Abstract
Epidemiology studies suggested that low birthweight was associated with a higher risk of hypertension in later life. However, little is known about the causality of such associations. In our study, we evaluated the causal association of low birthweight with adulthood hypertension following a standard analytic protocol using the study-level data of 183,433 participants from 60 studies (CHARGE-BIG consortium), as well as that with blood pressure using publicly available summary-level genome-wide association data from EGG consortium of 153,781 participants, ICBP consortium and UK Biobank cohort together of 757,601 participants. We used seven SNPs as the instrumental variable in the study-level analysis and 47 SNPs in the summary-level analysis. In the study-level analyses, decreased birthweight was associated with a higher risk of hypertension in adults (the odds ratio per 1 standard deviation (SD) lower birthweight, 1.22; 95% CI 1.16 to 1.28), while no association was found between genetically instrumented birthweight and hypertension risk (instrumental odds ratio for causal effect per 1 SD lower birthweight, 0.97; 95% CI 0.68 to 1.41). Such results were consistent with that from the summary-level analyses, where the genetically determined low birthweight was not associated with blood pressure measurements either. One SD lower genetically determined birthweight was not associated with systolic blood pressure (β = − 0.76, 95% CI − 2.45 to 1.08 mmHg), 0.06 mmHg lower diastolic blood pressure (β = − 0.06, 95% CI − 0.93 to 0.87 mmHg), or pulse pressure (β = − 0.65, 95% CI − 1.38 to 0.69 mmHg, all p > 0.05). Our findings suggest that the inverse association of birthweight with hypertension risk from observational studies was not supported by large Mendelian randomization analyses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hypertension, defined as high in systolic blood pressure, diastolic blood pressure, or both above normal levels, is a leading risk factor for mortality and morbidity. In 2015, high systolic blood pressure was associated with the heaviest disease burden among risk factors—more than either smoking or obesity [1]. Worldwide, the estimated rate of death attributable to high systolic blood pressure (140 mmHg or more) was 106.3/100,000 persons in 2015, and the number of disability-adjusted life-years was 7.8 million [2].
Over the past decades, epidemiology studies have provided emerging observational evidence for developmental origins for hypertension [3]. Low birthweight, a surrogate marker of intrauterine malnutrition and developmental stressors, has emerged as a potential risk factor for cardio-metabolic disorders, including hypertension in later life [4, 5]. Several lines of pathophysiological evidence have provided potential mechanisms including vascular dysfunction, reduced nephron numbers, sympathetic activation and neuroendocrine involved in the association of low birthweight with adulthood hypertension and blood pressure [6]. However, conventional observational studies are vulnerable to serious issues of confounding, reverse causality, inappropriate adjustment of current weight, and therefore are not able to make causal inference. Large-scaled meta-analyses of the observed associations between birthweight and hypertension in later life had reached controversial conclusions [5, 7]. Traditional clinical trials are unrealistic in such cases to assess the causality of these associations, necessitating other study designs.
Mendelian randomization (MR) is an emerging approach which takes advantage of genetic markers as instrumental variables (IVs) and therefore, potentially overcomes the limitations as mentioned above of observational studies and clinical trials. This approach exploits the fact that at meiosis individual genotypes are assigned randomly, and therefore, the effect of genetics on disease is free of confounding or reverse causality [8]. Birthweight has a significant genetic architecture, and approximately 15% of its variance can be attributed to fetal genetic variation [9], although the intrauterine environment also has considerable influence. Recent genome-wide association studies (GWAS) have identified seven variants [10] associated with birthweight, and such a list has expanded to 60 loci where fetal genotype was associated with birthweight [9]. These genetic variants can be used as a proxy for birthweight to examine whether low birthweight contributes causally to hypertension development.
In this study, we collected extensive study-level data from 60 studies with 183,433 participants (CHARGE-BIG consortium) and summary-level data from the Early Growth Genetics (EGG) consortium of 153,781 participants, the International Consortium of British Pensioners (ICBP) consortium and UK Biobank cohort (UKB) together of 757,601 participants, and explored the possible causal association of birthweight with adulthood blood pressure and hypertension using MR analyses. Because our study started earlier than the most recent published GWAS, which reported 60 loci of birthweight, we included the previous seven variants as the instrument variables in the analysis of study-level data, and 57 loci of birthweight in the analysis of summary-level data.
