetic structure and diversity among maize inbred lines as inferred from DNA microsatellites. Genetics 165:2117–2128
Lu H, Bernardo R (2001) Molecular marker diversity among current and historical maize inbreds. Theor Appl Genet 103:613–617
Melchinger AE (1999) Genetic diversity and heterosis. In: Coors JG, Pandey S (ed) The genetics and exploitation of heterosis in crops. ASA, CSSA & SSSA, Madison, WI
Moll RH, Lonnquist JH, Velez Fortuno J, Johnson C (1965) The relationship of heterosis and genetic divergence in maize. Genetics 52:139–144
Mumm RH, Dudley JW (1994) A classification of 148 U.S. maize inbreds: I. Cluster analysis based on RFLPs. Crop Sci 37:617–624
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York, NY
Nei M, Li WH (1979) Mathematical model of studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273
Olson PJ, Walster HL, Hopper TH (1927) Corn for North Dakota. ND Agr Exp Stn Bull 207
Pejic I, Ajmone-Marsan P, Morgante M, Kozumplick V, Castiglioni P, Taramino G, Motto M (1998) Comparative analysis of genetic similarity among maize inbred lines detected by RFLPs, RAPDs, SSRs and AFLPs. Theor Appl Genet 97:1248–1255
Richey FD (1931) Experiments on hybrid vigor and convergent improvement in corn. USDA Tec Bull 267
Rohlf FJ (1998) Numerical and Multivariate Analysis System (NTSYSpc version 2.0). Dep. of Ecology and Evolution, State Univ. of New York, Stony Brook, NY
Romero-Severson J, Smith JSC, Ziegle J, Hauser J, Joe L, Hookstra G (2001) Pedigree analysis and haplotype sharing within diverse groups of Zea mays L. inbreds. Theor Appl Genet 103:567–574
SAS (1989) SAS/STAT User's Guide. (4th ed). SAS Inst., Cary, NC
Senior ML, Murphy JP, Goodman MM, Stuber CW (1998) Utility of SSRs for determining genetic similarities and relationship in maize using agarose gel system. Crop Sci 38:1088–1098
Smith JSC, Chin ECL, Shu H, Smith OS, Wall SJ, Senior ML, Mitchell SE, Kresovich S, Ziegle J (1997) An evaluation of the utility of SSR loci as molecular markers in maize (Zea mays L.): comparisons with data from RFLPs and pedigree. Theor Appl Genet 95:163–173
Sprague GF, Tatum LA (1941) General vs. specific combining ability in single crosses of corn. J Am Soc Agron 34:923–932
Tautz D (1989) Hypervariability of simple sequences as a general source of polymorphic DNA markers. Nuclei Acid Res 17:6463–6471
Tivang JG, Nienhuis J, Smith OS (1994) Estimation of sampling variance of molecular-marker data using bootstrap procedure. Theor Appl Genet 89:259–264
Troyer AF (1999) Background of U.S. hybrid corn. Crop Sci 39:601–626
Troyer AF (2004) Background of U.S. hybrid corn II: breeding, climate, and food. Crop Sci 44:370–380
Wang D, Shi J, Carlson SR, Cregan PB, Ward RW, Diers BW (2003) A low-cost, high-throughput polyacrylamide gel electrophoresis system for genotyping with microsatellite DNA markers. Crop Sci 43:1828–1832
Warburton ML, Xianchun X, Crossa J, Franco J, Melchinger AE, Frisch M, Bohn M, Hoisington D (2002) Genetic characterization of CIMMYT inbred lines and open pollinated populations using large scale fingerprinting methods. Crop. Sci 42:1832–1840
Yu J, Lu H, Bernardo R (2001) Inconsistency between SSR groupings and genetic background of white corn inbreds. Maydica 46:133–139
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Barata, C., Carena, M.J. Classification of North Dakota maize inbred lines into heterotic groups based on molecular and testcross data. Euphytica 151, 339–349 (2006). https://doi.org/10.1007/s10681-006-9155-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-006-9155-y