Summary
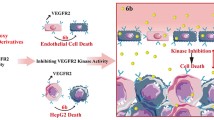
Herein, a novel series of dual histone deacetylase (HDAC) and vascular endothelial growth factor receptor (VEGFR) inhibitors were designed, synthesized and biologically evaluated based on previously reported pazopanib-based HDAC and VEGFR dual inhibitors. Most target compounds showed significant HDAC1, HDAC6 and VEGFR2 inhibition, which contributed to their potent antiproliferative activities against multiple cancer cell lines and significant antiangiogenic potencies in both human umbilical vein endothelial cell (HUVEC) tube formation assays and rat thoracic aorta ring assays. Further HDAC selectivity evaluations indicated that hydroxamic acids 5 and 9e possessed HDAC isoform selectivity profiles similar to that of the approved HDAC inhibitor suberoylanilide hydroxamic acid(SAHA), while hydrazide12 presented an HDAC isoform selectivity profilesimilar to that of the clinical HDAC inhibitor MS-275. The VEGFR inhibition profiles of 5, 9e and 12 were similar to that of the approved VEGFR inhibitor pazopanib. The intracellular target engagements of Compounds 5 and 12 were confirmed by western blot analysis. The metabolic stabilities of 5, 9e and 12 in mouse liver microsomes were inferior to that of pazopanib. These dual HDAC and VEGFR inhibitors provide lead compounds for further structural optimization to obtainpolypharmacological anticancer agents.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
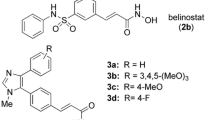
Targeting epigenetic aberrations is an important strategy for cancer treatment [1]. Among the various epigenetic enzymes, histone deacetylases (HDACs) are a family of validated anticancer targets with five inhibitors (vorinostat, romidepsin, belinostat, panobinostat and chidamide, Fig. 1) approved for the treatment of hematologic cancer [2].In addition to epigenetic regulation by removal of the acetyl groups from histones, HDACs also play important roles in posttranslational modification by deacetylating numerous nonhistones [3].
Kinases are one of the most intensively pursued targets in current pharmacological research, especially for cancer, due to their critical roles in regulating protein phosphorylation, one of the most important posttranslational modifications involved in signal transduction [4].Vascular endothelial growth factor receptors (VEGFRs) are a family of receptor tyrosine kinases that mediate the biological functions of VEGFs, thereby playing key roles in vascular development.Although many small molecular inhibitors targeting VEGFRs have been approved for the treatment of solid tumors [5], drug resistance and tumor relapse has occurred in most patients treated with VEGFR inhibitors,including pazopanib [6, 7] (Fig. 1).
It is worth noting that many preclinical studies havefound that combining HDAC inhibitors and pazopanib holds great promise for overcoming pazopanib resistance and enhancing antitumor efficacy [8,9,10]. More importantly, a recent phase I clinical study showed that targeting epigenetic modifications with the HDAC inhibitor abexinostat could enhance the response and reverse resistance to pazopanib in patients with many solid tumor malignancies [11].Based on the benefits of the HDAC inhibitor and VEGFR inhibitor combination, Zang et al. developed a series of pazopanib-based HDAC and VEGFR dual inhibitors, among which compounds JMC-13f and JMC-6d (Fig. 2) exhibited potent HDAC and VEGFR inhibitory activities, transforming their potent antiproliferative activities and antiangiogenic potencies [12]. Inspired by the pioneering work by Zang et al. [12], herein, a novel series of pazopanib derivatives were designed and synthesized via structural modification of compounds JMC-13f and JMC-6d in the hopes of obtaining novel dual HDAC and VEGFR inhibitors with promising antitumor potency (Fig. 2).
Materials and methods
Chemical reagents and solvents were purchased from commercial sources and used without further purification.1H NMR (Nuclear magnetic resonance, NMR) and 13C NMR spectra were obtained using a Bruker DRX spectrometer at 400 and 100 MHz, respectively. Chemical shifts are reported in parts per million (ppm). Multiplicity of the 1H NMR signals is reported as singlet (s), doublet (d), triplet (t), quartet (q), and multiplet (m). ESI–MS data were recorded on an API 4000 spectrometer. Melting points were determined using an open capillary on an uncorrected electrothermal melting point apparatus.
N-(2-Chloropyrimidin-4-yl)-N,2,3-trimethyl-2H-indazol-6-amine(1)and4-((4-((2,3-dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)benzoic acid (10) were synthesized according to previously reported methods [12].
4-((4-((2,3-Dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenol (2).To a solution of 1 (0.50 g,1.74 mmol) and 4-aminophenol (0.23 g, 2.09 mmol) inisopropanol (30 mL) was added 2 drops of concentrated HCl, and the mixture was heated to reflux with stirring for 4 h. The mixture was cooled to room temperature and the resulting precipitate was collected viafiltration and washed with ethyl acetate, affording Compound 2 as a white solid(0.43 g, 70%). ESI–MS m/z: 360.14 [M + H] + .