Methods
Study design and instruments
We use MR analyses to assess the causal association of birthweight with blood pressure and hypertension risk, under three assumptions [11]. First, genetic variants used as an instrument must be associated with birthweight. Second, genetic variants must not be associated with confounders. Third, genetic variants must not be associated with hypertension or blood pressure independent of birthweight. The above-mentioned second and third assumptions jointly refer to independence from pleiotropy.
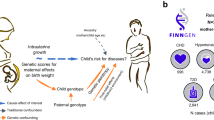
This study consisted of two parts (Fig. 1). First, we estimated the causal association of birth weight with hypertension risk using study-level data from the Cohorts for Heart and Aging Research in Genomic Epidemiology-Birth Gene (CHARGE-BIG) Study, which included 60 cross-sectional and prospective cohort studies with a total of 180,056 participants. The details of CHARGE-BIG study have been described before [12]. In brief, we analyzed the data within each study by standardized analytic methods using a genetic risk score (GRS) of the 7 single-nucleotide polymorphisms (SNPs) as an IV from an earlier GWAS of the EGG Consortium [10]. Second, we explored the causal association of birth weight with systolic blood pressure (SBP), diastolic blood pressure (DBP), and pulse pressure (PP) utilizing summary-level data from the EGG consortium (n = 153,781) [9], the UKB (n = 458,577) and the ICBP consortium (n = 299,024) [13]. Because neither UKB nor ICBP has hypertension as an existing categorical outcome in GWAS summary data, we included blood pressure measurements as the outcome variables in the summary-level analysis. A total of the available 57 SNPs or its proxies, a subset of the 60 SNPs reported by an updated result of EGG consortium [9], were used as the instrument for birth weight in the summary-level analysis.
Study design. The data sources included study-level data from the Cohorts for Heart and Aging Research in Genomic Epidemiology-Birth Gene (CHARGE-BIG) Study, which included 60 cross-sectional and prospective cohort studies, and summary-level data from the Early Growth Genetics (EGG) consortium, International Consortium of British Pensioners (ICBP) consortium and UK Biobank
All participants from CHARGE-BIG consortium provided written informed consent, and all participating studies received approval from local research ethics committees. The appendix (Supplemental Table 1) includes the description of all the included studies in CHARGE-BIG consortium in the analysis. Contributing studies received ethical approval from their respective institutional review boards.
Phenotypic measures
In the CHARGE-BIG consortium, Hypertension was defined as systolic blood pressure of 140 mmHg or higher, diastolic blood pressure of 90 mmHg or higher, or current use of antihypertensive medication. Birthweight was self-reported or collected from medical records, and information of covariates was collected in each study. The appendix (Supplemental Table 2) describes details about the methods used to collect information on birthweight and hypertension in each study. The detailed genome-wide analysis of blood pressure traits, including SBP, DBP and PP, among participants of European ancestry from UKB [14] and ICBP consortium [15, 16] have been described previously [13].
Selection of SNPs and genetic risk scores
In the study-level analyses, to create the GRS of low birthweight we selected 7 SNPs (CCNL1 rs900400, ADCY5 rs9883204, HMGA2 rs1042725, CDKAL1 rs6931514, 5q11.2 rs4432842, LCORL rs724577, ADRB1 rs1801253) based on findings from 69,308 participants of European descent by the EGG Consortium [10]. The genotyping information and the distribution of genotypes of these 7 SNPs in each study were described in Supplemental Tables 3 and 4. In a secondary analysis, we excluded 5 SNPs associated with blood pressure or significant confounders such as adult height and type 2 diabetes [10], and included the rest two SNPs in the GRS. We constructed an externally weighted low birthweight GRS, weighted by the effect estimates reported in EGG GWAS (β is the change in z score of birthweight per birthweight-lowering allele from linear regression, adjusted for sex and gestational age where available, assuming an additive genetic model) [10].
For the summary-level data analysis, a total of 60 SNPs were reported to be associated with birth weight by a more recent report from EGG consortium [9], of which 50 were available in UKB and ICBP consortium. For those SNPs that were not genotyped, we found proxies that are in high linkage disequilibrium with the corresponding SNP (r2 > 0.8) according to the information from 1000 Genomes Project. Ultimately, 57 SNPs were used as the instrument to assess the causal association of birth weight with blood pressure measurements.
