Abstract
Understanding the degree of genetic population differentiation is important in conservation genetics for inferring gene flow between populations and for identifying small and isolated threatened populations. We evaluated the genetic variation within and between three populations of the rock firefinch (Lagonosticta sanguinodorsalis), a range restricted firefinch endemic to Nigeria and Cameroon. The populations were closely located (c. 100 km apart) within the species’ core distribution in Central Nigeria. We found that the populations had similar levels of gene diversities (H E ) and low but significant inbreeding coefficients (F IS ). Despite the short distance between populations there was a weak but significant population structure, which indicates that the populations are somewhat isolated and affected by drift within the species’ core distribution in Nigeria. The knowledge of the genetic status of the rock firefinch will serve as a foundation to future studies to help understand population demography and for managing and maintaining viable populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Habitat fragmentation leads to reduction in total habitat area and separation into isolated patches, which in turn reduce total population size and may reduce gene flow between populations (Frankham et al. 2002). Thus, fragmented and isolated populations will experience a reduction in effective population size, and it has been shown that small populations are more prone to loss of genetic diversity through genetic drift and more vulnerable to harmful inbreeding effects due to loss of genetic variation (Allendorf 1986; Vilà et al. 2003). During population contractions, increased levels of inbreeding and loss of genetic variation can cause negative feedback loops on population size, which sometimes can lead to rapid drop in population size and even local extinction of populations (Westemeier et al. 1998; Madsen et al. 1999)—a scenario sometimes referred to as the extinction vortex (Fagan and Holmes 2006).
Maintaining high levels of genetic variation is crucial for the long-term persistence of populations also for other reasons than for avoiding inbreeding depression (Allendorf 1986; Keller et al. 1994). Most organisms inhabit constantly changing environments, and need to possess the genetic material necessary for adaptation. Populations that are genetically depleted will be unable to adapt to changing conditions and thus face a higher risk of extinction if the environment changes (Spielman et al. 2004). Therefore, maintaining dispersal possibilities between populations to ensure gene flow and genetic variation within populations is very important in wildlife management to sustain viable populations of vulnerable and threatened species (Page and Holmes 1998).
Naturally occurring as well as anthropogenic induced habitat heterogeneities structure the distribution of organisms. Species living in naturally fragmented habitats have however evolved as naturally fragmented populations, whereas other species have seen their distribution being fragmented due to recent anthropogenic impingements. The reed bunting (Emberiza schoeniclus; Mayer et al. 2009) and the great reed warbler (Acrocephalus arundinaceus; Hansson et al. 2000, 2002, 2008) for example, inhabit mosaics of wetlands, so it is likely that their population structure depended to a large extent on the size and connectivity of available wetlands prior to recent anthropogenic habitat alterations. In contrast, the red-cockaded woodpecker (Picoides borealis; Jackson 1978) and the chestnut-backed antbird (Myrmeciza exsul; Woltmann et al. 2012) are two of many examples of species that probably inhabited large continuous habitats, but currently exist in small and isolated pockets within their today strongly human-influenced range. In general, habitat specialists with preference for habitats with non-continuous distributions, such as wetlands, marshes, rocky outcrops, gallery forests, ponds, etc., are likely to have been evolving in fragmented populations, as they utilize suitable habitats which occur as scattered patches.
