Abstract
The aim of this study was to assess the genetic variation and population structure of the geophyte Leucojum aestivum L. across the Po river valley (N-Italy), to inform conservation management actions with the selection of most suitable source populations for translocation purposes. L. aestivum is self-incompatible and occurs in S-Europe in fragmented wetlands and lowland forests along rivers. The species is particularly interesting for habitat restoration practices for its simplicity of ex situ conservation and cultivation. AFLP analyses were carried out on 16 fragmented populations, using four primer combinations. Correlations between genetic variation and demographic and ecological traits were tested. AFLP produced a total of 202 bands, 95.5% of which were polymorphic. Our results suggest that L. aestivum holds low to moderate levels of genetic diversity (mean Nei’s genetic diversity: H = 0.125), mostly within-population. We found a gradient of two main biogeographic groups along western and eastern populations, while the STRUCTURE analysis found that the most likely number of clusters was K = 3, shaping a partially consistent pattern. We explain the unusual negative correlation between genetic variation and population size with the high rate of vegetative reproduction. The levels of population differentiation suggest that fragmentation in L. aestivum populations has occurred, but that an active gene flow between fragmented populations still exists, maintained by flooding events or pollinators. Conservation management actions should improve habitat connectivity, especially for pollinators that vehicle upstream gene flow. Moreover, the west–east structure due to the lithological composition of the gravel and sand forming the alluvial plain of the Po river, should be considered when selecting source populations for translocation purposes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Over the last centuries, habitat modifications and fragmentation as a result of human activity have caused a sharp decrease in many European herbaceous forest species, both in numbers of populations and individuals within populations (Petit et al. 2004; Alvarega and Pôrto 2007; Kolk and Naaf 2015). Habitat fragmentation reduced the availability of forest habitats and negatively affected the spatial arrangement of habitat patches, especially in lowland riparian forest of central and southern Europe (Tockner and Stanford 2002; Ezard and Travis 2006).
Several studies have shown that habitat fragmentation can negatively affect genetic diversity by decreasing species population size and interrupting the gene flow among populations (Hoehn et al. 2007; Dixo et al. 2009). However, historical and current gene flow should be considered separately. Indeed, Reisch et al. (2017) showed that current levels of genetic diversity may depend more on the historical landscape structure than on present fragmentation. Consequently, fragmented populations hold (or reach in time) low levels of genetic diversity and are subjected to high risks of inbreeding depression and/or genetic drift (Reed and Frankham 2003). In addition, they often have diminished evolutionary potential and chances of survival, due to higher effects of stochastic events in small than in large populations (Frankham 2005; Ortego et al. 2015). Effects of fragmentation are essentially species-specific, depending on factors such as the spatial structure of a population, the population size, the species dispersal ability (i.e. gene flow), the mating system, and the initial population genetic diversity (Luoy et al. 2007; Dixo et al. 2009). On the other hand, these intrinsic factors often change with both, local environmental and landscape characteristics (Mable and Adam 2007; Eckert et al. 2010). Ecosystem modifications related to fragmentation (i.e. edge effect, connectivity, spatial redistribution of pollinator and/or predators, etc.) may induce the shift of several reproductive traits and a change in the population dynamics (Jacquemyn et al. 2012; Gargano et al. 2017). It is well known how local environmental factors can exert a selective pressure on plant life traits, affecting individual survival under the new environment and/or promoting local adaptation (Ellis and Weis 2006).
Plant populations growing in different forest habitats can be exposed to different demographic and gene flow patterns (Shao et al. 2015). Populations located in old forest patches tend to have higher levels of genetic diversity than populations located in young forest patches (Jacquemyn et al. 2004). At the landscape level, species growing in riparian forests along rivers may be subject to unidirectional (downstream) dispersal and gene flow (Pollux et al. 2007). On the other hand, the individual age of the plants may be more important for the level of genetic diversity than the age of the habitat (Powolny et al. 2016).
For these reasons, investigating species biology, ecology and genetic patterns of fragmented populations may help envisage their fate and implement proper in situ species conservation and habitat management actions, including the development of conservation action plans at the global (e.g. for steno-endemic taxa) or regional levels. Specifically, restoration practices aiming to re-establish effective population sizes, adequate levels of genetic diversity and gene flow in fragmented populations and species contribute to their long-term survival (Menges 2008). It follows that source material for reinforcement/reintroduction should contain levels of genetic diversity similar to that of wild populations (Menges 2008; Godefroid et al. 2011; IUCN 2013). In addition, re-establishing natural levels of intra-population genetic diversity usually requires the restoration of habitat connectivity (McKay et al. 2005; Di Battista 2008).