Statistical analysis
Study-level analyses
In the study-level analyses, each of the CHARGE-BIG studies analyzed the data following a standard analytic protocol. Generalized linear regression models of the association between GRS and hypertension were adjusted with age, sex, body mass index (BMI), total energy intake, and principal components for population stratification if available. With respect to the phenotypic analyses, logistic regression models with hypertension as outcome and birthweight as exposure were adjusted with age, sex, BMI, and other risk factors of hypertension if available, such as smoking status (current vs. former/never), physical activity (MET h/day or hours) (quintiles), total energy intake (kcal) (quintiles), and alcohol consumption (quintiles). Concerning the genetic effects on birthweight, the effect allele was the birthweight-lowering allele, as established by the EGG consortium [10]. We tested for association of the GRS with birthweight using linear regression models, adjusting for sex, gestational age if available, and principal components for population stratification if available.
Within the CHARGE-BIG collaboration, formal MR analyses were conducted using the IV ratio method [17]. To assess the IV ratio for the effect of birthweight on hypertension, we divided the meta-analyzed association of birthweight GRS with hypertension by the association of birthweight GRS with birthweight. The variance for the IV ratio was estimated using a Taylor expansion.[18] The above analyses were repeated in the sex- and BMI (< 25 kg/m2, or ≥ 25 kg/m2)-stratified subgroups. To examine the strength of the GRS as an instrument, we calculated the F-statistic from the proportion of variation in the birthweight (R2) explained by the allele score, controlling for covariates (age, sex, and principal components for population stratification) in the Nurses’ Health Study (NHS) and the Health Professionals Follow-Up Study (HPFS) cohorts. An F statistic greater than 10 is evidence of a strong instrument [19].
To examine whether the SNPs for birthweight were associated with potential confounders, each birthweight-associated SNP was evaluated for pleiotropy associations with potential risk factors, including major lipids in 196,476 individuals (Global Lipids Genetics Consortium) [20], glycemic traits in 46,186 individuals without diabetes (Meta-Analyses of Glucose and Insulin-Related Traits Consortium) [21], type 2 diabetes in 110,452 individuals (Diabetes Genetics Replication and Meta-analysis) [22], BMI and waist-to-hip ratio adjusted for BMI in 224,459 individuals (Genetic Investigation of Anthropometric Traits) [23], and chronic kidney disease-defining traits in 175,579 individuals [24] (Supplemental Fig. 1).
In the presence of heterogeneity of association among studies, inverse variance-weighted random-effects models were used for meta-analyses; otherwise, fixed-effects models were used. Heterogeneity among studies was assessed with the I2 statistic.[25,26,27] We found non-negligible heterogeneity between studies, in particular among the birthweight-hypertension associations, but also for the association between low birthweight GRS and birthweight (I2 > 0.25).
Summary-level analyses
We extracted 57 beta-coefficients and standard errors of the SNP-birthweight associations from EGG consortium, and that of SNP-blood pressure associations from the ICBP consortium and UKB via GWAS catalog (https://www.ebi.ac.uk/gwas/downloads/summary-statistics). We computed individual MR estimates and standard errors by weighting the effect sizes based on the magnitude of the SNP-birthweight association [28]. We used the inverse variance-weighted (IVW) MR approach as the primary analysis, where the inverse variance weighted mean of ratio estimates from the multiple IVs is the IV estimate [28]. This approach assumes that IVs affect the outcome only through the exposure under consideration, and not via any alternative pathways [28]. Violation of this assumption implies horizontal pleiotropy of the IV, measured by the heterogeneity estimates of Cochran Q-derived p < 0.05, and it could bias the MR estimate. Thus, we further conducted several sensitivity analyses with different assumptions regarding the presence of pleiotropic genetic variants that may relate with the outcome independently of the exposure. For example, MR-Egger regression requires that the strengths of the instruments are independent of their direct associations with the outcome [11], and the weighted median method requires that at least half of the information for the MR analysis comes from valid instruments [29]. The intercept of the MR-Egger regression is a measure of directional pleiotropy (p < 0.05 was considered significant) [11].
We carried out all the analyses with R version 3.2.3 (https://www.r-project.org).
