Abstract
Background
Although several types of diet have been used in experimental steatohepatitis models, comparison of gut microbiota and immunological alterations in the gut among diets has not yet been performed.
Aim
We attempted to clarify the difference in the gut environment between mice administrated several experimental diets.
Methods
Male wild-type mice were fed a high-fat (HF) diet, a choline-deficient amino acid-defined (CDAA) diet, and a methionine-choline-deficient (MCD) diet for 8 weeks. We compared the severity of steatohepatitis, the composition of gut microbiota, and the intestinal expression of interleukin (IL)-17, an immune modulator.
Results
Steatohepatitis was most severe in the mice fed the CDAA diet, followed by the MCD diet, and the HF diet. Analysis of gut microbiota showed that the composition of the Firmicutes phylum differed markedly at order level between the mice fed the CDAA and HF diet. The CDAA diet increased the abundance of Clostridiales, while the HF diet increased that of lactate-producing bacteria. In addition, the CDAA diet decreased the abundance of lactate-producing bacteria and antiinflammatory bacterium Parabacteroides goldsteinii in the phylum Bacteroidetes. In CDAA-fed mice, IL-17 levels were increased in ileum as well as portal vein. In addition, the CDAA diet also elevated hepatic expression of chemokines, downstream targets of IL-17.
Conclusions
The composition of gut microbiota and IL-17 expression varied considerably between mice administrated different experimental diets to induce steatohepatitis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nonalcoholic steatohepatitis (NASH), a hepatic feature of metabolic syndrome, is characterized by excess fat deposition and inflammation in the liver. With the global increase in the obese population, the prevalence of patients with NASH is increasing in developed countries [1]. Because NASH can progress to cirrhosis and liver cancer, NASH is anticipated to be a major cause of liver cirrhosis and liver cancers in the near future. However, effective treatments have yet to be developed, and the molecular mechanisms of NASH remain largely unknown.
Recently, the gut–liver axis has attracted substantial attention regarding the molecular mechanism of NASH, because gut microbiota contribute to energy intake and are a source of harmful substances, such as toll-like receptor (TLR) ligands and ethanol. Firmicutes and Bacteroidetes are the major components of the gut microbiota in humans as well as rodents [2–4], and the phylum Firmicutes includes many bacteria that contribute to energy intake by generating short-chain fatty acids (SCFAs). In human studies, the Firmicutes/Bacteroidetes (F/B) ratio has been shown to be increased in obese subjects [2, 5], while the F/B ratio is decreased in lean subjects after Roux-en-Y gastric bypass and laparoscopic sleeve gastrectomy [6, 7]. The phylum Proteobacteria includes harmful bacteria such as Helicobacter [8] and Escherichia [9, 10], which are sources of lipopolysaccharide and ethanol. In contrast, Akkermansia muciniphila [3] and lactate-producing bacteria [11, 12] have beneficial effects on NASH. However, the pathological impact of these bacteria has not been consistently reproduced across models, likely due to altered gut microbiota composition. In addition, alteration of the gut microbiota can modulate the immune system in the gut, which may affect the severity of NASH; For instance, interleukin (IL)-17 is a key player modulating the immune system in the gut [13] as well as the progression of NAFLD/NASH [2, 14].
To clarify the molecular mechanisms of NASH, several diet models are used in mice, including a high-fat (HF) diet, a choline-deficient amino acid-defined (CDAA) diet, and a methionine-choline-deficient (MCD) diet [15–18]. While these diets all induce hepatic steatosis, the metabolic parameters vary greatly among models; For instance, HF diets induce insulin resistance with obesity, but the hepatic inflammation is weak [15, 18]. In contrast, MCD diets induce hepatic steatosis and inflammation, but insulin sensitivity is enhanced due to the loss of body weight [15–17]. CDAA diets can induce steatohepatitis as well as obesity and insulin resistance [19]. Although these diet models may alter the composition of the gut microbiota and immune response, detailed comparison has not yet been performed.
In the present study, we compared the severity of steatohepatitis, the composition of the gut microbiota, and the IL-17 expression in several experimental diet models, including an HF diet, a CDAA diet, and a MCD diet model. We found that the composition of gut microbiota and IL-17 expression varied considerably between mice fed different experimental diets, which may affect the severity of steatohepatitis.
Methods
Animals and Diets
Both male and female C57BL/6 wild-type mice were purchased from Japan SLC (Shizuoka, Japan) and were mated in the specific-pathogen-free room in the Animal Institute of Akita University. The male littermates were divided into four groups at 8 weeks of age and fed normal chow (NC: CE-7; CLEA Japan, Tokyo, Japan), HF diet (Oriental Yeast, Tokyo, Japan), CDAA diet (Oriental Yeast), or MCD diet (Oriental Yeast). The nutritional content of these diets is listed in Supplemental Table 1. After 8 weeks of feeding, 10 mice from each group were humanely killed, and their collected samples stored at −80 °C until further analyses. For gut sterilization, CDAA-fed mice were orally administered an antibiotics mixture containing ampicillin, neomycin, metronidazole, and vancomycin, according to published protocol [20]. The antibiotics treatment was continued for 8 weeks. The Institutional Review Board of Akita University Graduate School of Medicine approved all animal experiments in the present study.