The rock firefinch (Lagonosticta sanguinodorsalis) is a range-restricted species, endemic to Nigeria and Cameroon. It was described as recently as in 1998 (Payne 1998). It is a non-territorial monogamous, ground feeding seed eater. The species is sexually dimorphic, with adult males having a more brightly coloured plumage than adult females (Fry and Keith 2004). Studies in its core distribution on the Jos Plateau, Central Nigeria, have shown that it has a strong preference for rocky habitats (Wright and Jones 2005; Brandt and Cresswell 2008; Abalaka and Jones 2011), with population densities of 0.48 birds/ha in uncultivated areas and 0.62 birds/ha in cultivated areas (Abalaka and Jones 2011). Although the rock firefinch has a naturally patchy distribution due to its preference for rocky habitats, recent human activities (such as farming and blasting of rocks to establish settlements) have contributed to further isolating the populations. Being a range-restricted species, known to occur only in Nigeria and Cameroon, with high probability of losing subpopulations due to ongoing anthropogenic habitat fragmentation, the rock firefinch is of high conservation interest. Furthermore, it holds an important role in the life-history of another endemic species, the Jos Plateau indigobird (Vidua maryae)—a brood parasitic species specific to the rock firefinch (Payne 1998; Brandt and Cresswell 2008)—which makes the rock firefinch of additional conservation interest. In this study, we quantified the genetic diversity, differentiation and structure in three rock firefinch populations within its core distribution in Central Nigeria by screening individuals at a set of newly identified polymorphic microsatellite loci (Abakala and Hansson 2014). We also attempted to estimate the effective population size of the populations. Understanding the degree of genetic population differentiation is important for inferring gene flow between populations and for identifying small and isolated threatened populations. Knowledge of the genetic status of this species will also serve as a foundation to future studies to help understand its population demography and conservation biology.
Materials and methods
Study area
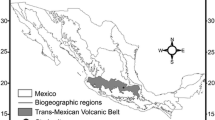
We selected three sites on the Jos Plateau, Central Nigeria: the Amurum Forest Reserve (Amurum from here on) (N09°52′44.75″, E08°58′55.75″), Pankshin (N09°21′36.23″, E09°25′52.67″) and Kagoro Boys Scout Camp (Kagoro from here on) (N09°34′27.20″, E08°22′54.39″; Fig. 1). The distance between these sites ranged from 74 to 118 km. In altitude, Amurum is 1,300 m.a.s.l., Pankshin is 1,200 m.a.s.l. and Kagoro is 700 m.a.s.l. Kagoro is situated on the south-eastern escarpment of the Jos Plateau as the plateau descends (which explains its comparatively low altitude), whereas the two other populations are located on the plateau. The habitat of all sites was fairly similar, with large rocky outcrops and seasonal streams. In Pankshin, the habitat was somewhat more degraded than at Kagoro and Amurum, as most of the gallery forest that grows along the stream has been totally removed and a major part of the land has been converted to farmlands. Amurum, a forest reserve of about 300 ha, is surrounded by farmlands and has been under protection with little human disturbance for over 10 years. Kagoro also has farmlands.
Sample collection
We collected 161 blood samples from the three populations of rock firefinch (Fig. 1) during the years 2006–2009 (Table 1). Birds were trapped using mist nets, and 20 µl of blood was collected by puncturing the wing vein (Wingfield and Farner 1976). Blood was preserved in 500 µl SET buffer (0.15 M NaCl, 0.05 M Tris, 0.001 M EDTA, pH 8.0) at room temperature until transported to the laboratory where they were store at −20 °C.
DNA extraction
We isolated DNA from the blood samples following Bensch et al. (1994). We added 13 µl of SDS (20 %) and 15 µl of Proteinase K (10 mg/ml) to blood samples and incubated overnight at 55 °C. DNA was purified by extracting with phenol/chloroform-isoamylalcohol. The dissolved DNA was precipitated with 0.1 volumes of 3 M sodium acetate and two volumes absolute ethanol, washed with 70 % ethanol and vacuum dried. Finally the DNA pellet was solved and stored in 50 µl 1× TE buffer.
Microsatellite genotyping
All 161 rock firefinches were genotyped at 11 autosomal microsatellite loci (Table 2; Abakala and Hansson 2014). The PCR-mix contained: 1 × Q-mix (Qiagen), 1.2 pmol each of forward and reverse primer, 10–25 ng DNA template and ddH2O in 6 µl reaction volume. The PCRs were initiated with a hotstart (95 °C for 15 min) followed by 35 cycles with 94 °C for 30 s, T a (see Table 2) for 90 s and 72 °C for 90 s; and a final extension at 72 °C for 10 min. Primers with the same PCR condition were labeled with different fluorescent dyes and were combined in the same PCR multiplex. The PCR products were separated and visualized using an ABI 3730 capillary sequencer (Applied Biosystems). The microsatellite data were analyzed in GeneMapper version 4 (Applied Biosystems) to determine the genotypes of individuals for each locus.
