Abstract
Background
microRNAs (miRNAs) are a class of non-coding, single-stranded RNA molecules that regulate gene expression at the posttranscriptional level. Methyl-CpG-binding domain proteins (MBPs) are transcription repressors through binding to methylated gene promoters. Recent studies have shown that the effect of miRNAs on DNA methylation by targeting DNA methyltransferase (DNMTs) and/or MBPs plays an important role in various human cancers.
Aims
This study focuses on the regulation of MBPs by miR-373 and its downstream effect in hilar cholangiocarcinoma.
Methods
miR-373 was investigated by TaqMan miRNA Assay; mRNA and protein of MBD1, MBD2, and Mecp2 were determined by QuantiTect® Primer Assays and Western blotting, respectively; RASSF1A mRNA was measured by SYBR-Green real-time PCR; The targeting at MBD2-3′UTR by miR-373 was evaluated by dual-luciferase reporter gene assay.
Results
miR-373 decreased and closely associated with poor cell differentiation, advanced clinical stage, and shorter survival in hilar cholangiocarcinoma; MBD2 exclusively over-expressed and reciprocally related to miR-373; precursor miR-373 inhibited the luciferase activity of MBD2-3′UTR construct; exogenous miR-373 suppressed the expression of MBD2 and enhanced RASSF1A mRNA in QBC939 cells; anti-miR-373 inhibitor up-regulated the expression of MBD2 and reduced RASSF1A mRNA in HIBEpic cells.
Conclusions
miR-373 is one negative regulator of MBD2. In hilar cholangiocarcinoma, down-expression of miR-373 leads to increase of MBD2, which in turn suppresses the methylation-mediated gene such as RASSF1A.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hilar cholangiocarcinoma, known as Klatskin tumor [1], is an uncommon cancer with an incidence of 0.01–0.2% per year [2]. Though it is relatively rare, hilar cholangiocarcinoma displays highly aggressive malignancy and is considered to be an incurable and rapidly lethal disease despite the recent progress in diagnostic and therapeutic techniques. The 5-year overall survival rate after curative resection ranges from 20 to 40% and 10-year survival is almost zero [3]. Furthermore, patients with inoperable, recurrent, or metastatic disease only can be treated with palliative therapy, such as endoscopic, percutaneous biliary drainage combination with radiotherapy, and chemotherapy. The median survival rate of these cases is only around 9 ~ 12 months [4].
MicroRNAs (miRNAs) are non-coding, single-stranded RNAs of 18–24 nucleotides in length that constitute a novel class of gene regulators [5]. In general, miRNA negatively regulates gene expression through targeting the three prime untranslated region (3′UTR) and consequently triggering mRNA degradation or translational suppression on the basis of complementarity value [6]. miRNAs have recently been validated to play important roles in cancers as miRNAs target about 60% of mammalian genes [7] and are abundant in many human cell types [8]. Depending on the target genes, miRNAs can function as tumor suppressor genes (TSG) or oncogenes [9, 10].
Aberrant DNA methylation is an essential mechanism in gene silencing. Once a given sequence becomes methylated, it either can directly represses transcription by impeding the recognition of transcriptional activators to DNA sequences [11] or by recruiting methyl-CpG-binding domain proteins (MBPs) to modify chromatin compaction [12]. The MBP family consists of five isoforms, including Mecp2, MBD1, MBD2, MBD3, and MBD4. With the exception of MBD4, which is primarily a thymine glycosylase involved in DNA repair [13], all MBPs are implicated in the transcriptional repression mediated by DNA methylation. Mecp2, MBD1, and MBD2 have been demonstrated to be involved in methylation-based gene repression and also affect chromatin structure [14–16]. MBD3 lacks a functional MBD but is an integral subunit of histone deacetylase complex the Mi2–NuRD complex that is recruited through MBD2 [17].
