Summary
Transcription factor Specificity protein 1 (Sp1) and its downstream target survivin (inhibitor of apoptosis protein), play major roles in the pathogenesis of various cancers. Ewing Sarcoma (ES) is a common soft tissue/bone tumor in adolescent and young adults. Overexpression of survivin is also linked to the aggressiveness and poor prognosis of ES. Small molecule Tolfenamic acid (TA) inhibits Sp1 and survivin in cancer cells. In this investigation, we demonstrate a strategy to target Sp1 and survivin using TA and positive control Mithramycin A (Mit) to inhibit ES cell growth. Knock down of Sp1 using small interfering RNA (siRNA) resulted in significant (p < 0.05) inhibition of CHLA-9 and TC-32 cell growth as assessed by CellTiter-Glo assay kit. TA or Mit treatment caused dose/time-dependent inhibition of cell viability, and this inhibition was correlated with a decrease in Sp1 and survivin protein levels in ES cells. Quantitative PCR results showed that Mit treatment decreased the mRNA expression of both survivin and Sp1, whereas TA diminished only survivin but not Sp1. Proteasome inhibitor restored TA-induced inhibition of Sp1 protein expression suggesting that TA might cause proteasome-dependent degradation. Gel shift assay using ES cell nuclear extract and biotinylated Sp1 consensus oligonucleotides confirmed that both TA and Mit decreased DNA-binding activity of Sp1. These results demonstrate that both Mit and TA reduce expression of Sp1 and survivin, disrupt Sp1 DNA-binding and inhibit ES cell proliferation. This investigation suggests that targeting Sp1 and survivin could be an effective strategy for inhibiting ES cell growth.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sarcomas are cancer arising from tissues of mesenchymal origin. According to the American Cancer Society’s estimation, approximately 12,000 cases and 5000 deaths occur annually in the United States due to these malignancies. Currently, more than 50 distinct subtypes of sarcomas are identified based on morphological, immunohistochemical and molecular characteristics [1–3]. Ewing sarcoma (ES) is the second most common bone and soft tissue sarcoma in the young and adolescent population. Approximately 30% of ES cases present with metastatic disease at diagnosis. The most common sites for this malignancy are the bones of lower extremity, pelvis, and chest wall (ribs) [1, 4]. The characteristic feature of ES is the presence of EWS-FLI1, a chimeric protein formed by the translocation between EWS and FLI1, in approximately 85% of cases [5], while 10–15% of ES cases harbor variants of translocation between EWS and ERG genes [6].
The introduction of chemotherapeutic agents in the 1960s was a paradigm shift in cancer treatment. However, chemotherapy used for ES is often associated with organ-specific side effects such as hormone imbalance and cardiotoxicity. Even though EWS-FLI1 is known to be associated with ES, only limited progress has been made in the field of drug development tailored towards targeting this protein. Therefore, there is a need to identify alternate targets and less toxic therapeutic agents. Upregulation of transcription factors, proteins that regulate gene expression and control cellular homeostasis, is implicated in several cancers, including sarcomas [7]. Specificity protein 1 (Sp1) is a ubiquitously expressed transcription factor which is critical in the regulation of cell cycle progression, cell proliferation, and apoptosis [8]. Sp1 is considered to be a prognostic factor for some cancers and targeting Sp1 can provide a novel mechanism for cancer treatment [9–11].
Sp1 regulates the expression of several genes associated with cancer including survivin (Baculoviral inhibitor of apoptosis repeat containing 5) a member of the Inhibitor of Apoptosis Protein family. Survivin regulates apoptosis and cell cycle in cancer cells. Overexpression of survivin has been linked to aggressiveness and poor prognosis in several cancers including sarcomas [12], and its inhibition is reported to sensitize ES cells to radiation treatment [13]. Disruption of survivin using siRNA has been shown to be effective in reducing tumor growth in pre-clinical models [14–16]. Therefore, it is believed that targeting survivin provides a promising strategy for cancer treatment.
