Abstract
Once widely abundant, most subpopulations of the endangered Atlantic sturgeon are now estimated to be only 1–10% of their historical levels. The Edisto River has been sampled for a long period and extensively for juvenile Atlantic sturgeon from separate spring- and fall-spawned cohorts. Our objectives are to characterize the genetic diversity, stability, adaptive potential, and potential genetic structure of Atlantic sturgeon in the Edisto River and to identify any past bottlenecks experienced by this species, as well as to conduct forward simulation modeling of the population under multiple population trajectories. Our results indicate that fall- and spring-spawned Atlantic sturgeon in the Edisto River are genetically distinct (overall \({{F}_{ST}}\) = 0.092) with little gene flow or admixture between groups, both of which are diverse from a neutral genetic marker standpoint. Genetic diversity of both groups is on the higher end of published population diversity values. A lack of inbreeding and recent bottlenecks also bode well for these two groups of sturgeon, although future projections indicate a loss of allelic richness and genetic diversity even with population stability. Our effective population size estimates are moderate compared to published estimates for other Atlantic sturgeon populations. The most significant finding of our research is the genetic distinctness of the fall- and spring-spawned Atlantic sturgeon in the Edisto River, which may have several important ramifications for management of the species, including re-evaluating the demarcation of distinct population segments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
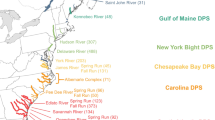
Atlantic sturgeon (Acipenser oxyrinchus oxyrinchus) are representatives of an ancient actinopterygiian lineage whose life history makes the species susceptible to anthropogenic impacts. Once widely abundant, most subpopulations are now estimated to be only 1–10% of their historical levels (Atlantic Sturgeon Status Review team, ASSRT 2007). Atlantic sturgeon have faced many threats to recovery, including overharvest for meat and caviar, dams that block access to historic spawning grounds, habitat degradation, and bycatch mortality. Given the precipitous decline in abundance range-wide, the species was originally petitioned for listing under the endangered species act (ESA) in 1998. In 2009, the National Marine Fisheries Service (NMFS) received a petition to list Atlantic sturgeon as endangered under the ESA and to be delineated by five distinct population segments (DPSs) based on the ASSRT (2007). In 2012, NMFS made the final determination to list one of the DPSs as threatened and the other four as endangered, including the two southernmost (Carolina and South Atlantic), under the ESA (NMFS 2012a, b). The South Atlantic DPS comprises seven river systems: ACE Basin (Ashepoo, Combahee, and Edisto Rivers), Savannah River, Ogeechee River, Altamaha River, Satilla River, St. Mary’s River, and St. Johns River (ASSRT 2007; NMFS 2012b). These DPS delineations are largely based on genetic data (Wirgin et al. 2000, 2002; King et al. 2001; Waldman et al. 2002; Grunwald et al. 2008) and were supported by latitudinal differences in life history strategies and movement patterns (ASSRT 2007).
The complex life history and long-distance movements of Atlantic sturgeon make understanding the evolutionary relatedness and patterns of gene flow among and within river systems and DPSs challenging. For instance, in southern river systems, sub-adult Atlantic sturgeon (~750 to 1500 mm total length (TL), 2+ years old) begin to migrate to coastal waters and undertake life-time long-distance movements (Murawski and Pacheco 1977; Smith 1985; Bain 1997). Prior to this stage, Atlantic sturgeon are considered river-resident juveniles that have not yet left their natal river. As adults, Atlantic sturgeon may move coastally throughout their range before migrating into river systems, mostly homing to natal estuaries based on low gene flow patterns (Grunwald et al. 2008), to spawn. However, the frequency of individual spawning migrations is not clearly understood, but does not appear to be annual, especially for females (ASSRT 2007). Additionally, in some single river systems spawning migrations have been documented in both the spring and fall (Collins et al. 2000; McCord et al. 2007; Balazik et al. 2012; Balazik and Musick 2015; Smith et al. 2015). Adult Atlantic sturgeon are known to make large-scale coastal movements (ASSRT 2007; Dunton et al. 2012; Wirgin et al. 2012), potentially as far as from the Hudson River, NY to the Altamaha River, GA within a period of a year (Erickson et al. 2011). Therefore, the capture of adult Atlantic sturgeon within a given estuary does not guarantee that the individual is in or even near its natal system or in spawning condition. Examining the natal river-resident juveniles of a river system is the best approach to genetically characterizing a given population.
The Edisto River is one of the longest undammed blackwater rivers in the United States and drains approximately 8000 km2 (Beasley et al. 1996). It contains about 60 km of tidally influenced river, with the saltwater-freshwater interface occurring roughly 32 km upstream of the mouth (Blake et al. 2001). Additionally, the Edisto River has been sampled since 1994 with many juveniles being captured over that time frame (McCord et al. 2007). Previous research identified ripe adults during both the fall and spring in the Edisto River (Collins et al. 2000). Based on a bimodal size distribution, McCord et al. (2007) were able to identify separate spring- and fall-spawned cohorts. Despite the long sampling history, the Edisto River population has not been previously characterized genetically, which is one needed area of research (ASSRT 2007).
