Abstract
The Lao People’s Democratic Republic (PDR) may have the largest Asian elephant population in Indochina. However, elephants on Lao PDR’s Nakai Plateau are potentially threatened by the construction of a hydropower dam that will flood important habitat. We conducted a non-invasive genetic study of elephants in this region to provide baseline data on genetic diversity and social structure prior to dam construction. For the 102 elephants we detected, values of observed heterozygosity (0.711) and allelic diversity (8.0 alleles/locus) at microsatellite loci were higher than those found in elephant populations in India and Vietnam, while mitochondrial diversity (haplotype diversity 0.741; nucleotide diversity 0.011) was similar to that reported for the Lao/Vietnam region. Six mitochondrial haplotypes were detected, representing both major clades previously reported in this species. Relatedness estimates between females and young detected near each other are consistent with familial relationships, and relatedness estimates between adult males and females suggest male locational dispersal. Since family group structure appears to be intact in the Nakai region, these elephants will likely move as relatively large family groups in response to habitat disturbance. These results have positive implications for the viability of the elephant population in this region, demonstrate its conservation significance, and will be valuable for predicting and monitoring the effects of the hydropower dam over time.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Asian elephants (Elephas maximus) are struggling to survive in many parts of their range, and outside of the Indian subcontinent it is mainly small populations in habitat fragments that persist (Leimgruber et al. 2003; Hedges 2006). The primary threats to Asian elephants are illegal hunting, increasing human encroachment into elephant habitat, and other forms of habitat loss and degradation (Duckworth and Hedges 1998; Leimgruber et al. 2003; Hedges 2006). The loss of habitat often leads to human–elephant conflict (HEC), which erodes support for elephant conservation and can lead to the retaliatory killing of elephants or the removal of elephants from the wild. As efforts to maintain Asian elephants in the wild progress, it is important to understand where the population strongholds are and to develop a better understanding of their ecology and their demographic and genetic parameters. The Lao People’s Democratic Republic (PDR) is thought to harbor the largest Asian elephant populations in Indochina (Duckworth and Hedges 1998). Within Lao PDR, however, possibly the largest population of Asian elephants (Duckworth et al. 1999) is experiencing habitat loss as a result of the construction of the NT2 dam on the Nam Theun river.
The ultimate goal of most conservation strategies is to protect viable and ecologically functional populations, and one of the most important factors to consider for long-term population viability is genetic diversity. Because many of the remaining populations of Asian elephants are small and isolated, they are threatened with the loss of genetic diversity through inbreeding and/or genetic drift. Genetic diversity can also be influenced by social structure, which affects how animals interact and breed (Archie et al. 2008). Although Asian elephants are not believed to form social groups of maternally related females as large as those formed by African savanna elephants (Loxodonta africana africana), smaller groups of related females and their young have been observed (Fernando and Lande 2000; Vidya and Sukumar 2005b).
Mating systems also influence genetic diversity. In elephants, as in many mammals, males are the dispersing sex (Sukumar 1989). Dispersal can be locational, in which individuals move permanently away from their natal areas at sexual maturity, or social, in which they remain in their natal areas but move long distances for mating. Two studies suggest non-random locational dispersal in Asian elephants (Vidya and Sukumar 2005a, b). However, with the exception of a study in Vietnam (Vidya et al. 2007), most of what is known about the social structure and mating strategies of Asian elephants comes from larger populations in the Indian subcontinent. It is unclear whether the patterns detected there will be found in other parts of the Asian elephant’s range where most populations are now smaller and more fragmented.
The overall goals of this project were to provide baseline data about the size, genetic diversity, and social structure of the elephant population on the Nakai Plateau, Lao PDR, and to compare this population with larger populations on the Indian subcontinent. Here we sought to document the overall genetic diversity of the elephants on the Nakai Plateau, to examine the genetic relatedness of female elephants found in the same place at the same time, and to determine whether adult males in the population exhibit evidence of locational dispersal. Asian elephants live in forests in most of the areas where they still occur, which makes it difficult to directly observe them and to obtain the behavioral data necessary to understand social structure. Therefore, we used non-invasively collected samples to address these objectives. Because this population is threatened with habitat loss from the construction of a dam on the Nam Theun River, we also examined the patterns of genetic diversity and sexual segregation across the landscape. These are important baseline data that can be used to predict the consequences of future range reductions for the population and that can also be used to assess its conservation significance.
Methods
Study area
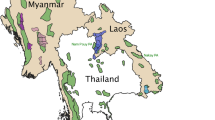
Our study was conducted in the Nakai-Nam Theun (NNT) National Protected Area (NPA; 3,532 km²) and adjoining protected areas, in the Khammouane and Bolikhamxay Provinces (Fig. 1). This region represents the Lao PDR’s largest and most diverse natural and protected forest area, and it has been suggested that the NNT NPA contains one of the two largest elephant populations in the country (Duckworth et al. 1999). In and around the NNT NPA, elephants occur on the Nakai Plateau (1,250 km²) and range into the Nakai-Nam Theun-Phou Hin Poun (NNT-PHP) Corridor (738 km²) that links the NNT NPA with the Phou Hin Poun NPA.