In the Po Valley (Italy) the C-S-European/W-Asiatic geophyte Leucojum aestivum L. subsp. aestivum (Amaryllidaceae) grows in fragmented floodplain habitats such as riparian forests with Alnus glutinosa and Salix alba, sedge banks, reed communities and wet grasslands. As a result of anthropogenic habitat fragmentation and degradation, L. aestivum is facing a strong population decline across Europe and an increased fragmentation of its range. Although not globally threatened (Lansdown 2014), the species is protected in many European regions and countries (Parolo et al. 2011) and vulnerable in Italy (Orsenigo S., in verbis). Thus, this species is particularly important for the restoration of lowland riparian habitats, because of its high conservation value and because it is relatively easy to reproduce ex situ and to reintroduce in the wild (Abeli et al. 2016). However, any reintroduction or reinforcement activities should select suitable source material, of known origin and with adequate levels of genetic diversity (Godefroid et al. 2011) and considering the species-specific reproductive traits that might affect the reproductive performance in reinforced/reintroduced populations. L. aestivum is self-incompatible and insect pollinated. Reproductive performance is density-dependent, with dense populations usually producing higher fruit-set and seed-set than less dense stands due to higher pollinator visitation rates (Abeli et al. 2016). Fruits ripen between May and June and are often dispersed at long distances by river flooding events. The plant also exhibits vegetative reproduction through secondary bulbs developing from main bulbs, resulting in clumps of several shoots growing very close to each other (Parolo et al. 2011). Despite many aspects of the biology and the ecology of L. aestivum being well-known, the lack of information on the population genetic structure and patterns of gene flow make safe translocations difficult.
Amplified fragment length polymorphism (AFLP) molecular markers have been successfully used to estimate the genetic diversity of species belonging to the Amaryllidaceae family (Medrano et al. 2014) and of other endangered or fragmented species in the Po Valley (e.g. Bruni et al. 2013; Orsenigo et al. 2016). In the case of L. aestivum AFLP may provide valuable information for its conservation management (Zaya et al. 2017).
The aim of this study was to assess the genetic structure and the level of genetic diversity within and between natural populations of L. aestivum growing in the Po Valley. This information will favour the conservation management of this species (e.g. development of a science-based action plan and to assess which populations are best suitable as source populations for reinforcement or reintroduction). Specific aims of this study were: (1) to investigate the relationship between population genetic variation, demography and reproductive traits; (2) to investigate the relationship between population genetic diversity and the ecological characteristics of the sites of occurrence (i.e. soil physical and chemical traits; see below). We expect to find: (i) a positive relationship between genetic diversity and population size; (ii) higher genetic variation in downstream populations due to water-mediated unidirectional dispersal; (iii) low between-population differentiation, basically coinciding with the main forest types of the Po Valley, due to recent fragmentation and supposed continuous gene flow due to the dispersal strategy of the species.
Materials and methods
Sampling and population parameters
In May 2009 we collected seeds of L. aestivum from 16 wild populations in the Po Valley (Italy; Fig. 1; Table 1). A single fruit from 15 to 25 clumps per populations was collected and the seeds of each fruit were cultivated as different seed families from each population at the Botanical Garden of the University of Pavia. This collection method was designed to avoid the sampling of clones. Young fresh leaves used for AFLP analyses were sampled from these ex situ cultivated plants. A total of 226 accessions for the AFLP analysis were obtained.
Sampled populations could be assigned to two distinct biogeographic districts characterized by two main European forest types which meet in the Po Valley (EEA 2006; Blasi 2010; Table 1), (a) forests on neutral-acidic soils belonging to the phytosociological alliance Carpinion betuli, which include mesophyllous hornbeam forests of western Europe (and the western parts of the Po Valley); (b) forests on basic soils belonging to the alliance Erythronio-Carpinion, which include mesophyllous oak-hornbeam forests of eastern Europe (and the eastern parts of the Po Valley; Adorni 2016).
Demographic data and germination tests
Demographic and ecological data were collected by the University of Pavia (TA, GP, GR) in April 2009. For each population, the total area occupied, and the perimeter of the population were determined with a differential GPS (Leica™ GX1230) with sub-metric precision. The total number of flowering ramets (reproductive population size) in each population was estimated by counting the number of flowering stems in 3–8 rectangular plots (1 × 0.5 m2), multiplied by the total area. For three very small populations, the total number of flowering individuals was counted. Flowering stem density was estimated by dividing the reproductive population size by the area occupied. In May 2009 when fruits were ripe and ready to disperse the seeds, 20–70 fruiting stems per population were collected. The number of flowers and fruits were counted, and the fruit set estimated as the ratio between the number of developed fruits and the total number of produced flowers (as those flowers which do not develop into fruits were still visible on fruiting stems). Developed fruits were opened and the seed set estimated as ratio between the number of developed seeds and the total number of ovules per fruit.
Germination tests were performed sowing seeds in three replicates of 30 seeds each into 90 mm Petri dishes filled with 1% distilled water-Agar. The Petri dishes were placed in temperature and light-controlled incubators (LMS Ltd, Sevenoaks, UK) with a 12 h daily photoperiod. A temperature move-along experiment was chosen to simulate the seasonal conditions to which seeds of L. aestivum are exposed in the wild after dispersal (for further details see Parolo et al. 2011). The tests started with 20 °C for 21 weeks (= summer conditions). At monthly intervals, the temperature was then reduced to 15, 10 and 4 °C (= autumn and winter conditions) and increased again to 10, 15, 20 and 25 °C at a 4-week interval (= spring and late spring conditions of a second year) and finally decreased to again 20 °C for 11 weeks (= summer conditions of a second year). Germination events were recorded weekly until the end of the test, 60 weeks after sowing. Seed germination could not be tested in populations D and UP due to low seed availability. Additionally, one soil sample per plot was collected in each population and analysed for the following variables: pH, sand, silt, clay, total calcium carbonate (CaCO3), organic carbon (C), organic matter, total nitrogen (N) and available phosphorous (P2O5) following the MIPAAF (2000) standard protocol (for further detail on the methodology see Parolo et al. 2011). A total of 76 soil samples was collected by removing the upper layer containing undissolved organic matter and up to − 10 cm depth.