Many previous genetic studies on Atlantic sturgeon utilized analyses of mitochondrial DNA (mtDNA) to assess populations (Waldman et al. 1996, 2002; Wirgin et al. 2000, 2002; Grunwald et al. 2008; Dunton et al. 2012); however, mutation rates of mitochondrial genes are often too slow to detect more recent genetic changes in population structure due to spatial separation and restrictions to gene flow (Anders et al. 2011). Co-dominant, nuclear microsatellite loci have relatively high mutation rates and thus are more useful for genetic estimates of population structure due to more recent and smaller-scale genetic changes (Tranah et al. 2004). Microsatellites have been developed and used to genotype some Atlantic sturgeon populations (May et al. 1997; King et al. 2001; Henderson-Arzapalo and King 2002; Wirgin et al. 2012). However, microsatellite data is lacking from many river systems within the two southernmost DPSs, creating a genetic data gap between the Savannah River and Albemarle sound. The incorporation of genetic data from the missing river systems may improve the definition of these DPSs, thereby providing an even more powerful tool for understanding the complex among-river and among-DPS interactions of Atlantic sturgeon.
The use of microsatellites also affords the ability to estimate genetic metrics such as allelic diversity, heterozygosity (\({{H}_{O}}\)), inbreeding (\({{F}_{IS}}\)), gene flow (\({{F}_{ST}}\)), and effective population size (\({{N}_{e}}\)). \({{N}_{e}}\) is a key parameter in conservation genetics because it determines the rate of change in the composition of a population caused by genetic drift and inbreeding and is important in determining the level of variability in a population (Charlesworth 2009). Low effective population sizes have been shown to negatively impact population stability and resilience to environmental change (Newman and Pilson 1997; Saccheri et al. 1998; Nielsen and Hansen 2008). By tracking temporal changes in \({{N}_{e}}\) and other measures of genetic diversity (allelic diversity and heterozygosity), it is possible to generate population data that can independently corroborate (or refute) trends seen in independent population and stock assessments. Furthermore, measures of genetic diversity reveal vital characteristics of the population that cannot be revealed through traditional surveys, yet are important for conservation and management, especially when a stock has been depleted. Therefore, these measurements are important baseline genetic information to monitor and assess recovery of a population over time and compare genetic diversity among populations. Estimates of these parameters for Atlantic sturgeon populations are lacking for many of the river systems in the Carolina and South Atlantic DPSs.
Our objectives are to characterize the genetic diversity, stability, and to test for potential genetic structure of Atlantic sturgeon in the Edisto River. In addition, the genetic data was analyzed to identify and understand the influences of past bottlenecks experienced by this species, as well as to conduct forward simulation modeling of the population under multiple population trajectories. The combination of long lifespan and overlapping generations, as seen in Atlantic sturgeon, has been shown to provide a masking effect in terms of genetic diversity (Kuo and Janzen 2003; Darden and Tarpey 2014). Understanding the degree of masking afforded by their life history as well the influence of potential future population trends on their capability of maintaining future genetic diversity is important information for effective management. Therefore, knowledge of the current status and future trajectory of the genetic diversity of a subpopulation is a significant variable in managing the viability and persistence of Atlantic sturgeon (Anders et al. 2011).
Materials and methods
Sample processing
Our sample set conservatively was limited to only river-resident juvenile Atlantic sturgeon (age 0 or 1, <530 mm TL), as recommended by the ASSRT (2007), to avoid any potential confounding influences from the high degree of coastal movements documented in both subadult and adult Atlantic sturgeon. The restriction of samples to the juvenile life stage allowed for an unambiguous genetic characterization of Atlantic sturgeon from these river systems as these fish are still residents of their natal river. Atlantic sturgeon have been collected in the Edisto River since 1994 as part of a long-term monitoring project. Sampling efforts targeted juvenile sturgeon by deploying drifted gillnets (92 m long, 7.5 m deep, with 12.5 and 14.0 cm stretched-mesh monofilament) between 4 and 6 days per month from January–May 1994–1995, January–December 1996–2000, March–December 2001–2002, May–December 2003–2005, March–September 2006, and May–October 2007–2015. The sampling area was consistently near Jehossee Island (rkm 28.2) each year with the gear deployed in areas that were free of debris (Fig. 1). The genetic tissue archival database was queried for river-resident juvenile Atlantic sturgeon captured in the Edisto River, SC; a total of 1387 samples from the 1996 to 2011 year classes (YCs) were identified. Captured fish were assigned a YC and a spawning season (spring or fall) based on TL at day of year (Fig. 2) following published length data of juvenile Atlantic sturgeon in the South Atlantic DPS (Peterson et al. 2000; Schueller and Peterson 2010) and assuming similar growth rates among all fish within the Edisto River. The few outliers which did not fall within one of the size distribution modes were designated as unknown and removed from all analyses.
Sturgeon DNA samples were stored in three different preservatives: sarcosyl urea, SDS-urea, and ethanol (EtOH). Isolation techniques for samples stored in urea based preservatives were identical, but a different method was used for samples stored in EtOH. Urea based preservatives digest most of the tissue during storage; to complete the tissue digestion and cell lysis, 100 µl of liquid from each sample storage tube was added to a 1.5 ml microcentrifuge tube containing 50 µl of lysis buffer (Wizard SV Genomic DNA Purification System; Promega, Madison, WI) and 20 µl of Proteinase K (20 mg/ml) and incubated at 55 °C for at least 2.5 h. The total 170 µl of lysate was transferred to a 96-well round-bottom plate, and 13 µl of Sera-Mag SpeedBeads (GE Healthcare Life Sciences, Pittsburgh, PA) was added. The lysate and SpeedBeads were then mixed with cold, 100% isopropanol by pipetting. The 96-well plate was placed on a magnet for at least 10 min and the supernatant was then discarded. A series of five EtOH washes were performed by removing the 96-well plate from the magnet, mixing the SpeedBeads with cold, 95% EtOH by pipetting and returning the plate to the magnet for ~1 min before discarding the supernatant. After the EtOH washes, the SpeedBeads and DNA were allowed to dry at room temperature until all EtOH evaporated (~1 h). DNA was then eluted from the beads by mixing with 40 µl of autoclaved Milli-Q water (EMD Millipore, Billerica, MA) and stored at −20 °C until amplification.