Results
The study-level results
In the study-level analysis, the analytic sample included 183,433 individuals from 60 cohort and case–control studies (Fig. 2, Supplemental Table 5). Twenty-four studies (51,568 participants) reported the GRS-birthweight associations; and 33 studies (109,735participants) reported the GRS-hypertension associations. A total of 70,874 hypertensive participants and 61,933 normotensive controls provided hypertension-related data, and 50,626 participants provided GRS-birthweight associations only. The majority of participants were of European (86%) and Asian (14%) ancestry (Supplemental Table 5).
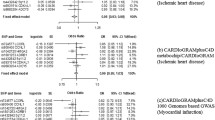
Large scale GWAS consortia did not suggest that the seven SNPs were associated with potential hypertension risk factors, including circulating major lipids, fasting glucose and insulin, type 2 diabetes, BMI, waist-to-hip ratio, and chronic kidney disease (Supplemental Fig. 1). The low birthweight GRS was inversely associated with birthweight (Fig. 3a, each risk allele was associated with 0.02 standard deviation (SD) lower birthweight, and there was evidence for heterogeneity in such an association (I2 = 78%, p < 0.01). The F-statistics for the score were both > 18 using data from the NHS and the HPFS (Supplemental Table 6), indicating the GRS is a strong composite instrument.
Meta-analysis of associations of low birth weight genetic risk score with birth weight (a) and hypertension (b) using the study-level data from CHARGE-BIG consortium. Betas were the associations of low birth weight genetic risk score with outcome, per risk allele for low birth weight. CI, confidence interval
In the meta-analysis of the CHARGE-BIG studies, lower birthweight was associated with a higher risk of hypertension in adults (Table 1 and Fig. 4, odds ratio (OR) per 1 SD lower birthweight, 1.22, 95% CI 1.16 to 1.28). There was no significant association of the low birthweight GRS with hypertension risk (Table 1 and Fig. 3b, OR per 1 risk allele of low birthweight: 1.00, 95% CI 0.99 to 1.01). The relationships of lower birthweight and low birthweight GRS with the risk of hypertension in both sexes and BMI status were consistent with those in the overall population (Table 1).
In the formal MR analysis, genetically instrumented birthweight was not associated with risk of hypertension (Table 1 and Fig. 4, instrumental OR for causal effect per 1 SD lower birthweight: 0.97, 95% CI 0.68 to 1.41). Again, no association was seen in each sex or BMI status group (Table 1). The secondary analysis using two SNPs conservatively either showed no association between genetically instrumented birthweight and risk of hypertension (instrumental OR 1.12, 95% CI 0.66 to 1.89, Supplemental Fig. 2).
The summary-level results
In the random-effect IVW MR analyses using the 57 SNPs as the IVs, one SD lower genetically instrumented birth weight showed a trend of association with 0.76 mmHg lower SBP (95% CI − 2.45 to 1.08 mmHg), 0.06 mmHg lower DBP (95% CI − 0.93 to 0.87 mmHg), and 0.65 mmHg lower PP (95% CI 95% CI − 1.38 to 0.69 mmHg), however, none of these associations was significant (all p > 0.05, Table 2). No presentation for directional pleiotropy effects was detected by the MR-Egger intercept (SBP, p = 0.73; DBP, p = 0.64; PP, p = 0.90; Table 2). Although there was evidence for horizontal pleiotropy of the IV (Cochran Q derived p < 0.05), the results from MR-Egger method and weighted median based method were consistent with that from IVW MR method for SBP, DBP and PP (Table 2). We further excluded 14 previously reported SNPs for blood pressure or hypertension, or used the 7 SNPs only as sensitivity analyses in order to be consistent with the study-level analyses, and in either situation low birthweight remained not associated with blood pressure measurements (Supplemental Table 7).
Discussion
Numerous nutritional interventions have been effective in reducing the short-term risk of low birthweight and prematurity. Understanding the potential long-term benefits of such interventions is crucial to inform policy decisions to interrupt the developmental programming cycle and stem the growing epidemics of hypertension worldwide. With low birthweight related genetic loci as the IV, the results of our MR analysis provide evidence for a non-causal effect of low birthweight on a higher risk of hypertension and blood pressure measurements, suggesting that low birthweight might not be a casual risk factor for development of hypertension.