Statistical analyses
All loci were tested for presence of null alleles, large allele dropout and scoring errors due to stutter peaks in MICRO-CHECKER 2.2.3 (Oosterhout et al. 2004). Mean number of alleles (A), observed and expected heterozygosity (H O and H E ) were calculated in GENALEX 6.5 (Peakall and Smouse 2006) and inbreeding coefficient (F IS ) and associated P values were computed in FSTAT 2.9.3.2 (1,000 permutations; Goudet 2001).
Levels of differentiation among populations were evaluated using both population level approach (F ST ) and individual-based clustering approach (STRUCTURE; Pritchard et al. 2000). Global and pairwise F ST were determined as described by Weir and Cockerham (1984) in the FSTAT 2.9.3.2 (P values were calculated using 1,000 permutations). Given the recent debate surrounding the use of F ST in resolving population differentiation (Jost 2008; Meirmans and Hedrick 2011), both global and pairwise Jost’s measure of differentiation (D est ) were also calculated (in GENALEX 6.5; P values were calculated using 999 permutations).
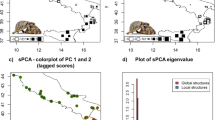
The Bayesian clustering method and Markov Chain Monte Carlo (MCMC) simulation implemented in STRUCTURE 2.3.4 (Pritchard et al. 2000) was used to corroborate the population level approaches. The STRUCTURE analyses were run by using an admixture model and correlated allele frequencies with a burn-in period of 50,000 replicates and sampling period of 100,000 replicates for number of clusters (K) from 1 to 6. Thirty independent runs were performed for each K. Two sets of STRUCTURE analyses were run: one without using sample location as prior, one using sample location as prior. To determine the number of genetic clusters (K), we used the ΔK method of Evanno et al. (2005) based on the second-order rate of change in log Pr (X|K) as implemented by the program STRUCTURE HARVESTER (Earl and vonHoldt 2012). Replicate cluster analyses were aligned with CLUMPP 1.1.2 (Jakobsson and Rosenberg 2007) and visualizes in DISTRUCT 1.1 (Rosenberg 2004). Lastly, the pattern of genetic differentiation between the three populations was plotted using principal coordinates analysis (PCA) implemented in GENALEX 6.5.
We calculated effective population size using the moment-based temporal approach based on (Pollak 1983) implemented in NEESTIMATOR 1.3 (Peel et al. 2004), which uses genotypic data of individuals collected over different years (Table 1).
Results
We successfully genotyped 153 out 161 individuals at 11 polymorphic microsatellite loci, with number of alleles per locus ranging from 2 at WBSW7 to 18 at Lsa127. TG13-017 was found to have null alleles in all three populations (estimated null allele frequencies: 0.143 in Amurum, 0.133 in Pankshin and 0.180 in Kagoro) and was therefore excluded from further analyses.
The number of alleles, and observed and expected heterozygosities, were similar among populations at most loci (Table 3). The F IS within the three populations were low to moderate (Amurum: F IS = 0.054, P = 0.037; Pankshin: F IS = 0.024, P = 0.155; Kagoro: F IS = 0.038, P = 0.105), but did not differ significantly from zero after Bonferroni correction (P critical = 0.017). However, the global F IS value (0.036) was statistically significant (P = 0.008), indicating a slight deviation from Hardy–Weinberg equilibrium (HWE).
A weak but statistically significant population differentiation was detected between the three populations for both F ST -based statistics (F ST = 0.015, P < 0.001; 95 % CI 0.008–0.024;) and Jost’s measure of population differentiation (D est ) analysis (D est = 0.021, P = 0.002; 95 % CI 0.009–0.045). D est values and F ST values were similar for most loci (Table 3). Both F ST -based analysis and D est -differentiation analysis detected significant pairwise differentiation between all population pairs (Table 4).
