Abstract
Twelve bacteriophage isolates of Erwinia amylovora, the causal agent of fire blight, were isolated from blighted apple, pear and quince trees from different sites in Hungary. According to morphological characteristics they were assigned to the order Caudovirales, two isolates belonging to the Podoviridae and ten to the Myoviridae families. Examining plaque morphology, host range and molecular characterization by PCR established that these phages are not identical neither to the three North American strains used as references nor the earlier isolated Hungarian Siphoviridae strains. Studying the efficacy of selected phages in apple blossoms and green pear fruit slices it was found that a combination of three phage isolates (ΦEaH2A, ΦEaH5K and ΦEaH7B) significantly reduced bacterial multiplication and fire blight symptoms as compared to untreated controls. Combined application of these new E. amylovora-specific phages as biocontrol agents may contribute to a better control of E. amylovora under field conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fire blight is the most devastating bacterial disease of Rosaceae plants (van der Zwet and Keil 1979; van der Zwet and Beer 1991). It is caused by the phytopathogenic bacterium Erwinia amylovora (Burrill) Winslow et al. (1920), inducing huge economic losses in pome fruits (van der Zwet and Keil 1979; Vanneste 2000). Currently disease control is challenging since use of the most effective pesticide, the antibiotic streptomycin applied on open blossoms has been limited due to human and plant health concerns.
Several reviews have previously been published, highlighting the possibilities and limitations of phage therapy in plant disease control (Gill and Abedon 2003; Jones et al. 2007; Balogh et al. 2010; Nagy et al. 2012; Doffkay et al. 2015). Till now bacteriophages (i.e. the viruses of bacteria) have been found to be effective for the control of several phytobacteria including xanthomonads (Civerolo and Keil 1969; Saccardi et al. 1993; Flaherty et al. 2000, 2001; McNeil et al. 2001; Balogh et al. 2003, 2008, 2010; Obradovic et al. 2004; Lang et al. 2007; Iriarte et al. 2012; Dömötör et al. 2016), pseudomonads (Munsch and Olivier 1995; Rombouts et al. 2016), Ralstonia solanacearum (Tanaka et al. 1990), Streptomyces scabies (McKenna et al. 2001) and Pectobacterium carotovorum (Ravensdale et al. 2007). However, the probability that bacteria mutate and become resistant to individual phages could be a real concern when considering the application of phages as biological control agents. This concern arose already in the 1930s (Katznelson 1937) and was expressed later in review articles by Okabe and Goto (1963) and Vidaver (1976) as a major limiting factor for the use of phage infections to control phytopathogenic bacteria. In the 1980s a strategy was developed to address the problem of phage-resistance in natural mutants of bacteria. It was found that by preparing mixtures of wild type and host range mutant phages (h-mutants), bacteria resistant to the original, wild type parent phages are also lysed (Jackson 1989). Therefore, commercially used phage preparations usually include two or more different phage strains to avoid development of phage resistance in bacteria and confer a broader host-range.
A number of different E. amylovora-phages have been isolated, characterized and tested for their biocontrol efficacy worldwide (Billing 1960; Okabe and Goto 1963; Civerolo 1972; Erskine 1973; Ritchie and Klos 1977; Schnabel et al. 1999; Kim and Geider 2000; Schnabel and Jones 2001; Gill et al. 2003; Kim et al. 2004; Svircev et al. 2006; Lehman 2007; Müller et al. 2011a; Schwarczinger et al. 2011; Boulé et al. 2011; Nagy et al. 2012, 2015; Roach et al. 2013; Born et al. 2014, 2015; Samoilova and Leclerque 2014). Moreover complete genomes of some of these phages have become available (Lehman et al. 2009; Müller et al. 2011b; Born et al. 2011; Yagubi et al. 2014; Lagonenko et al. 2015). On the other hand, in Hungary only two E. amylovora-specific phage species – both belonging to the Siphoviridae family – have been characterized so far (Dömötör et al. 2012; Meczker et al. 2014). These two phages are the biocontrol agents of the biopesticide ERWIPHAGE FORTE that has been available in Hungary since 2012 and seems to have a promising protective effect against fire blight (http://biotechnologia.enviroinvest.hu/).
In order to identify other bacteriophage isolates that may broaden the spectrum of potential biocontrol agents against fire blight of pome fruits we aimed to isolate and characterize additional E. amylovora-specific phages from Hungary and to study the efficacy of the phage treatments on E. amylovora.
Materials and methods
Bacterial strains, reference bacteriophage strains
A list of bacterial strains used in this work is listed in Supplemental Table S1. Strains of Erwinia, Tatumella and Pantoea spp. were cultured on Luria-Bertani agar (LBA) or broth (LB) (Difco) and incubated at 28 °C, except for P. agglomerans MB96 and P. stewartii ssp. stewartii that were grown on casamino-acid peptone glucose (CPG) media. For culturing a spontaneous streptomycin mutant strain of E. amylovora (Ea 1/79Sm) LB broth and LBA media supplemented with 500 mg L−1 streptomycin–sulphate (Duchefa Biochemie) were used. Strains of Pseudomonas spp., Rhizobium radiobacter, Allorhizobium vitis and Xanthomonas campestris pv. zinniae were grown on nutrient agar (NA) and incubated at 28 °C. Escherichia coli was grown on LBA and incubated at 37 °C. Strains were multiplied in the appropriate liquid broth with constant agitation, and were stored at −70 °C supplemented with 15% (v/v) glycerol. The pathogenicity of Ea1/79Sm and Ea1/79 was established using the plant hypersensitive reaction (HR) test in tobacco leaves (Klement 1963). Three phage strains; ΦEa1(h) (Ritchie and Klos 1979), ΦEa100 and ΦEa116C (Schnabel and Jones 2001) originally isolated in the USA were used as reference strains (Supplemental Table S1).