Genomic DNA was extracted and isolated from tissue stored in EtOH using the standard Purification of Genomic DNA from Lysates Using a Microcentrifuge protocol for the Wizard SV Genomic DNA Purification System (Promega). Approximately 5 mm2 of tissue, when possible, was used with the digestion solution. Isolated DNA was stored at −20 °C until amplification.
Genetic data were generated for the following twelve microsatellite loci: LS19, LS39, LS54, LS68 (May et al. 1997), Aox23, Aox45, Aox12 (King et al. 2001), AoxD44, AoxD165, AoxD170, AoxD188, and AoxD241 (Henderson-Arzapalo and King 2002). These loci were previously used by Wirgin et al. (2012) because they could be reliably scored, were in Hardy–Weinberg and linkage equilibrium in previous studies and were effective in population/DPS assignment (King et al. 2001; Henderson et al. 2005).
Optimized multiplex reactions contained 1× Hotmaster PCR buffer (5 Primer Inc., Gaithersburg, MD), 0.25 mM dNTPs, 2.0 mM MgCl2, 0.3 µM forward and reverse primers, 0.06 U Hotmaster Taq DNA polymerase (5 Primer Inc.), and 1 µl sample isolated DNA. Specific ratios and fluorescent Well-RED dye labels (Sigma–Aldrich, St. Louis, MO) varied for each primer and multiplex group. Amplifications occurred in 11 µl reaction volumes using iCycler thermocyclers (Bio-Rad Laboratories, Hercules, CA) under these conditions: initial 2 min denaturing at 94 °C; 35 cycles of 94 °C denaturing for 30 s, 58 °C annealing for 30 s, and 64 °C extension for 1 min; and a final 64 °C extension for 1 h.
PCR products were separated and visualized by capillary gel electrophoresis on a Beckman CEQ 8000 Genetic Analysis System (Beckman Coulter, Brea, CA) with a 400BP size standard kit. Chromatograms were scored with CEQ Fragment Analysis software using the frag3/PA version 1 analysis algorithm (Beckman Coulter) to determine allele size. Two independent readers scored each chromatogram for quality assurance and quality control, and these scores were compared for agreement.
As the microsatellite data currently incorporated into the baseline Atlantic sturgeon DPS genetic data set were generated at the United States Geological Survey Leetown Science Center, a validation test was conducted to avoid intra-facility variation in the data. A total of 96 Atlantic sturgeon samples were independently processed and genotyped at both the Leetown Science Center and the South Carolina Department of Natural Resources (SCDNR). Scores for all samples were compared and differences were reconciled in conference to develop a microsatellite scoring conversion table. By scoring all samples in the same manner, despite different facilities and equipment, the microsatellite data generated here can now be added to the coastwide baseline Atlantic sturgeon data set to further refine assignments to DPSs.
Data analysis
Preliminary locus verification was conducted by testing for adherence to Hardy–Weinberg equilibrium (HWE), degree of linkage disequilibrium (LD), and the presence of genotyping artifacts. Examinations for deviations from HWE and LD between loci pairs were performed using the program GenePop 4.3 (Raymond and Rousset 1995) with default parameters (10,000 dememorizations, 100 batches, and 5000 iterations per batch). The frequency of any null alleles segregating at each locus was evaluated in CERVUS 3.0.7 (Kalinowski et al. 2007). Significance levels for all simultaneous analyses were adjusted using a Bonferroni correction (Holm 1979; Rice 1989). MicroChecker 2.2.3 (Van Oosterhout et al. 2004) was used to check for large-allele dropout.
COLONY 2.0.6.2 (Jones and Wang 2010) was used to identify and remove full siblings from the data prior to analysis with Structure because having many closely related individuals can result in an overestimation of the number of clusters (Anderson and Dunham 2008; Rodríguez-Ramilo and Wang 2012). Fall- and spring-spawned fish were analyzed separately with three short-length runs using the full likelihood and pairwise-likelihood combined method (FPLS) with a polygamous breeding system, medium precision, weak priors, updating allele frequencies, and no genotyping errors assumed. Each short run was given a different random seed. Conservatively, we removed a full sibling from a pair or family only if it was identified in all three runs with a probability of ≥0.9 and if the samples were within the same YC.
The Bayesian clustering program Structure 2.3.4 (Pritchard et al. 2000) was used to observe individual genetic clustering over a range of populations (K). Run parameters were set at 10,000 burn-in repetitions followed by 10,000 Markov chain Monte-Carlo repetitions, with and without location information included as a prior, with all samples separated by season and YC, and with K varied 1–18 and three independent runs per K. Each season also was run under the same conditions independently by YC to determine if there was any underlying structure present. Structure Harvester (Earl and vonHoldt 2012) was used to assess the results and determine the number of groups that best fit the data (Evanno et al. 2005).