Mithramycin A (Mit) is an anti-neoplastic antibiotic isolated from Streptomyces plicatus. Mit has been shown to bind with GC-rich DNA and inhibit transcription factors Sp1 and Sp3 and induce anti-cancer activity [17]. It is also evident that Mit downregulates survivin via modulating Sp transcription factors [18]. Several preclinical studies demonstrated similar activity of an anti-cancer non-steroidal anti-inflammatory drug (NSAID), Tolfenamic acid (TA), to target Sp proteins and survivin. We have previously investigated the anti-proliferative effect of TA in pediatric cancer models. These studies demonstrated that TA inhibits medulloblastoma and neuroblastoma cell growth by targeting Sp1 and survivin [19, 20]. TA has not been tested in preclinical/clinical models for human sarcomas.
In this study, we examined the anti-proliferative activity of TA in ES cells and the association of Sp1 and survivin in its anti-cancer activity using Mit as a positive control. Initially, multiple ES cell lines were screened for the expression of Sp1 and survivin. Two selected ES cell lines were used for all subsequent assays. Knocking down of Sp1 using small interference RNA (siRNA) was used to test the significance of Sp1 in ES cell growth. Both transcriptional and post-transcriptional modifications were tested by evaluating mRNA expression and proteasome-dependent degradation using a proteasome inhibitor, Lactacystin, to understand the inhibitory effect of TA on Sp1 protein expression. The ability of Mit and TA to disrupt the Sp1 DNA-binding was also assessed using gel shift assay. This study tests the hypothesis that both Sp1 and survivin could serve as reliable targets for inhibiting ES cell growth and identifies TA as a promising candidate to target Sp1 and survivin in ES cells.
Material and methods
Cell lines and cell culture
ES cell lines were obtained from the bio-repository of Children’s Oncology Group (Lubbock, TX). CHLA-9, CHLA-10, TC-71, TC-32, CHLA-32, and CHLA-99 cells were grown in Iscove’s Modified Dulbecco’s Media (supplemented with 4 mM L-Glutamine, 5 μg/mL Insulin, 5 μg/mL Transferrin and 5 ng/mL Selenous Acid) and fetal bovine serum. Cells were maintained at 37 °C and 5% CO2.
Chemicals and reagents
TA, Mit, dimethyl sulfoxide (DMSO), Lactacystin, and Anti-actin antibody were purchased from Sigma-Aldrich (St. Louis, MO). Sp1 antibody was obtained from Santa Cruz Biotechnology (Santa Cruz, CA), and Survivin was purchased from R & D Systems (Minneapolis, MN). Dulbecco’s phosphate-buffered saline was procured from Hyclone Laboratories (Logan, Utah). ITS premix was purchased from Corning (Bedford, MA). Cell Titer-Glo kit was obtained from Promega (Madison, WI). Lipofectamine RNAi MAX reagent and Cell lysis buffer were procured from Invitrogen (Carlsbad, CA). Bicinchoninic acid (BCA) protein assay kit and Supersignal West Dura were purchased from Pierce (Rockford, IL).
Cell viability assay
Viability of CHLA-9 and TC-32 cells was assessed using CellTiter-Glo kit (Promega, Madison). Briefly, cells were plated in 96-well plates and treated with DMSO (Control) or TA or Mit. After 24 or 48 h post-treatment, CellTiter-Glo reagent was added and luminescence was measured as per the manufacturer’s instructions using SYNERGY HT microplate reader (BioTek Instruments, Inc., Winooski, VT).
Western immunoblotting
CHLA-9 and TC-32 cells were plated in 10 cm culture dish and treated with DMSO or TA (10 or 15 μg/ml) or Mit (15 nM). After 24 or 48 h, cells were harvested, protein extracts were prepared, and Western blot analysis was performed as described earlier [20].
Small interfering RNA (siRNA) transfection
CHLA-9 and TC-32 cells were transfected with 25 pmol of siRNA1 (catalogue no: J-026959-07) and siRNA2 (J-026959-08) (Dharmacon, Lafayette, CO) that target Sp1, using Lipofectamine RNAi MAX reagent. Cells were harvested; 48 h post-transfection and whole cell lysates were prepared. Expression of Sp1 protein was determined using western immunoblot analysis. Cell viability was measured with CellTiter-Glo assay kit as described above.