N2-(4-Aminophenyl)-N4-(2,3-dimethyl-2H-indazol-6-yl)-N4-methylpyrimidine-2,4-diamine(3). To a solution of 1 (0.50 g,1.74 mmol) and benzene-1,4-diamine (0.23 g, 2.09 mmol), inisopropanol (30 mL) was added to 2 drops of concentrated HCl, and the mixture was heated to reflux with stirring for 4 h. The mixture was cooled to room temperature, and the resulting precipitate was collected via filtration and washed with ethyl acetate, affording Compound 3 as a white solid (0.41 g, 65%). ESI–MS m/z: 360.05 [M + H] + .
Methyl-8-((4-((4-((2,3-dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenyl)amino)-8-oxooctanoate(4).To a solution of 3 (0.46 g, 1.29 mmol) in DMF (10 mL) in an ice bath, 2-(1H-benzotriazole-1-yl)-1,1,3,3-tetramethyluronium tetrafluoroborate (TBTU, 0.50 g, 1.54 mmol) was added, followed by Et3N (0.16 g,1.54 mmol). Thirty minutes later, suberic acid monomethyl ester (0.29 g, 1.54 mmol) was added. Twelve hours later, the solution was diluted with water and extracted with ethyl acetate. The combined organic extracts were washed with saturated NaHCO3and brine and dried over Na2SO4overnight, and the solvent was evaporated under vacuum. The crude product was purified by silica gel column chromatography (MeOH/CH2Cl2, 1/50 to 1/20) to affordCompound 4 as a white solid (0.40 g, 58% yield). ESI–MS m/z: 530.14 [M + H] + .
N1-(4-((4-((2,3-Dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenyl)-N8-hydroxyoctanediamide (5).KOH (28.55 g, 509 mmol) and NH2OH·HCl (23.84 g, 343 mmol) were dissolved in 70 mL and 120 mL of MeOH to obtain solution A and solution B, respectively. Then, solution A was added dropwise to solution B. After filtering the precipitated KCl, an NH2OK solution was obtained. Compound 4(0.25 g, 0.47 mmol) was dissolved in 30 mL of NH2OK solution and stirred for 2 h. After the reaction was complete, the solvent was evaporated under vacuum. The residue was acidified by the addition of 1 M HCl to pH 5–6.The resulting precipitate was collected by filtration and dried to affordCompound5 as a white solid (0.11 g, 43% yield). 1H NMR (400 MHz, DMSO-d6) δ 10.34 (s, 1H), 9.68 (s, 1H), 9.04 (s, 1H), 8.65 (s, 1H), 7.82 (d, J = 5.9 Hz, 1H), 7.75 (d, J = 8.8 Hz, 1H), 7.61 (d, J = 9.0 Hz, 2H), 7.43 (d, J = 1.7 Hz, 1H), 7.37 (d, J = 8.9 Hz, 2H), 6.88 (dd, J = 8.8, 1.8 Hz, 1H), 5.76 (d, J = 6.0 Hz, 1H), 4.06 (s, 3H), 3.46 (s, 3H), 2.63 (s, 3H), 2.25 (t, J = 7.4 Hz, 2H), 1.94 (t, J = 7.4 Hz, 2H), 1.64 – 1.40 (m, 4H), 1.34 – 1.21 (m, 4H). 13C NMR (101 MHz, DMSO-d6) δ 171.25, 169.62, 162.89, 158.01, 153.25, 147.39, 142.05, 135.81, 134.08, 132.71, 122.30, 120.07, 119.97, 119.91, 114.48, 96.64, 38.61, 37.86, 36.76, 32.75, 28.89, 25.60, 25.52, 9.89. HRMS (AP-ESI) m/z calcd for C28H35N8O3 [M + H] + 531.2832, found 531.2882.
Methyl-6-((4-((4-((2,3-dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenyl)amino)hexanoate (6).To a solution of 3 (0.40 g, 1.11 mmol) in DMF (10 mL), potassium carbonate (K2CO3, 0.18 g, 1.33 mmol) was added, followed by methyl 6-bromohexanoate (0.28 g,1.33 mmol). The reaction mixture was stirred at 70 °C. Twelve hours later, the solution was diluted with water and extracted with ethyl acetate. The combined organic extracts were washed with saturated NaHCO3 and brine and dried over Na2SO4 overnight, and the solvent was evaporated under vacuum. The crude product was purified by silica gel column chromatography (MeOH/CH2Cl2, 1/50 to 1/20) to afford Compound 6 asa white solid (0.21 g, 39% yield). ESI–MS m/z: 488.25 [M + H] + .