Evidence from observational studies of low birthweight and a higher risk of hypertension constitutes some most robust finding supporting the fetal origins of adult disease [30]. Barker et al. were the first to report that low birthweight was associated with a higher risk of cardiovascular disease [31]. Subsequently, Brenner and colleagues proposed that developmental programming in the kidney may reduce nephron number, which may result in a limited filtration surface area and reduced sodium excretion, and eventually development of hypertension [32]. Our observed inverse association of birthweight with hypertension risk was consistent with traditional observational studies, which were largely from Caucasians [4, 33,34,35,36]. In Chinese populations, intrauterine exposure to famine was related to a higher risk of hypertension in adults [37, 38], and such findings were indirectly consistent with our observational findings.
In our study, we did not observe an association of genetically determined birthweight with hypertension risk or blood pressure measurements during adulthood. Our result is in line with that from the recent MR analysis from UKB [39], which also reported a null association of birthweight with blood pressure and hypertension risk. However, the UKB analysis exclusively studied the Caucasian population in the UK, and our analysis included samples of Caucasians and Asians from diverse populations and countries. It is worth mentioning the genetic correlation analyses of birthweight with hypertension from the recent GWAS for birthweight [9]. This GWAS is in line with our findings that it suggested a lack of genetic association between birthweight and blood pressure from linkage-disequilibrium score regression, indicating that birthweight is not causal for hypertension risk and blood pressure as well. Consistently, a recent MR study with a smaller sample size (n = 5000) selecting instruments according this GWAS did not found significant causal association between birth weight and hypertension either [40, 41]. Our study suggested a lack of association of the genetic instruments of birthweight, and this observation did not implicate that a lack of association of the intrauterine malnutrition and developmental stressors with hypertension risk. It is possible that the environment determined lower birth weight might have an effect on the risk of hypertension, though it is beyond the scope of the current analysis. Our findings should not be interpreted as to undermine the critical value of interventions improving birthweight in order to lower the hypertension risk in later life.
Our study has several strengths. First, we carried out an IV analysis on the causality of birthweight on hypertension and blood pressure using large and diverse populations. The large sample size might provide us with sufficient power to estimate the causal effect of low birthweight on hypertension and blood pressure, and the diverse source of data allows decent generalizability. Second, we used a standardized analysis protocol to collect study-level statistics within CHARGE-BIG consortium, and it minimized the potential bias from different data analyses methods. Our data should be interpreted with caution, and several limitations were related to the validity of the assumptions underlying the causal interpretation of MR studies. First, for the instrument variable, in the study-level analyses we only used seven SNPs related with low birthweight instead of the 60 SNPs from the most recent GWAS [9], however, in our summary-level analyses, we included 57 available SNPs. The results were consistent in study-level and summary-level analyses, as well as in different sensitivity analyses, providing further support for the noncausal association of birthweight with blood pressure and hypertension risk. Second, though we have minimized the horizontal pleiotropic effects using exiting large consortia data and different MR sensitivity analysis methods, future studies are warranted to take into consideration other essential factors that may be causatively related with intrauterine growth restriction. Such factors may include prenatal factors such as gestational week and postnatal behaviors such as breastfeeding. Third, we did not include the maternal genetic background in the analysis, which may affect the intrauterine environment and therefore, birthweight. Recent GWAS suggested that several maternal genetic variants influence fetal birthweight independently of the fetal genome [42]. Therefore, future MR studies with IVs from both maternal and fetal aspects of adult hypertension risk and blood pressure would provide new insights. Fourth, we did not collect blood pressure measurements from individual studies in the study-level analysis. Blood pressure may have a more significant measurement error, and the estimated association with blood pressure may be weaker compared that with hypertension [43]. Nevertheless, we used the blood pressure measurements in the summary-level analyses and reached consistent conclusion. Fifth, in the study-level analyses, we defined hypertension according to the previous definition [44] not the one currently proposed [45] by the American Heart Association, as the study was designed and conducted before the new definition issued. Canalization is one possible explanation for our results, because the low birthweight allele score might have led to biological adaptations during development [8]. Furthermore, we assumed that the association of genetically determined birthweight with hypertension risk and blood pressure is linear; however, such assumption may not be correct because both the extreme low or high birthweights influence hypertension risk [46].
The associations of low birthweight, as an indicator of intrauterine growth restriction, with a higher hypertension risk and blood pressure measurements in adults from observational studies were not supported by our MR analyses. These findings suggest that the observational association of birthweight with hypertension risk in later life could be the result of confounding.