Quantitative polymerase chain reaction (qPCR)
CHLA-9 and TC-32 were treated with DMSO (Control) or TA (15 μg/ml) or Mit (100 nM). Cells were harvested at 12 and 24 h post-treatment. RNA was extracted using TRIzol RNA extraction protocol (Invitrogen, Carlsbad, CA), and cDNA was synthesized using Superscript III reverse transcriptase (Invitrogen, Carlsbad, CA). The mRNA expression was evaluated by Roche thermal cycler using TaqMan primer-probes specific for Sp1 and Survivin purchased from Thermo Fisher (Waltham, MA).
Proteasome inhibition assay
CHLA-9 and TC-32 cells were treated with two doses of TA (10 μg/ml and 15 μg/ml) in the presence of 2 μM Lactacystin (proteasome inhibitor). Cells were harvested 48 h post-treatment. Whole cell lysates were prepared and the expression of Sp1 and β-Actin (loading control) was determined by Western blot analysis as described above.
DNA binding gel-shift assay
Sp1 oligonucleotide [5 ́-ATTCGATCGGGGCGGGGCGAGC-3 ́(biotin); (biotin)3 ́ TAAGCTAGCCCCGCCCCGCTCG-5′ IDT ID: 129,054,347] were obtained from Integrated DNA Technologies (Coralville, IA) and annealed to each other in annealing buffer. Binding reaction was carried out with TC-32 nuclear extract, binding buffer (10 mM Tris, 50 μM KCl and 10 nM DTT pH 7.5), and Sp1 oligo (biotin labelled) in the presence of DMSO (Control) or TA (50 μM) or Mit (50 μM) or positive control (100 μM of EDTA). Unlabeled oligo was used as a competitor. Sp1 oligo-protein complex was resolved on 6% polyacrylamide gel at 100 V and transferred to positively charged 0.2 μm nylon membrane using iBlot DNA transfer stacks (Invitrogen, Carlsbad, CA). The membranes were developed with Chemiluminescent Nucleic Acid Detection Module Kit (Thermo Fisher, Waltham, MA), and the signal detected using UVP GelDoc imager.
Statistical analysis
All experiments were performed in triplicates and the data were presented as mean ± SEM (standard error of mean). Data were analyzed using Graphpad Prism software V6.0 (La Jolla, California, USA) following standard methods (student t-test or one-way ANOVA; p-value <0.05 considered as statistically significant).
Results
ES cell lines express Sp1 and survivin
The expression of key markers (Sp1, survivin) in ES cells was determined to understand their association with this malignancy. Several (six) ES cell lines were screened by western immunoblot to examine the expression of Sp1 and survivin. All tested cell lines expressed Sp1 and survivin (Fig. 1). Interestingly, the expression of Sp1 and survivin show similar patterns suggesting a correlation between these candidates.
Sp1 knockdown inhibits cell viability of ES cells
In order to study the effect of Sp1 inhibition on ES cell growth, we performed siRNA knockdown of Sp1 transcription factor in ES cells and measured the cell viability. Our results showed that both siRNAs (siRNA1 and siRNA2) inhibited the expression of Sp1 protein which correlated with a decrease in cell viability in both CHLA-9 and TC-32 cell lines (Fig. 2).