Methyl-2-(4-((4-((2,3-dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenoxy)acetate (8a). To a solution of 2 (0.48 g, 1.33 mmol) in DMF (10 mL), cesium carbonate (Cs2CO3, 0.52 g, 1.60 mmol) was added, followed by methyl bromoacetate (0.24 g,1.60 mmol). The reaction mixture was stirred at 80 °C. Six hours later, the solution was diluted with water and extracted with ethyl acetate. The combined organic extracts were washed with saturated NaHCO3 and brine and dried over Na2SO4 overnight, and the solvent was evaporated under vacuum. The crude product was purified by silica gel column chromatography (MeOH/CH2Cl2, 1/50 to 1/20) to affordCompound8a as a white solid (0.42 g, 73% yield). ESI–MS m/z: 433.23 [M + H] + .
Compounds 8b-8e were prepared from Compound 2 in a similarmanner to that described for Compound 8a.
Methyl-4-(4-((4-((2,3-dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenoxy)butanoate (8b). White solid. 70% yield. ESI–MS m/z: 461.21 [M + H] + .
Methyl-5-(4-((4-((2,3-dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenoxy)pentanoate (8c). White solid. 67% yield. ESI–MS m/z: 475.19 [M + H] + .
Methyl-6-(4-((4-((2,3-dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenoxy)hexanoate (8d). White solid. 65% yield. ESI–MS m/z: 489.32 [M + H] + .
Methyl-7-(4-((4-((2,3-dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenoxy)heptanoate (8e). White solid. 75% yield. ESI–MS m/z: 503.31 [M + H] + .
Compounds 7 and 9a-9e were prepared from Compounds 6 and 8a-8e,respectively, in a similarmanner to that described for Compound 5.
6-((4-((4-((2,3-Dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2yl)amino)phenyl)amino)-N-hydroxyhexanamide (7).White solid. 60% yield. 1H NMR (400 MHz, DMSO-d6) δ 10.33 (s, 1H), 8.80 (s, 1H), 8.64 (s, 1H), 7.74 (t, J = 7.7 Hz, 2H), 7.43 (s, 1H), 7.36 (d, J = 8.7 Hz, 2H), 6.90 – 6.83 (m, 1H), 6.45 (d, J = 8.6 Hz, 2H), 5.69 (d, J = 6.1 Hz, 1H), 4.06 (s, 3H), 3.44 (s, 3H), 2.94 (t, J = 6.8 Hz, 2H), 2.62 (s, 3H), 1.96 (t, J = 7.4 Hz, 2H), 1.53 (p, J = 7.2 Hz, 4H), 1.39 – 1.28 (m, 2H). HRMS (AP-ESI) m/z calcd for C26H33N8O2 [M + H] + 489.2726, found 489.2749.
2-(4-((4-((2,3-Dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenoxy)-N-hydroxyacetamide (9a). white solid (0.16 g, 50% yield). 1H NMR (400 MHz, DMSO-d6) δ 10.78 (s, 1H), 9.07 (s, 1H), 8.93 (s, 1H), 7.80 (d, J = 6.1 Hz, 1H), 7.76 (d, J = 8.7 Hz, 1H), 7.61 (d, J = 8.9 Hz, 2H), 7.44 (s, 1H), 6.88 (dd, J = 8.8, 1.5 Hz, 1H), 6.81 (d, J = 8.9 Hz, 2H), 5.76 (d, J = 6.0 Hz, 1H), 4.39 (s, 2H), 4.06 (s, 3H), 3.46 (s, 3H), 2.63 (s, 3H). 13C NMR (101 MHz, methanol-d4) δ 166.56, 163.04, 158.33, 153.41, 152.56, 147.46, 142.86, 133.94, 133.43, 121.64, 121.54, 119.95, 119.81, 114.53, 113.54, 96.08, 66.48, 37.41, 36.21, 8.28. HRMS (AP-ESI) m/z calcd for C22H24N7O3 [M + H] + 434.1941, found434.1922.
4-(4-((4-((2,3-Dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenoxy)-N-hydroxybutanamide (9b). White solid. 55% yield.1H NMR (400 MHz, DMSO-d6) δ 10.40 (s, 1H), 8.99 (s, 1H), 8.69 (s, 1H), 7.80 (d, J = 6.0 Hz, 1H), 7.75 (d, J = 8.7 Hz, 1H), 7.59 (d, J = 8.9 Hz, 2H), 7.43 (d, J = 1.8 Hz, 1H), 6.87 (dd, J = 8.8, 1.8 Hz, 1H), 6.79 – 6.74 (m, 2H), 5.75 (d, J = 6.0 Hz, 1H), 4.06 (s, 3H), 3.89 (t, J = 6.3 Hz, 2H), 3.45 (s, 3H), 2.63 (s, 3H), 2.12 (t, J = 7.4 Hz, 2H), 1.91 (p, J = 6.7 Hz, 2H). 13C NMR (101 MHz, DMSO-d6) δ 169.17, 162.88, 159.52, 155.41, 153.55, 147.45, 142.41, 134.56, 132.61, 122.15, 120.86, 120.20, 119.94, 114.73, 114.39, 96.35, 67.44, 38.33, 37.84, 29.27, 25.42, 9.88. HRMS (AP-ESI) m/z calcd for C24H28N7O3 [M + H] + 462.2254, found 462.2276.