Histological Examination
Hematoxylin and eosin (H&E) staining, Oil Red O staining, and immunohistochemistry for IL-17 (catalog no. ab79056; Abcam, Tokyo, Japan), neutrophil elastase (catalog no. ab68672; Abcam), F4/80 (catalog no. 14-4801-82; eBioscience, San Diego, CA), and lysozyme (catalog no. ab108508; Abcam) were performed according to manufacturer instructions. The NAFLD activity score was evaluated according to the published report [21].
Quantitative Real-Time PCR Analysis
RNA was extracted from the liver and macrophages using TRIzol (Life Technologies Japan, Tokyo, Japan). The extracted RNA was converted to complementary DNA (cDNA) using reverse transcription. The cDNA was then subjected to polymerase chain reaction (PCR) using the listed primers (Supplemental Table 2) and the LightCycler 480 SYBR Green I Master device (Roche Diagnostics, Basel, Switzerland). Gene expression was normalized to that of 18S or β-actin RNA as internal control.
Measurement of Hepatic Lipid, Serum Alanine Transaminase (ALT), and IL-17A
Hepatic lipids were isolated as previously described [19]. Total cholesterol, free cholesterol, triglyceride, and free fatty acids were measured using cholesterol E (catalog no. 439-17501; Wako Pure Chemical Industries, Osaka, Japan), free cholesterol E (catalog no. 435-35801; Wako Pure Chemical Industries), triglyceride E (catalog no. 432-40201; Wako Pure Chemical Industries), and NEFA C (catalog no. 279-75401; Wako Pure Chemical Industries), respectively, according to manufacturer instructions. Serum ALT levels and IL-17A concentration in portal vein were measured using transaminase CII (catalog no. 431-30901; Wako Pure Chemical Industries) and the mouse IL-17A enzyme-linked immunosorbent assay (ELISA) kit (catalog no. 88-7371-88; eBioscience), respectively, according to manufacturer instructions.
Gut Microbiota Analysis
Stool was harvested from the colon when the mice were killed at 16 weeks of age, and three or four stool pellets were collected from each mouse. These stool samples from three or four mice per group were separately cryopreserved until further analyses. DNA was isolated from each stool sample using the MORA-EXTRACT kit (COSMO BIO, Tokyo, Japan). We used two high-throughput sequencing technologies: the MiSeq system (Illumina, San Diego, CA) and GS Junior system (Roche Diagnostics). The MiSeq system provides a larger number of reads with a relatively short read length, while the GS Junior system provides a longer read length [22]. In the present study, the MiSeq system was used for primary analyses and the GS Junior system when bacteria could not be classified by MiSeq-based analysis.
For the MiSeq system (n = 4 each), we performed high-throughput sequencing according to the instructions (http://www.illumina.com). Amplicon PCR was performed using the following thermal conditions: denaturing at 95 °C for 3 min, 25 cycles of amplification at 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s, and final extension at 72 °C for 5 min. The size was then checked using the Bioanalyzer with High Sensitivity DNA kit (Agilent Technology, Santa Clara, CA). The amplicons were purified using the Ampure XP (NIPPON Genetics, Tokyo, Japan), then subjected to index PCR following manufacturer instructions. After index PCR, the libraries were quantified using a fluorometric quantification method following manufacturer instructions. Paired-end sequencing of bacterial 16S ribosomal RNA (rRNA) gene amplicons was conducted using the MiSeq platform. MiSeq Reporter (version 2.5.1.3) bioinformatics software was used for metagenome analysis. Quantitative Insights Into Microbial Ecology, an open-source bioinformatics pipeline, was also used to perform principal component analysis.
For the GS Junior system, DNA mixtures were analyzed, with an equivalent amount of DNA from three mice of the same group mixed. These DNA samples were amplified between the V3 and V4 regions of the 16S rRNA gene by PCR with the fusion primers listed in Supplemental Table 3. PCR was performed using the following thermal conditions: denaturing at 94 °C for 5 min, 20 cycles of amplification at 94 °C for 1 min, 65 to 55 °C for 1 min (decrease of 1 °C every two cycles), and 72 °C for 1 min, 10 cycles of amplification at 94 °C for 1 min, 55 °C for 1 min, and 72 °C for 1 min, and final extension at 72 °C for 3 min. The amplicons were purified from the agarose gel after electrophoresis, then size and purity were checked using the Bioanalyzer with High Sensitivity DNA kit (Agilent Technology), quantified by Quant-iT PicoGreen dsDNA reagent (Invitrogen, Carlsbad, CA), then subjected to emulsion PCR (Roche Diagnostics) following manufacturer instructions. After emulsion PCR, bacterial tag-encoded FLX amplicon pyrosequencing was conducted with the GS Junior system at Akita Prefectural University. The obtained FASTA files were subjected to online software analyses using the DECIPHER web tool (http://decipher.cee.wisc.edu/index.html) [23].
Macrophage Isolation
Hepatic macrophages were isolated from normal mice as previously reported [24]. A total of 1 × 106 cells were treated with recombinant IL-17A (catalog no. 421-ML; R&D Systems, Minneapolis, MN) at the indicated concentration and harvested at 8 h after stimulation.