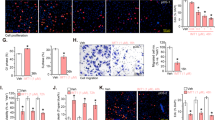
Sp1 silencing using siRNA inhibits Sp1 protein expression and decreases cell viability in ES cells. (a) CHLA-9 and (b) TC-32 cells were transfected with siRNA 1 and siRNA 2 specific for Sp1. After 48 h cell viability assay was performed using CellTiter Glo reagent. One-way ANOVA test was performed for statistical analysis. Bars are the mean of percentage of viable cells ± SEM of three independent replicates. “*” represent p-value <0.05. (c) CHLA-9 and (d) TC-32 cells were transfected with two different Sp1 siRNAs. 48 h post-transfection Sp1 protein expression was determined by western immunoblotting. β-Actin was used as a loading control
TA and Mit treatment inhibit cell viability of ES cells
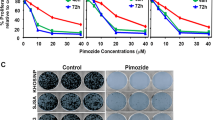
The effect of TA and Mit on the proliferation of CHLA-9 and TC-32 cells was evaluated. Cells were treated with TA (0–30 μg/ml) or Mit (0–80 nM) and cell viability was measured at 24 & 48 h posttreatment. Both Mit and TA significantly inhibited cell viability following a clear dose/time-dependent pattern in both CHLA-9 and TC-32 cells. The IC50 values at 48 h were ranging from 7 to 13 nM (CHLA-9: 12.92 nM; TC-32: 7.21 nM) for Mit, and 11–15 μg/ml (CHLA-9: 11.26 μg/ml; TC-32: 14.79 μg/ml) for TA as calculated by nonlinear curve fitting (Fig. 3).
TA and Mit inhibited cell viability of ES cells. (a) CHLA-9 and (b) TC-32 cells were treated with DMSO (Control) or 0–30 μg/ml) of TA. (c) CHLA-9 and (d) TC-32 cells were treated with DMSO or 0–80 nM of Mit. Cell viability was assessed at 24 and 48 h post-treatment using CellTiter Glo assay kit. Dose-response curves were plotted as log(inhibitor) vs. normalized response -- variable slope using non-linear regression
TA and Mit inhibits Sp1 and survivin protein expression
CHLA-9 and TC-32 cells were treated with vehicle (DMSO) or Mit (15 nM) or TA (15 μg/ml) and cells were harvested at 24 and 48 h post-treatment. Sp1 and survivin protein expression was determined by western immunoblot analysis. TA and Mit decreased expression of Sp1 and survivin in both (tested) cell lines (Fig. 4).
TA and Mit treatment decreased Sp1 & survivin protein expression. (a) CHLA-9 and (b) TC-32 cells were treated with DMSO or TA (15 μg/ml) or Mit (15 nM). At 24 and 48 h posttreatment cells were harvested and Sp1 & survivin protein expression was assayed (Western immunoblot analysis). β-Actin was a loading control
Effect of TA and Mit on Sp1 and survivin mRNA expression
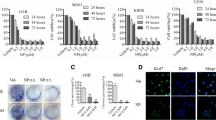
Levels of Sp1 and survivin mRNA were evaluated in CHLA-9 and TC-32 cells following 12 and 24 h treatment with Mit (100 nM) or TA (15 μg/ml) using qPCR. Mit treatment significantly decreased Sp1 and survivin mRNA expression in both cell lines at 12 h post-treatment (Fig. 5a & b). TA inhibited survivin mRNA expression at 24 h post-treatment; however, it did not change Sp1 mRNA expression (Fig. 5c & d).
Effect of TA and Mit treatment on Sp1 and survivin mRNA. (a, c) CHLA-9 and (b, d) TC-32 cells were treated with DMSO, TA or Mit. Total RNA was isolated with TRIzol RNA extraction protocol. Sp1 and survivin mRNA expression was measured using qPCR (data was normalized to GAPDH). Student t-test was performed for statistical analysis. Bars (mean ± SEM) of three independent replicates. “*” represent p value <0.05
TA induces proteasome-dependent Sp1 degradation
Previously it was reported that TA induced the activation of proteasome dependent degradation of Sp1 in pancreatic cancer cell lines [21] but not in esophageal cancer cells [22]. In this study we investigated whether TA treatment induces the activation of proteasome-mediated Sp1 degradation in ES cells. CHLA-9 and TC-32 cells were treated with vehicle (DMSO) or TA in the presence or absence of Lactacystin, a proteasome inhibitor. Cells were harvested after 48 h and whole cell lysates were prepared. The expression of Sp1 protein was analyzed by Western immunoblot. Results revealed that TA-induced inhibition of Sp1 protein expression was blocked following co-treatment with Lactacystin (Fig. 6).