5-(4-((4-((2,3-Dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenoxy)-N-hydroxypentanamide (9c). White solid. 50% yield.1H NMR (400 MHz, DMSO-d6) δ 10.37 (s, 1H), 9.11 (s, 1H), 8.67 (s, 1H), 7.79 (d, J = 6.2 Hz, 1H), 7.76 (d, J = 8.8 Hz, 1H), 7.60 – 7.55 (m, 2H), 7.44 (d, J = 1.7 Hz, 1H), 6.88 (dd, J = 8.8, 1.8 Hz, 1H), 6.81 – 6.75 (m, 2H), 5.76 (d, J = 6.1 Hz, 1H), 4.06 (s, 3H), 3.90 (t, J = 5.9 Hz, 2H), 3.46 (s, 3H), 2.63 (s, 3H), 2.01 (t, J = 6.7 Hz, 2H), 1.71 – 1.59 (m, 4H). 13C NMR (101 MHz, DMSO-d6) δ 169.42, 162.85, 158.81, 154.27, 153.90, 147.41, 142.21, 134.03, 132.65, 122.23, 121.16, 120.08, 120.00, 114.73, 114.42, 96.42, 67.66, 38.45, 37.85, 32.41, 28.76, 22.30, 9.88.HRMS (AP-ESI) m/z calcd for C25H30N7O3 [M + H] + 476.2410, found 476.2489.
6-(4-((4-((2,3-Dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenoxy)-N-hydroxyhexanamide (9d). White solid. 47% yield. 1H NMR (400 MHz, DMSO-d6)δ 10.35 (s, 1H), 8.93 (s, 1H), 8.67 (s, 1H), 7.80 (d, J = 6.0 Hz, 1H), 7.75 (dd, J = 8.7, 0.8 Hz, 1H), 7.60 (d, J = 9.1 Hz, 2H), 7.42 (dd, J = 1.8, 0.8 Hz, 1H), 6.87 (dd, J = 8.8, 1.8 Hz, 1H), 6.75 (d, J = 9.1 Hz, 2H), 5.73 (d, J = 5.9 Hz, 1H), 4.06 (s, 3H), 3.87 (t, J = 6.4 Hz, 2H), 3.45 (s, 3H), 2.63 (s, 3H), 1.97 (t, J = 7.3 Hz, 2H), 1.68 (p, J = 6.6 Hz, 2H), 1.55 (p, J = 7.4 Hz, 2H), 1.43 – 1.33 (m, 2H).HRMS (AP-ESI) m/z calcd for C26H32N7O3 [M + H] + 490.2567, found 490.2517.
7-(4-((4-((2,3-Dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)phenoxy)-N-hydroxyheptanamide (9e). White solid53% yield. 1H NMR (400 MHz, DMSO-d6) δ 10.33 (s, 1H), 8.93 (s, 1H), 8.65 (s, 1H), 7.80 (d, J = 6.0 Hz, 1H), 7.75 (d, J = 8.7 Hz, 1H), 7.60 (d, J = 9.0 Hz, 2H), 7.44 – 7.41 (m, 1H), 6.87 (dd, J = 8.8, 1.7 Hz, 1H), 6.75 (d, J = 9.0 Hz, 2H), 5.73 (d, J = 5.9 Hz, 1H), 4.06 (s, 3H), 3.88 (t, J = 6.5 Hz, 2H), 3.45 (s, 3H), 2.63 (s, 3H), 1.99 – 1.91 (m, 2H), 1.67 (dt, J = 14.8, 6.8 Hz, 2H), 1.51 (dt, J = 14.8, 6.6 Hz, 2H), 1.39 (m, 2H), 1.34 – 1.27 (m, 2H). HRMS (AP-ESI) m/z calcd for C27H34N7O3 [M + H] + 504.2723, found 504.2746.
4-((4-((2,3-Dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)benzohydrazide (11).To a solution of 10 (0.40 g, 1.03 mmol) in dichloromethane (10 mL) in an ice bath, 2-(1H-benzotriazole-1-yl)-1,1,3,3-tetramethyluronium tetrafluoroborate (TBTU; 0.40 g, 1.24 mmol) was added, followed by Et3N (0.13 g,1.24 mmol). Thirty minutes later, hydrazine hydrate (0.06 g, 1.24 mmol) was added. Twelve hours later, the solution was diluted with water and extracted with dichloromethane. The combined organic extracts were washed with saturated NaHCO3 and brine and dried over Na2SO4 overnight, and the solvent was evaporated under vacuum. The crude product was purified by silica gel column chromatography (MeOH/CH2Cl2, 1/50 to 1/20) to affordCompound11 as a white solid (0.22 g, 52% yield). ESI–MS m/z: 403.21 [M + H] + .