Isolation of phages
Bacteriophage isolates of Erwinia amylovora were collected from various sites in Hungary between 2006 and 2007. Phages were isolated from aerial parts of apple, pear and quince trees exhibiting fire blight symptoms. Three bacterial host strains (Ea12, Ea18, Ea26) were used in the initial isolation and enrichment process in LB. Phage enrichment and isolation have been made according to procedures of Crosse and Hingorani (1958), modified by Gill et al. (2003). Phage detection, purification and titre assessment were conducted with spot tests and Adams’ double agar overlay method (Adams 1959). Phages were diluted and stored in SM buffer with gelatine (100 mM NaCl, 8 mM MgSO4∙7H2O, 50 mM Tris-HCl (pH 7.5), 0.002% (w/v) gelatine) at 4 °C, or supplemented with 15% (v/v) glycerol for long term storage at −70 °C.
Plaque morphology
Bacteriophage isolates were distributed on LBA top agar layers supplemented with 1% (w/v) sucrose and containing the test bacterium (E. amylovora EaCFBP1430 strain) according to Adams’ double agar overlay method (Adams 1959). Following incubation for one day at 28 °C phage isolates were visually characterized based on plaque size and width of halos surrounding the plaques.
Virion morphology
Samples containing purified phage lysates were assayed by a Morgagni 268D type transmission electron microscope (TEM) following a negative staining procedure according to Gill et al. (2003).
Host range tests
Phages were tested for lysis efficacy on bacterial species and strains belonging to the genera Erwinia, Pantoea, Pseudomonas, Tatumella, Rhizobium, Escherichia and Xanthomonas (Supplemental Table S1). Susceptibility of test bacteria (108 CFU mL−1) to phages (106 PFU mL−1) was determined by Adams’ spot test (Adams 1959).
PCR assay
DNA extraction for PCR was carried out by adding 50 μL 2X sodium-azide (NaN3) solution (2% Triton X-100, 0.5% NaN3, 0.1 M Tris-HCl buffer, pH 8.0) to 50 μL of fresh phage lysate originating from a single plaque. The mixture was incubated at 99 °C for 10 min, cooled down, and then centrifuged at 13500 rpm for 10 min at 4 °C. The supernatants transferred to new tubes served as DNA templates.
During PCR assays, nine sets of primer pairs specific to characteristic E. amylovora phage DNA sequences were used (Supplemental Table S2). The first two primer pairs are specific for genes coding holin (Bläsi and Young 1996) and lysozyme (Kim et al. 2004) enzymes from ΦEa1(h), respectively. The next primer pairs are specific for given regions of terminase (Black 1995), peptidase and tape measure protein coding genes from ΦEa116C. PEa1A/B primers target a sequence of ΦEa1(h) encoding a portion of HNH endonuclease and a hypothetic protein (HNH endonuclease-like) (Gill et al. 2003; Müller et al. 2011a) The Ea100-F/R primer pair was designed for a given region (10337–10,662 bp) of ΦEa100 encoding a portion of HNH DNase and a hypothetic protein (HNH Dnase-like). The 1hcap-F/R primer pair is specific for the capsid-encoding gene of ΦEa1(h) while H2cap-F/R primers are specific for the capsid-encoding sequence of the phage ΦEaH2 (Dömötör et al. 2012). PCR assays were carried out (final volume of 18 μl) with 1 μl of template DNA/fresh phage lysate, 9 μL of Thermo Fisher Scientific (Waltham, MA, USA) 2X PCR Master Mix (0.05 U/μL Taq DNA polymerase, reaction buffer, 4 mM MgCl2, and 4 mM of each dNTP) and 4–4 μl (2.5 pmol μL−1) of each primer. The reaction mixtures were incubated in an MJ Research PTC-200 Peltier Thermal Cycler (GMI, Ramsey, MN, USA). PCR was carried out by using the MM2 or MM3 programs (Supplemental Table S2). Ten microlitres of each amplification mixture was electrophoresed on 1% agarose (Invitrogen) gels prepared in 1X TAE buffer and pre-casted with GelRed 10,000X (Biotium, Fremont, CA, USA) solution in water.
DNA sequencing and sequence analysis
Our investigations have focused on direct sequence analysis of partial nucleotide sequences of two phage isolates (ΦEaH2A, ΦEaH5K). In case of ΦEaH2A, DNA fragments for sequencing were PCR-amplified with primers specific for the genes encoding peptidase, tape measure protein and terminase, while in case of ΦEaH5K, primers specific for tape measure protein and terminase genes were used (Supplemental Table S2). 50 μl of each PCR product was cleaned by using the PCR-M Clean Up System (Viogene, New Taipei City, Taiwan) Kit according to the manufacturers’ protocol. Nucleotide sequences were determined by Eurofins Genomics (Ebersberg, Germany). Sequences obtained by automated DNA sequencing were analysed and compared to homologous nucleotide sequences in international databases (http://www.ncbi.nlm.nih.gov/nucleotide). The DNA sequences were also analysed with the sequence analysis program BioEdit Biological Sequence Alignment Editor (www.mbio.ncsu.edu/bioedit.html). Database searches were performed on the Internet with the nucleotide BLAST program of NCBI (National Center for Biotechnology Information) (http://blast.ncbi.nlm.nih.gov/Blast.cgi).
Flower assay
A combination of three selected phages (ΦH2A, ΦH5K and ΦH7B) was tested for their capability to reduce bacterial numbers in apple flowers similarly as described by Müller et al. (2011a). Four apple cultivars differentially susceptible to fire blight (Malus x domestica Borkh. ‘Idared’, ‘Golden Delicious Reinders’, ‘Gala Schniga’, ‘Pinova’) and one quince (Cydonia oblonga Mill.) cultivar (‘Berecki’) were used as test plants. Flowers were collected in the balloon phenophase and placed individually into small glass vials filled with 1% (w/v) sucrose. Within 12 h, flowers opened and 20 μL of a 1:1 mixture of phage lysate (1010 PFU mL−1) and bacterial suspension (Ea1/79Sm, 105 CFU mL−1) was pipetted onto pistils (MOI = 105). To obtain concentrated, fresh phage lysates the plate lysing method was applied as described earlier (Nagy et al. 2015). The flowers (15 flowers /treatment in two replications) were incubated in a climate chamber at a relative humidity of 80% (16 h/8 h day/night cycles at 24 °C/21 °C). After 4 days petals and stems of the flowers were removed, and the flowers were incubated in one mL sterile distilled water for 10 min and bacterial cells were extracted by centrifugation (3 min, 13,500 rpm). From each extract, 50 μL of a 10,000 x dilution was plated on LBA medium plates containing 500 mg L−1 streptomycin-sulphate and 50 mg L−1 cycloheximid. Results were evaluated following incubation for 2 days in the dark at 28 °C based on bacterial colony numbers.