Lactacystin blocked TA induced proteasome-dependent degradation of Sp1 protein. (a) CHLA-9 and (b) TC-32 cells were treated with DMSO or TA (15 μg/ml) or Lactacystin (2 μM) + TA (15 μg/ml). Cells were harvested at 48 h post-treatment, and Sp1 and β-Actin (loading control) protein expression was analyzed by western immunoblot
TA and Mit disrupted the DNA-binding activity of Sp1
Since Sp1 is a transcription factor, its ability to bind to DNA is critical for its functional activity. We tested the ability of Sp1 to bind to the Sp1 consensus oligo, in the presence of TA or Mit, by gel shift assay. ES cell nuclear extract was used as the source for Sp1 protein. The gel shift assay results demonstrated that both Mit and TA inhibited the Sp1 DNA-binding, which was consistent with the positive control (EDTA) (Fig. 7, lane 3, 4, & 5). The specificity of Sp1 DNA-binding was confirmed by the competition assay, where the addition of excess unlabeled Sp1 oligo competed with biotin-labeled oligo and completely knocked down the binding of Sp1 (from ES cell nuclear extract) to biotin labeled oligo (Fig. 7, lane 6).
TA and Mit inhibited DNA-binding activity of Sp1 transcription factor. Sp1 DNA-binding assay was performed with TC-32 nuclear extract and biotin labeled Sp1 oligo. Nuclear extracts were incubated with TA (50 μM), or Mit (50 μM), or EDTA (100 μM), or 10X concentration of unlabeled Sp1 oligo (competitor), and gel shift assay was performed to examine perturbations in the Sp1 DNA-binding
Discussion
Sp1 is one of the first identified mammalian transcription factors. It regulates critical cellular processes such as cell cycle progression and apoptosis and impacts cell proliferation. Studies have also demonstrated the regulation of cell proliferation and apoptosis by Sp1 via modulating the expression of survivin, vascular endothelial growth factor, and cyclin D1 [23, 24]. Although Sp1 is expressed in ES and viewed as a “hallmark of cancer”, the strategies to target Sp1 are still lacking [9]. High expression of survivin is associated with aggressiveness and poor prognosis in various cancer types including ES [12, 25]. In this study we investigated the anti-proliferative activity of TA and Mit in association with their efficacy to target Sp1 and survivin in ES cells.
Sp1 and survivin are considered as prognostic markers in several malignancies [9–16], and recently, the association of these candidates in ES is under investigation [26, 27]. In this study, six ES cell lines obtained from COG were screened to evaluate the presence of Sp1 and survivin. All six cell lines showed moderate to high levels of expression for both Sp1 and survivin (Fig. 1). Silencing of Sp1 in CHLA-9 and TC-32 cells using siRNA technology resulted in significant decrease in cell viability correlating with the inhibition of Sp1 protein levels (Fig. 2). These results strongly suggest that inhibition of Sp1 can induce an anti-proliferative response in ES cells. Mit is an antibiotic with antineoplastic properties which was shown to target Sp proteins and inhibit cell proliferation and tumor growth in some pre-clinical models [28–30]. TA also showed similar effects and inhibited cancer cell growth by inhibiting Sp1 and survivin protein expression in preclinical studies [21, 31].
TA and Mit treatment caused a time/dose-dependent decrease in cell viability (Fig. 3) that was accompanied by a decrease in Sp1 and survivin protein as determined by western immunoblot (Fig. 4). Both TA and Mit have been shown to inhibit Sp1 protein expression in several cancers. Hence, the growth inhibition of ES cells by TA and Mit could be attributed to the effect of these agents on Sp1 and survivin. We also investigated the effect of Mit and TA on mRNA expression of Sp1 & survivin. Both TA and Mit treatment caused significant inhibition of survivin mRNA expression (Fig. 5). Mit significantly decreased the mRNA expression of Sp1 (Fig. 5a & b), however, TA did not cause any change in Sp1 expression levels (Fig. 5c & d), suggesting involvement of post-translational effects. Lactacystin, a proteasome inhibitor, binds and blocks the 20S subunit of proteasome complex. In this study, Lactacystin blocked TA-induced Sp1 inhibition (Fig. 6) confirming the proteasome-dependent degradation of Sp1 by TA. These results are in agreement with an earlier study which demonstrated that TA caused proteasome-dependent degradation of Sp1 in pancreatic cancer cell lines [21].