DNA extraction
Genomic DNA from about 0.1 g of frozen young leaves was isolated using the DNeasy Plant Mini Kit (Qiagen, Hilden, Germany). Quality and quantity of the isolated DNA were determined by absorbance measurements. The DNA was stored at − 20 °C until used.
AFLP and data scoring
For each sample, genomic DNA (ca. 100 ng) was digested for 2 h at 37 °C with EcoRI (1 U) and MseI (1 U), and ligated using a T4 DNA ligase (1 U; Promega, Madison, USA) with MseI-(50 pMol) and EcoRI-adapters (5 pMol). M01 (10 µM) and E01 (10 µM) were used as primer pairs in the pre-selective PCR reaction (psPCR). The psPCR was performed using 5 µl of the restriction and ligation product (diluted 1:5) combined with a reaction mix containing 0.5 µl of 1.25 mM EcoRI- and MseI preselective primers each, 200 µM dNTPs (Invirtogen, Carlsbad, USA), 1.5 µl 10 × PCR buffer (Applied Biosystems, Carlsbad, USA), 0.7 μ AmpliTaq Gold DNA polymerase (Applied Biosystems) and 9.75 µl H2O. The thermocycler protocol was 2 min at 72.0 °C followed by 25 cycles of 20 s at 94.0 °C, 30 s at 56.0 °C and 2 min at 72.0 °C and a final extension of 30 min at 60.0 °C. The product of the psPCR was diluted 1:10.
To detect EcoRI/MseI genomic restriction/ligation fragments, selective PCR reactions were carried out using four different primer combinations (chosen after a screening of 20 different combinations of MseI/EcoRI primers) having three selective nucleotides (Online Resource 1). The selective amplification (selPCR) was performed using 2.5 µl of the psPCR product combined with 1 µl PCR× buffer (Applied Biosystems), 0.6 µl MgCl2 (2.5 mM), 0.25 µl fluorescent labeled EcoRI (1.25 mM) and 0.30 µl MseI (1.25 mM) selective primers (Invitrogen; Carlsbad, USA), 0.7 μ AmpliTaq Gold DNA polymerase (Applied Biosystems) and 4.35 µl H2O. The EcoRI primers were fluorescently labelled 5′-end with 6-carboxyfluorescein (6-FAM). Amplifications were performed using a Mastercycler Gradient thermal cycler (Eppendorf, Hamburg, Germany) with the following cycle profile, 30 s at 94 °C, 1 min at 65 °C and 1 min at 72 °C. The annealing temperature of the first cycle (65 °C) was then reduced by 0.7 °C at each cycle for the subsequent 12 cycles, and kept at 56 °C for the last 25 cycles.
To detect fluorescently labelled DNA fragments, 1 µl of PCR product were mixed with 0.2 µl of GeneScan® LIZ Size Standard (Applied Biosystems, Carlsbad, USA) and 8.8 µl of Hi-Di formamide (Applied Biosystems, Carlsbad, USA). Fragment analysis was performed on a 3730xl DNA Analyzer sequencer (Applied Biosystems, Carlsbad, USA).
The reproducibility of the analysis (from DNA extraction to capillary electrophoresis) was assessed repeating the protocol for 25 samples (10% of the total). The error rate of the analyses was estimated as the total number of loci differences relative to the total number of loci comparisons, and subsequently averaged over the four combinations (Bonin et al. 2007). AFLP electropherograms were analysed and scored with the internal size standard using the RawGeno package, an R CRAN library (Arrigo et al. 2009), following the parameter setting suggested by Arrigo et al. (2012). Only peaks in the 100–800 bp size range were scored. All scoring data were then validated by visual peak inspection.
Genetic diversity and population structure
The binary matrix generated after the scoring was analysed to calculate genetic diversity parameters. The number and proportion of polymorphic loci (P%) were calculated using AFLP-SURV version 1.0 (Vekemans 2002). The parameters, HT (gene diversity in the overall sample), GST (genetic differentiation among populations) and gene flow (Nm) were calculated to estimate genetic variation using Nei’s statistics (Nei 1973, 1977), with the software POPGENE v. 1.31. The H Nei’s (gene diversity) and the effective allele number (ne) were determined using GenAlEx 6.5 (Peakall and Smouse 2006).
In order to investigate population structure and degree of genetic differentiation within populations, among populations and among biogeographic districts, analysis of molecular variance (AMOVA) was performed using the GenAIEx version 6.5 software (Peakall and Smouse 2006). The significance of the estimates was tested through 999 data replications. To visualize the spatial relationships among populations the AFLP binary matrix was also subjected to principal coordinates analyses (PCoA) in PAST software, version 3.09.