References
GBD 2015 Risk Factors Collaborators. Global, regional, and national comparative risk assessment of 79 behavioural, environmental and occupational, and metabolic risks or clusters of risks, 1990–2015: a systematic analysis for the Global Burden of Disease Study 2015. Lancet. 2016;388(10053):1659–724. https://doi.org/10.1016/S0140-6736(16)31679-8.
Forouzanfar MH, Liu P, Roth GA, et al. Global burden of hypertension and systolic blood pressure of at least 110 to 115 mmHg, 1990–2015. JAMA. 2017;317(2):165–82. https://doi.org/10.1001/jama.2016.19043.
Luyckx VA, Brenner BM. Birth weight, malnutrition and kidney-associated outcomes—a global concern. Nat Rev Nephrol. 2015;11(3):135–49. https://doi.org/10.1038/nrneph.2014.251.
Curhan GC, Chertow GM, Willett WC, et al. Birth weight and adult hypertension and obesity in women. Circulation. 1996;94(6):1310–5.
Mu M, Wang SF, Sheng J, et al. Birth weight and subsequent blood pressure: a meta-analysis. Arch Cardiovasc Dis. 2012;105(2):99–113. https://doi.org/10.1016/j.acvd.2011.10.006.
Bruno RM, Faconti L, Taddei S, Ghiadoni L. Birth weight and arterial hypertension. Curr Opin Cardiol. 2015;30(4):398–402. https://doi.org/10.1097/hco.0000000000000180.
Huxley R, Neil A, Collins R. Unravelling the fetal origins hypothesis: is there really an inverse association between birthweight and subsequent blood pressure? Lancet. 2002;360(9334):659–65. https://doi.org/10.1016/S0140-6736(02)09834-3.
Smith GD, Ebrahim S. 'Mendelian randomization': can genetic epidemiology contribute to understanding environmental determinants of disease? Int J Epidemiol. 2003;32(1):1–22.
Horikoshi M, Beaumont RN, Day FR, et al. Genome-wide associations for birth weight and correlations with adult disease. Nature. 2016;538(7624):248–52. https://doi.org/10.1038/nature19806.
Horikoshi M, Yaghootkar H, Mook-Kanamori DO, et al. New loci associated with birth weight identify genetic links between intrauterine growth and adult height and metabolism. Nat Genet. 2013;45(1):76–82. https://doi.org/10.1038/ng.2477.
Bowden J, Davey Smith G, Burgess S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol. 2015;44(2):512–25.
Group B-GSW, Huang T, Wang T, et al. Association of birth weight with type 2 diabetes and glycemic traits a mendelian randomization study. JAMA Netw Open. 2019;2(9):e1910915.
Evangelou E, Warren HR, Mosen-Ansorena D, et al. Genetic analysis of over 1 million people identifies 535 new loci associated with blood pressure traits. Nat Genet. 2018;50(10):1412–25. https://doi.org/10.1038/s41588-018-0205-x.
Sudlow C, Gallacher J, Allen N, et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 2015;12(3):e100779.
Wain Louise V, Vaez A, Jansen R, et al. Novel blood pressure locus and gene discovery using genome-wide association study and expression data sets from blood and the kidney. Hypertension. 2017;70(3):e4–e19. https://doi.org/10.1161/HYPERTENSIONAHA.117.09438.
Ehret GB, Munroe PB, Rice KM, et al. Genetic variants in novel pathways influence blood pressure and cardiovascular disease risk. Nature. 2011;478(7367):103–9. https://doi.org/10.1038/nature10405.
Lawlor DA, Harbord RM, Sterne JA, Timpson N, Davey SG. Mendelian randomization: using genes as instruments for making causal inferences in epidemiology. Stat Med. 2008;27(8):1133–63. https://doi.org/10.1002/sim.3034.
Thomas DC, Lawlor DA, Thompson JR. Re: Estimation of bias in nongenetic observational studies using "Mendelian triangulation" by Bautista et al. Ann Epidemiol. 2007;17(7):511–3. https://doi.org/10.1016/j.annepidem.2006.12.005.
Burgess S, Thompson SG. Avoiding bias from weak instruments in Mendelian randomization studies. Int J Epidemiol. 2011;40(3):755–64. https://doi.org/10.1093/ije/dyr036.