Statistical Analysis
Statistical analyses were performed using one-way analysis of variance (ANOVA) or Mann–Whitney U test (SPBS software, version 9.67); P values <0.05 were considered to be statistically significant.
Results
CDAA Diet Induced the Most Severe Steatohepatitis
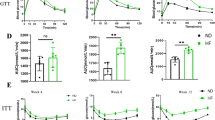
Male wild-type mice on NC, HF, and CDAA diet gained roughly 23, 75, and 25 % of their initial weight after 8 weeks of feeding, respectively, whereas mice on MCD diet lost 35 % of their initial weight after feeding (Table 1). The hepatic steatosis as examined by H&E staining and Oil Red O staining was most severe in the CDAA diet group, followed by the HF diet and MCD diet groups (Fig. 1a). The hepatic contents of triglycerides, total cholesterol, free cholesterol, and free fatty acids are presented in Table 1. Inflammatory cell infiltration and hepatocyte ballooning were frequently observed in mice fed the CDAA diet, followed by those fed the MCD diet (Fig. 1a, b). As a result, the NAFLD activity score was highest in the CDAA diet group (Fig. 1b). Serum ALT levels and hepatic gene expression of proinflammatory cytokines were highest in the CDAA diet group (Fig. 1c, d). Serum ALT levels were elevated in the MCD diet group, but showed only slight elevation in the HF diet group (Fig. 1c). Although none of the diets induced histological liver fibrosis after 8 weeks of feeding, gene expression of fibrogenic factors, including collagen 1α-1 and TIMP-1, was elevated in the CDAA diet group compared with the other groups (Fig. 1d). These results indicate that the CDAA diet induced the most severe steatohepatitis, followed by the MCD diet, in the present study. The HF diet induced steatosis without inflammation, namely simple steatosis.
CDAA diet induces the most severe steatohepatitis. a H&E staining and Oil Red O staining of liver sections. Mice were fed normal chow (NC), high-fat (HF) diet, choline-deficient amino acid-defined (CDAA) diet, or methionine-choline-deficient (MCD) diet. Boxed areas for CDAA and MCD diets show foci of inflammation. Scale bars indicate 1000 µm (upper panel) or 200 µm (middle and lower panels). b NAFLD activity score. c Serum ALT level. d Hepatic gene expression of TNFα, IL-1β, collagen 1α-1, and TIMP-1. Gene expression was normalized to 18S RNA as internal control. b–d Data presented as mean ± SD (n = 10 each, one-way ANOVA). #a Significantly different from NC group. #b Significantly different from HF and MCD diet groups. #c Significantly different from HF diet group
Composition of Gut Microbiota Varied Markedly Among Experimental NASH Models
Principal component analysis showed that the gut microbiota compositions obtained from mice in the same diet group were aggregated at similar positions. In contrast, the composition of the gut microbiota varied markedly among diets (Fig. 2a). We obtained an average of 133,618, 95,418, 120,789, and 123,583 reads in the NC, HF, CDAA, and MCD diet groups, respectively. Of the seven phyla identified, the most abundant bacteria belonged to the phylum Firmicutes, followed by Bacteroidetes and Proteobacteria in the NC group (Fig. 2b). Over 85 % of gut microbiota belonged to the Firmicutes and Bacteroidetes phyla, which largely comprise Gram-positive and Gram-negative bacteria, respectively. Consistent with previous reports [2, 5], the F/B ratio was increased in the HF diet group (Fig. 2c), in which obesity was induced. In addition, the F/B ratio was increased in the CDAA diet group, which showed an increase in body weight similar to that in the NC group (Fig. 2c). Interestingly, the F/B ratio was also increased in the MCD diet group, although the body weight decreased after feeding (Fig. 2c).
Composition of gut microbiota varied markedly among experimental NASH models. a Principal component analysis of gut microbiota (n = 4 each). Three- and two-dimensional analyses are shown. b Composition of gut microbiota at phylum level. c Firmicutes-to-Bacteroidetes ratio. #a significantly different from NC group. #d significantly different from CDAA and MCD diet groups
CDAA Diet Results in Decreased Prevalence of Lactate-Producing Bacteria in Phylum Firmicutes
We obtained an average of 72,871, 84,984, 72,505, and 80,369 reads belonging to the phylum Firmicutes in the NC, HF diet, CDAA diet, and MCD diet groups, respectively. The members of the phylum Firmicutes include Clostridiales and Lactobacillales at order level, which can produce SCFAs and lactate, respectively. The abundance of Firmicutes was increased in the HF diet group (Fig. 3a). However, detailed analysis revealed that the composition varied markedly among diets at order level in the phylum Firmicutes. Clostridiales was increased in the CDAA diet group, while Lactobacillales was increased in the HF diet group. At genus level, lactate-producing bacteria, including Lactobacillus and Lactococcus, are reported to exert beneficial effects on the host by inhibiting expansion of pathogenic bacteria [11]. Although Lactobacillus was a major component of Firmicutes in the NC group, the prevalence of Lactobacillus was decreased in the CDAA diet and MCD diet groups at genus level. In the HF group, the increased Lactobacillales was due to increased Lactococcus at genus level. Because the MiSeq-based microbial community analyses failed to classify the bacteria belonging to the order Clostridiales, we examined the same samples using the GS Junior system. The abundance of Clostridium was increased in the CDAA diet group. At species level, the increase in genus Clostridium in the CDAA diet group was due to the increased number of Clostridium phytofermentans (Supplemental Figure 1), which can produce ethanol [25, 26] and toxic aldehyde intermediates [27].