The population structure of L. aestivum at the regional level was inferred and individuals assigned to supposed ancestral populations by using the software STRUCTURE v. 2.3.4. which allows the use of dominant markers such as AFLP (Pritchard et al. 2000; Falush et al. 2007). Allele frequencies of the L. aestivum populations were supposed to be correlated, which is a representative model for populations that are expected to be similar due to shared migration events and/or ancestry. To calculate the number of clusters, 20 independent runs of K (K = 1–16) were performed with an admixture model (LOCPRIOR option; estimate λ) at 50,000 runs of burn-in period and 500,000 Markov chain Monte Carlo iterations. To determine the number of clusters we used ΔK, the second-order rate of change in lnP(X|K) for successive values of K (Evanno et al. 2005).
GenAlEx software allowed the calculation of the ΦST values, an estimation of FST for dominant data (Peakall and Smouse 2006).
To assess possible multivariate relationships between geographic, genetic and demographic distances among populations, we calculated pairwise correlations between the correspondent distance matrixes, applying Mantel tests (Mantel 1967) with 999 permutations in ARLEQUIN version 3.5 (Excoffier and Lischer 2010). In particular, to investigate demographic variation across populations, we used demographic data (number of flowering individuals, flowering stem density and seed germination; see Table 1) to generate a distance matrix between population pairs, calculating the Euclidean distance.
To investigate univariate relationships between genetic (H Nei, P%, Na), demographic (population size, density, germination) and ecological (soil) variables linear regression analyses were performed. In addition, to test our second hypothesis a linear regression was performed to identify a relationship between Nei’s genetic variation and population position (i.e. longitude). Finally, t-test was used to investigate differences in ecological variables between the two biogeographic districts (eastern and western Po Valley). Non-normal variables were log-transformed.
Results
Demographic data and germination tests
The reproductive population sizes of the 16 studied populations ranged from 34 to more than 77,000 flowering stems, and density ranged from 0.04 to 35.63 flowering stems/m2 (Table 1). Fruit set and seed set were moderately high ranging from 29 to 86% (mean ± SD: 56 ± 18.03%) and from 22 to 57% (33.75 ± 9.37%), respectively (Table 1), while mean seed weight varied slightly among populations and ranged from 0.07 to 0.15 g (0.09 ± 0.02 g; Table 1). Final germination percentages were high in most populations (86.28 ± 10.76%), being higher than 80% in all but populations O, Q and S and reaching 100% in populations C and L.
AFLP
The four AFLP primer combinations produced a total of 202 reproducible bands ranging from 100 to 700 bp, 193 of which were polymorphic (error-rate of 3.1% across all replicated samples). The most informative AFLP primer pair was E32/M40 (E-ACC/M-AGC) with the production of 56 polymorphic bands (96.55%; Online Resource 1). At the population level, the percentages of polymorphic loci ranged from 25.25% in population C to 84.65% in population R; the number of observed alleles scored the same trend. Populations T exhibited the highest genetic diversity (H = 0.217) (Fig. 2; Table 2). The mean value of H Nei across the 16 populations was 0.125, while the overall genetic diversity (Ht) and gene flow were 0.163 and 1.762, respectively.
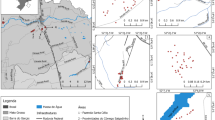
A Spatial genetic structure and population clusters of L. aestivum inferred by Bayesian clustering implemented in STRUCTURE. At each location, pie charts in the map indicate mean proportion of membership of individuals for K = 3 genetic groups; B results of the ΔK calculation (see "Materials and methods" for details); C in the bar diagram different colours (q values) represent the proportion of ancestry in each of the K populations. (Color figure online)
Population genetic structure
The genetic relationships among the L. aestivum populations were assessed by a PCoA analysis (based on Nei’s distance) and performed at the individual level. In the PCoA the first three coordinates explained about 30% of the molecular variance. This analysis allowed us to identify a gradient of the two main biogeographic groups (populations in the western and eastern Po Valley) along axis 3 of the 3D-scatterplot (Fig. 3).
The STRUCTURE analysis found that the most likely number of clusters was K = 3 [highest mean log likelihood: ln P(D) (– 24,769.79)], indicating that populations of L. aestivum are subdivided into three different genetic clusters (Fig. 2). The results of STRUCTURE were set on an admixture model which admits that individuals may have mixed ancestry. The analysis showed a modest degree of structure in L. aestivum populations in the main geographic groups. A slight west/east cline is observable in a higher frequency of the green colour in the eastern Po Valley (pop C, D, E, L, O, S, and T). Assuming a K value = 2, such a cline is still present (Online Resource 2).
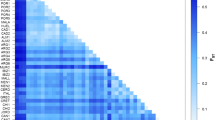
The overall genetic differentiation among populations according to Nei’s statistics was GST = 0.221. The genetic differentiation according to the ΦST value (analogous to FST) estimated with AMOVA was 0.216. The ΦST pairwise comparisons between populations ranged from 0.024 (B vs. FG) and 0.628 (T vs. FG). AMOVA analyses indicated that 78% (estimated variance = 15.45; p < 0.001) of the total genetic variation is ascribed to individuals within populations, while 13% (estimated variance = 2.61; p < 0.001) and 9% (estimated variance = 1.69; p < 0.001) are ascribed to differences among populations and between regions, respectively (Table 3).