Willer CJ, Schmidt EM, Sengupta S, et al. Discovery and refinement of loci associated with lipid levels. Nat Genet. 2013;45(11):1274–83. https://doi.org/10.1038/ng.2797.
Dupuis J, Langenberg C, Prokopenko I, et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat Genet. 2010;42(2):105–16. https://doi.org/10.1038/ng.520.
Mahajan A, Go MJ, Zhang W, et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat Genet. 2014;46(3):234–44. https://doi.org/10.1038/ng.2897.
Shungin D, Winkler TW, Croteau-Chonka DC, et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature. 2015;518(7538):187–96. https://doi.org/10.1038/nature14132.
Pattaro C, Teumer A, Gorski M, et al. Genetic associations at 53 loci highlight cell types and biological pathways relevant for kidney function. Nat Commun. 2016;7(1):10023. https://doi.org/10.1038/ncomms10023.
Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003;327(7414):557–60. https://doi.org/10.1136/bmj.327.7414.557.
Ioannidis JP, Patsopoulos NA, Evangelou E. Uncertainty in heterogeneity estimates in meta-analyses. BMJ. 2007;335(7626):914–6. https://doi.org/10.1136/bmj.39343.408449.80.
Higgins JP, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med. 2002;21(11):1539–58. https://doi.org/10.1002/sim.1186.
Burgess S, Butterworth A, Thompson SG. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol. 2013;37(7):658–65. https://doi.org/10.1002/gepi.21758.
Bowden J, Davey Smith G, Haycock PC, Burgess S. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet Epidemiol. 2016;40(4):304–14. https://doi.org/10.1002/gepi.21965.
Lenfant C. Low birth weight and blood pressure. Metabolism. 2008;57(Suppl 2):S32–S3535. https://doi.org/10.1016/j.metabol.2008.07.013.
Barker DJ, Osmond C. Infant mortality, childhood nutrition, and ischaemic heart disease in England and Wales. Lancet. 1986;1(8489):1077–81.
Brenner BM, Garcia DL, Anderson S. Glomeruli and blood pressure. Less of one, more the other? Am J Hypertension. 1988;1(4 Pt 1):335–47.
Curhan GC, Willett WC, Rimm EB, Spiegelman D, Ascherio AL, Stampfer MJ. Birth weight and adult hypertension, diabetes mellitus, and obesity in US men. Circulation. 1996;94(12):3246–50.
Bergvall N, Iliadou A, Johansson S, et al. Genetic and shared environmental factors do not confound the association between birth weight and hypertension: a study among Swedish twins. Circulation. 2007;115(23):2931–8. https://doi.org/10.1161/CIRCULATIONAHA.106.674812.
Li Y, Ley SH, Vander Weele TJ, et al. Joint association between birth weight at term and later life adherence to a healthy lifestyle with risk of hypertension: a prospective cohort study. BMC Med. 2015;13:175. https://doi.org/10.1186/s12916-015-0409-1.
Johansson S, Iliadou A, Bergvall N, Tuvemo T, Norman M, Cnattingius S. Risk of high blood pressure among young men increases with the degree of immaturity at birth. Circulation. 2005;112(22):3430–6. https://doi.org/10.1161/CIRCULATIONAHA.105.540906.
Li Y, Jaddoe VW, Qi L, et al. Exposure to the Chinese famine in early life and the risk of hypertension in adulthood. J Hypertens. 2011;29(6):1085–92. https://doi.org/10.1097/HJH.0b013e328345d969.
Huang C, Li Z, Wang M, Martorell R. Early life exposure to the 1959–1961 Chinese famine has long-term health consequences. J Nutr. 2010;140(10):1874–8. https://doi.org/10.3945/jn.110.121293.
Zanetti D, Tikkanen E, Gustafsson S, Priest JR, Burgess S, Ingelsson E. Birthweight, type 2 diabetes mellitus, and cardiovascular disease: addressing the barker hypothesis with Mendelian randomization. Circ Genomic Precis Med. 2018;11(6):e002054. https://doi.org/10.1161/CIRCGEN.117.002054.
Zeng P, Zhou X. Causal association between birth weight and adult diseases: evidence from a Mendelian randomisation analysis. Front Genet. 2019;10:618.
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature. 2007;447(7145):661.