Pear slice assay
Effects of the phage combination (ΦH2A + ΦH5K + ΦH7B) were tested on unripe fruit slices of two pear cultivars (Pyrus x communis L.’Conference’, ‘Jules Guyot Dr.’). The 0.5 cm thick pear slices (six slices/treatment) have been placed into glass Petri dishes and soaked in either of the following solutions: 10 mL phage suspension (1010 PFU mL−1), water or 100 mg L−1 streptomycin-sulphate. Both sides of the slices were soaked for 5 min each. Afterwards, briefly dried slices were inoculated with 10 μL (105 CFU mL−1) of E. amylovora wild type strain (Ea1/79) according to Müller et al. (2011a) (MOI = 105) and incubated in close Petri dishes at 28 °C in the dark for 4 days. Pears were rated for symptoms according to a bonitation scale from 0 to 6 as following: (0) symptomless; (1) browning of the middle part of slices, around the inoculation site, with mucus; enhanced mucus production accompanied by browning of (2) 1/8 of the slice; (3) 1/4 of the slice; (4) 1/2 of the slice; (5) 3/4 of the slice; (6) the whole slice. To ensure impartiality and avoid errors arising from bias a blind experiment was employed.
Results
Phage isolation
Twelve phage isolates were characterized. Eight phages (ΦEaH1A, ΦEaH11, ΦEaH2A ΦEaH2B, ΦEaH5B, ΦEaH5K, ΦEaH4A, ΦEaH4B) were isolated from blighted quince trees, two (ΦEaH7A and ΦEaH7B) from apple shoots and two (ΦEaH9B and ΦEaH12B) from pear shoots (Supplemental Table S3).
Plaque morphology
Our phage isolates formed plaques of different sizes, with a diameter of 0.5–7.1 mm on the soft agar layer containing the test bacterium (Table 1, Supplemental Fig. S1). The halo around plaques – when present – had a diameter between 0.1–5.0 mm. The smallest plaques were formed by ΦEaH5K (0.5–0.7 mm), being smaller (including halos) than plaques of ΦEa116C (Supplemental Table S4, Supplemental Fig. S1). ΦH7B had one of the largest plaques with a much broader halo than those of ΦEa100, a reference strain giving the largest plaques in our assays (Supplemental Table S4, Supplemental Fig. S1). This indicates a higher lytic activity of ΦH7B.
Virion morphology
The E. amylovora phages studied were assigned to the order Caudovirales (morphotypes C1 and A1), to the Podoviridae and Myoviridae families (Ackermann 2007) (Fig. 1, Table 1 and Supplemental Table S4). Among these, the smallest is ΦH11, smaller than phages from the USA (Müller et al. 2011a, Supplemental Table S4). The largest is ΦH4A having a larger size than reference phages (Müller et al. 2011a, Table 1 and Supplemental Table S4). Isolates assigned to Podoviridae have a head diameter of 60 nm, while those belonging to Myoviridae have a head diameter of ca. 70 nm. Our phage isolates markedly differ from the two previously described Hungarian isolates belonging to Siphoviridae [ΦEaH1 (Meczker et al. 2014), ΦEaH2 (Dömötör et al. 2012)].
PCR assays
Based on PCR assays isolates were separated into two main groups (I, II) and subdivided into five subgroups (A-E) (Table 2). Phages assigned to the first group (ΦEaH5B, ΦEaH4B, ΦEaH4A, ΦEaH9B) were positive for holin, lysozyme, terminase, peptidase and HNH endonuclease-like genes. Phages in the second group did not give a PCR product by the primers used for genes encoding holin and lysozyme, but were positive for terminase and tape measure protein sequences, similarly as reference strain ΦEa116C.
DNA sequencing
The partial regions coding for peptidase in ΦEaH2A, and for terminase and phage tail tape measure protein in ΦEaH2A and ΦEaH5K display a high similarity with the corresponding sequences of E. amylovora phages: 99% with vB_EamM-M7 (Born et al. 2011), and 85% with ΦEa21–4 (Lehman et al. 2009) and ΦEa104 (Müller et al. 2011b). Partial nucleotide sequences of the genes encoding peptidase, tape measure protein and terminase were deposited in the NCBI nucleotide sequence database (GenBank accession numbers: KT881239, KT881240, KT881241, KT881242, KT907049).
Host range analysis
Bacterial susceptibility was characterized based on purity of plaques in the upper agar layer containing indicator bacteria (Table 3). The host ranges of phages were determined by the ability to form plaques on test bacteria. Clear plaques indicated high host sensitivity, turbid plaques indicated partial lysis, and no plaques indicated a nonhost (Roach et al. 2013). Out of the 12 studied phage isolates ΦEaH2A, ΦEaH2B, ΦEaH4A, ΦEaH7A, and ΦEaH7B, have lysed most tested bacterial strains, while - among the reference phage strains - ΦEa116C had the broadest host range. The tested phage isolates were capable of lysing not only Hungarian E. amylovora strains but also those derived from other geographical areas. On the other hand all of the Hungarian E. amylovora isolates were susceptible to all of the phages tested, irrespective of their origin. Phage sensitivity profile of the bacterium EaDS05 isolated in Germany from quince is nearly the same as of the Hungarian E. amylovora strains. However, four E. amylovora strains (EaOR1/07, Ea 1/79, Ea1/79del100, EaDS02) are susceptible to only ca. half of the tested phages. Among other Erwinia species E. billingiae Eb661 was the most susceptible to the tested phages. On the other hand, Erwinia tasmaniensis was resistant to all tested phages. This result is similar to those published by Müller et al. (2011a). Pantoea species, closely related to E. amylovora, displayed phage sensitivity profiles similar to those of the most phage-susceptible E. amylovora strains, except P. vagans C9–1. In line with our results, Gill et al. (2003) and Lehman (2007) also found that certain tested Pantoea agglomerans strains were susceptible to E. amylovora phages, some of which belong to the Podoviridae family. In fact, Adriaenssens et al. (2011) isolated two additional Podoviridae phages that are able to form clear plaques on Pantoea agglomerans strains, however, either cannot lyse E. amylovora, or form only turbid plaques on this bacterium.