Though the regulations act on mRNA 3′UTR and DNA promoters, respectively, both miRNA and methylation are reversible alteration and are considered as potential therapeutic target [18, 19], so an increasing number of studies have focused on the interaction between miRNA and DNA methylation. Evidence of DNA methylation-mediated down-regulation of miRNAs, which harboring CpG island in promoter region or being embedded in CpG Island have been reported in numbers of groups [20]. Moreover, miRNA interferes with DNA methylation through mediating DNA methyltransferases (DNMTs) 3a, 3b, and DNMT1 has been observed [21, 22].
miR-373 is a tumor-related miRNA in which expression has been reported to be altered in various human cancers including malignant cholangiocytes [23]. In this study, we investigated the aberrant expression of miR-373 in solid tumors of hilar cholangiocarcinoma. Furthermore, we explored the effect of miR-373 on DNA methylation through regulating over-expressed MBPs, MBD2, and its influence on RASSF1A expression in hilar cholangiocarcinoma. We illuminated that miR-373 is a negative regulator of MBD2. Our finding provides evidence on the interaction between miRNAs and DNA methylation.
Materials and Methods
Patients and Tissues
A total 48 patients (Table 1) with both tumors and normal bile duct tissues successfully obtained from operation were involved in this study at Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology (China) from January 2005 to December 2008. Written informed consent was obtained from each patient before sample collection. This study was approved by the Review Board of Tongji Medical College and Hospital.
Cell Culture and Transfection
QBC939 cell line originated from human common bile duct adenocarcinoma was kindly provided by Dr. Shuguang W. from Southwest Hospital of the Third Military Medical University (China) [24]. The human intrahepatic biliary epithelial cell line redesigned as HIBEpic was purchased from Chinese Academy of Sciences Cell bank (China). Cells are cultured in RPMI 1640 medium supplemented with 10% fetal bovine serum (Gibco-BRL). Transfections were performed with Lipofectamine™ LTX and Plus Reagent (Invitrogen) in accordance with the manufacturer’s instructions. Double-stranded miR-373 precursor, single-stranded miR-373 inhibitor, or their relative negative control RNA at a final concentration of 100 nM was introduced into cells.
Quantitative Real-Time PCR for Detection of miR-373, MBPs, and RASSF1A Transcripts
To quantify miR-373, RNAs were extracted using mirVana™ miRNA Isolation Kit (Applied Biosystems). Reverse transcription and qPCR were performed according to the TaqMan MicroRNA Assay protocol (ABi). U6 (RUN6B) was assessed as endogenous control. For determination of MBD1, MBD2, and Mecp2 mRNA levels, RNA was extracted from samples and cells using the RNeasy® mini kit (Qiagen), and cDNA was generated with a reverse-transcription system kit (Invitrogen). And QuantiTect® Primer Assays was used [25] (Qiagen: QT00066843, QT00007084, QT00039361). Values were normalized to 18S RNA expression. For RASSF1A transcripts. Real-time PCR was performed with a standard SYBR-Green PCR kit protocol. β-Actin was used as an endogenous control to normalize the amount of total mRNA in each sample. The primer sequences used were as follows [26]: for RASSF1A5 ′ -CGCGCATTGCAAGTTCAC -3 ′ (forward) and 5 ′ -AGGCTCGTCCACGTTCGT-3 ′ (reverse); for GAPDH, 5 ′ -GGAGTCAAC GGATTTGGTC -3 ′ (forward) and 5 ′ -TGGGTGGAATCATATT GGAACAT-3 ′ (reverse). The real-time PCR reactions were performed in triplicate including no-template controls (NTC). The relative expression was calculated with the comparative Ct method, and fold change was calculated to indicate the alteration clearly [27].
Vectors
Precursor miR-373 clones (miRNA Accession: MI0000781) and scrambled control clone, Anti-miR™ miRNA Inhibitor (anti-miR-373; AM0000781) and negative control (anti-neg) were bought from Ambion. Wild-type MBD2 3 ′ UTR vector pEZX-MBD2-3UTR (Gene Accession: NM_004992.3) and scrambled control were constructed at the company GeneCopoeia. The potential binding sequences of miR-373 at the MBD2 3′ UTR were mutated using the QuikChange™ Site-Directed Mutagenesis Kit (Stratagene) using the following primers: 5 ′ - TGCAATCTACTGGAAAC GGAACGCTTACGTAAAAC -3 ′ and 5 ′ - CAAAATGCATTCGCAAGGCAAAGGTCATCTAA CGT-3 ′ , mutated vector was confirmed by sequencing.