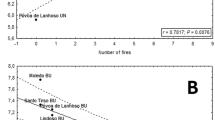
The STRUCTURE analyses, without using location as prior revealed no structure (Fig. 2a) whereas moderate clustering was detected in models using location as a prior (Fig. 2b). The ΔK method implemented in STRUCTURE HARVESTER identified three clusters (K = 3) as the most probable number of clusters for the later model (Fig. 3), but it should be noted that the estimated likelihood of K (Ln P(K)) was very similar for K = 1–3 (Fig. 3).
Individual Bayesian assignment probabilities for K = 2 and 3 for three populations of the rock firefinch. Individuals are represented by thin vertical bars, which are partitioned into K shaded segments representing each individual’s estimated membership fraction. The black lines separate sampling sites. Results for (a) without location as prior and (b) with location as prior are given
The first three axes from the PCA explained a total of 57.9 % of the variation in the data (1st axis = 21.2 %, 2nd axis = 18.9 % and 3rd axis = 17.9 %). A plot of coordinate 1 and 2 (Fig. 4) shows that although there is substantial overlap between populations, there is also slight structuring. This agrees with the weak differentiation detected in the population level and individual-based approaches above.
Analysis of the effective population size (Ne) using temporal sampling indicated relatively small Ne:s but the estimates were unfortunately associated with wide confidence intervals in two of the populations (Amurum: 15.6, 95 % CI 5.3–189.2; Pankshin: ∞, 95 % CI 61.9–∞; Kagoro: 27.0, 95 % CI 7.9–∞).
Discussion
We have found weak but significant population differentiation between three rock firefinch populations within its core distribution in Central Nigeria. The F IS within populations were low to moderate (significant at the global level but non-significant after Bonferroni correction in the population specific analyses), which indicates some level of inbreeding. The estimated effective population sizes indicate relatively low population sizes but the estimates were associated with wide confidence intervals in two populations and are therefore difficult to interpret.
Both the population level and individual-based population differentiation analyses, and the PCA plot, support a slight population differentiation among the examined rock firefinch populations on the Jos Plateau. The results from the STRUCTURE analysis (Fig. 2b; K = 2) further suggested that the populations at Amurum and Pankshin are somewhat more closely genetically related to each other than to the Kagoro population (this was however not supported by pairwise F ST and D est results; Table 4). Kagoro is not located further away than any of the other populations (Fig. 1), but its slightly more pronounced genetic isolation may be related to its low-altitude location (c. 700 m.a.s.l.) compared to the other populations (>1,000 m.a.s.l.).
Ecological studies have found that rock firefinches exhibit high site fidelity (Brandt and Cresswell 2008), and do not move far from their natal place. The present analysis partly supports this view by suggesting that even closely located rock firefinch populations are to some extent isolated and affected by drift and inbreeding within its core distribution in Nigeria. However, the site fidelity behavior could also be as a result of a recent resistance to dispersal driven by recent habitat fragmentation and isolation.
Most species that originally inhabit continuous habitats are, when faced with fragmentation, forced into smaller remaining patches. This leads to overall reduction in population size and reduced dispersal among patches (Frankham et al. 2002). The expected long-term consequences of restricted gene flow between patches are differentiation and substructuring of populations, which has for example been reported in the chestnut-backed antbird in human influenced rain forests (Woltmann et al. 2012) and in the red-cockaded woodpecker, in fragmented pine forests (Stangel et al. 1992). Conversely, species that have existed as naturally fragmented populations may be well adapted to the patchy nature of their distribution and may be able to maintain gene flow between the patches by dispersal, thus avoiding genetic substructuring between populations. This may for example apply to the great reed warbler, which occurs in patches of wetlands and maintains gene flow and lack small-scaled population structure in spite of its patchy distribution (Hansson et al. 2000, 2002, 2008). Nevertheless, with continuous destruction and modification of landscapes, both organisms that have evolved in continuous distributions and those that evolved with fragmented distributions may be affected eventually.