Selection of phages for biocontrol tests
Three phages of the Myoviridae family (ΦEaH2A, ΦEaH5K and ΦEaH7B) were selected for biocontrol tests, based on plaque morphology, host range and activity towards E. amylovora in liquid culture. We found that ΦEaH5K produces the smallest, while ΦEaH7B the largest plaques. Host range tests revealed that ΦEaH2A and ΦEaH7B have a broad, almost identical host range, although ΦEaH2A can produce clear plaques on a higher number of bacterial strains. ΦEaH5K, however, has a narrower host range, not being able to lyse all tested E. amylovora strains. The selected phages could significantly decrease optical density values in liquid cultures of E. amylovora (Ea1/79Sm) at the end of the incubation period (24 h), similarly as described earlier by Schnabel and Jones (2001) (data not shown). Considering that phages may have an increased efficacy in combination than alone (Schnabel et al. 1999) and in order to prevent the possible development of phage resistance in bacteria, we decided to use a triple combination of the selected phages (ΦEaH2A, ΦEaH5K and ΦEaH7B) in our in planta assays testing biocontrol efficacy on E. amylovora.
Flower assays
The flower assay is the best method to select the most effective phage candidates for biocontrol, because the main strategy for controlling fire blight with biocontrol agents is preventing the accumulation of E. amylovora populations on nutrient-rich stigmatic surfaces of blossoms in the spring (Thomson 1986; Johnson and Stockwell 1998; Müller et al. 2011a). The triple phage combination reduced multiplication of E. amylovora significantly (by 65–84%) as compared to untreated controls in case of all apple and quince cultivars, although this difference translates to a reduction in bacterial concentrations of only ca. 0.5–1 order of magnitude (Fig. 2a). A correlation between plant susceptibility to fire blight and phage effects was not observed, since the best results were obtained on E. amylovora-resistant apple cv. ‘Pinova’ and susceptible quince cv. ‘Bereczki’ flowers. In case of cv. ‘Pinova’ phage-treatment reduced the recovered pathogen by 84% compared to untreated control flowers, a difference close to 1 order of magnitude. Similar results were shown by Müller et al. (2011a) applying individual phages. Samples recovered from the most susceptible ‘Idared’ flowers were assayed by quantitative PCR as well. Bacterial concentrations were determined by real time qPCR using primers specific for the pEA29 plasmid of E. amylovora (Salm and Geider 2004). The same trend in bacterial multiplication as obtained by colony counting could be observed (data not shown).
Influence of phage combination on multiplication of E. amylovora on apple flowers and on unripe pear slices. Columns show the concentration of re-isolated bacteria (CFU mL-1) on flowers of different apple cultivars (Fig. 2a). Treatments included a combination of three phages (ΦEaH2A, ΦEaH5K and ΦEaH7B) and Ea1/79Sm (MOI = 105), water (without phages) or Ea1/79Sm cells only. No bacteria were detectable from water controls. Re-isolation of bacteria (Ea1/79Sm) was done 4 days after initial treatments. Values are average titres of the re-isolated bacterial suspensions (mean CFU / mL−1 ± SE) from two independent biological experiments (n = 15 / treatment). Figure 2b shows reduction of E. amylovora symptoms by phage treatments on immature pear slices (6 slices/ treatment). Pear slices were treated with the triple phage combination (1010 PFU mL−1) and inoculated with 10 μL (105 CFU mL−1) of the E. amylovora wild type strain Ea1/79. Results are evaluated based on the average severity of symptoms by a predefined bonitation scale (symptom illustration on left side of Fig. 2b). No symptoms were observed on water controls or streptomycin sulphate treated, Ea1/79-inoculated pear slices. Asterisks indicate statistically significant differences between phage-treated and untreated control samples using Student’s t-test at P ≤ 0.01
Pear slice assay
The effect of the phage combination was also tested on unripe pear slices (Fig. 2b). The immature pear slice assay provides a general and useful prediction of antagonist activity on plant surfaces (Wilson et al. 1990). The same method was also used in another study on phage biocontrol effects (Müller et al. 2011a). This experimental approach provides an overview not only on the effect of phages on bacterial symptoms but is also suitable to compare this effect to that of streptomycin. Symptom severity was reduced on both cultivars (‘Conference’, ‘Jules Guyot Dr.’) compared to positive controls. On the other hand, the effect of phages was markedly lower than that conferred by streptomycin sulphate in all experiments, because streptomycin-treated pears remained symptomless. Phage treatments of pear slices were less efficient than that of flowers, similarly as found by Müller et al. (2011a).
Discussion
Bacteriophages from the Podoviridae and Myoviridae families that infect E. amylovora, the bacterium causing fire blight of pome fruits have been isolated in Hungary and are characterized in this study. Their virion morphology is considerably different from the two E. amylovora-phages previously reported from Hungary (Dömötör et al. 2012; Meczker et al. 2014). Some of the studied isolates exhibit similar host ranges to the reference strains (Müller et al. 2011a; this study), while others have an even broader cross-genera infectivity. A broader host range of E. amylovora-phages raises the possibility of alternative biocontrol applications. Phage sensitive Pantoea agglomerans strains (i.e. MB96, NB2) can be potentially used as carrier organisms for the propagation of phages and subsequent delivery to orchards for the control of fire blight, as described earlier (Svircev et al. 2006; Lehman 2007; Boulé et al. 2011). These authors used P agglomerans, an orchard epiphyte, to deliver and sustain phages on the blossom surface. The lytic ability of bacteriophages and the additional biological control activity of P. agglomerans provided effective and stable control of the fire blight pathogen with an efficacy comparable to the antibacterial effect of streptomycin. Importantly, our newly isolated phages might also be useful biocontrol agents against a diverse group of phytopathogenic bacteria including E. persicina, E. rhapontici, P. stewartii ssp. stewartii, T. citrea comb. Nov or P. syringae pv. syringae.