Gel shift assay revealed that the both Mit and TA caused perturbations in DNA-binding activity of Sp1 (Fig. 7). It is possible that TA could be working as a chelating agent disrupting the zinc finger motif thereby decreasing the DNA-binding activity of Sp1 to its consensus sequence (Fig. 7). Therefore, Mit and TA could affect the activity of Sp1 and potentially impact the expression of genes that are regulated by Sp1.
Survivin is implicated in the pathogenesis of several cancers including sarcomas. Sp1 transcription factor regulates survivin and is known to be associated with cancer. Both of these candidates are linked to poor prognosis and considered as potential targets in cancer treatment. However, there have been no specific agents identified to specifically target Sp1 and/or survivin. In this investigation, we demonstrate that inhibition of Sp1 transcription factor using TA and Mit effectively decreased CHLA-9 and TC-32 cell growth. We also show that inhibiting Sp1 resulted in inhibition of survivin protein and mRNA expression. At the molecular level, both TA and Mit treatment significantly inhibited Sp1 DNA-binding activity. We also show that TA induced activation of proteasome-dependent Sp1 degradation, whereas Mit treatment caused transcriptional inhibition of Sp1. Overall, results of this study demonstrate that ES cells express Sp1 and survivin, and both TA and Mit significantly decrease ES cell viability which correlated with an inhibition of Sp1 and survivin protein expression. However TA and Mit vary in their mechanism of action to modulate Sp1 activity.
EWS-FLI1 is overexpressed in a majority of ES tumors. High throughput screening identified Mit as an effective agent to target EWS-FLI1 [32] and the Federal Drug Administration (FDA) approved its testing on ES patients. Since there are some concerns about Mit side effects, it is important to explore alternative targets and less toxic agents for clinical testing. Non-steroidal anti-inflammatory drugs are being widely tested for their anti-cancer activity [33–37]. TA has been used for treating migraine headaches and is well tolerated by patients. It is a well-studied NSAID with limited toxicity among several other commonly used NSAIDs such as ibuprofen. Therefore, testing this agent will serve as a robust and cost-effective strategy for diseases such as ES. Collectively, our results provide evidence that supports the significance of targeting Sp1 transcription factor and survivin for the treatment of ES and suggest that TA could serve as an effective agent in such therapy.
References
Jo VY, Fletcher CD (2014) Who classification of soft tissue tumours: an update based on the 2013 (4th) edition. Pathology 46(2):95–104
Jo VY, Doyle LA (2016) Refinements in sarcoma classification in the current 2013 world health organization classification of tumours of soft tissue and bone. Surg Oncol Clin N Am 25(4):621–643
Ushigome S, Machinami R, Sorensen PH (2002) World health organization classification of tumours: Pathology and genetics of tumours of soft tissue and bone. In: IARCPress, 69008 Lyon, France, pp 297–300
Burchill SA (2003) Ewing’s sarcoma: diagnostic, prognostic, and therapeutic implications of molecular abnormalities. J Clin Pathol 56(2):96–102
Zucman J, Melot T, Desmaze C, Ghysdael J, Plougastel B, Peter M, Zucker JM, Triche TJ, Sheer D, Turc-Carel C et al (1993) Combinatorial generation of variable fusion proteins in the Ewing family of tumours. EMBO J 12(12):4481–4487