The rock firefinch on the Jos Plateau has naturally existed in habitat patches and, therefore, the effect of fragmentation may not be as evident in this species. In line with this reasoning, we expect that rock firefinch populations should behave genetically to a large extent as a single large population. However, the result of this study has shown evidence for some degree of isolation and drift, but these effects are considered to be weak when compared to populations whose genetic diversity have been heavily structured by fragmentation. It may be that, the population structure detected in this study results from a recent resistance to gene flow, which may reflect that the rock firefinch populations on the Jos Plateau have reacted to increased impact of anthropogenic activities on their habitats. Thus, there is a need for further studies to detect and prevent any anthropogenic activities that may become a hindrance to efficient dispersal between populations.
References
Abakala J, Hansson B (2014) Microsatellite markers for population genetic study of the rock firefinch, Lagonosticta sanguinodorsalis. Afr Zool 49
Abalaka JI, Jones PJ (2011) Population densities of the Rock Firefinch Lagonosticta sanguinodorsalis and some other estrildine and viduine finches on the Jos Plateau, Nigeria. In: Harebottle DM, Craig AJFK, Anderson MD, Rakotomanana H, Muchai M (eds) Proceedings of the 12th Pan-African Ornithological Congress, 2008. Cape Town, Animal Demography Unit, pp 131–134
Allendorf FW (1986) Genetic drift and the loss of alleles versus heterozygosity. Zoo Biol 5:18–190
Bensch S, Hasselquist D, von Schantz T (1994) Genetic similarity between parents predicts hatching failure: nonincestuous inbreeding in the great reed warbler. Evolution 48:317–326
Brandt M, Cresswell W (2008) Breeding behaviour, home range and habitat selection in rock firefinches Lagonosticta sanguinodorsalis in the wet and dry season in Central Nigeria. Ibis 150:495–507
Dawson DA, Horsburgh GJ, Kupper C, Stewart IRK, Ball AD, Durrant KL, Hansson B, Bacon I, Bird S, Klein A, Krupa AP, Lee JW, Martin-Galvez D, Simeoni M, Smith G, Spurgin LG, Burke T (2010) New methods to identify conserved microsatellite loci and develop primer sets of high cross-species utility: as demonstrated for birds. Mol Ecol Resour 10:475–494
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Fagan FW, Holmes EE (2006) Quantifying the extinction vortex. Ecol Lett 9:51–60
Frankham R, Briscoe DA, Ballou JD (2002) Introduction to conservation genetics. University of Cambridge Press, Cambridge
Fry CH, Keith S (2004) Birds of Africa, Vol. VII: sparrows to buntings, Christopher Helm, London
Goudet J (2001) FSTAT, a program to estimate and test gene diversities and fixation indices, version 2.9.3. http://www2.unil.ch/popgen/softwares/fstat.htm
Hansson B, Bensch S, Hasselquist D, Lillandt B-G, Wennerberg L, von Schantz T (2000) Increase of genetic variation over time in a recently founded population of great reed warblers (Acrocephalus arundinaceus) revealed by microsatellites and DNA fingerprinting. Mol Ecol 9:1529–1538
Hansson B, Bensch S, Hasselquist D, Nielsen B (2002) Restricted dispersal in a long-distance migrant bird with patchy distribution, the great reed warbler. Oecologia 130:536–542
Hansson B, Hasselquist D, Tarka M, Zehtindjiev P, Bensch S (2008) Postglacial colonisation patterns and the role of isolation and expansion in driving diversification in a passerine bird. PLoS ONE 3:e2794
Jackson JA (1978) Analysis of the distribution and population status of the red-cockaded wood pecker. In: Odum RR, Landers L (eds) Proceedings of the rear and endangered wildlife symposium Technical Bulletin WL4, Georgia Department of Natural Resources, Game and Fish Division, Athens, Georgia, pp 101–110
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Jost L (2008) GST and its relatives do not measure differentiation. Mol Ecol 17:4015–4026