Field methods
The DNA samples for the study were obtained non-invasively from dung collected during a capture–recapture dung survey. That project was carried out in two phases: a pilot phase and a formal survey phase. The pilot phase (16 September 2004 to 3 March 2005) aimed to assess whether ‘fresh’ (<48 h old, moist with odor) and ‘reasonably fresh’ (boli mostly intact and not obviously degraded or damaged by climate, fungus or insects, believed to be no older than 2 weeks) dung-piles could be found in adequate numbers if survey teams searched in likely elephant ‘hotspots’ (Hedges and Lawson 2006). It also aimed to collect preliminary samples to test preservation methods and select loci that were informative for this population. The second phase of fieldwork (21 February 2006 to 1 May 2006) comprised a formal capture–recapture survey with a design that followed Hedges and Lawson (2006). The survey focused on 16 blocks containing one or more ‘hotspots’, which are places where elephant dung had been found reliably in the past or places known to be used by elephants including salt licks, waterholes, and areas of HEC (Fig. 1). An additional design constraint resulted from the need to have no ‘gaps’ in the coverage of the site where elephants could move without any chance of the teams collecting their dung (i.e., every elephant’s home range should contain at least 2–3 sample collection locations). The blocks varied in size from 2 to 8 km² and were located predominantly along the banks of the Nam Theun River and in other areas of the plateau where crop raiding by elephants had been reported (Fig. 1). Each block was sampled approximately once per month. Sample sizes depended on the availability and detectability of fresh dung, but the experience of the field teams was used to maximize sampling rates.
For each sample, approximately 10 g of fresh (<48 h old) dung were placed in 40 ml polypropylene tubes, boiled for 15 min to destroy potential pathogens, and preserved in Queens College Buffer (20% DMSO, 100 mM Tris pH 7.5, 0.25 M EDTA, saturated with NaCl; Amos et al. 1992). Samples were initially stored in the dark at room temperature at the field station and then moved to Vientiane where they were stored at 4°C before being shipped to the USA under USDA permit number 48529.
Genetic methods
DNA was extracted from dung samples following the methods outlined in Eggert et al. (2005). To minimize the possibility of contamination, extractions were performed in a lab designated exclusively for the extraction of DNA from non-invasively collected samples. Each group of extractions was accompanied by controls for all reagents, which were tested for contamination in the first set of amplifications of all loci.
All samples were genotyped at ten dinucleotide microsatellite loci. Nine of the loci were developed for Asian elephants (EMU03, EMU04, EMU07, EMU10, EMU12, EMU13, EMU14, EMU15, EMU17; Kongrit et al. 2008), and one locus was redesigned from a locus originally developed for African elephants (FH94R; Eggert et al. 2008). Samples were amplified in 25 μl reactions containing 0.5 U AmpliTaq Gold DNA Polymerase (Applied Biosystems), 1× PCR Gold Buffer (Applied Biosystems), 0.4 μM fluorescently labeled forward primer, 0.4 μM unlabeled reverse primer, 2 mM MgCl2, 0.2 mM each dNTP, 1.5 μl 10× BSA, and 3 μl of the DNA extract. PCR profiles followed Kongrit et al. (2008). PCR products were visualized in a 2% agarose gel stained with Gel Star (Lonza) to verify amplification of samples and non-amplification of negative controls. Fragment analysis was performed in an ABI 3730 DNA Analyzer, and genotypes were scored using GeneMarker v1.6 (Soft Genetics LLC). To minimize the probability of genotyping error, we used the “comparative approach” (Frantz et al. 2003; Hansen et al. 2008), in which matching heterozygotes were scored at least twice, and matching homozygotes at least three times. The computer program Microchecker (van Oosterhout et al. 2004) was used to test for evidence of large allele dropout and null alleles, and to estimate the frequency of nulls at loci where they were detected.
Genotypes were compared in the Excel Microsatellite Toolkit (Park 2001), a utility for Microsoft Excel, and all samples that differed by four or less alleles were identified. Because DNA extracts from non-invasively collected samples are dilute and contain degraded DNA, each of these genotypes were rechecked for possible problems with allelic dropout. Those that differed at two or less alleles but matched in sex and had very similar bolus circumferences were considered to be the same individual. This conservative approach was taken to avoid scoring samples as individuals when they are actually erroneous genotypes. Scoring erroneous genotypes as captures in the capture–recapture analysis could positively bias the estimate of population size (Creel et al. 2003).
Molecular sexing was used to detect the sexes of individuals. PCR reactions were performed using the methods and primers of Munshi-South et al. 2008, which amplify a 141 bp fragment of the X- and Y-linked zinc finger protein (ZFX/ZFY) genes. 7 μL of the amplification products were digested with BamHI (New England Biolabs) for 2 h, after which products were visualized in a 3% agarose gel stained with Gel Star (Lonza). Because the restriction site is on the Y-chromosome, single bands were scored as females and three bands were scored as males. Sexes were confirmed by repeating the process once for each sample.