Delattre O, Zucman J, Plougastel B, Desmaze C, Melot T, Peter M, Kovar H, Joubert I, de Jong P, Rouleau G et al (1992) Gene fusion with an ets DNA-binding domain caused by chromosome translocation in human tumours. Nature 359(6391):162–165
Darnell JE Jr (2002) Transcription factors as targets for cancer therapy. Nat Rev Cancer 2(10):740–749
Wang YT, Yang WB, Chang WC, Hung JJ (2011) Interplay of posttranslational modifications in sp1 mediates sp1 stability during cell cycle progression. J Mol Biol 414(1):1–14
Beishline K, Azizkhan-Clifford J (2015) Sp1 and the ‘hallmarks of cancer. FEBS J 282(2):224–258
Liu L, Ji P, Qu N, Pu WL, Jiang DW, Liu WY, Li YQ, Shi RL (2016) The impact of high co-expression of sp1 and hif1alpha on prognosis of patients with hepatocellular cancer. Oncol Lett 12(1):504–512
Hu J, Hu H, Hang JJ, Yang HY, Wang ZY, Wang L, Chen DH, Wang LW (2016) Simultaneous high expression of pld1 and sp1 predicts a poor prognosis for pancreatic ductal adenocarcinoma patients. Oncotarget
Hingorani P, Dickman P, Garcia-Filion P, White-Collins A, Kolb EA, Azorsa DO (2013) Birc5 expression is a poor prognostic marker in Ewing sarcoma. Pediatr Blood Cancer 60(1):35–40
Greve B, Sheikh-Mounessi F, Kemper B, Ernst I, Gotte M, Eich HT (2012) Survivin, a target to modulate the radiosensitivity of ewing’s sarcoma. Strahlenther Onkol 188(11):1038–1047
Khan Z, Khan AA, Prasad GB, Khan N, Tiwari RP, Bisen PS (2016) Growth inhibition and chemo-radiosensitization of head and neck squamous cell carcinoma (hnscc) by survivin-sirna lentivirus. Radiother Oncol 118(2):359–368
Habib R, Akhtar J, Taqi M, Yu C, Zhang C (2015) Lentiviral vector-mediated survivin shrna delivery in gastric cancer cell lines significantly inhibits cell proliferation and tumor growth. Oncol Rep 34(2):859–867
Liu S, Huang W, Jin MJ, Fan B, Xia GM, Gao ZG (2016) Inhibition of murine breast cancer growth and metastasis by survivin-targeted sirna using disulfide cross-linked linear pei. Eur J Pharm Sci 82:171–182
Blume SW, Snyder RC, Ray R, Thomas S, Koller CA, Miller DM (1991) Mithramycin inhibits sp1 binding and selectively inhibits transcriptional activity of the dihydrofolate reductase gene in vitro and in vivo. J Clin Invest 88(5):1613–1621
Esteve PO, Chin HG, Pradhan S (2007) Molecular mechanisms of transactivation and doxorubicin-mediated repression of survivin gene in cancer cells. J Biol Chem 282(4):2615–2625
Eslin D, Lee C, Sankpal UT, Maliakal P, Sutphin RM, Abraham L, Basha R (2013) Anticancer activity of tolfenamic acid in medulloblastoma: a preclinical study. Tumour Biol 34(5):2781–2789
Eslin D, Sankpal UT, Lee C, Sutphin RM, Maliakal P, Currier E, Sholler G, Khan M, Basha R (2013) Tolfenamic acid inhibits neuroblastoma cell proliferation and induces apoptosis: a novel therapeutic agent for neuroblastoma. Mol Carcinog 52(5):377–386
Abdelrahim M, Baker CH, Abbruzzese JL, Safe S (2006) Tolfenamic acid and pancreatic cancer growth, angiogenesis, and sp protein degradation. J Natl Cancer Inst 98(12):855–868
Papineni S, Chintharlapalli S, Abdelrahim M, Lee SO, Burghardt R, Abudayyeh A, Baker C, Herrera L, Safe S (2009) Tolfenamic acid inhibits esophageal cancer through repression of specificity proteins and c-met. Carcinogenesis 30(7):1193–1201
Fuchs B, Inwards CY, Janknecht R (2004) Vascular endothelial growth factor expression is up-regulated by ews-ets oncoproteins and sp1 and may represent an independent predictor of survival in ewing’s sarcoma. Clin Cancer Res 10(4):1344–1353