Luciferase Reporter Gene Assay
For 3′UTR Luciferase reporter assay, QBC939 cells were cultured in 96-well plates and each was transfected with 100 ng MBD2-3′UTR or MBD2-3′UTR-mut together with 100 nM Pre-miR-373 or Pre-neg. Forty-eight hours after transfection, the cells were harvested and assayed with the Dual-Luciferase Reporter Assay Kit (Promega) according to the manufacturer’s instructions.
Western Blotting
Proteins were analyzed by Western blotting with primary antibody against Mecp2 (1:500, Sigma–Aldrich), MBD1(1:1,000, Millipore) and MBD2 (1:1,000, Millipore), which produce a signal of size of ~75, ~61, and ~50 kDa, respectively. GAPDH (1:5,000, abcam) was used as a loading control.
Statistical Analysis
Data analysis was achieved using SPSS for Windows, version 14.0 (Chicago, USA). Student’s t test, one-way ANOVA, and Pearson's Chi-square test was used according to the data characteristic. The duration of recurrence of hilar cholangiocarcinoma and death measured from the date of surgery was referenced against disease-free survival and overall survival time. Survival duration was calculated via the Kaplan–Meier method. Log-rank test was employed for comparison of cumulative survival rate and disease-free survival in the patient group. P values of <0.05 were deemed statistically significant.
Results
miR-373 Decreased and Associated with Poor Cell Differentiation, Advanced Clinical Stage, and Shorter Survival in Hilar Cholangiocarcinoma
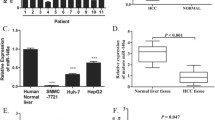
To assess the alteration of miR-373 in hilar cholangiocarcinoma, we measured the expression of miR-373 using TaqMan miRNA assay. Compared to normal bile duct tissues, miR-373 was significantly down-regulated in 35 (72.92%) tumors including seven samples in which miR-373 was undetectable (p < 0.01; Fig. 1a). Fold-change analysis showed a 2.94-fold decrease in tumors than control (p < 0.01, Fig. 1b). In QBC939 cells, expression of miR-373 decreased dramatically in comparison with HIBEpic cells (Fig. 1a).
Expression of miR-373 and its association with clinicopathological factors in patients with hilar cholangiocarcinoma. a Representative expression of miR-373 decreased in tumor and in QBC939 detected by RT–PCR (*p < 0.05). b TaqMan microRNA assay of miR-373 displayed 2.94-fold down-regulation in the tumor group (**p < 0.01). c Relationship between miR-373 expression and overall survival in the patients with hilar cholangiocarcinoma. The median overall survival time was 16.3 and 29.7 months in the low- and high- miR-373 groups, respectively (p < 0.05). d Kaplan–Meier disease-free survival. The median disease-free survival times were 11.2 and 23.4 months in the low- and high-miR-373 groups, respectively, *p < 0.05
In identification of the correlation between miR-373 expression and clinicopathological factors, miR-373 showed low expression in specimens with poor cell differentiation (p = 0.031) and advanced clinical stages (stage III, IV vs. I, II; p = 0.017; Table 1) while no association was observed with age, gender, tumor size, different pathological type, bismuth classification, or lymphatic metastasis (p > 0.05). For further evaluation of the role of miR-373 in hilar cholangiocarcinoma, we executed a small survival analysis according to the miR-373 expression level. Kaplan–Meier analysis showed that down-regulation of miR-373 was relevant to decreased overall survival (Fig. 1c, p < 0.05, log-rank test) and disease-free survival (Fig. 1d, p < 0.05).
MBD2 Exclusively Over-Expressed and Reciprocally Related to Down-Regulation of miR-373 in Hilar Cholangiocarcinoma
Among the MBPs, MBD1, MBD2, and Mecp2 have been established to be involved in the methylation-dependent repression of transcription. Therefore, we explored the expression of MBD1, MBD2, and Mecp2 at the level of mRNA and protein with QuantiTect® Primer Assays and Western blotting. A dramatic increase of MBD2 was found at both the level of mRNA and protein in hilar cholangiocarcinoma compared to normal bile duct tissues. Though MBD1 and Mecp2 displayed upward and downward tendency, respectively, no significance was revealed in statistical analysis (Fig. 2a, b).