Genetic population structure within the Edisto River (spring-spawned versus fall-spawned fish by YC and spring versus fall overall) was evaluated using pairwise \({{F}_{ST}}\), a measure of genetic distance, as calculated in ARLEQUIN 3.5.2.1 (Excoffier et al. 2005) with 100 permutations to assess significance. \({{{F}'}_{ST}}\), a measure of genetic distance that is adjusted to the maximum value of \({{F}_{ST}}\) for a given suite of markers (Meirmans 2006; Meirmans and Hedrick 2011), was computed for the same pairwise comparisons (excluding any YC with a single sample) using GenAlEx 6.502 (Peakall and Smouse 2006, 2012) with 999 standard permutations and individual analysis suppressed. The \({{F}_{ST}}\) and \({{{F}'}_{ST}}\) analyses were complemented by exact G tests to compare allele frequency distributions between every YC from each fall and spring season. The exact G tests were performed with a Markov chain randomization method (10,000 dememorizations, 20 batches, and 5000 iterations per batch) using GenePop.
Molecular diversity indices for the Edisto River were calculated using ARLEQUIN, including number of alleles per locus (\({{N}_{a}}\)), allelic size range, and heterozygosity (\({{H}_{E}}\)) (Nei 1987). Allelic richness (A, the number of alleles corrected for sample size) was calculated using FSTAT 2.9.3 (Goudet 1995); inbreeding coefficients (\({{F}_{IS}}\)) (Weir and Cockerham 1984) were calculated using GenePop. Effective population size (\({{N}_{e}}\)) was calculated using both contemporary methods in LDNe 1.31 (Waples 2006) and long-term heterozygosity-based methods. Contemporary (parental generation) estimates of \({{N}_{eb}}\) (effective number of breeding adults) were estimated using the single-sample program LDNe (Waples 2006). Genetic drift generates non-random associations among unlinked loci; LDNe analyzes this linkage disequilibrium between a set of loci to estimate contemporary \({{N}_{eb}}\) for a single time point. Minimal allele frequencies for inclusion were set at default values (0.01, 0.02, and 0.05) assuming a random mating model. The \({{N}_{eb}}\) estimates were corrected for overlapping generations according to Waples et al. (2014), by using the two trait formula with a 30 year life span and 9 year age at maturity (Smith 1985). COLONY was also used to calculate \({{N}_{e}}\) for a single time point based on sibling relationships among individuals. Long-term estimates were calculated using heterozygosity-based methods (Ohta and Kimura 1973) with both a step-wise mutation model (SMM) and an infinite alleles model (IAM). The SMM predicts that at mutation-drift equilibrium, \({{N}_{e}}\) is represented by [(1/1−HE)2−1]/8 µ. Although the SMM is likely the most appropriate model for microsatellite data, it is sensitive to mutational modeling; therefore, \({{N}_{e}}\) was also estimated under the IAM using the equation \({{N}_{e}}\) = H/4 µ(1−H). The most commonly used microsatellite mutation rate in fishes, µ = 5 × 10−4 (Estoup and Angers 1998), was used for both model estimations. The comparison of \({{N}_{eb}}\) estimates for the Edisto River population across YCs was used to determine the degree of stability in the contribution to the reproductive pool over nearly a decade (eight YCs).
BOTTLENECK 1.2.02 (Cornuet and Luikart 1996; Piry et al. 1999) was used to test for recent (2−4\({{N}_{e}}\) generations) reductions in Atlantic sturgeon population size through the evaluation of a population’s heterozygosity excess based on that expected at mutation-drift equilibrium. Although a two-phase mutational model (TPM) with 95% single step mutations and 12% variance among mutational steps is typically recommended for dinucleotide microsatellite repeat loci (Piry et al. 1999), we evaluated all loci under both the TPM and SMM. All three tests of population bottlenecks were performed; however, interpretation was primarily based on the Wilcoxon signed rank test as the sign test suffers from low statistical power and the standardized differences test requires at least 20 polymorphic loci for robust results (Cornuet and Luikart 1996).
The M-ratio (Garza and Williamson 2001) was also calculated to identify long term population reductions, indicated by M < critical M (\({{M}_{C}}\)). M and \({{M}_{C}}\) were calculated using the M_P_Val and Critical_M software packages (https://swfsc.noaa.gov/textblock.aspx?Division=FED&id=3298). A range of \({{M}_{C}}\) values were calculated through 10,000 distribution simulations; input parameters of the average size of multistep mutations (\({{\Delta }_{g}}\): 2.8 and 3.5) and the proportion of one-step mutations (p s : 95, 90, 85, and 80%) were varied.
Many conservation goals aim to maintain genetic diversity of a population to help prevent extinction (Frankham 2005). We used BOTTLESIM v2.6 (Kuo and Janzen 2003) to simulate the evolution of genetic diversity over time for Atlantic sturgeon, taking into account their long lifespan and overlapping generations, to estimate the sustainable population size needed to meet this conservation goal. All simulations were conducted for a 200 year period incorporating a stable population size with dioecious reproduction, random mating, 1:1 sex ratio, 30 year lifespan and an age at maturity of nine years (in the southern part of their range, male sturgeon mature by 8 years of age and females by 10 years of age; Smith 1985), with 1000 iterations. Calculated \({{N}_{e}}\) estimates were used as the basis for population sizes for the fall- and spring-spawned sturgeon. As the contemporary estimates are realistically somewhere in between the effective number of breeders and the true effective population size, we used population sizes ranging from the worst case scenario (\({{N}_{e}}\) is the true population size) to 5 × \({{N}_{e}}\).