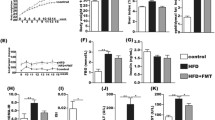
Composition of gut microbiota. The abundance of gut microbiota is shown for NC, HF, CDAA, and MCD diets. The composition of the gut microbiota is shown at phylum, order, genus, and species level. a Composition of gut microbiota in phylum Firmicutes. b Composition of gut microbiota in phylum Bacteroidetes. B and P indicate Bacteroides and Parabacteroides, respectively. c Alteration of gut microbiota in phylum Proteobacteria. #a Significantly different from NC group. #b Significantly different from HF and MCD diet groups. #c Significantly different from HF diet group. #d Significantly different from CDAA and MCD diet groups. #e Significantly different from HF and CDAA diet groups
Prevalence of Phylum Bacteroidetes and Parabacteroides goldsteinii Decreased in All Experimental Diet Groups
We obtained an average of 46,767, 1037, 8568, and 15,898 reads belonging to the phylum Bacteroidetes in the NC, HF diet, CDAA diet, and MCD diet groups, respectively. The abundance of the phylum Bacteroidetes was decreased across all diet groups, as a result of decrease in the order Bacteroidales (Fig. 3b). Bacteroides and Parabacteroides were major components at genus level in the order Bacteroidales. A small increase in the genus Bacteroides was observed for the CDAA and MCD diets, due to the increase of Bacteroides acidifaciens. In contrast, the abundance of genus Parabacteroides was decreased across all diet groups, corresponding to the alteration of phylum Bacteroidetes. At species level, the prevalence of P. goldsteinii was reduced, which is associated with anti-inflammation in HF diet and alcohol liver injury [28, 29].
Prevalence of Lipopolysaccharide-Producing Bacteria in Phylum Proteobacteria Increased in CDAA and MCD Diet Groups
The phylum Proteobacteria includes pathogenic bacteria, and an increase in Proteobacteria has been reported in patients with liver cirrhosis [30]. We obtained an average of 1547, 872, 802, and 1970 reads belonging to the phylum Proteobacteria in the NC, HF diet, CDAA diet, and MCD diet groups, respectively. In the present study, Proteobacteria had a relatively small population (Fig. 2b). The proportion of Proteobacteria increased in response to the MCD diet (Fig. 3c). Although the phylum Proteobacteria includes Helicobacter, Desulfovibrio, and Escherichia at genus level, further classification by MiSeq-based study was unsuccessful. In contrast, the proportions of lipopolysaccharide-producing bacteria, including Desulfovibrio vulgaris and Helicobacter hepaticus, were increased in the CDAA and MCD diet groups at species level in the GS Junior-based study (Supplemental Figure 1). Although the Escherichia family includes ethanol-producing bacteria [10], this population was very small in the present study.
IL-17A Levels Are Increased in Small Intestine and Portal Vein in Mice Fed CDAA Diet
Alterations in the gut microbiota can modulate the immune system in the gut. Thus, we examined the expression of IL-17, because IL-17 is a gut microbiota-mediated cytokine [31]. In mice fed the NC and HF diets, IL-17-positive cells were not observed in small intestine, including ileum (Fig. 4a). In contrast, populations of IL-17-positive cells were increased in mice fed the CDAA diet and similarly increased but to a lesser degree in those fed the MCD diet (Fig. 4a). In these mice, IL-17-positive cells were observed in ileum but not in jejunum (data not shown). These IL-17-positive cells expressed lysozyme (Fig. 4a), indicating that IL-17-producing cells are Paneth cells. Gut sterilization by an antibiotics mixture decreased the expression of IL-17 in the Paneth cells (Fig. 4a). Furthermore, the IL-17A levels in portal vein were significantly higher in the CDAA diet group than in any other diet group, and these levels were decreased by gut sterilization (Fig. 4b).