Expression of methyl-CpG-binding proteins (MBPs) and the correlation of MBD2 with miR-373 in hilar cholangiocarcinoma. a MBPs mRNA expression in tumor and control tissues. mRNAs were determined using QuantiTect® Primer Assays, and data were normalized to 18S RNA and related to control tissue to show fold change. b MBPs protein in tumor and control tissues were measured by Western blotting, relative optical density was normalized to GAPDH to indicate protein content *p < 0.05. c Down-expression of miR-373 versus up-regulation of MBD2 in patients with hilar cholangiocarcinoma. The expression of miR-373 was measured with TaqMan microRNA assay. After Ct values were determined and normalized to internal control, 2−ΔΔCt was calculated and related to normal tissues was used to indicate the expression. d The miR-373 and MBD2 expression levels were inversely correlated. 2−ΔΔCt values of miR-373 and MBD2 mRNA were subjected to a Pearson correlation analysis (r = −0.493, p < 0.05, Pearson’s correlation)
Next, we assessed the effect of miR-373 on MBD2 using data obtained from quantitative PCR. A significantly inverse correlation was observed between MBD2 mRNA and miR-373 (n = 48, r = −0.462, p = 0.02, Pearson’s correlation; Fig. 2c, d). These data showed the reciprocal regulation of miR-373 and MBD2, which suggest that down-regulation of miR-373 may play a causal role in MBD2 overexpression in hilar cholangiocarcinoma.
MBD2-3′UTR Is a Functional Target of miR-373
MBD2 was predicted as one of high-scoring candidate genes of miR-373 targets by four algorithms (TargetScan, PicTar, miRanda, and miRbase Target). As shown in Fig. 3a, the MBD2-encoded mRNA contains a 3′-UTR element ranged from dinucleotide 295–301 bp, which is stringently complementary to seed sequence of miR-373, and it indicates that miR-373 would directly target this site. To investigate whether miR-373 directly recognizes the MBD2-3′UTR, we transfected the wild-type MBD2-3′UTR construct into QBC939 cells in combination with pre-miR-373 or miR-neg control. As result, miR-373 led to a 45.8% decrease of reporter activity of MBD2-3′UTR construct compared to miR-neg. After the conserved targeting regions for miR-373 recognition were mutated, the relative luciferase activity of the reporter gene was also restored (Fig. 3b, p < 0.01). Furthermore, experiments were carried out using anti-miR-373, which bound to the endogenous miR-373 and thereby antagonized its activity. When HIBEpic cells (high endogenous miR-373 expression) were co-transfected with anti-miR-373 and MBD2-3′UTR construct, a significant increase in activity of the wild-type reporter was observed (Fig. 3c). Take together, these observations suggest that the predicted complementary sequence in MBD2-3′UTR is a functional element of miR-373.
MBD2 is a target of miR-373. a Bioinformatics predicted interactions of miR-373 and their binding sites at the 3′UTR of MBD2 (TargetScan 4.0). b Luciferase reporter assay revealed a reduction of luciferase activity following the cotransfection of MBD2-3′UTR construct together with pre-miR-373 in QBC939 cells (low endogenous miR-373 expression). c The increase of luciferase activity following the cotransfection of MBD2-3′UTR construct together with anti-miR-373 in HIBEpic cells (high endogenous miR-373 expression). Relative luciferase activities are normalized by the ratio of firefly and Renilla luciferase activities, data represent the mean ± SEM of the three independent experiments performed in triplicate, *p < 0.05, **p < 0.01
miR-373 Regulates MBD2 Expression In Vitro
To validate that MBD2 expression was down-regulated by miR-373 in hilar cholangiocarcinoma, we transfected miR-373 precursor or scrambled negative control into QBC939 cells, and anti-miR-373 inhibitor or miRNA inhibitor negative control into HIBEpic cells. After 48 or 72 h of transfection, we measured the mRNA and protein expression levels of MBD2, respectively. Our results showed that enforced miR-373 expression led to a reduction of MBD2 expression at both the mRNA and protein levels in comparison with wild-type QBC939 cells (Fig. 4a, b). On the contrary, the inhibition of miR-373 in HIBEpic cells increased MBD2 expression (Fig. 4c, d).