4-((4-((2,3-Dimethyl-2H-indazol-6-yl)(methyl)amino)pyrimidin-2-yl)amino)-N'-propylbenzohydrazide (12). Compound 11 (0.60 g, 1.49 mmol) and propionaldehyde (0.10 g, 1.79 mmol) were added to 15 mL of anhydrous methanol, and then p-toluenesulfonic acid (0.025 g, 0.15 mmol) was added at room temperature. Eight hours later, the reaction solution was filtered and concentrated. The obtained residue was dissolved in 15 mL of anhydrous methanol, and NaBH3CN (0.14 g, 2.24 mmol) was added. The pH of the solution was adjusted to 5 with concentrated HCl/MeOH (v:v = 1:1). Twelve hours later, the pH of the solution was adjusted to 8 with saturated NaHCO3. The organic phase was collected and evaporated, and the residual was extracted with ethyl acetate. The combined organic extracts were washed with saturated NaHCO3 and brine and dried over Na2SO4 overnight, and the solvent was evaporated under vacuum. The crude product was purified by silica gel column chromatography (MeOH/CH2Cl2, 1/100 to 1/45) to afford Compound 12 asa white solid (0.21 g, 31% yield). 1H NMR (400 MHz, DMSO-d6) δ 9.81 (s, 1H), 9.47 (s, 1H), 7.90 (d, J = 6.0 Hz, 1H), 7.82 (d, J = 8.5 Hz, 2H), 7.77 (d, J = 8.8 Hz, 1H), 7.68 (d, J = 8.5 Hz, 2H), 7.47 (d, J = 1.7 Hz, 1H), 6.90 (dd, J = 8.8, 1.7 Hz, 1H), 5.86 (d, J = 6.0 Hz, 1H), 5.10 (s, 1H) 4.07 (s, 3H), 3.50 (s, 3H), 2.74 (t, J = 7.1 Hz, 2H), 2.64 (s, 3H), 1.47 (q, J = 7.3 Hz, 2H), 0.92 (t, J = 7.4 Hz, 3H)0.13C NMR (101 MHz, DMSO-d6) δ 165.69, 162.90, 159.59, 156.18, 147.48, 144.41, 142.38, 132.62, 128.07, 125.25, 122.20, 120.20, 119.97, 117.84, 114.43, 97.38, 53.72, 38.44, 37.83, 21.32, 12.15, 9.88. HRMS (AP-ESI) m/z calcd for C24H29N8O1 [M + H] + 445.2464, found 445.2478.
In vitro HDAC inhibition assay
In vitro HDAC inhibition assays were conducted according to reported methods [12]. Briefly, 10 μL of enzyme solution (HDAC1, HDAC4, HDAC6 or HDAC11) was mixed with different concentrations of the tested compounds (50 μL). The mixture was incubated at 37 °C for 5 min, followed by the addition of 40 μL of fluorogenic substrate (Boc-Lys(acetyl)-AMC for HDAC1 and HDAC6, Boc-Lys(trifluoroacetyl)-AMC for HDAC4, and Ac-Leu-GlyLys(Ac)-AMC for HDAC11). After incubation at 37 °C for 30 min, the mixture was quenched by adding 100 μL of developer containing trichostatin A (TSA) and trypsin. After another 20 min of incubation at 37 °C, the fluorescence intensity was measured using a microplate reader at excitation and emission wavelengths of 390 and 460 nm, respectively. The inhibition ratios were calculated from the fluorescence intensity readings of the test wells relative to those of the control wells, and the IC50 values were calculated using nonlinear regression with a normalized dose–response curve using GraphPad Prism software.
In vitro VEGFR inhibition assay
TheVEGFR1, VEGFR2 and VEGFR3 inhibitory activities were measured using a Kinase-GloTM Luminescent Kinase Assay from HUAWEI PHARMA (Ji’nan, China). In brief, the tested compounds, kinases, substrate, and ATP were diluted in kinase buffer to the indicated concentrations, added to the assay plate and incubated at room temperature for 40 min. Then, Kinase-Glo reagent was added. After an additional 15 min of incubation, the luminescence was measured with a microplate reader (SpectraMax M5). The IC50values were calculated using nonlinear regression with a normalized dose–response curve using GraphPad Prism software.
In vitro antiproliferation assay
All cell lines were maintained in RPMI 1640 medium containing 10% FBS at 37 °C in a 5% CO2 humidified incubator. Antiproliferation was determined by the MTT (3- [4,5-dimethyl-2-thiazolyl]-2,5-diphenyl-2H-tetrazolium bromide) method. Briefly, cells were passaged the day before seeding into a 96-well plate, allowed to grow for 12 h, and then treated with different concentrations of compound for 48 h. A 0.5% MTT solution was added to each well. After incubation for another 4 h, the formazan formed from MTT was extracted by adding 200 µL of DMSO. The absorbance was then determined using an ELISA reader at 570 nm.