CDAA diet increases IL-17 levels and chemokine expression in liver. a Immunohistochemistry for IL-17 and lysozyme in ileum. IL-17-positive cells are located at crypts in ileum of mice, particular in the CDAA diet group. Serial sections showed that IL-17-expressing cells were positive for lysozyme, indicating that IL-17-expressing cells were Paneth cells. Antibiotics treatment decreased the expression of IL-17. Boxed areas show magnification of Paneth cells. Scale bars indicate 200 µm. b IL-17A levels in portal vein. The CDAA diet increased the IL-17 levels that had been decreased by antibiotics treatment (Anti). Data presented as mean ± SD (n = 4–8, Mann–Whitney U test, *P < 0.05). c Hepatic gene expression of chemokines and their receptors. Gene expression was normalized to β-actin as internal control. Data presented as mean ± SD (n = 10 each, one-way ANOVA). d Immunohistochemistry for neutrophil elastase (scale bars 100 µm) and F4/80 (scale bars 200 µm) in liver sections. e Gene expression of chemokines and their receptors in hepatic macrophages (n = 4 each, one-way ANOVA). Gene expression was normalized to β-actin as internal control. Data are presented as mean ± SD, *significantly different (P < 0.05)
IL-17A has been reported to increase expression of chemokines, including CXCL1, CXCL2, and CCL2 that recruit neutrophils and macrophages [32, 33]. Hepatic expression of CXCL2 and CCL2 was significantly increased in the CDAA diet group (Fig. 4c). In addition, gene expression of CXCR3 was also increased in the CDAA diet group (Fig. 4c). These increased genes were suppressed by gut sterilization (Fig. 4c). Indeed, the number of infiltrating neutrophils and macrophages in liver was significantly increased by the CDAA diet (Fig. 4d), and the number of inflammatory cells was decreased by gut sterilization (data not shown). We also examined chemokine expression in isolated macrophages in response to IL-17A. IL-17A increased the messenger RNA (mRNA) expression of CXCL2 and CCL2 (Fig. 4e), which recruit neutrophils and macrophages. These data suggest that gut-derived IL-17A may increase the levels of chemokines in the liver, which promotes progression of steatohepatitis by recruiting inflammatory cells.
Discussion
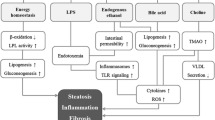
To the best of the authors’ knowledge, this is the first report to demonstrate large differences in both gut microbiota and IL-17 levels between experimental steatohepatitis models induced by different diets. In the most severe steatohepatitis model, induced by the CDAA diet, the prevalence of ethanol-producing bacteria in the phylum Firmicutes was increased. On the other hand, the CDAA diet decreased the prevalence of lactate-producing bacteria in the phylum Firmicutes and the anti-inflammatory bacterium P. goldsteinii in the phylum Bacteroidetes. In addition, IL-17 levels were elevated in ileum as well as portal vein, which may subsequently promote progression of steatohepatitis by stimulating chemokine production in the liver.
In this study, we sought to determine the key microbiota involved in NASH pathogenesis. In line with previous studies [2, 5], the F/B ratio and the abundance of Firmicutes were increased in mice fed the HF diet. Interestingly, the CDAA and MCD diets also increased the F/B ratio, regardless of body weight. These results suggest that the composition of the gut microbiota is affected by not only obese conditions but also dietary components. Although the abundance of the phylum Firmicutes was increased for all diets inducing steatosis, we found that the bacteria in this phylum varied markedly between the CDAA and HF diet groups. The abundance of Clostridiales was increased in the CDAA group, while Lactobacillales was increased in the HF diet group. Clostridiales includes many species of fermentation-associated bacteria, leading to production of SCFAs and ethanol by fermenting indigestible carbohydrates. Additional analyses showed that the prevalence of C. phytofermentans was markedly higher in the CDAA diet group than in the other diet groups. C. phytofermentans, a member of Clostridium cluster XIV, is known to produce ethanol [25, 26] and toxic aldehyde intermediates [27], rather than anti-inflammatory SCFAs. In addition, the abundance of C. phytofermentans was higher in the CDAA diet group than in the MCD diet group. Thus, increased number of ethanol-producing bacteria may exacerbate the severity of steatohepatitis in the CDAA diet. On the other hand, the CDAA and MCD diets reduced the prevalence of Lactobacillus, which has been shown to exert beneficial effects on the host in human and animal studies [11, 34]. The decreased abundance of lactate-producing bacteria may also have contributed to the deterioration of steatohepatitis in the CDAA and MCD diet groups.
In the phylum Bacteroidetes, P. goldsteinii was decreased, corresponding to the alteration observed at phylum level. P. goldsteinii is associated with anti-inflammation in HF-diet-induced obesity and alcohol liver injury [28, 29]. Using the GS Junior system, we observed a decrease in Parabacteroides distasonis but not P. goldsteinii. P. distasonis is close to P. goldsteinii [35] and exerts anti-inflammatory effects in the colon with reduction of tumor necrosis factor-alpha (TNFα) production in macrophages and IL-17 expression in colonic tissue [36, 37]. In line with previous reports, P. distasonis was more prominent in mice fed the NC diet, with relatively low prevalence in the CDAA and MCD diet groups. These data suggest that the CDAA and MCD diets may have little benefit on the Parabacteroides population in the present study. Although we noticed a small increase in B. acidifaciens, the role of these bacteria has not been well characterized.
The phylum Proteobacteria includes various harmful Gram-negative bacteria such as Helicobacter spp., Desulfovibrio spp., and Escherichia spp. and is associated with development of NASH [34, 38, 39]. The prevalence of Proteobacteria was increased in the MCD diet group. Further analyses using the GS Junior system showed that the population of H. hepaticus was significantly increased in the CDAA and MCD diet groups in the phylum Proteobacteria. H. hepaticus can increase expression of TLR4, TNFα, and IL-1β [39]. In addition, H. hepaticus is associated with chronic hepatic and intestinal inflammation and hepatobiliary cancers in mice [39]. The prevalence of D. vulgaris was also significantly higher in the CDAA and MCD diet groups according to the GS Junior-based analyses. The Desulfovibrionaceae family including D. vulgaris, the sulfate-reducing bacteria, is a potential source of lipopolysaccharide and can injure the barrier function of the intestine [3, 40]. Because TLR4 signaling promotes progression of NASH [3, 41], H. hepaticus and D. vulgaris may contribute to the development of steatohepatitis through TLR4 in the CDAA and MCD diet groups.