The Mantel test revealed a significant correlation between the genetic and geographical pairwise distance matrixes (rxy = 0.197; p < 0.05); the same test detected a statistically not significant relation between the population differentiation (according to ΦST) and geographical pairwise distance matrixes (rxy = − 0.045; p > 0.05).
Correlation between ΦST, genetic, demographic and geographic data
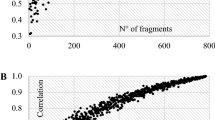
Applying a Mantel test to ΦST, genetic demographic and geographic distance matrixes, we investigated possible patterns of correlation and isolation by distance between all population pairs (n = 16) of L. aestivum. In particular: (a) genetic distance according to Nei and geographic distance matrixes between population were correlated (Mantel Rxy = 0.197; p < 0.044); (b) demographic and geographic distance matrix between populations were negatively correlated (Mantel Rxy − 0.403; p < 0.018). Distance matrixes are reported in Online Resource 3. Linear regression between genetic, demographic and ecological variables showed a weak significant negative relationship between Nei’s genetic diversity and the reproductive population size (R2 = 0.290; F = 5.711; n = 16; p = 0.031), a significant negative relationship between Nei’s genetic diversity and soil pH (R2 = 0.504; F = 14.225; n = 16; p = 0.002) and between the proportion of polymorphic loci and soil pH (R2 = 0.639; F = 24.731; n = 16; p < 0.001). Nei’s genetic variation was not related to longitude (p = 0.213). T test revealed only a small significant difference (p = 0.055) in soil pH between populations in the eastern (mean pH ± st. dev. = 7.15 ± 0.43) and western (6.52 ± 0.79) Po Valley.
Discussion
Our genetic analyses of fragmented populations of L. aestivum across the Po Valley revealed low to moderate within-population genetic diversity and significant (medium to high) levels of between-population differentiation. This pattern is not entirely consistent with the mating system of L. aestivum as an outcrossing, self-incompatible plant species (Parolo et al. 2011; Leimu et al. 2006; Reisch and Bernhardt-Römermann 2014). Within the Amaryllidaceae family, similar patterns of genetic diversity have been found in populations of other species with similar reproductive and ecological traits; (a) Narcissus section pseudonarcissi (gene diversity from 0.022 to 0.118; FST = 0.35) using AFLP (Medrano et al. 2014); (b) Allium oleraceum (gene diversity from 0.113 to 0.204; GS ≥ 0.45) using allozyme (Duchoslav and Staňková 2015). On the other hand, our results are also in countertendency with respect to other species belonging to the Amaryllidaceae family. For instance, Jordàn-Pla et al. (2009) observed high within-population genetic variation (Nei’s gene diversity from 0.490 to 0.756) and low between-population differentiation in Leucojum valentinum Pau in Spain, using RAPD markers. Similarly, Sanaa et al. (2010) found high within-population genetic variation in Pancratium maritimum L., using isozyme markers, but low population differentiation.
The genetic pattern found in L. aestivum (i.e. low to medium within genetic diversity and medium to high between-population differentiation) can be attributed to different intrinsic or extrinsic mechanisms. Primarily, habitat loss, and population isolation and fragmentation may have had a role in the low values of genetic variability detected in some populations (e.g. population C, D and E with genetic diversity values lower than 0.1). Results showed that gene flow has occurred across populations, however the moderate to high inter-population differentiation suggests that the transfer of genetic variation likely mediated by rivers, especially during recurrent flooding events that in the study area can be severe, was higher in the past and is now reduced (Genovese et al. 2007; AdBPo 2009). In contrast to our second hypothesis, historical/current gene flow of L. aestivum is not unidirectional and is likely affected by pollinators and not only by the water as the dispersal agent, because the observed genetic variation is not higher in downstream populations. Another possible explanation for some low values of within-population genetic diversity can be found in the ability of the species to reproduce vegetatively. Indeed, our collecting strategy was designed to avoid sampling clones, since we sampled individuals originating from seeds. However, we cannot exclude that some seeds were produced by mother plants related to each other, which may have contributed to a reduced allelic and genetic diversity in some populations. It is known that high levels of clonality and spatial isolation of populations may have a detrimental effect on genetic diversity within populations and favour genetic differentiation among populations (Despres et al. 2002). In northern Italy, about 30% of L. aestivum individuals remain vegetative (Parolo et al. 2011). A certain degree of clonality may also explain the observed weak (R2 = 0.291) negative relationship between flowering population size and Nei’s H. This pattern contrasts with our first hypothesis and with the general observation of a positive relationship between population size and genetic variation (Ellstrand and Elam 1993), which is generally stronger in self-incompatible species (Leimu et al. 2006).