Results
Four Atlantic sturgeon were identified as being recaptured based on identical genotypes (Table 1); one from each pair was removed for subsequent analyses. Locus LS39 was dropped from further analysis due to substantial problems with its amplification. Deviations from HWE and LD were observed in the dataset, partially due to family structure being present in the data. Data were reanalyzed after removing full siblings (see COLONY results). With full siblings removed and after Bonferroni correction, two loci deviated from HWE (Aox23 and AoxD241) in the fall sample set and three loci deviated from HWE (AoxD188, Aox45, and LS19) in the spring sample set (Supplementary Table 1). Given the lack of consistency in deviations from HWE at particular loci between sample sets, these loci were retained. However, LD was still observed after removing full siblings. Given the lack of evidence of linkage in other studies (King et al. 2001; Henderson-Arzapalo and King 2002) and the Albemarle Sound and Savannah River populations from our research (unpublished data), true physical linkage of loci is unlikely, and the observed LD possibly is a result of our conservative approach to full sibling removal and latent relatedness. There was no evidence of null alleles (frequency < 0.05) at any loci (except LS39, which was already dropped) or large-allele dropout.
Each COLONY run produced minor variations in relatedness, but consistently identified families of three or more individuals. For the fall dataset, COLONY identified 61 full siblings that were removed: 21 of 1996 YC, 27 of 1997 YC, seven of 1998 YC, and six of 1999 YC. The largest family in the fall samples comprised 14 members. For the spring dataset, COLONY identified 45 full siblings that were removed: two of 1998 YC and 43 of 2003 YC. The largest family in the spring samples also comprised 14 members. The resulting data contained 1117 samples of fall-spawned sturgeon and 164 samples of spring-spawned sturgeon and was used to infer the number of genetic clusters present with Structure analyses.
The results from Structure and Structure Harvester (ΔK = 1020.33) indicated that there were two population clusters in the Edisto River, with distinct signatures assigned to sturgeon from fall spawning events and spring spawning events (Fig. 3). When fall- and spring-spawned sturgeon were run independently, both resulted in K = 1, indicating that there was no further population or family structure present. The same results were obtained with and without location used as a prior.
Pairwise \({{F}_{ST}}\) estimates between each YC and season (excluding YCs with <10 samples) varied from 0.0 to 0.109 (Table 2). When all YCs were combined and separated by season spawned, the comparison between fall- and spring-spawned sturgeon resulted in an \({{F}_{ST}}\) estimate of 0.092, which was significantly different than 0 (p < 0.05). All \({{{F}'}_{ST}}\) results were consistent with \({{F}_{ST}}\) results on a YC pairwise basis and the overall comparison between fall- and spring-spawned sturgeon resulted in \({{{F}'}_{ST}}\) = 0.316. The exact G tests resulted in significant differences in allele frequencies at each locus and overall from the fall- and spring-spawned sturgeon, corroborating the \({{F}_{ST}}\) and \({{{F}'}_{ST}}\) results.
The number of alleles per locus (\({{N}_{a}}\)) was higher for fall-spawned sturgeon than for spring-spawned sturgeon (11.2 vs. 9.4, respectively; Table 3). Allelic richness (8.3 fall; 8.5 spring) and size range (6–36 fall; 6–32 spring) were similar for both spawning periods. Heterozygosity was moderate overall for both fall (\({{H}_{O}}\) = 0.70, \({{H}_{E}}\) = 0.69) and spring (\({{H}_{O}}\) = 0.77, \({{H}_{E}}\) = 0.74), and no inbreeding was detected for either group (\({{F}_{IS}}\) = −0.018 fall; \({{F}_{IS}}\) = −0.044 spring).
Overall, LDNe estimated fall sturgeon \({{N}_{e}}\) = 48.0 (44.7–51.5; 95% CI) and spring sturgeon \({{N}_{e}}\) = 13.3 (12.1–14.6) (Table 3). The three COLONY estimates of \({{N}_{e}}\) were an order of magnitude higher than LDNe estimates for the fall sturgeon (~328), but were similar for spring sturgeon (~38). Long-term estimates of \({{N}_{e}}\) were 2301.8 and 3419.9 for fall and spring, respectively, under the SMM and 1097.4 and 1415.7 for fall and spring, respectively, under the IAM. Estimates of \({{N}_{eb}}\) by YC were roughly similar, mostly near \({{N}_{eb}}\) = 30, with some individual YCs burdened by large confidence intervals or negative estimates due to small sample sizes (Table 4). The highest and lowest YC \({{N}_{eb}}\) estimates were 60.8 for fall 2004 and 9.5 for spring 2003. No signs of a recent bottleneck were detected under either the TPM or SMM for fall- or spring-spawned sturgeon with the heterozygosity excess method (all Wilcoxon signed rank test p > 0.8). The M-ratio test indicated evidence of a bottleneck as fall (M = 0.731) and spring (M = 0.708) groups of sturgeon were both below the entire range of calculated \({{M}_{C}}\) values (0.773–0.915).
Similar patterns were seen between fall and spring populations in the simulations (Fig. 4). Allelic richness decreased faster over time than observed heterozygosity for both spawning groups. The worst case scenario (N eb is the true population size) for fall resulted in a ~40% reduction in allelic richness and a <20% reduction in observed heterozygosity after 200 years. The worst case scenario for spring resulted in a ~60% reduction in allelic richness and a ~40% reduction in observed heterozygosity after 200 years. However, the best case scenario (5 × \({{N}_{eb}}\) is the true population size) for fall resulted in a ~20% reduction in allelic richness and a <5% reduction in observed heterozygosity after 200 years. The best case scenario for spring resulted in a ~35% reduction in allelic richness and a < 10% reduction in observed heterozygosity after 200 years. These results assume that the Edisto River is a closed population. Some of the losses of genetic diversity could be substantially countered by mutation and immigration from other populations, which is not estimated in these simulations; therefore, these results should be considered very conservative.