In the present study, we unexpectedly had to use two high-throughput sequencing technologies, the MiSeq and GS Junior systems, because the maintenance service of the GS Junior system ended during the present study. The use of two high-throughput sequencing systems and their related bioinformatics yielded some different results at species level; For instance, P. goldsteinii was not detected in the GS Junior-based analyses, whereas P. distasonis was not detected in the MiSeq-based analyses. This often happens when different bioinformatics pipelines are used [42]. Therefore, one should pay attention to the sequencing platforms and bioinformatics used when comparing our results with those obtained in other studies. Although the GS Junior has an advantage in its ability to obtain a longer read length, the MiSeq system has benefits in term of cost, speed of analysis, and number of reads [22]. Thus, the MiSeq system is widely used for analyses of gut microbiota.
We also found that IL-17 levels were increased in the CDAA diet group compared with the other diet groups. IL-17 functions as a proinflammatory cytokine in NAFLD and NASH [14, 43, 44]. To the best of the authors’ knowledge, this is the first report to show increased IL-17 levels in ileum and portal vein in severe steatohepatitis. Paneth cells in small intestine are a source of IL-17 [45, 46]. Consistent with these reports, IL-17 expression was observed in Paneth cells of ileum, particular in CDAA-fed mice. IL-17 can induce production of chemokines that recruit inflammatory cells, which leads to progression of NASH by producing inflammatory cytokines [47]. In the present study, we showed a positive correlation between IL-17 levels and chemokine expression. Although it remains unclear whether the dietary components directly modified IL-17 expression, the CDAA diet altered the gut microbiota composition and increased the IL-17 levels. In addition, gut sterilization reduced the IL-17 levels in gut and portal vein, indicating that the gut microbiota regulate the IL-17 expression. Among bacteria in the gut, lactate-producing bacteria protect the host from IL-17-induced injury [48], suggesting that lactate-producing bacteria may suppress IL-17 expression. Lipopolysaccharide derived from the phylum Proteobacteria increases expression of IL-17 [49]. The abundance of lipopolysaccharide-producing bacteria was altered in the CDAA diet group, suggesting that the gut microbiota component induced by the CDAA diet may increase the expression of IL-17. Segmented filamentous bacteria have been reported to increase IL-17 [50, 51] but were not detected in our analyses. Although we did not identify the bacteria that promote IL-17 production, our data do suggest that the gut microbiota contribute to IL-17 expression.
In conclusion, the composition of gut microbiota and the immune response vary considerably between mice administrated different experimental diets to induce steatohepatitis.
Abbreviations
- ALT:
-
Alanine transaminase
- CDAA:
-
Choline-deficient amino acid-defined
- F/B :
-
Firmicutes/Bacteroidetes
- HF:
-
High-fat
- H&E:
-
Hematoxylin and eosin
- IL:
-
Interleukin
- MCD:
-
Methionine-choline-deficient
- NAFLD:
-
Nonalcoholic fatty liver disease
- NASH:
-
Nonalcoholic steatohepatitis
- NC:
-
Normal chow
- SCFAs:
-
Short-chain fatty acids
- TLR:
-
Toll-like receptor
References
Okanoue T, Umemura A, Yasui K, et al. Nonalcoholic fatty liver disease and nonalcoholic steatohepatitis in Japan. J Gastroenterol Hepatol. 2011;1:153–162.
Mouzaki M, Comelli EM, Arendt BM, et al. Intestinal microbiota in patients with nonalcoholic fatty liver disease. Hepatology. 2013;58:120–127.
Miura K, Ohnishi H. Role of gut microbiota and toll-like receptors in nonalcoholic fatty liver disease. World J Gastroenterol. 2014;20:7381–7391.
Bäckhed F, Ley RE, Sonnenburg JL, et al. Host-bacterial mutualism in the human intestine. Science. 2005;307:1915–1920.
Ley RE, Turnbaugh PJ, Klein S, et al. Microbial ecology: human gut microbes associated with obesity. Nature. 2006;444:1022–1023.
Mathur R, Barlow GM. Obesity and the microbiome. Expert Rev Gastroenterol Hepatol. 2015;9:1087–1099.
Damms-Machado A, Mitra S, Schollenberger AE, et al. Effects of surgical and dietary weight loss therapy for obesity on gut microbiota composition and nutrient absorption. Biomed Res Int. 2015;2015:806248.
Huang Y, Fan XG, Wang ZM, et al. Identification of helicobacter species in human liver samples from patients with primary hepatocellular carcinoma. J Clin Pathol. 2004;57:1273–1277.
Ferolla SM, Armiliato GN, Couto CA, et al. The role of intestinal bacteria overgrowth in obesity-related nonalcoholic fatty liver disease. Nutrients. 2014;6:5583–5599.