However, in perennials with the ability to reproduce vegetatively (like L. aestivum), genetic variation due to intrinsic factors (ecological or life history traits due to biotic factors) may be a less important cause of differentiation than ecological clines or biogeographic patterns (geography, geology, or climatic history; see Jacquemyn et al. 2004; Papadopoulou and Knowles 2016). Indeed, consistent with our third hypothesis, Leucojum populations are differentiated along a W–E cline. This last hypothesis is supported by several results of this study. The correlation between demographic and geographic distances seems to arise from the higher values of flowering stems and lower density of western populations than eastern ones. Moreover, the results from the AMOVA show that the two biogeographic districts (corresponding to the two main forest types occurring in the Po Valley), accounted for 9% of molecular variance, while PCoA clearly subdivided the western and the eastern populations. On the other hand, this trend was less evident in STRUCTURE for the individuated clusters (best K = 3) and was not evident when deliberately assuming two clusters (K = 2). This last analysis highlighted the fact that admixture between regional populations via gene flow occurs as the species prominently dispersed its seeds by water (during flooding events that are frequent along the Po Valley, Parolo et al. 2011). The W–E genetic cline could also reflect different post-glacial (re-) colonization events from west (central and western Europe) and east (Balkans) or a directional selection in the two areas (Hewitt 1999).
From a geological point of view, the lithological composition of the gravel and sand forming the alluvial plain of the Po river are also different. Limestone-dolomitic lithologies prevail in the eastern Po Valley, magmatic and metamorphic lithologies prevail in the western Po Valley, as a result of the complex lithology of the Alpine chain (Ruffo 2002 and references therein). These differences also result in dissimilar soil patterns (Ruffo 2002). It is known that substrate and soil types in the Alpine region are among the major drivers shaping genetic structure of plant populations (Alvarez et al. 2009). In fact, although the differences in soil pH in populations belonging to the two biogeographic regions is not statistically significant in our results, we found a clear negative relationship between soil pH, genetic variation and percentage of polymorphic loci. This result is in line with previous studies that demonstrated that a complex network of factors such as landscape parameters (i.e. isolation and fragmentation), population history, habitat history and environmental characters can drive genetic diversity and genetic differentiation of species’ populations (Reisch et al. 2017; Alvarez et al. 2009).
Conservation implications
Knowledge on population genetics is essential to address adequate and successful conservation actions like reconnecting fragmented populations, population management and reinforcement or reintroduction of individual plant species. In this latter respect, two main views dominate the debate. On one side, it is often suggested to mix different provenances to increase the genetic variation of a new combined population and reduce the risk of inbreeding depression (Godefroid et al. 2011; Bupp et al. 2017), while on the other side a second view is that original population genetic identity should be maintained (especially in species reinforcement activities) in order to avoid outbreeding depression (Huff et al. 2011; Orsenigo et al. 2016). Both approaches are valid, but applicable under different circumstances. Conservation genetic studies like the one presented in this paper drive the choice of the best source populations. In the specific case of L. aestivum, within-population genetic diversity is very important as the species is self-incompatible. The establishment of new populations or reinforcement of existing populations should consider within-population genetic diversity, more than a mix between substantially similar populations. Nevertheless, the distinction between eastern and western populations and the relationship between genetic diversity and soil pH suggests some caution when moving individuals far from the original source area. In such cases, ecological similarity other than genetic variation and differentiation should be considered to select source material for reintroduction and reinforcement (Lawrence and Kaye 2011). Importantly, the current gene flow between populations may also suggest that range connectivity may not be necessary if pollinators and rivers will continue to vector inter-population pollen and seed exchanges. It is therefore crucial that the role of low-impact farming systems and of suitable ecological corridors for pollinators are maintained (Paracchini et al. 2015).
References
Abeli T, Cauzzi P, Rossi G, Adorni M, Vagge I, Parolo G, Orsenigo S (2016) Restoring population structure and dynamics in translocated species: learning from wild populations. Plant Ecol 217:183–192
AdBPo (2009). Il rischio alluvionale sui fiumi di pianura - Stato dell’arte in materia di valutazione e gestione del rischio alluvioni. Diabasis - Autorità di Bacino Fiume Po, Parma