miR-373 regulates MBD2 expression in vitro. Total RNA was extracted at 48 h and proteins were lysed from cells at 72 h after transfection. MBD2 mRNA was measured using QuantiTect® Primer Assay, and data were normalized to 18S RNA and presented with fold change to wild-type cells. MBD2 protein was examined by Western blotting, and data were related to GAPDH and presented with relative optical density. Transfection was performed with three independent experiments in triplicate, *p < 0.05. a, b QBC939 cells were transfected with 100 nM pre-miR-373 or pre-miR-neg. c, d HIBEpic cells were transfected with 100 nM anti-miR-373 or anti-miR-neg, *p < 0.05, **p < 0.01
miR-373 Reduces RASSF1A Expression by Targeting MBD2
To test the downstream effect of MBD2 down-regulation by miR-373, we investigated whether the enforced expression of miR-373 could functionally result in RASSF1A reactivation. QBC939 cells were transfected with miR-373 precursor or HIBEpic cells were transfected with miR-373 inhibitor as described above, respectively. RASSF1A mRNA was measured at 72 h after transfection. We observed a 5.9-fold increase of RASSF1A following MBD2 suppression in QBC939 cells treated with miR-373 precursor in comparison to wild-type cells (Fig. 5a) and a 4.3-fold reduction following MBD2 activation in HIBEpic cells treated with miR-373 inhibitors (Fig. 5b).
Effect of miR-373 on RASSF1A expression. Total RNA was extracted at 72 h after transfection. RASSF1A mRNA was determined by SYBR Green real-time PCR, GAPDH was used as internal control. Data were obtained from three independent transfections in triplicate and presented with fold change to wild-type cells, **p < 0.01. a QBC939 cells were transfected with 100 nM pre-miR-373 or pre-miR-neg. b HIBEpic cells were transfected with 100 nM anti-miR-373 or anti-miR-neg
Discussion
The miR-373 gene was reported to display a controversial feature in different cancers. In testicular germ cell tumors [28], esophageal cancer [29], and breast cancer [30], miR-373 was identified as a novel oncogene. In contrast, miR-373 behaves as a TSG in prostate cancer [31] and malignant cholangiocytes including extrahepatic cholangiocarcinoma cell lines [23]. Despite the complexities of different conclusions, function of miR-373 has been established well to be implicated in the process of tumorgenesis, invasion, and metastasis. In the present study, we delineated that miR-373 is dramatically down-regulated in hilar cholangiocarcinoma and correlated closely with poor cell differentiation, advanced clinical stages, and shorter survival. Our finding is in agreement with the last two reports. How to explain these directly opposite results remains unclear. The expression of individual miRNA with pattern of strict specificity to tissue, developmental stage and clinical features [32]; or the different regulation network involving distinct target gene in particular cancer could be considered as the cause of discrepant conclusions.
Methyl-CpG-binding domain proteins (MBPs) are critical modificator in methylation-mediated silencing gene. In this study, MBD2 was detected as the exclusive MBP, which was expressed aberrantly in hilar cholangiocarcinoma. The reciprocal correlation of expression between miR-373 and MBD2 encourage us to explore whether miR-373 is a negative regulator of MBD2. We first predict the alignment of miR-373 with MBD2 3′UTR using four algorithms. Culminated in consensus, the seed region of miR-373 matches with nucleotides 295–301 of the MBD2-3′UTR, which suggests MBD2 is a direct target of miR-373. However, not all miRNAs identified in this manner are likely to be functional, since factors such as steric hindrance may render them inaccessible to the mRNA [33]. Hence, functional validation experiments including transfection of miR-373 precursors and anti-miR-373 inhibitor were performed. As described in the Results section, the miR-373 precursor suppressed the relative luciferase activity of MBD 3′UTR construct and MBD2 expression in QBC939 cell lines. On the contrary, anti-miR-373 inhibitor up-regulated the expression of MBD in HIBEpic cells. These findings indicate that miR-373 functionally regulates the expression of MBD2 by targeting to 3′UTR.