Beaumont RN, Warrington NM, Cavadino A, et al. Genome-wide association study of offspring birth weight in 86,577 women identifies five novel loci and highlights maternal genetic effects that are independent of fetal genetics. Hum Mol Genet. 2018;27(4):742–56. https://doi.org/10.1093/hmg/ddx429.
Vimaleswaran KS, Cavadino A, Berry DJ, et al. Association of vitamin D status with arterial blood pressure and hypertension risk: a mendelian randomisation study. Lancet Diab Endocrinol. 2014;2(9):719–29. https://doi.org/10.1016/S2213-8587(14)70113-5.
Carretero OA, Oparil S. Essential hypertension: part I: definition and etiology. Circulation. 2000;101(3):329–35.
Whelton PK, Carey RM, Aronow WS, et al. 2017 ACC/AHA/AAPA/ABC/ACPM/AGS/APhA/ASH/ASPC/NMA/PCNA guideline for the prevention, detection, evaluation, and management of high blood pressure in adults: a report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. J Am Coll Cardiol. 2018;71(19):e127–e248.
Gamborg M, Byberg L, Rasmussen F, et al. Birth weight and systolic blood pressure in adolescence and adulthood: meta-regression analysis of sex- and age-specific results from 20 Nordic studies. Am J Epidemiol. 2007;166(6):634–45. https://doi.org/10.1093/aje/kwm042.
Acknowledgements
We thank the staff and the participants of the studies for their valuable contributions for their important contributions. We thank the following individuals for their assistance with data collection and analysis: Andre G. Uitterlinden (University of Leipzig, Leipzig, Germany), Chiea-Chuen Khor (Agency for Science Technology and Research, Singapore), Christian Hengstenberg (Technische Universität München, Germany), Fernando Rivadeneira (University Medical Center Rotterdam, the Netherlands), Jianjun Liu (Agency for Science Technology and Research, Singapore), JianMin Yuan (University of Pittsburgh Cancer Institute, Pittsburgh, USA), Lise Tarnow (University of Copenhagen, Hillerød, Denmark), Markus Loeffler (University of Leipzig, Leipzig, Germany), Oluf Pedersen (University of Copenhagen, Denmark), Oscar Franco (Erasmus MC University Medical Center, Rotterdam, the Netherlands), Paul Franks (Lund University, Skåne University Hospital Malmö, Malmö, Sweden), ShuPei Tan (Singapore National Eye Centre, Singapore), Thomas Meitinger (Helmholtz Zentrum München–German Research Center for Environmental Health, Germany), Tine Marie Pedersen (University of Copenhagen, Denmark), Wanting Zhao (Singapore National Eye Centre, Singapore), Yik-Ying Teo (National University of Singapore, Singapore), Yoichiro Kamatani (RIKEN Center for Integrative Medical Sciences, Japan), Yuan Shi (Singapore National Eye Centre, Singapore), Ilja Demuth (Charité - Universitätsmedizin Berlin, Germany), Lars Bertram (Technische Universität München, Germany), Linda S. Adair (University of North Carolina, USA), Cyrus Cooper (University of Southampton, Southampton General Hospital, Southampton, UK), Hazel Inskip (University of Southampton, Southampton General Hospital, Southampton, UK), Sarah Crozier (University of Southampton, Southampton General Hospital, Southampton, UK), Elaine Dennison (MRC Lifecourse Epidemiology Unit, Southampton, UK), and Karen Jameson (MRC Lifecourse Epidemiology Unit, Southampton, UK)
Funding
This work was supported by Shanghai Municipal Science and Technology Major Project (Grant No. 2017SHZDZX01) and the National Key Research and Development Program of China (Grant No. 2016YFC1304801). YZ was supported by the Program for Professor of Special Appointment (Eastern Scholar) at Shanghai Institutions of Higher Learning. For the funding information of each study within CHARGE-BIG consortium, please see Supplemental Table 8.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
There are no relevant conflicts of interest on the part of any study authors. There are no relevant financial, personal or professional relationships with other people or organizations to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zheng, Y., Huang, T., Wang, T. et al. Mendelian randomization analysis does not support causal associations of birth weight with hypertension risk and blood pressure in adulthood. Eur J Epidemiol 35, 685–697 (2020). https://doi.org/10.1007/s10654-020-00638-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10654-020-00638-z