Molecular characterization of phages with PCR revealed that these newly characterized isolates can be classified into two larger groups and five subgroups. Phages from the second group are similar to ΦEa116C. Based on preliminary experiments three phages were selected for biocontrol tests. All three phage isolates belong to the Myoviridae family, ΦEaH5K producing the smallest, while ΦEaH7B the largest plaques. Regarding their host range, ΦEaH2A and ΦEaH7B have a broad, almost identical host range, although ΦEaH2A is capable of producing clear plaques on a higher number of bacterial strains. On the contrary, ΦEaH5K has a narrower host range, not being able to lyse all E. amylovora strains tested. A common feature of these three phages is that all of them proved to be positive for terminase and tape measure protein genes in PCR assays with specific primers. Furthermore, ΦEaH2A and ΦEaH7B also contain the peptidase gene. Sequencing of appropriate gene portions in two of the characterized phage isolates (ΦEaH2A and ΦEaH5K) revealed high (85–99%) similarity with corresponding DNA sequences of vB_EamM-M7 (Born et al. 2011), ΦEa21–4 (Lehman et al. 2009) and ΦEa104 phage strains (Müller et al. 2011b). Testing the biocontrol efficacy of newly isolated phages against E. amylovora on apple blossoms and on green pear fruit slices it was found that a combination of these three phages (ΦEaH2A, ΦEaH5K and ΦEaH7B) effectively limits bacterial multiplication or development of fire blight symptoms, similarly as shown by Müller et al. (2011a).
Phage treatments and E. amylovora inoculations were applied simultaneously implying that phage treatments prior to bacterial exposure of host plants might even enhance phage efficacy. This is suggested by previous studies demonstrating that phage treatments within 24 h before bacterial inoculation can also effectively inhibit E. amylovora or Xanthomonas pruni (Civerolo and Keil 1969; Nagy et al. 2012). It is possible that, under optimal conditions, such phage pre-treatments enhance phage penetration and translocation into plants, providing an improved biocontrol of bacteria like E. amylovora (Rao and Srivastava 1970; Ward and Mahler 1982; Iriarte et al. 2012; Nagy et al. 2015). On the other hand, phage application prior to bacterial exposure, as compared to co-application, could also reduce the efficacy of biocontrol. This could be due to suboptimal conditions (e.g. extreme heat, high UV radiation, drought, etc.) that phages may often encounter on plant surfaces, especially in the field (Balogh et al. 2003; Ishimaru et al. 1988; Nagy et al. 2012).
One of the main hurdles of successfully controlling bacterial diseases with bacteriophages is the appearance of phage-resistant bacterial strains (Okabe and Goto 1963; Vidaver 1976; Schnabel and Jones 2001; Jones et al. 2007; Roach et al. 2008; Doffkay et al. 2015). This disadvantage can be circumvented by the application of phages in combination (Schnabel et al. 1999; Svircev et al. 2010; Nagy et al. 2012). In contrast, observations of Roach et al. (2015) suggest that while lysogeny is possible in E. amylovora, it could be rare or absent in certain natural populations, with a minimal risk of lysogenic conversion (i.e. emergence of phage-resistance in these bacteria) and transduction by Erwinia spp. phages. Furthermore P. agglomerans isolates from different geographical areas also did not show the presence of any prophage, a likely indication of the absence of phage resistance (Roach et al. 2015). Nevertheless, commercially used phage preparations usually include several different phage strains to avoid development of phage resistance in bacteria and confer a broader host-range. For instance, the biopesticide ListShield is an aqueous phage preparation designed to attack a very broad spectrum of Listeria monocytogenes strains containing six Listeria-specific bacteriophages (http://www.intralytix.com), while the biocontrol preparation ERWIPHAGE FORTE contains two E. amylovora-specific phage strains (http://biotechnologia.enviroinvest.hu/). Another important factor that should be considered when planning a phage-based control of fire blight is the dependence of phage host range on the extracellular polysaccharide (EPS) content of E. amylovora. For example, it is known that the virulence of Podoviridae phages depends on high EPS levels (specifically, amylovoran contents) of their host (see e.g. Müller et al. 2011a; Roach et al. 2013). In line with these findings our results show that the Podoviridae phages characterized by us (EaH9B és EaH11) can efficiently lyse a high amylovoran producer strain (Ea1/79Sm) but not bacteria containing low levels of amylovoran or no amylovoran at all (Ea 1/79 and Ea1/79-del 100, respectively). Interestingly, however, we found that the Podoviridae phages investigated could also lyse most of the tested Pantoea species, which are not known to produce amylovoran, results similar to those of Gill et al. (2003) and Lehman (2007). Furthermore, Adriaenssens et al. (2011) isolated two additional Podoviridae phages that are able to form clear plaques on Pantoea agglomerans strains, but not E. amylovora. Therefore, it seems that amylovoran may not be the sole factor determining the phage sensitivity of these bacteria, at least in case of Pantoea spp.
For the above mentioned reasons, new and well characterized phage isolates are desperately needed as alternative sources of phage-based biological control. In the present study we have characterized several newly isolated E. amylovora-specific phages that may serve as potentially effective biocontrol agents for the management of fire blight in pome fruits and contribute to the effort to minimize the emergence of phage-resistant E. amylovora strains.
References
Ackermann, H. W. (2007). 5500 Phages examined in the electron microscope. Archives of Virology, 152(2), 227–243.