Giorgi C, Boro A, Rechfeld F, Lopez-Garcia LA, Gierisch ME, Schafer BW, Niggli FK (2015) Pi3k/akt signaling modulates transcriptional expression of ews/fli1 through specificity protein 1. Oncotarget
Fukuda S, Pelus LM (2006) Survivin, a cancer target with an emerging role in normal adult tissues. Mol Cancer Ther 5(5):1087–1098
Yan S, Li Z, Thiele CJ (2013) Inhibition of stat3 with orally active jak inhibitor, azd1480, decreases tumor growth in neuroblastoma and pediatric sarcomas in vitro and in vivo. Oncotarget 4(3):433–445
Giorgi C, Boro A, Rechfeld F, Lopez-Garcia LA, Gierisch ME, Schafer BW, Niggli FK (2015) Pi3k/akt signaling modulates transcriptional expression of ews/fli1 through specificity protein 1. Oncotarget 6(30):28895–28910
Shin JA, Jung JY, Ryu MH, Safe S, Cho SD (2013) Mithramycin a inhibits myeloid cell leukemia-1 to induce apoptosis in oral squamous cell carcinomas and tumor xenograft through activation of bax and oligomerization. Mol Pharmacol 83(1):33–41
Wang L, Guan X, Zhang J, Jia Z, Wei D, Li Q, Yao J, Xie K (2008) Targeted inhibition of sp1-mediated transcription for antiangiogenic therapy of metastatic human gastric cancer in orthotopic nude mouse models. Int J Oncol 33(1):161–167
Choi ES, Jung JY, Lee JS, Park JH, Cho NP, Cho SD (2013) Myeloid cell leukemia-1 is a key molecular target for mithramycin a-induced apoptosis in androgen-independent prostate cancer cells and a tumor xenograft animal model. Cancer Lett 328(1):65–72
Konduri S, Colon J, Baker CH, Safe S, Abbruzzese JL, Abudayyeh A, Basha MR, Abdelrahim M (2009) Tolfenamic acid enhances pancreatic cancer cell and tumor response to radiation therapy by inhibiting survivin protein expression. Mol Cancer Ther 8(3):533–542
Grohar PJ, Woldemichael GM, Griffin LB, Mendoza A, Chen QR, Yeung C, Currier DG, Davis S, Khanna C, Khan J, McMahon JB et al (2011) Identification of an inhibitor of the ews-fli1 oncogenic transcription factor by high-throughput screening. J Natl Cancer Inst 103(12):962–978
Gately S, Li WW (2004) Multiple roles of cox-2 in tumor angiogenesis: a target for antiangiogenic therapy. Semin Oncol 31(2 Suppl 7):2–11
Jacoby RF, Seibert K, Cole CE, Kelloff G, Lubet RA (2000) The cyclooxygenase-2 inhibitor celecoxib is a potent preventive and therapeutic agent in the min mouse model of adenomatous polyposis. Cancer Res 60(18):5040–5044
Tarnawski AS, Jones MK (2003) Inhibition of angiogenesis by nsaids: molecular mechanisms and clinical implications. Journal of molecular medicine (Berlin, Germany) 81(10):627–636
Juni P, Reichenbach S, Egger M (2005) Cox 2 inhibitors, traditional nsaids, and the heart. BMJ 330(7504):1342–1343
Koki AT, Masferrer JL (2002) Celecoxib: a specific cox-2 inhibitor with anticancer properties. Cancer Control 9(2 Suppl):28–35
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors (SS, UTS, WPW, MW, AR and RB) of this manuscript declare no conflict of interest.
Funding
This work is partially supported by the Institute for Cancer Research of UNT Health Science Center to RB and the Young Investigator research award from ‘Hyundai Hope on Wheels’ awarded to AR.
Ethical approval
Research presented in this manuscript does not contain studies with human participants or animals.
Rights and permissions
About this article
Cite this article
Shelake, S., Sankpal, U.T., Paul Bowman, W. et al. Targeting specificity protein 1 transcription factor and survivin using tolfenamic acid for inhibiting Ewing sarcoma cell growth. Invest New Drugs 35, 158–165 (2017). https://doi.org/10.1007/s10637-016-0417-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10637-016-0417-9