Adorni M (2016) La vegetazione legnosa in Emilia. Censimento e analisi delle fitocenosi arboree e arbustive. Istituto per i Beni Artistici Culturali della Regione Emilia-Romagna, Bologna
Alvarenga LDP, Pôrto KC (2007) Patch size and isolation effects on epiphytic and epiphyllous bryophytes in the fragmented Brazilian Atlantic Forest. Biol Conserv 134:415–427
Alvarez N, Thiel-Egenter C, Tribsch A et al (2009) History or ecology? Substrate type as a major driver of spatial genetic structure in Alpine plants. Ecol Lett 12:632–640
Arrigo N, Holderegger R, Alvarez N (2012) Automated scoring of AFLPs using RawGeno v 2.0, a free R CRAN library. Methods Mol Biol 888:155–175
Arrigo N, Tuszynski JW, Ehrich D, Gerdes T, Alvarez N (2009) Evaluating the impact of scoring parameters on the structure of intra–specific genetic variation using RawGeno, an R package for automating AFLP scoring. BMC Bioinform 10:33
Blasi C (2010) La vegetazione d’Italia con carta delle serie di vegetazione in scala 1:500.000. Palombi Editore, Roma
Bonin A, Ehrich D, Manel S (2007) Statistical analysis of amplified fragment length polymorphism data: a toolbox for molecular ecologists and evolutionists. Mol Ecol 16:3737–3758
Bruni I, Gentili R, De Mattia F, Cortis P, Rossi G, Labra M (2013) A multi-level analysis to evaluate the extinction risk and the conservation strategy of the aquatic fern Marsilea quadrifolia L. in Europe. Aquat Bot 111:35–44
Bupp G, Ricono A, Peterson CL, Pruett CL (2017) Conservation implications of small population size and habitat fragmentation in an endangered lupine. Conserv Genet 18:77–88
Despres L, Loriot S, Gaudeul M (2002) Geographic pattern of genetic variation in the European globeflower Trollius europaeus L. (Ranunculaceae) inferred from amplified fragment length polymorphism markers. Mol Ecol 11:2337–2347
Di Battista JD (2008) Patterns of genetic variation in anthropogenically impacted populations. Conserv Genet 9:141–156
Dixo M, Metzger JP, Morgante JS, Zamudio KR (2009) Habitat fragmentation reduces genetic diversity and connectivity among toad populations in the Brazilian Atlantic Coastal Forest. Biol Conserv 142:1560–1569
Duchoslav M, Staňková H (2015) Population genetic structure and clonal diversity of Allium oleraceum (Amaryllidaceae), a polyploid geophyte with common asexual but variable sexual reproduction. Folia Geobot 50:123–136
Eckert CG, Kalisz S, Geber MA et al (2010) Plant mating systems in a changing world. Trends Ecol Evol 25:35–43
EEA (2006) European forest types categories and types for sustainable forest management reporting and policy. EEA technical reports, 9, European Environment Agency, Copenhagen
Ellis AG, Weis AE (2006) Coexistence and differentiation of ‘flowering stones’: the role of local adaptation to soil microenvironment. J Ecol 94:322–335
Ellstrand NC, Elam DR (1993) Population genetic consequences of small population size: implications for plant conservation. Annu Rev Ecol Syst 24:217–242
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Ezard THG, Travis JMJ (2006) The impact of habitat loss and fragmentation on genetic drift and fixation time. Oikos 114:367–375
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7:574–578
Frankham R (2005) Genetics and extinction. Biol Conserv 26:131–140
Gargano D, Fenu D, Bernardo L (2017) Local shifts in floral biotic interactions in habitat edges and their effect on quantity and quality of plant offspring. AoB Plants. https://doi.org/10.1093/aobpla/plx031
Genovese E et al (2007) An assessment of weather-related risk in Europe: maps of flood and drought risks. EC Joint Research Center, Ispra
Godefroid S, Piazza C, Rossi G et al (2011) How successful are plant species reintroductions? Biol Conserv 144:672–682
Hewitt GM (1999) Post-glacial re-colonization of European biota. Biol J Linn Soc 68:87–112
Hoehn M, Sarre D, Henle K (2007) The tales of two geckos: does dispersal prevent extinction in recently fragmented populations? Mol Ecol 16:3299–3312
Huff DD, Miller LM, Chizinski CJ, Vondracek B (2011) Mixed-source reintroductions lead to outbreeding depression in second-generation descendants of a native North American fish. Mol Ecol 20:4246–4258
IUCN (2013) Guidelines for reintroductions and other conservation translocations. Version 1.0. Gland. IUCN Species Survival Commission, Switzerland
Jacquemyn H, Honnay O, Galbusera P, Roldán-Ruiz I (2004) Genetic structure of the forest herb Primula elatior in a changing landscape. Mol Ecol 13:211–219
Jacquemyn H, De Meester L, Jongejans E, Honnay O (2012) Evolutionary changes in plant reproductive traits following habitat fragmentation and their consequences for population fitness. J Ecol 100:76–87
Jordán-Pla A, Estrelles E, Boscaiu M, Soriano P, Vicente O, Mateu-Andrés I (2009) Genetic variability in the endemic Leucojum valentinum. Biol Plantarum 53:317–320
Kolk J, Naaf T (2015) Herb layer extinction debt in highly fragmented temperate forests—completely paid after 160 years? Biol Conserv 182:164–172
Lansdown RV (2014) Leucojum aestivum. The IUCN red list of threatened species 2014: T164488A45461549. https://doi.org/10.2305/IUCN.UK.2014-1.RLTS.T164488A45461549.en. Accessed on 26 Jan 2017
Lawrence BA, Kaye TN (2011) Reintroduction of Castilleja levisecta: effects of ecological similarity, source population genetics, and habitat quality. Restor Ecol 19:166–176