Simulated loss of genetic diversity in fall (top, a and b panels) and spring (bottom, c and d panels) populations of Atlantic sturgeon in the Edisto River over a 200-year time period with various population sizes kept constant over time. Evolution of allelic richness (OA) is shown on the left for each river (a and c panels) and observed heterozygosity (\({{H}_{O}}\)) is shown on the right (b and d panels; note the contracted y-axis scale)
Discussion
The Edisto River contained two genetically distinct groups of Atlantic sturgeon, fall- and spring-spawned fish, both of which are diverse from a neutral genetic marker standpoint. The lack of inbreeding also bodes well for these two groups of sturgeon, though there was evidence of recent and historic bottlenecks in both groups and future projections indicate a loss of allelic richness and genetic diversity even with population stability. Our \({{N}_{e}}\) estimates (48.0 for fall and 13.3 for spring) seem moderate compared to published estimates for populations such as the James River (\({{N}_{e}}\) = 62), Delaware River (\({{N}_{e}}\) = 109), and the Hudson River (\({{N}_{e}}\) = 198) (O’Leary et al. 2014); however, given that our samples comprised several cohorts but not a complete generation these estimates are likely biased downward. Likewise, our YC N eb estimates are typically around 30, which seem realistic compared to cohort estimates from the Altamaha River population (73–138; Moyer et al. 2012). However, these previous studies did not separate samples by spawning seasons, which could have upwardly biased their \({{N}_{e}}\) and \({{N}_{eb}}\) estimates if subpopulation structure is present in those river systems.
The most significant finding of our research is the genetic distinctness of the fall- and spring-spawned Atlantic sturgeon. Some individuals appear to be outliers within the fall or spring groupings (Fig. 3), and unfortunately the underlying explanation for this is unknown, but individuals may have been categorized to the wrong group initially due to clerical errors, abnormal growth in some individuals, or infrequent straying between spawning seasons. The high values of \({{F}_{ST}}\) and \({{{F}'}_{ST}}\) and the lack of ancestral mixture from Structure indicate there is little gene flow or admixture between the two spawning groups of sturgeon in the Edisto River. Although this is a new finding for Atlantic sturgeon, it is not unknown among fishes. For example, chinook salmon from different seasonal spawning runs in central California have a mean \({{F}_{ST}}\) of 0.082 (Banks et al. 2000). Population structure within a river system has been suggested based on differential spawning behaviors of Atlantic sturgeon (Balazik et al. 2012; Balazik and Musick 2015), but has not been documented genetically. The findings here raise several important questions as to the evolutionary history of the species. For example, would this pattern of seasonal spawning differentiation also be observed in other river systems? If so, does this differentiation between seasons arise within river systems or across river systems? The latter possibility would indicate a deeper evolutionary divergence between fall- and spring- spawning Atlantic sturgeon, which can be assessed by analyzing data from other populations.
By most metrics, the spring-spawned sturgeon appear to be more diverse and have higher long-term \({{N}_{e}}\) estimates than the fall-spawned sturgeon in the Edisto River, though the sample size was smaller and contemporary \({{N}_{e}}\) estimates were lower. These seemingly conflicting results may be attributable to a higher historical population of spring sturgeon that were disproportionally targeted by fisheries for their meat and roe (Smith et al. 1984). Balazik and Musick (2015) observed that the spring spawning group appears less abundant than the fall-spawning group in the James River, Virginia, although this may be due to a sampling bias.
Previous studies that identified bimodal size distributions as different cohorts should be re-evaluated to determine if they are instead representative of separate fall- and spring-spawned cohorts. We documented better modal separation by plotting TL by day of year, which allowed us to categorize individual sturgeon to either a fall or spring spawning event. These categorizations and growth rates were supported by the genetic identification of four recaptures whose growth between captures mirrored that of the entire groups of sturgeon. The two distinct spawning events also raise a question as to whether previous age estimates based on pectoral fin spines are accurate and reliable if they did not account for multiple spawning events in a single year.
Our results also have several important ramifications for management of Atlantic sturgeon. Currently, the species is managed in five DPSs across the entire range. Management on a sub-DPS level should be considered if population structure as strong as that observed in the Edisto River also occurs in other parts of the species’ range since this was not accounted for in the previous analyses. Fall- and spring-spawning Atlantic sturgeon may warrant separate management as well since various commercial fisheries may differentially encounter the fish during fall and spring spawning migrations. The population genetics analysis of this rich dataset provides a robust evaluation of the current status of Atlantic sturgeon in the Edisto River and fulfills one area identified as needed research by the ASSRT (2007). These metrics can be used as benchmarks from which to measure recovery of the species.
References
Anders PJ, Drauch-Schreier A, Rodzen J, Powell MS, Narum S, Crossman JA (2011) A review of genetic evaluation tools for conservation and management of North American sturgeons: roles, benefits, and limitations. J Appl Ichthyol 27:3–11
Anderson EC, Dunham KK (2008) The influence of family groups on inferences made with the program Structure. Mol Ecol Resour 8:1219–1229
ASSRT (2007) Status review of Atlantic sturgeon (Acipenser oxyrinchus oxyrinchus). Report to National Marine Fisheries Service, Northeast Regional Office Februrary, 23, 174.