HUVEC tube formation assay
The HUVEC tube formation assay was conducted according to reported methods [12]. Briefly, Matrigel (100 μL; BD Biosciences, NJ) was added to the test wells of 96-well plates and then allowed to polymerize for 0.5 h at 37 °C. HUVECs were trypsinized and seeded at a density of 40,000 per well in M199 (5% FBS) containing DMSO or test compounds for 6 h at 37 °C in a CO2 incubator. Morphological changes in cell and tube formation were observed under a phase-contrast microscope (OLYMPUS IX51) and photographed at 200 × magnification. Experiments were repeated at least two times.
Rat thoracic aorta ring (TAR) assay
The TAR assay was conducted according to previously reported methods [12]. Briefly, Matrigel (100 μL; BD Biosciences, NJ) was added to the test wells of 96-well plates and then allowed to polymerize for 0.5 h at 37 °C. Sprague–Dawley rats (4 to 6 weeks old) were sacrificed, and the aortas were harvested. Each aorta was cut into 1-mm slices and embedded in an additional 100 μL of Matrigel in 96-well plates. After that, the rings were incubated for 30 min at 37 °C with 5% CO2. Aortic rings were treated with vehicle or the test compounds each day for 6 days and photographed on the 7th day at 200 × magnification. Experiments were repeated at least two times.
Western blot analysis
A549 or HUVECs were treated with compounds or DMSO for a specified period of time. Then, the cells were washed twice with cold PBS and lysed in ice-cold RIPA buffer. Lysates were cleared by centrifugation. Protein concentrations were determined using the BCA assay. Equal amounts of cell extracts were then resolved by SDS–PAGE, transferred to nitrocellulose membranes and probed with an ac-histone H4 antibody, an ac-α-tubulin antibody, a β-actin antibody, a phosphorylated VEGFR2 antibody or a total VEGFR-2 antibody. Blots were imaged using an enhanced chemiluminescence system.
In vitro liver microsomal stability assay
Mouse liver microsomes containing the test compounds were incubated with NADPH at 37 °C. At specific time points, acetonitrile was added to the samples to terminate the reaction, and then the samples were subjected to vortex mixing for 5 min and stored in a freezer at -80 °C. Before analysis, the samples were centrifuged at 4000 rpm for 15 min. The amount of remaining test compounds in the supernatants were analyzed by LC–MS/MS. The t1/2 values were calculated using the Equation t1/2 = -0.693/k, where k is the slope of the linear fit of the natural logarithm of the remaining fraction of test compounds vs. incubation time.
Results and discussion
Compound design and synthesis
Compounds5, 7, and 9a-9e were designed by replacing the amide-connecting unit of ZYJ-13f with a reverse amide, secondary amine, and ether, respectively (Fig. 2). In addition, the N-acyl o-diaminobenzene zinc binding group of JMC-6d was changed to a hydrazide due to the good metabolic stability of hydrazide [13], leading to Compound 12 (Fig. 2).
The procedures to synthesize the target Compounds 5,7 and 9a-9e are outlined in Scheme 1. Compound 1, which was obtained according to previous methods [12], was treated with 4-aminophenol and benzene-1,4-diamine to produce key intermediates 2and 3, respectively. Intermediate 3 reacted with suberic acid monomethyl ester by TBTU-mediated amide condensation to afford intermediate 4, which was transformed into hydroxamic acid5.In addition, intermediate 3 could also react with methyl 6-bromohexanoate by nucleophilic substitution to obtainCompound 6, which could be transformed into hydroxamic acid7.Intermediate 2 reacted with various methyl ω-bromoalkanoates by nucleophilic substitution to obtainCompounds 8a-8e, which were further converted into target hydroxamic acids 9a-9e.
The procedures to synthesize the target Compound 12 are outlined in Scheme 2. Compound10, obtained according to previous methods [12], was reacted with hydrazine hydrate by TBTU-mediated amide formation to afford intermediate 11. Then, reductive amination of 11 afforded target hydrazide12.
In vitro HDAC and VEGFR inhibition assay
The HDAC and VEGFR inhibitory potencies of all target compounds were preliminarily tested by determining the HDAC1, HDAC6 and VEGFR2 inhibition rates at 0.5 μM. The approved pan-HDAC inhibitor SAHA, the clinical class I selective HDAC inhibitor MS275, and the approved VEGFR inhibitor pazopanib were used as the positive controls. The results in Table 1 revealed that most of the target compounds could effectively inhibit all three enzymes with inhibition rates greater than 50% at 0.5 μM.Generally, hydroxamates with linkers containing more thanthree methylenes (5, 7, 9c, 9d, 9e) were more potent HDAC inhibitors than compounds with shorter linkers (9a, 9b).These results indicatedthat hydrazide Compound 12 exhibited selective HDAC1 inhibition over HDAC6, which was similar to the positive control MS275.