Leimu R, Mutikainen P, Koricheva J, Fischer M (2006) How general are positive relationships between plant population size, fitness and genetic variation? J Ecol 94:942–952
Luoy D, Habel JC, Schmitt T, Assmann T, Meyer M, Müller P (2007) Strongly diverging population genetic patterns of three skipper species: the role of habitat fragmentation and dispersal ability. Conserv Genet 8:671–681
Mable BK, Adam A (2007) Patterns of genetic diversity in outcrossing and selfing populations of Arabidopsis lyrata. Mol Ecol 16:3565–3580
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
McKay K, Christian CE, Harrison S, Rice KJ (2005) How local is local? A review of practical and conceptual issues in the genetics of restoration. Restor Ecol 13:432–440
Medrano M, López-Perea E, Herrera CM (2014) Population genetics methods applied to a species delimitation problem: endemic trumpet daffodils (Narcissus section pseudonarcissi) from the southern Iberian Peninsula. Int J Plant Sci 175:501–517
Menges ES (2008) Restoration demography and genetics of plants: when is a translocation successful? Aust J Bot 56:187–196
Ministero delle Politiche Agricole Alimentari e Forestali (MiPAAF) (2000) Metodi di analisi chimica del suolo. Franco Angeli, Milano
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nei M (1977) F-statistics and analysis of gene diversity in subdivided populations. Ann Hum Genet Lond 41:225–233
Orsenigo S, Gentili R, Smolders AJP, Efremov A, Rossi G, Ardenghi NMG, Citterio S, Abeli T (2016) Reintroduction of a dioecious aquatic macrophyte (Stratiotes aloides L.) regionally extinct in the wild. Interesting answers from genetics. Aquatic Conserv 27:10–23
Ortego J, Aguirre MP, Noguerales V, Cordero PJ (2015) Consequences of extensive habitat fragmentation in landscape-level patterns of genetic diversity and structure in the Mediterranean esparto grasshopper. Evol Appl 8:621–632
Papadopoulou A, Knowles LL (2016) Toward a paradigm shift in comparative phylogeography driven by trait-based hypotheses. Proc Natl Acad Sci USA 113:8018–8024
Paracchini ML, Bulgheroni C, Borreani G, Tabacco E, Banterle A, Bertoni D, Rossi G, Parolo G, Origgi R, De Paola C (2015) A diagnostic system to assess sustainability at a farm level, the SOSTARE model. Agric Syst 133:35–53
Parolo G, Abeli T, Rossi G, Dowgiallo G, Matthies D (2011) Biological flora of Central Europe: Leucojum aestivum L. Perspect Plant Ecol 13:319–330
Peakall R, Smouse PE (2006) GenAlEx 6: genetic analysis in Excel. Population genetic for teaching and research. Mol Ecol Notes 6:288–295
Petit S, Griffiths L, Smart SS, Smith GM, Stuart RC, Wright SW (2004) Effects of area and isolation of woodland patches on herbaceous plant species richness across Great Britain. Landsc Ecol 19:463–471
Pollux BJA, Jong MDE, Steegh A, Verbruggen E, van Groenendael JM, Ouborg NJ (2007) Reproductive strategy, clonal structure and genetic diversity in populations of the aquatic macrophyte Sparganium emersum in river systems. Mol Ecol 16:313–325
Powolny M, Poschlod P, Reisch C (2016) Genetic variation in Silene acaulis increases with population age. Botany 94:241–247
Pritchard JK, Stephens M, Donnelly O (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Reed DH, Frankham R (2003) Correlation between fitness and genetic diversity. Conserv Biol 17:230–237
Reisch C, Bernhardt-Römermann M (2014) The impact of study design and life history traits on genetic variation of plants determined with AFLPs. Plant Ecol 115:1493–1511
Reisch C, Schmidkonz S, Meier K, Schöpplein Q, Meyer C, Hums C, Putz C, Schmid C (2017) Genetic diversity of calcareous grassland plant species depends on historical landscape configuration. BMC Ecol 17:19. https://doi.org/10.1186/s12898-017-0129-9
Ruffo S (ed) (2002) Woodlands of the Po plain. Italian Ministry of the Environment and Territory Protection, Friuli Museum of Natural History, Graphic lines, Udine
Sanaa A, Zouaghi O, Boussaid M, Ben Fadhel N (2010) Genetic diversity and population structure of tunisian Pancratium maritimum L. (Amaryllidaceae). Acta Hortic 853:61–68. https://doi.org/10.17660/ActaHortic.2010.853.6
Shao J-W, Wang J, Xu Y-N, Pan Q, Shi Y, Kelso S, Lv G-S (2015) Genetic diversity and gene flow within and between two different habitats of Primula merrilliana (Primulaceae), an endangered distylous forest herb in eastern China. Bot J Linn Soc 179:172–189
Tockner C, Stanford JA (2002) Riverine flood plains: present state and future trends. Environ Conserv 29:308–330
Vekemans X (2002) AFLP-SURV version 1.0. Laboratoire de Génétique et Ecologie, Végétale. Université Libre de Bruxelles, Belgium
Zaya DN, Molano-Flores B, Feist MA, Koontz JA, Coons J (2017) Assessing genetic diversity for the USA endemic carnivorous plant Pinguicula ionantha R.K. Godfrey (Lentibulariaceae). Conserv Genet 18:171–180
Acknowledgements
This study was funded by University of Milano-Bicocca, University of Pavia, and by the project “CORINAT”, D.G. Agricoltura, Regione Lombardia.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gentili, R., Abeli, T., Parolo, G. et al. Genetic structure of Leucojum aestivum L. in the Po Valley (N-Italy) drives conservation management actions. Conserv Genet 19, 827–838 (2018). https://doi.org/10.1007/s10592-018-1057-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-018-1057-8