Bain MB (1997) Atlantic and shortnose sturgeons of the Hudson River: common and divergent life history attributes. Sturgeon biodiversity and conservation. Springer, The Netherlands. pp. 347–358
Balazik MT, Musick JA (2015) Dual annual spawning races in Atlantic sturgeon. PloS ONE 10:e0128234. doi:10.0128210.0121371/journal.pone.0128234
Balazik MT, Garman GC, Van Eenennaam JP, Mohler J, Woods LC (2012) Empirical evidence of fall spawning by Atlantic sturgeon in the James River, Virginia. Trans Am Fish Soc 141:1465–1471
Banks MA, Rashbrook VK, Calavetta MJ, Dean CA, Hedgecock D (2000) Analysis of microsatellite DNA resolves genetic structure and diversity of chinook salmoon (Oncorhynchus tshawytscha) in California’s central valley. Can J Fish Aquat Sci 57:915–927
Beasley BR, Marshall WD, Miglarese AH, Scurry JD, Vanden Houten C (1996) Managing resources for a sustainable future: the Edisto River Basin Project report. South Carolina Department of Natural Resources, Columbia, SC
Blake A, Kineke G, Milligan T, Alexander C (2001) Sediment trapping and transport in the ACE Basin, South Carolina. Estuaries 24:721–733
Charlesworth B (2009) Effective population size and patterns of molecular evolution and variation. Nat Rev Genet 10:195–205
Collins MR, Smith TI, Post WC, Pashuk O (2000) Habitat utilization and biological characteristics of adult Atlantic sturgeon in two South Carolina rivers. Trans Am Fish Soc 129:982–988
Cornuet JM, Luikart G (1996) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144:2001–2014
Darden T, Tarpey C (2014) Genetic characterization of the Savannah and Pee Dee River populations of robust redhorse (Moxostoma robustum) with conservation implications. Copeia 2014:70–78
Dunton KJ, Chapman D, Jordaan A, Feldheim K, O’Leary SJ, McKown KA, Frisk MG (2012) Genetic mixed-stock analysis of Atlantic sturgeon Acipenser oxyrinchus oxyrinchus in a heavily exploited marine habitat indicates the need for routine genetic monitoring. J Fish Biol 80:207–217
Earl DA, vonHoldt BM (2012) Structure harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Res 4:359–361
Erickson DL, Kahnle A, Millard MJ, Mora EA, Bryja M, Higgs A, Mohler J, DuFour M, Kenney G, Sweka J, Pikitch EK (2011) Use of pop-up satellite archival tags to identify oceanic-migratory patterns for adult Atlantic sturgeon, Acipenser oxyrinchus oxyrinchus Mitchell, 1815. J Appl Ichthyol 27:356–365
Estoup A, Angers B (1998) Theoretical and empirical considerations. Adv. Mol Ecol 306:55
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47
Frankham R (2005) Genetics and extinction. Biol Conserv 126:131–140
Garza JC, Williamson EG (2001) Detection of reduction in population size using data from microsatellite loci. Mol Ecol 10:305–318
Goudet J (1995) FSTAT (version 1.2): a computer program to calculate F-statistics. J Hered 86:485–486
Grunwald C, Maceda L, Waldman J, Stabile J, Wirgin I (2008) Conservation of Atlantic sturgeon Acipenser oxyrinchus oxyrinchus: delineation of stock structure and distinct population segments. Conserv Genet 9:1111–1124
Henderson A, Spidle A, King T (2005) Genetic diversity, kinship analysis, and broodstock management of captive Atlantic sturgeon for population restoration. American Fisheries Society Symposium 2005:621–633
Henderson-Arzapalo A, King T (2002) Novel microsatellite markers for Atlantic sturgeon (Acipenser oxyrinchus) population delineation and broodstock management. Mol Ecol Notes 2:437–439
Holm S (1979) A simple sequentially rejective multiple test procedure. Scand J Stat 65–70
Jones OR, Wang J (2010) Colony: a program for parentage and sibship inference from multilocus genotype data. Mol Ecol Resour 10:551–555
Kalinowski ST, Taper ML, Marshall TC (2007) Revising how the computer program cervus accommodates genotyping error increases success in paternity assignment. Mol Ecol 16:1099–1106
King T, Lubinski B, Spidle A (2001) Microsatellite DNA variation in Atlantic sturgeon (Acipenser oxyrinchus oxyrinchus) and cross-species amplification in the Acipenseridae. Conserv Genet 2:103–119
Kuo CH, Janzen FJ (2003) Bottlesim: a bottleneck simulation program for long-lived species with overlapping generations. Mol Ecol Notes 3:669–673
May B, Krueger CC, Kincaid HL (1997) Genetic variation at microsatellite loci in sturgeon: primer sequence homology in Acipenser and Scaphirhynchus. Can J Fish Aquat Sci 54:1542–1547
McCord JW, Collins MR, Post WC, Smith TI (2007) Attempts to develop an index of abundance for age-1 Atlantic sturgeon in South Carolina, USA. In: American Fisheries Society Symposium, p. 397. American Fisheries Society.
Meirmans PG (2006) Using the AMOVA framework to estimate a standardized genetic differentiation measure. Evolution Int J org Evolution 60:2399–2402
Meirmans PG, Hedrick PW (2011) Assessing population structure: \({{F}_{ST}}\) and related measures. Mol Ecol Resour 11:5–18
Moyer GR, Sweka JA, Peterson DL (2012) Past and present processes influencing genetic diversity and effective population size in a natural population of Atlantic sturgeon. Trans Am Fish Soc 141:56–67
Murawski SA, Pacheco AL (1977) Biological and fisheries data on Atlantic sturgeon, Acipenser oxyrhynchus (Mitchill). Sandy Hook Laboratory, Northeast Fisheries Center, National Marine Fisheries Service, National Oceanic and Atmospheric Administration, US Department of Commerce.