In vitro antiproliferation assay
Considering their promising HDAC inhibitory activities, Compounds 5, 7 and 9c, 9d, 9e and 12 were further tested in antiproliferative assays against five solid tumor cell lines. The results in Table 2 show that Compounds 5, 9e and 12 were the three most potent compounds, each with IC50 values lower than 5 μM against all tested cancer cell lines. Remarkably, the overall antiproliferative activities of 5, 9e and 12 were even more potent than those of the two clinical HDAC inhibitors SAHA and MS275. Consistent with previously reported results [12], the VEGFR inhibitor pazopanib possessed negligible cytotoxicity.
In vitro HUVEC tube formation assay
Compounds 5, 9e and 12 were subjected to anin vitroHUVEC tube formation assay to evaluate their antiangiogenic activities. The test concentration of compounds was set to 0.5 μM, which is lower than their antiproliferative IC50 values, to avoid cytotoxicity to HUVECs.It was demonstrated that 5, 9e and 12 could significantly inhibit HUVEC tube formation, which was similar to the positive control pazopanib (Fig. 3).
Ex vivo rat thoracic aorta ring (TAR) assay
An ex vivo rat TAR assay was carried out to further validate the antiangiogenic activities of Compounds 5, 9e and 12, and the results clearly showed that Compounds 5, 9e and 12as well as pazopanib could almost completely inhibit microvessel outgrowth (Fig. 4).
HDAC and VEGFR selectivity profiling
The Zn2+-dependent HDAC family contains 11 isoforms, which can be categorized into class I (HDACs 1, 2, 3, and 8), class IIa (HDACs 4, 5, 7, and 9), class IIb (HDACs 6 and 10) and class IV (HDAC11) [14]. To profile the selectivity of our dual HDAC and VEGFR inhibitors, the IC50 values of Compounds5, 9e and 12 against HDAC1, HDAC4, HDAC6 and HDAC11 were determined with SAHA and MS275 as the positive controls (Table 3). Compared with SAHA, hydroxamates 5 and 9e showed comparable or even slightly better inhibitory activity against HDAC1 and HDAC6, the representative isoforms of class I and class IIb HDAC, respectively. Similar to SAHA,hydroxamates 5 and 9e were not potent class IIa (isoform HDAC4) and class IV (isoform HDAC11) inhibitors. Similar to MS275, hydrazide 12 exhibited high selectivity for HDAC1 over the other tested isoforms, indicating class I selectivity. Moreover, Compounds 5, 9e and 12 were tested against VEGFR1, VEGFR2 and VEGFR3 to profile their VEGFR isoform selectivity.In addition to pazopanib, Compounds 5, 9e and 12 exhibited potent pan-VEGFR inhibition with no significant discrimination between the VEGFR family members (Table 3).
Western blot analysis
Western blot analysis was performed to validate the intracellular target engagement of Compounds 5 and 12. The results showed that both 5 and 12 could remarkably increase the levels of acetyl-histone H4 (Ac-HH4), which is the intracellular substrate of class I HDACs (Fig. 5A). In addition, Compound 5 could increase the levels of acetyl-α-tubulin(Ac-Tub), the substrate of HDAC6, while Compound 12 showed no effects on Ac-Tub. These results were consistent with their HDAC isoform selectivity presented in Table 3. The inhibition of intracellular VEGFR by Compounds12 and 5 was confirmed by the decreased levels of phosphorylated VEGFR2 (p-VEGFR2) in HUVECs (Fig. 5B).
A. A549 cells were treated with DMSO or compounds(2 μM) for 5 h. The levels of the indicated proteins were determined by immunoblotting. β-Actin was used as a loading control. B. HUVECs were treated with DMSO or compounds (0.5 μM) for 2 h and then stimulated with VEGF (50 ng/ml). The levels of p-VEGFR2 were determined by immunoblotting. β-Actin and total VEGFR-2 were used as loading controls
In vitro liver microsomal stability assay
Considering their promising in vitro activities, the metabolic stabilities of Compounds 5, 9e and 12 in mouse liver microsomeswere determined and compared with that of pazopanib. Unfortunately, no compounds possessed superior metabolic stability relative to pazopanib (Table 4).
Conclusion
A novel series of pazopanib analogs were developed as dual HDAC and VEGFR inhibitors. Compared with pazopanib, many of the new dual HDAC and VEGFR inhibitors exhibited superior cytotoxicity against multiple solid tumor cell lines, which could be ascribed to their potent HDAC inhibition. Moreover, Compounds 5, 9e and 12 exhibited uncompromised VEGFR inhibitory activity and antiangiogenic capacity relative to pazopanib. Subsequent work should be focused on structural optimization of these analogs to improve their pharmacokinetic properties, which will lead to multitargeted compounds within vivo antitumor activity.