Adams, M. H. (1959). Bacteriophages. New York: Interscience Publishers.
Adriaenssens, E. M., Ceyssens, P. J., Dunon, V., Ackermann, H. W., Van Vaerenbergh, J., Maes, M., De Proft, M., & Lavigne, R. (2011). Bacteriophages LIMElight and LIMEzero of Pantoea agglomerans, belonging to the “phiKMV-like viruses”. Applied and Environmental Microbiology, 77(10), 3443–3450.
Balogh, B., Jones, J. B., Momol, M. T., Olson, S. M., Obradovic, A., King, P., & Jackson, L. E. (2003). Improved efficacy of newly formulated bacteriophages for management of bacterial spot on tomato. Plant Disease, 87(8), 949–954.
Balogh, B., Canteros, B. I., Stall, R. E., & Jones, J. B. (2008). Control of citrus canker and citrus bacterial spot with bacteriophages. Plant Disease, 92(7), 1048–1052.
Balogh, B., Jones, J. B., Iriarte, F. B., & Momol, M. T. (2010). Phage therapy for plant disease control. Current Pharmaceutical Biotechnology, 11(1), 48–57.
Billing, E. (1960). An association between capsulation and phage sensitivity in Erwinia amylovora. Nature, 186, 819–820.
Black, L. W. (1995). DNA packaging and cutting by phage terminases: control in phage T4 by a synaptic mechanism. Bioessays, 17(12), 1025–1030.
Bläsi, U., & Young, R. (1996). Two beginnings for a single purpose: the dual - start holins in the regulation of phage lysis. Molecular Microbiology, 21(4), 675–682.
Born, Y., Fieseler, L., Marazzi, J., Lurz, R., Duffy, B., & Loessner, M. J. (2011). Novel virulent and broad-host-range Erwinia amylovora bacteriophages reveal a high degree of mosaicism and a relationship to Enterobacteriaceae phages. Applied and Environmental Microbiology, 77(17), 5945–5954.
Born, Y., Fieseler, L., Klumpp, J., Eugster, M. R., Zurfluh, K., Duffy, B., & Loessner, M. J. (2014). The tail - associated depolymerase of Erwinia amylovora phage L1 mediates host cell adsorption and enzymatic capsule removal, which can enhance infection by other phage. Environmental Microbiology, 16(7), 2168–2180.
Born, Y., Bosshard, L., Duffy, B., Loessner, M. J., & Fieseler, L. (2015). Protection of Erwinia amylovora bacteriophage Y2 from UV-induced damage by natural compounds. Bacteriophage, 5(4), e1074330.
Boulé, J., Sholberg, P. L., Lehman, S. M., O'gorman, D. T., & Svircev, A. M. (2011). Isolation and characterization of eight bacteriophages infecting Erwinia amylovora and their potential as biological control agents in British Columbia, Canada. Canadian Journal of Plant Pathology, 33(3), 308–317.
Civerolo, E. L. (1972). Interaction between bacteria and bacteriophage on plant surfaces and in plant tissues. In H. P. Maas Geesteranus (Ed.), Proc. Of the Third International Conference on Plant Pathogenic Bacteria (pp. 25–37). Wageningen: PUDOC.
Civerolo, E. L., & Keil, H. L. (1969). Inhibition of bacterial spot of peach foliage by Xanthomonas pruni bacteriophage. Phytopathology, 59, 1966–1977.
Crosse, J. E., & Hingorani, M. K. (1958). A method for isolating Pseudomonas mors-prunorum phages from the soil. Nature, 181, 60–61.
Doffkay, Z., Dömötör, D., Kovács, T., & Rákhely, G. (2015). Bacteriophage therapy against plant, animal and human pathogens. Acta Biologica Szegediensis, 59(Suppl. 2), 291–302.
Dömötör, D., Becságh, P., Rákhely, G., Schneider, G., & Kovács, T. (2012). Complete genomic sequence of Erwinia amylovora phage PhiEaH2. Journal of Virology, 86(19), 10899–10899.
Dömötör, D., Frank, T., Rákhely, G., Doffkay, Z., Schneider, G., & Kovács, T. (2016). Comparative analysis of two bacteriophages of Xanthomonas arboricola pv. juglandis. Infection. Genetics and Evolution, 43, 371–377.
Erskine, J. M. (1973). Characteristics of Erwinia amylovora bacteriophage and its possible role in the epidemiology of fire blight. Canadian Journal of Microbiology, 19(7), 837–845.
Flaherty, J. E., Jones, J. B., Harbaugh, B. K., Somodi, G. C., & Jackson, L. E. (2000). Control of bacterial spot on tomato in the greenhouse and field with H-mutant bacteriophages. Horticultural Science, 35(5), 882–884.
Flaherty, J. E., Harbaugh, B. K., Jones, J. B., Somodi, G. C., & Jackson, L. E. (2001). H-mutant bacteriophages as a potential biocontrol of bacterial blight of geranium. Horticultural Science, 36(1), 98–100.
Gill, J., & Abedon, S. T. (2003). Bacteriophage ecology and plants. APSnet Feature. doi:10.1094/APSnetFeature-2003-1103.
Gill, J. J., Svircev, A. M., Smith, R., & Castle, A. J. (2003). Bacteriophages of Erwinia amylovora. Applied and Environmental Microbiology, 69(4), 2133–2138.
Iriarte, F. B., Obradović, A., Wernsing, M. H., Jackson, L. E., Balogh, B., Hong, J. A., et al. (2012). Soil-based systemic delivery and phyllosphere in vivo propagation of bacteriophages: two possible strategies for improving bacteriophage persistence for plant disease control. Bacteriophage, 2(4), 215–224.
Ishimaru, C. A., Klos, E. J., & Brubaker, R. R. (1988). Multiple antibiotic production by Erwinia herbicola. Phytopathology, 78(6), 746–750.
Jackson, L. E. (1989). Bacteriophage prevention and control of harmful plant bacteria. US Patent No.4828999. 1989.