Furthermore, we select RASSF1A as an effect gene to assess downstream influence involved in the regulation of MBD2 by miR-373. RASSF1A is a well-studied tumor suppressor gene that has been concluded to be silenced or suppressed by promoter methylation and MBPs recruitment in various human cancers [34]. Our previous studies have also proved that RASSF1A was hypermethylated and repressed in hilar cholangiocarcinoma samples [35], so RASSF1A is an applicable module for effect analysis regarding methylation-mediated. In the present study, RASSF1A expression decreased following suppression of MBD2 by miR-373 in QBC939 cells, and increased following activation of MBD2 by anti-miR-373 inhibitor in HIBEpic cells. The consistency of RASSF1A fluctuation with regulation of MBD2 by miR-373 indirectly demonstrates that the MBD2 is a functional target of miR-373.
It should be explained two cell lines used in this study, QBC939 and HIBEpic. The QBC939 cell originated from human common bile duct adenocarcinoma, which regarding biological characteristics, is the same as hilar cholangiocarcinoma, and the expression of miR-373 and MBD2 is coincident with hilar cholangiocarcinoma. The HIBEpic cell line originated from human intrahepatic biliary epithelial cells in which the miR-373 and MBD2 expression is similar to normal bile duct tissues. We finally chose it as the control for QBC939.
In conclusion, in this study we demonstrate that miR-373 is one negative regulator of MBD2. In hilar cholangiocarcinoma, down-expression of miR-373 led to an increase of MBD2, which in turn suppresses methylation-mediated genes such as RASSF1A.
References
Klatskin G. Adenocarcinoma of the hepatic duct at its bifurcation within the portal hepatic. An unusual tumor with distinctive clinical and pathological features. Am J Med. 1965;38:241–256.
Akoad M, Jenkins R. Proximal biliary malignancy. Surg Clin North Am. 2008;88(6):1409–1428.
Jarnagin WR, Fong Y, DeMatteo RP, et al. Staging, resectability, and outcome in 225 patients with hilar cholangiocarcinoma. Ann Surg. 2001;234:507–517.
Aljiffry M, Walsh MJ, Molinari M. Advances in diagnosis, treatment and palliation of cholangiocarcinoma: 1990–2009. World J Gastroenterol. 2009;15(34):4240–4262.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297.
Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136:215–233.
Friedman RC, Farh KK, Burge CB, Bartel DP. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009;19:92–105.
Lim LP, Lau NC, Weinstein EG, et al. The microRNAs of Caenorhabditis elegans. Genes Dev. 2003;17:991–1008.
Esquela-Kerscher A, Slack FJ. Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer. 2006;6:259–269.
He L, Thomson JM, Hemann MT, et al. A microRNA polycistron as a potential human oncogene. Nature. 2005;435:828–833.
Watt F, Molloy PL. Cytosine methylation prevents binding to DNA of a HeLa cell transcription factor required for optimal expression of the adenovirus major late promoter. Genes. 1988;2:1136–1143.
Bird AP, Wolffe AP. Methylation-induced repression—belts, braces, and chromatin. Cell. 1999;99:451–454.
Hendrich B, Hardeland U, Ng HH, Jiricny J, Bird A. The thymine glycosylase MBD4 can bind to the product of deamination at methylated CpG sites. Nature. 1999;401:301–304.
Nan X, Campoy FJ, Bird A. MeCP2 is a transcriptional repressor with abundant binding sites in genomic chromatin. Cell. 1997;88(4):471–481.
Fujita N, Takebayashi S, Okumura K, et al. Methylation-mediated transcriptional silencing in euchromatin by methyl-CpG binding protein MBD1 isoforms. Mol Cell Biol. 1999;19:6415–6426.
Ng HH, Zhang Y, Hendrich B, et al. MBD2 is a transcriptional repressor belonging to the MeCP1 histone deacetylase complex. Nat Genet. 1999;23:58–61.