Zhu L, Baker SS, Gill C, et al. Characterization of gut microbiomes in nonalcoholic steatohepatitis (NASH) patients: a connection between endogenous alcohol and NASH. Hepatology. 2013;57:601–609.
Okubo H, Sakoda H, Kushiyama A, et al. Lactobacillus casei strain Shirota protects against nonalcoholic steatohepatitis development in a rodent model. Am J Physiol Gastrointest Liver Physiol. 2013;305:G911–G918.
Naito E, Yoshida Y, Makino K, et al. Beneficial effect of oral administration of Lactobacillus casei strain Shirota on insulin resistance in diet-induced obesity mice. J Appl Microbiol. 2011;110:650–657.
Ivanov II, Frutos Rde L, Manel N, et al. Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe. 2008;4:337–349.
Tang Y, Bian Z, Zhao L, et al. Interleukin-17 exacerbates hepatic steatosis and inflammation in nonalcoholic fatty liver disease. Clin Exp Immunol. 2011;166:281–290.
Ibrahim SH, Hirsova P, Malhi H, et al. Animal models of nonalcoholic steatohepatitis: Eat, delete, and inflame. Dig Dis Sci. 2016;61:1325–1336.
Yamaguchi K, Yang L, McCall S, et al. Inhibiting triglyceride synthesis improves hepatic steatosis but exacerbates liver damage and fibrosis in obese mice with nonalcoholic steatohepatitis. Hepatology. 2007;45:1366–1374.
Hebbard L, George J. Animal models of nonalcoholic fatty liver disease. Nat Rev Gastroenterol Hepatol. 2011;8:35–44.
Ito M, Suzuki J, Tsujioka S, et al. Longitudinal analysis of murine steatohepatitis model induced by chronic exposure to high-fat diet. Hepatol Res. 2007;37:50–57.
Miura K, Kodama Y, Inokuchi S, et al. Toll-like receptor 9 promotes steatohepatitis by induction of interleukin-1beta in mice. Gastroenterology. 2010;139:e7.
Miura K, Ishioka M, Minami S, et al. Toll-like receptor 4 on macrophage promotes the development of steatohepatitis-related hepatocellular carcinoma in mice. J Biol Chem. 2016;27:11504–11517.
Brunt EM, Kleiner DE, Wilson LA, et al. NASH clinical research network (CRN). Nonalcoholic fatty liver disease (NAFLD) activity score and the histopathologic diagnosis in NAFLD: distinct clinicopathologic meanings. Hepatology. 2011;53:810–820.
Loman NJ, Misra RV, Dallman TJ, et al. Performance comparison of benchtop high-throughput sequencing platforms. Nat Biotechnol. 2012;30:434–439.
Wright ES, Yilmaz LS, Noguera DR. DECIPHER, a search-based approach to chimera identification for 16S rRNA sequences. Appl Environ Microbiol. 2012;78:717–725.
Kitani H, Takenouchi T, Sato M, Yoshioka M, Yamanaka N. A simple and efficient method to isolate macrophages from mixed primary cultures of adult liver cells. J Vis Exp JoVE. 2011;e0000.
Petit E, LaTouf WG, Coppi MV, et al. Involvement of a bacterial microcompartment in the metabolism of fucose and rhamnose by Clostridium phytofermentans. PLoS ONE. 2013;8:e54337.
Collins MD, Lawson PA, Willems A, et al. The phylogeny of the genus Clostridium: proposal of five new genera and eleven new species combinations. Int J Syst Bacteriol. 1994;44:812–826.
Tuck LR, Altenbach K, Ang TF, et al. Insight into Coenzyme A cofactor binding and the mechanism of acyl-transfer in an acylating aldehyde dehydrogenase from Clostridium phytofermentans. Sci Rep. 2016;6:22108.
Chang CJ, Lin CS, Lu CC, et al. Ganoderma lucidum reduces obesity in mice by modulating the composition of the gut microbiota. Nat Commun. 2015;6:7489.
Neyrinck AM, Etxeberria U, Taminiau B, et al. Rhubarb extract prevents hepatic inflammation induced by acute alcohol intake, an effect related to the modulation of the gut microbiota. Mol Nutr Food Res. 2016. doi:10.1002/mnfr.201500899.
Chen Y, Yang F, Lu H, et al. Characterization of fecal microbial communities in patients with liver cirrhosis. Hepatology. 2011;54:562–572.
Chinen T, Rudensky AY. The effects of commensal microbiota on immune cell subsets and inflammatory responses. Immunol Rev. 2012;245:45–55.
Lemmers A, Moreno C, Gustot T, et al. The interleukin-17 pathway is involved in human alcoholic liver disease. Hepatology. 2009;49:646–657.
Zhang Y, Chen L, Gao W, et al. IL-17 neutralization significantly ameliorates hepatic granulomatous inflammation and liver damage in Schistosoma japonicum infected mice. Eur J Immunol. 2012;42:1523–1535.
Delzenne NM, Cani PD. Interaction between obesity and the gut microbiota: relevance in nutrition. Annu Rev Nutr. 2011;31:15–31.