Johnson, K. B., & Stockwell, V. O. (1998). Management of fire blight: a case study in microbial ecology. Annual Review of Phytopathology, 36(1), 227–248.
Jones, J. B., Jackson, L. E., Balogh, B., Obradovic, A., Iriarte, F. B., & Momol, M. T. (2007). Bacteriophages for Plant Disease Control. Annual Review of Phytopathology, 45, 245–262.
Katznelson, H. (1937). Bacteriophage in relation to plant diseases. Botanical Review, 3, 499–421. doi:10.1007/BF02870486.
Kim, W. S., & Geider, K. (2000). Characterization of a viral EPS-depolymerase. a potential tool for control of fire blight. Phytopathology, 90, 1263–1268.
Kim, W. S., Salm, H., & Geider, K. (2004). Expression of bacteriophage ϕEa1h lysozyme in Escherichia coli and its activity in growth inhibition of Erwinia amylovora. Microbiology, 150(8), 2707–2714.
Klement, Z. (1963). Method for the rapid detection of the pathogenicity of phytopathogenic Pseudomonas. Nature, 199, 299–300.
Lagonenko, A. L., Sadovskaya, O., Valentovich, L. N., & Evtushenkov, A. N. (2015). Characterization of a new Vil-like Erwinia amylovora bacteriophage phiEa2809. FEMS Microbiology Letters, 362(7), fnv031. doi:10.1093/femsle/fnv031.
Lang, J. M., Gent, D. H., & Schwartz, H. F. (2007). Management of Xanthomonas leaf blight of onion with bacteriophages and a plant activator. Plant Disease, 91, 871–878.
Lehman, S. M. (2007). Development of a bacteriophage-based biopesticide for fire blight. Dissertation, Brock University. St. Catharines. Ontario.
Lehman, S. M., Kropinski, A. M., Castle, A. J., & Svircev, A. M. (2009). Complete genome of the broad-host-range Erwinia amylovora phage ΦEa21–4 and its relationship to Salmonella phage Felix O1. Applied and Environmental Microbiology, 75(7), 2139–2147.
McKenna, F., El-Tarabily, K. A., Hardy, G. S., & Dell, B. (2001). Novel in vivo use of a polyvalent Streptomyces phage to disinfest Streptomyces scabies-infected seed potatoes. Plant Pathology, 50(6), 666–675.
McNeil, D. L., Romero, S., Kandula, J., Stark, C., Stewart, A., & Larsen, S. (2001). Bacteriophages: a potential biocontrol agent against walnut blight (Xanthomonas campestris pv juglandis). In Proc. of the New Zealand Plant Protection Conference (pp. 220–224). 1998. New Zealand: Plant Protection Society.
Meczker, K., Dömötör, D., Vass, J., Rákhely, G., Schneider, G., & Kovács, T. (2014). The genome of the Erwinia amylovora phage PhiEaH1 reveals greater diversity and broadens the applicability of phages for the treatment of fire blight. FEMS Microbiology Letters, 350(1), 25–27.
Müller, I., Lurz, R., Kube, M., Quedenau, C., Jelkmann, W., & Geider, K. (2011a). Molecular and physiological properties of bacteriophages from North America and Germany affecting the fire blight pathogen Erwinia amylovora. Microbial Biotechnology, 4(6), 735–745.
Müller, I., Kube, M., Reinhardt, R., Jelkmann, W., & Geider, K. (2011b). Complete genome sequences of three Erwinia amylovora phages isolated in North America and a bacteriophage induced from an Erwinia tasmaniensis strain. Journal of Bacteriology, 193(3), 795–796.
Munsch, P., & Olivier, J. M. (1995). Biocontrol of bacterial blotch of the cultivated mushroom with lytic phages: some practical considerations. Mushroom Science, 14, 595–602.
Nagy, J. K., Király, L., & Schwarczinger, I. (2012). Phage therapy for plant disease control with a focus on fire blight. Central European Journal of Biology, 7(1), 1–12.
Nagy, J. K., Schwarczinger, I., Künstler, A., Pogány, M., & Király, L. (2015). Penetration and translocation of Erwinia amylovora-specific bacteriophages in apple-a possibility of enhanced control of fire blight. European Journal of Plant Pathology, 142(4), 815–827.
Obradovic, A., Jones, J. B., Momol, M. T., Balogh, B., & Olson, S. M. (2004). Management of tomato bacterial spot in the field by foliar applications of bacteriophages and SAR inducers. Plant Disease, 88(7), 736–740.
Okabe, N. O. R. I. O., & Goto, M. A. S. A. O. (1963). Bacteriophages of plant pathogens. Annual Review of Phytopathology, 1(1), 397–418.
Rao, Y. P., & Srivastava, D. N. (1970). Application of phages in investigation of epidemiology of bacterial blight disease of rice. Proceedings of the Indian National Science Academy, 37, 314–321.
Ravensdale, M., Blom, T. J., Gracia-Garza, J. A., Svircev, A. M., & Smith, R. J. (2007). Bacteriophages and the control of Erwinia carotovora subsp. carotovora. Canadian Journal of Plant Pathology, 29(2), 121–130.
Ritchie, D. F., & Klos, E. J. (1977). Isolation of Erwinia amylovora bacteriophage from aerial parts of apple trees. Phytopathology, 67, 101–104.
Ritchie, D. F., & Klos, E. J. (1979). Some properties of Erwinia amylovora bacteriophages. Phytopathology, 69, 1078–1083.
Roach, D. R., Castle, A. J., Svircev, A. M., & Tumini, F. A. (2008). Phage-based biopesticides characterization of phage resistance and host range for sustainability. Acta Horticulturae, 793, 397–401.
Roach, D. R., Sjaarda, D. R., Castle, A. J., & Svircev, A. M. (2013). Host exopolysaccharide quantity and composition impact Erwinia amylovora bacteriophage pathogenesis. Applied and Environmental Microbiology, 79(10), 3249–3256.