Keller LF, Arcese P, Smith JNM, Hochachka WM, Stearns SC (1994) Selection against inbred song sparrows during a natural population bottleneck. Nature 372:356–357
Madsen T, Shrine R, Olsson M, Wittzell H (1999) Restoration of an inbred adder population. Nature 402:34–35
Mayer C, Schiegg K, Pasinelli G (2009) Patchy population structure in a short distance migrant: evidence from genetic and demographic data. Mol Ecol 18:2353–2364
McRae SB, Amos W (1999) Characterization of hypervariable microsatellites in the cooperatively breeding white-browed sparrow weaver Plocepasser mahali. Mol Ecol 8:903–904
Meirmans PG, Hedrick PW (2011) Assessing population structure: FST and related measures. Mol Ecol Resour 11:5–18
Oosterhout CV, Hutchinson WF, Wills DPM, Shipley P (2004) Micro-Checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Page RDM, Holmes EC (1998) Molecular Evolution: a phylogenetic approach. Blackwell, Oxford
Payne RB (1998) A new species of firefinch Lagonosticta from northern Nigeria and its association with the Jos Plateau Indigobird Vidua maryae. Ibis 140:368–381
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Peel D, Ovenden JR, Peel SL (2004) NEESTIMATOR: software for estimating effective population size, Version 1.3. Queensland Government, Department of Primary Industries and Fisheries, St Lucia, Queensland. http://www.dpi.qld.gov.au/28_6908.htm.
Pollak E (1983) A new method for estimating the effective population size from allele frequency changes. Genetics 104:531–548
Pritchard JK, Stephens M, Donelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Slate J, Hale MC, Birkhead TR (2007) Simple sequence repeats in zebra finch (Taeniopygia guttata) expressed sequence tags: a new resource for evolutionary genetic studies of passerines. BMC Genome 8:52
Spielman D, Brook BW, Frankham R (2004) Most species are not driven to extinction before genetic factors impact them. Proc Natl Acad Sci USA 101:15261–15264
Stangel PW, Lennartz RM, Smith MH (1992) Genetic variation and population structure of red-cockaded Woodpeckers. Conserv Biol 6:283–292
Vilà C, Sundqvist A, Flagstad Ø, Seddon J, Björnerfeldt S, Kojola I, Casulli A, Sand H, Wabakken P, Ellegren H (2003) Rescue of a severely bottlenecked wolf (Canis lupus) population by a single immigrant. Proc R Soc Lond B 270:91–97
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Westemeier RL, Brawn JD, Simpson SA, Esker TL, Jansen RW, Walk JW, Kershner EL, Bouzat JL, Paige KN (1998) Tracking the long-term decline and recovery of an isolated population. Science 282:1695–1698
Wingfield JC, Farner DS (1976) Avian endocrinology: field investigations and methods. Condor 78:570–573
Winker K, Glenn TC, Graves GR (1999) Dinucleotide microsatellite loci in a migratory wood warbler (Parulidae: Limnothlypis swainsonii) and amplification among other songbirds. Mol Ecol 8:1553–1556
Woltmann S, Kreiser BR, Sherry TW (2012) Fine-scale genetic population structure of an understory rainforest bird in Costa Rica. Conserv Genet 13:925–935
Wright D, Jones P (2005) Population densities and habitat associations of the range-restricted Rock Firefinch Lagonosticta sanguinodorsalis on the Jos Plateau, Nigeria. Bird Conserv Int 15:287–295
Acknowledgments
This study was supported by Grants from the British Ecological Society (to JA), the A. P. Leventis foundation (to JA), Swedish Ornithological Society (to JA), Swedish Research Council (to BH) and EU FP7 (Marie Curie IRSES, project Avian Genomics 295276) (to BH). We also thank the A. P. Leventis Ornithological Research Institute (APLORI) for logistical support. Arin Izang was a great assistant during the fieldwork. This is APLORI publication no. 80.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Abalaka, J., Hudin, N.S., Ottosson, U. et al. Genetic diversity and population structure of the range restricted rock firefinch Lagonosticta sanguinodorsalis . Conserv Genet 16, 411–418 (2015). https://doi.org/10.1007/s10592-014-0667-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-014-0667-z