Sakamoto M. Reclassification of Bacteroides distasonis, Bacteroides goldsteinii and Bacteroides merdae as Parabacteroides distasonis gen. nov., comb. nov., Parabacteroides goldsteinii comb. nov. and Parabacteroides merdae comb. nov. Int J Syst Evol Microbiol. 2006;56:1599–1605.
Pfalzer AC, Nesbeth PD, Parnell LD, et al. Diet- and genetically-induced obesity differentially affect the fecal microbiome and metabolome in Apc1638N mice. PLoS ONE. 2015;10:e0135758.
Kverka M, Zakostelska Z, Klimesova K, et al. Oral administration of Parabacteroides distasonis antigens attenuates experimental murine colitis through modulation of immunity and microbiota composition. Clin Exp Immunol. 2011;163:250–259.
Murphy EF, Cotter PD, Healy S, et al. Composition and energy harvesting capacity of the gut microbiota: relationship to diet, obesity and time in mouse models. Gut. 2010;59:1635–1642.
Segura-López FK, Güitrón-Cantú A, Torres J. Association between Helicobacter spp. infections and hepatobiliary malignancies: a review. World J Gastroenterol. 2015;21:1414–1423.
Goldstein EJ, Citron DM, Peraino VA, et al. Desulfovibrio desulfuricans bacteremia and review of human Desulfovibrio infections. J Clin Microbiol. 2003;41:2752–2754.
Imajo K, Fujita K, Yoneda M, et al. Hyperresponsivity to low-dose endotoxin during progression to nonalcoholic steatohepatitis is regulated by leptin-mediated signaling. Cell Metab. 2012;16:44–54.
D’Argenio V, Casaburi G, Precone V, Salvatore F. Comparative metagenomic analysis of human gut microbiome composition using two different bioinformatic pipelines. BioMed Res Int. 2014;2014:1–10.
Rolla S, Alchera E, Imarisio C, et al. The balance between IL-17 and IL-22 produced by liver-infiltrating T-helper cells critically controls NASH development in mice. Clin Sci (Lond). 2016;130:193–203.
Takahashi N, Vanlaere I, de Rycke R, et al. IL-17 produced by Paneth cells drives TNF-induced shock. J Exp Med. 2008;205:1755–1761.
Park SW, Kim M, Brown KM, et al. Paneth cell-derived interleukin-17A causes multiorgan dysfunction after hepatic ischemia and reperfusion injury. Hepatology. 2011;53:1662–1675.
Jin W, Dong C. IL-17 cytokines in immunity and inflammation. Emerg Microbes Infect. 2013;2:e60.
Miura K, Yang L, van Rooijen N, et al. Hepatic recruitment of macrophages promotes nonalcoholic steatohepatitis through CCR2. Am J Physiol Gastrointest Liver Physiol. 2012;302:G1310–G1321.
Gomes AC, Bueno AA, de Souza RG, et al. Gut microbiota, probiotics and diabetes. Nutr J. 2014;13:60.
Haileselassie Y, Johansson MA, Zimmer CL, et al. Lactobacilli regulate Staphylococcus aureus 161:2-induced pro-inflammatory T-cell responses in vitro. PLoS ONE. 2013;8:e77893.
Geem D, Medina-Contreras O, McBride M, et al. Specific microbiota-induced intestinal Th17 differentiation requires MHC class II but not GALT and mesenteric lymph nodes. J Immunol. 2014;193:431–438.
Harley IT, Stankiewicz TE, Giles DA, et al. IL-17 signaling accelerates the progression of nonalcoholic fatty liver disease in mice. Hepatology. 2014;59:1830–1839.
Acknowledgments
We thank Daichi Nakagawa for excellent technical assistance and the Biotechnology Center, Faculty of Bioresource Sciences, Akita Prefectural University, for assistance with the GS Junior instrument for pyrosequencing analyses.
Funding
This study was supported by a Grant-in-Aid for Scientific Research from the Japan Society for the Promotion of Science (K.M.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors have no conflicts to disclose.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplemental Figure 1
Composition of gut microbiota examined by the GS Junior system. The relative abundance of gut microbiota is shown for HF, CDAA, and MCD diets, compared with the NC group. First, the percentage of the gut microbiota was calculated in the total count reads (NC: 10,829, HF: 13,374, CDAA: 26,009, MCD: 22,601 reads). Red indicates increased abundance, while green indicates decreased abundance, compared with the NC group. The composition of the gut microbiota is shown at phylum, order, genus, and species level. (A) Composition of gut microbiota in phylum Firmicutes. “C” indicates Clostridium. (B) Composition of gut microbiota in phylum Bacteroidetes. “B” and “P” indicate Bacteroides and Parabacteroides, respectively. (C) Composition of gut microbiota in phylum Proteobacteria. “D” and “H” indicate Desulfovibrio and Helicobacter, respectively. (TIFF 2927 kb)
Rights and permissions
About this article
Cite this article
Ishioka, M., Miura, K., Minami, S. et al. Altered Gut Microbiota Composition and Immune Response in Experimental Steatohepatitis Mouse Models. Dig Dis Sci 62, 396–406 (2017). https://doi.org/10.1007/s10620-016-4393-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-016-4393-x