Saito M, Ishikawa F. The mCpG-binding domain of human MBD3 does not bind to mCpG but interacts with NuRD/Mi2 components HDAC1 and MTA2. J Biol Chem. 2002;277:35434–35439.
Li C, Feng Y, Coukos G, Zhang L. Therapeutic microRNA strategies in human cancer. AAPS J. 2009;11:747–757.
Jean-Pierre J. DNA methylation as a therapeutic target in cancer. Clin Cancer Res. 2007;13:1634–1637.
Wu L, Zhou H, Zhang Q, et al. DNA methylation mediated by a microRNA pathway. Mol Cell. 2010;38:465–475.
Fabbri M, Garzon R, Cimmino A, et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci USA. 2007;104:15805–15810.
Garzon R, Liu S, Fabbri M, et al. MicroRNA-29b induces global DNA hypomethylation and tumor suppressor gene reexpression in acute myeloid leukemia by targeting directly DNMT3A and 3B and indirectly DNMT1. Blood. 2009;113:6411–6648.
Meng F, Henson R, Lang M, et al. Involvement of human micro-RNA in growth and response to chemotherapy in human cholangiocarcinoma cell lines. Gastroenterology. 2006;130:2113–2129.
Li D, Chen J, Gao Z, et al. 67-kDa laminin receptor in human bile duct carcinoma. Eur Surg Res. 2009;42:168–173.
Careen K, Nicole S, Ad G, et al. Role of DNA methylation and methyl-DNA binding proteins in the repression of 5-lipoxygenase promoter activity. Biochim Biophys Acta. 2010;1801:49–57.
Vera NS, Ekaterina AA, Tatiana TK, et al. Simultaneous down-regulation of tumor suppressor genes RBSP3/CTDSPL, NPRL2/G21 and RASSF1A in primary non-small cell lung cancer. BMC Cancer. 2010;10:75–87.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–408.
Voorhoeve PM, le Sage C, Schrier M, et al. A genetic screen implicates miRNA-372 and miRNA-373 as oncogenes in testicular germ cell tumors. Cell. 2006;124:1169–1181.
Lee KH, Goan YG, Hsiao M, et al. MicroRNA-373 (miR-373) post-transcriptionally regulates large tumor suppressor, homolog 2 (LATS2) and stimulates proliferation in human esophageal cancer. Exp Cell Res. 2009;315:2529–2538.
Huang Q, Gumireddy K, Schrier M, et al. The microRNAs miR-373 and miR-520c promote tumour invasion and metastasis. Nat Cell Biol. 2008;10:202–210.
Yang K, Handorean AM, Iczkowski KA. MicroRNAs 373 and 520c are downregulated in prostate cancer, suppress CD44 translation and enhance invasion of prostate cancer cells in vitro. Int J Clin Exp Pathol. 2009;2:361–369.
Foteini C, Florian R, Raju T, et al. Ancient animal microRNAs and the evolution of tissue identity. Nature. 2010;463:1084–1088.
Lai KW, Koh KX, Loh M, et al. MicroRNA-130b regulates the tumour suppressor RUNX3 in gastric cancer. Eur J Cancer. 2010;46(8):1456–1463.
Dammann R, Li C, Yoon JH, Chin PL, Bates S, Pfeifer GP. Epigenetic inactivation of a RAS association domain family protein from the lung tumour suppressor locus 3p21.3. Nat Genet. 2000;25:315–319.
Chen YJ, Tang QB, Zou SQ. Inactivation of RASSF1A, the tumor suppressor gene at 3p21.3 in extrahepatic cholangiocarcinoma. World J Gastroenterol. 2005;11:1333–1338.
Acknowledgments
This work was supported in part by grants from the Doctoral Program of Higher Education of China (No. 20070487114), and the Natural Science Foundation of Hubei Province, China (No. 2008CDB159).
Conflicts of interest
No potential conflicts of interest exist.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chen, Y., Luo, J., Tian, R. et al. miR-373 Negatively Regulates Methyl-CpG-Binding Domain Protein 2 (MBD2) in Hilar Cholangiocarcinoma. Dig Dis Sci 56, 1693–1701 (2011). https://doi.org/10.1007/s10620-010-1481-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-010-1481-1