Roach, D. R., Sjaarda, D. R., Sjaarda, C. P., Ayala, C. J., Howcroft, B., Castle, A. J., & Svircev, A. M. (2015). Absence of lysogeny in wild populations of Erwinia amylovora and Pantoea agglomerans. Microbial Biotechnology, 8(3), 510–518. doi:10.1111/1751–7915.12253.
Rombouts, S., Volckaert, A., Venneman, S., Declercq, B., Vandenheuvel, D., Allonsius, C. N., et al. (2016). Characterization of novel bacteriophages for biocontrol of bacterial blight in leek caused by Pseudomonas syringae pv. porri. Frontiers in Microbiology, 7, 279–294. doi:10.3389/fmicb.2016.00279.
Saccardi, A., Gambin, E., Zaccardelli, M., Barone, G., & Mazzucchi, U. (1993). Xanthomonas campestris pv. pruni control trials with phage treatments on peaches in the orchard. Phytopathologia Mediterranea, 32(3), 206–210.
Salm, H., & Geider, K. (2004). Real-time PCR for detection and quantification of Erwinia amylovora, the causal agent of fireblight. Plant Pathology, 53(5), 602–610.
Samoilova, A. V., & Leclerque, A. (2014). PCR-based Identification of Erwinia amylovora bacteriophages isolated in the Republic of Moldova. Journal of Virology and Microbiology. http://www.ibimapublishing.com/journals/JVM/AIP/293991/293991.pdf.
Schnabel, E. L., & Jones, A. L. (2001). Isolation and characterization of five Erwinia amylovora bacteriophages and assessment of phage resistance in strains of Erwinia amylovora. Applied and Environmental Microbiology, 67(1), 59–64.
Schnabel, E. L., Fernando, W. G. D., Meyer, M. P., Jones, A. L., & Jackson, L. E. (1999). Bacteriophage of Erwinia amylovora and their potential for biocontrol. Acta Horticulturae, 489, 649–654.
Schwarczinger, I., Kiss, E., Süle, S., Tóth, M., & Hevesi, M. (2011). Control of fire blight by bacteriophages on apple flowers. Acta Horticulturae, 896, 457–462.
Svircev, A. M., Lehman, S. M., Kim, W. S., Barszcz, E., Schneider, K. E., & Castle, A. J. (2006). Control of the fire blight pathogen with bacteriophages. In W. Zeller & C. Ullrich (Eds.), Proceedings of the 1st international symposium on biological control of bacterial plant diseases (pp. 259–261). Berlin: Arno Brynda.
Svircev, A. M., Castle, A. J., & Lehman, S. M. (2010). Bacteriophages for control of phytopathogens in food production system. In P. M. Sabour & M. W. Griffith (Eds.), Bacteriophages in the control of food- and waterborne pathogens (pp. 79–103). Washington DC: ASM Press.
Tanaka, H., Negishi, H., & Maeda, H. (1990). Control of tobacco bacterial wilt by an avirulent strain of Pseudomonas solanacearunm M4S and its bacteriophage. Annals of the Phytopathological Society of Japan, 56, 243–246.
Thomson, S. V. (1986). The role of stigma in fire blight infections. Phytopathology, 76, 476–482.
van der Zwet, T., & Beer, S. V. (1991). Fire blight, its nature, prevention and control - a practical guide to integrated disease management. Washington: Agriculture information bulletin. U.S. Department of Agriculture.
van der Zwet, T., & Keil, H. L. (1979). Fire blight, a bacterial disease of Rosaceous plants. Agriculture Handbook, Science and Education Administration. Washington: U.S. Department of Agriculture.
Vanneste, J. L. (2000). Fire blight, the disease and its causative agent, Erwinia amylovora. New York: CABI.
Vidaver, A. K. (1976). Prospects for control of phytopathogenic bacteria by bacteriophages and bacteriocins. Annual Review of Phytopathology, 14(1), 451–465.
Ward, R. L., & Mahler, R. J. (1982). Uptake of bacteriophage f2 through plant roots. Applied and Environmental Microbiology, 43(5), 1098–1103.
Wilson, M., Sigee, D. C., & Epton, H. A. S. (1990). Erwinia amylovora, infection of hawthorn blossom: III. The Nectary. Journal of Phytopathology, 128(1), 62–74.
Winslow, C. E., Broadhurst, J., Buchanan, R. E., Krumwiede Jr., C., Rogers, L. A., & Smith, G. H. (1920). The families and genera of the bacteria: final report of the committee of the Society of American Bacteriologists on characterization and classification of bacterial types. Journal of Bacteriology, 5(3), 191.
Yagubi, A. I., Castle, A. J., Kropinski, A. M., Banks, T. W., & Svircev, A. M. (2014). Complete genome sequence of Erwinia amylovora bacteriophage vB EamM Ea35- 70. Genome Announcements, 2(4), e00413–e00414.
Acknowledgements
Research has been funded by the Hungarian National Research, Development and Innovation Office (OTKA K 104730, OTKA PD 75280) and the Bolyai Scholarship (BO 609 12), which are gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
K. Geider has passed away on 03.03.2016.
Electronic supplementary material
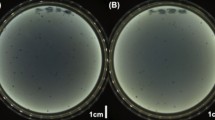
Fig. S1
Plaque morphology of Erwinia amylovora phages. Plaques in the upper agar layer containing the indicator bacterium EaCFBP1430 formed by ΦEaH5K (a) ΦEa116 (b) ΦEaH7B (c) and ΦEa100 (d). Pictures show plaques of different sizes produced by Hungarian (a, c) and North-American (b, d) phages investigated in this study. (JPEG 108 kb)
Supplemental Tables S1-S4
(DOCX 49.8 kb)
Rights and permissions
About this article
Cite this article
Schwarczinger, I., Kolozsváriné Nagy, J., Künstler, A. et al. Characterization of Myoviridae and Podoviridae family bacteriophages of Erwinia amylovora from Hungary - potential of application in biological control of fire blight. Eur J Plant Pathol 149, 639–652 (2017). https://doi.org/10.1007/s10658-017-1214-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-017-1214-9