Availability of data and materials
All data generated or analyzed during this study are included in this published article.
References
Bates SE (2020) Epigenetic Therapies for Cancer. N Engl J Med 383:650–663
Zagni C, Floresta G, Monciino G, Rescifina A (2017) The search for potent, small-molecule HDACIs in cancer treatment: a decade after Vorinostat. Med Res Rev 37:1373–1428
Ali I, Conrad RJ, Verdin E, Ott M (2018) Lysine Acetylation Goes Global: From Epigenetics to Metabolism and Therapeutics. Chem Rev 118:1216–1252
Wu P, Nielsen TE, Clausen MH (2015) FDA-approved small-molecule kinase inhibitors.Trends Pharmacol Sci 36:422–439
Musumeci F, Radi M, Brullo C, Schenone S (2012) Vascular endothelial growth factor (VEGF) receptors: drugs and new inhibitors. J Med Chem 55:10797–10822
Choueiri TK, Escudier B, Powles T, Mainwaring PN, Rini BI, Donskov F, Hammers H, Hutson TE, Lee JL, Peltola K, Roth BJ, Bjarnason GA, Géczi L, Keam B, Maroto P, Heng DY, Schmidinger M, Kantoff PW, Borgman-Hagey A, Hessel C, Scheffold C, Schwab GM, Tannir NM, Motzer RJ, Investigators METEOR (2015) Cabozantinib versus Everolimus in Advanced Renal-Cell Carcinoma. N Engl J Med 373:1814–1823
Vyse S, McCarthy F, Broncel M, Paul A, Wong JP, Bhamra A, Huang PH (2018) Quantitative phosphoproteomic analysis of acquired cancer drug resistance to pazopanib and dasatinib. J Proteomics 170:130–140
Chan D, Zheng Y, Tyner JW, Chng WJ, Chien WW, Gery S, Leong G, Braunstein GD, Koeffler HP (2013) Belinostat and panobinostat (HDACI): in vitro and in vivo studies in thyroid cancer. J Cancer Res Clin Oncol 139:1507–1514
Booth L, Roberts JL, Sander C, Lee J, Kirkwood JM, Poklepovic A, Dent P (2017) The HDAC inhibitor AR42 interacts with pazopanib to kill trametinib/dabrafenib-resistant melanoma cells in vitro and in vivo. Oncotarget 8:16367–16386
Tavallai S, Hamed HA, Grant S, Poklepovic A, Dent P (2014) Pazopanib and HDAC inhibitors interact to kill sarcoma cells. Cancer Biol Ther 15:578–585
Aggarwal R, Thomas S, Pawlowska N, Bartelink I, Grabowsky J, Jahan T, Cripps A, Harb A, Leng J, Reinert A, Mastroserio I, Truong TG, Ryan CJ, Munster PN (2017) Inhibiting Histone Deacetylase as a Means to Reverse Resistance to Angiogenesis Inhibitors: Phase I Study of Abexinostat Plus Pazopanib in Advanced Solid Tumor Malignancies. J Clin Oncol 35(11):1231–1239. https://doi.org/10.1200/JCO.2016.70.5350
Zang J, Liang X, Huang Y, Jia Y, Li X, Xu W, Chou CJ, Zhang Y (2018) Discovery of Novel Pazopanib-Based HDAC and VEGFR Dual Inhibitors Targeting Cancer Epigenetics and Angiogenesis Simultaneously. J Med Chem 61:5304–5322
McClure JJ, Zhang C, Inks ES, Peterson YK, Li J, Chou CJ (2016) Development of Allosteric Hydrazide-Containing Class I Histone Deacetylase Inhibitors for Use in Acute Myeloid Leukemia. J Med Chem 59:9942–9959
Yoshida M, Kudo N, Kosono S, Ito A (2017) Chemical and structural biology of protein lysine deacetylases. Proc Jpn Acad Ser B Phys Biol Sci 93:297–321
Acknowledgements
Not applicable.
Funding
This study was funded by a grant from the National Natural Science Foundation of China (grant no. 82073872).This study was also supported by the Natural Science Foundation of Shandong Province (ZR2020MH001).
Author information
Authors and Affiliations
Contributions
XX and ZXG designed the project. LLN and LY performed the enzymatic screening.LYX, ZYJ and ZJ synthesized the molecules. WYJ, SDQ and LY, JW performed the in vitro experiments. XX analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Xue, X., Zhang, Y., Liao, Y. et al. Design, synthesis and biological evaluation of dual HDAC and VEGFR inhibitors as multitargeted anticancer agents. Invest New Drugs 40, 10–20 (2022). https://doi.org/10.1007/s10637-021-01169-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10637-021-01169-4