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Newman D, Pilson D (1997) Increased probability of extinction due to decreased genetic effective population size: experimental populations of Clarkia pulchella. Evolution, 354–362
Nielsen EE, Hansen MM (2008) Waking the dead: the value of population genetic analyses of historical samples. Fish Fish 9:450–461
NMFS (2012a) Endangered and threatened wildlife and plants; threatened and endangered status for distinct population segments of Atlantic sturgeon in the Northheast Region. Fed Reg 77:5880–5912
NMFS (2012b) Endangered and threatened wildlife and plants; final listing determinations for two distinct population segments of Atlantic sturgeon (Acipenser oxyrinchus oxyrinchus) in the Southeast. Fed Reg 77:5914–5982
O’Leary SJ, Dunton KJ, King TL, Frisk MG, Chapman DD (2014) Genetic diversity and effective size of Atlantic sturgeon, Acipenser oxyrhinchus oxyrhinchus river spawning populations estimated from the microsatellite genotypes of marine-captured juveniles. Conserv Genet 15:1173–1181
Ohta T, Kimura M (1973) A model of mutation appropriate to estimate the number of electrophoretically detectable alleles in a finite population. Genet Res 22:201–204
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539
Peterson DL, Bain MB, Haley N (2000) Evidence of declining recruitment of Atlantic sturgeon in the Hudson River. N Am J Fish Manage 20:231–238
Piry S, Luikart G, Cornuet J-M (1999) BOTTLENECK: a program for detecting recent effective population size reductions from allele data frequencies. J Hered 90:502–503
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rice WR (1989) Analyzing tables of statistical tests. Evolution, 223–225
Rodríguez-Ramilo ST, Wang J (2012) The effect of close relatives on unsupervised Bayesian clustering algorithms in population genetic structure analysis. Mol Ecol Resour 12:873–884
Saccheri I, Kuussaari M, Kankare M, Vikman P, Fortelius W, Hanski I (1998) Inbreeding and extinction in a butterfly metapopulation. Nature 392:491–494
Schueller P, Peterson DL (2010) Abundance and recruitment of juvenile Atlantic sturgeon in the Altamaha River, Georgia. Trans Am Fish Soc 139:1526–1535
Smith TI (1985) The fishery, biology, and management of Atlantic sturgeon, Acipenser oxyrhynchus, in North America. Environ Biol Fish 14:61–72
Smith TI, DE Marchette, GF Ulrich (1984) The Atlantic sturgeon fishery in South Carolina. N Am J Fish Manage 4:164–176
Smith JA, Flowers HJ, Hightower JE (2015) Fall spawning of Atlantic sturgeon in the Roanoke River, North Carolina. Trans Am Fish Soc 144:48–54
Tranah G, Campton DE, May B (2004) Genetic evidence for hybridization of pallid and shovelnose sturgeon. J Hered 95:474–480
Van Oosterhout C, Hutchinson WF, Wills DP, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Waldman JR, Hart JT, Wirgin II (1996) Stock composition of the New York Bight Atlantic sturgeon fishery based on analysis of mitochondrial DNA. Trans Am Fish Soc 125:364–371
Waldman J, Grunwald C, Stabile J, Wirgin I (2002) Impacts of life history and biogeography on the genetic stock structure of Atlantic sturgeon Acipenser oxyrinchus oxyrinchus, Gulf sturgeon A. oxyrinchus desotoi, and shortnose sturgeon A. brevirostrum. J Appl Ichthyol 18:509–518
Waples RS (2006) A bias correction for estimates of effective population size based on linkage disequilibrium at unlinked gene loci. Conserv Genet 7:167–184
Waples RS, Antao T, Luikart G (2014) Effects of overlapping generations on linkage disequilibrium estimates of effective population size. Genetics 197:769–780
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution, 1358–1370
Wirgin I, Waldman JR, Rosko J, Gross R, Collins MR, Rogers SG, Stabile J (2000) Genetic structure of Atlantic sturgeon populations based on mitochondrial DNA control region sequences. Trans Am Fish Soc 129:476–486
Wirgin I, Waldman J, Stabile J, Lubinski B, King T (2002) Comparison of mitochondrial DNA control region sequence and microsatellite DNA analyses in estimating population structure and gene flow rates in Atlantic sturgeon Acipenser oxyrinchus. J Appl Ichthyol 18:313–319
Wirgin I, Maceda L, Waldman JR, Wehrell S, Dadswell M, King T (2012) Stock origin of migratory Atlantic sturgeon in Minas Basin, Inner Bay of Fundy, Canada, determined by microsatellite and mitochondrial DNA analyses. Trans Am Fish Soc 141:1389–1398
Acknowledgements
We would like to thank the assistance of J. Carter and National Oceanic and Atmospheric Administration (NOAA) Northwest Fisheries Science Center (NWFSC) Forensics Laboratory (formerly in Charleston, SC) for the archiving and access to Atlantic sturgeon genetic samples. We would also like to thank the Diadromous Fishes Research team at SCDNR for the collection of the samples, the Hollings Marine Laboratory for laboratory space, and the Fish Population Genetics research group at SCDNR for lab assistance, especially T. O’Donnell for laboratory optimization. This project was funded by a Sect. 6 Species Recovery Grants to States from the National Marine Fisheries Service (Grant No. NA13NMF4720044). This manuscript represents publication number 755 from the SCDNR Marine Resources Research Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Farrae, D.J., Post, W.C. & Darden, T.L. Genetic characterization of Atlantic sturgeon, Acipenser oxyrinchus oxyrinchus, in the Edisto River, South Carolina and identification of genetically discrete fall and spring spawning. Conserv Genet 18, 813–823 (2017). https://doi.org/10.1007/s10592-017-0929-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-017-0929-7