Once individuals had been determined based on their unique microsatellite genotypes, a 600 bp fragment of the left domain of the mitochondrial control region was amplified for all but four of them (which had insufficient sample remaining) using the primers MDL3 and MDL5 (Fernando and Lande 2000) and the conditions outlined in Eggert et al. (2002). PCR products were sequenced in both directions using Big Dye Terminator cycle sequencing chemistry in an ABI 3730 DNA Analyzer (Applied Biosystems). Sequence alignment and editing was completed in SEQUENCHER v4.5 (Gene Codes Corporation), and sequences that differed by at least one nucleotide were designated as different haplotypes.
Data analysis
For microsatellite data, GENEPOP (Raymond and Rousset 1995) was used to test for deviations from the expected genotype frequencies under Hardy–Weinberg equilibrium (HWE) and linkage disequilibrium, as well as to calculate the allelic diversity and observed and expected heterozygosities for each locus. We tested the population for a signature of a recent bottleneck using a test for heterozygosity excess (Cornuet and Luikart 1996) as well as a test for an allele distribution mode shift (Luikart et al. 1998) using the program Bottleneck v1.2.02 (Cornuet and Luikart 1996). Relatedness between individuals was estimated using Kinship v1.3.1 (Goodnight and Queller 1999).
MtDNA haplotype and nucleotide diversity was calculated using Arlequin v2.0 (Schneider et al. 2000). Published Asian elephant haplotypes for this region of mtDNA were downloaded from GenBank (accession #s AF368903, AF317518-317535, AF324827-324828, AY245823, AY365432-365433, AY589512-589516) and aligned with the Nakai haplotypes in SEQUENCHER to create a NEXUS file. This file was analyzed in TCS (Clement et al. 2000) to create a haplotype network.
The GPS coordinates recorded for each dung pile along with the date collected were imported into ArcMap 9.2 (Environmental Systems Research Institute Inc.) to identify the individuals detected within the same sampling block at approximately the same time. All dung-piles sampled were estimated to be no older than 48 h, and sampling blocks were generally sampled in 1 or 2 days. We assumed that the presence of females in the same sampling block at approximately the same time indicated a possible association between them, and calculated the average pairwise relatedness between these individuals. All females and juvenile males (assumed to be dependent on their mother) were included in the initial analysis. The age class of an individual was assigned using the criteria of Tyson et al. (2002): briefly, a mean dung bolus circumference of ≤30 cm was considered a neonate or juvenile, 30–42 cm was considered a subadult, and >42 cm was considered an adult.
The same procedure was used to examine the relatedness between subadult and adult males detected in the same sampling block at approximately the same time. The analysis was then broken down to examine females and juvenile males that shared the same haplotype. Only males with a known bolus circumference (and therefore an estimated age class) were included in the analysis. Average pairwise relatedness was calculated for all individuals in the groups of interest. We report the average pairwise relatedness, the standard error, and 95% confidence intervals for all calculations.
Results
Evidence of recent elephant activity and fresh dung samples were found and collected in only five of the 16 sampling blocks (Fig. 2). From blocks 5, 8, 9, 10, and 12, a total of 267 samples were collected. Of these, 215 (80.5%) were successfully genotyped and a total of 102 individuals [out of an estimated population of 132 (95% CI 120–149) individuals, Hedges et al. unpublished] were identified by their unique genotypes. Using molecular sexing, 73 females and 29 males were detected (Table 1). The age structure of the population included 40 adults, 31 subadults, 21 juveniles, and 10 elephants whose age could not be estimated due to the lack of bolus circumference measurements.
Genetic diversity
The level of genetic diversity, as indicated by microsatellite genotypes, was high in this population. The average number of alleles over ten loci was 8.1 (range: 5–15), and the average observed heterozygosity was 0.748 (Table 2). After applying a Bonferroni correction for multiple tests, three of the ten loci (EMU10, EMU13, EMU17) did not conform to Hardy–Weinberg expectations, with lower than expected heterozygosities (Table 2). Analysis in Microchecker did not detect any evidence of large allele dropout, and estimates of null allele frequencies ranged from 0.097 to 0.134 (Table 2). Excluding these three loci, the average number of alleles was 8.0 and average heterozygosity was 0.711. Five of 45 pairs of loci exhibited linkage disequilibrium (EMU04-EMU07, EMU10-EMU12, EMU07-EMU15, EMU10-EMU03, EMU15-FH94R). While it is possible that the five pairs of loci are actually linked, we believe it is more likely that these problems are the result of the inclusion of a relatively high percentage of closely related individuals in our analyzes. Further support for this can be found in the fact that linkage between these loci was not seen in a study of elephants from Cambodia (Pollard et al. 2008).
Our analyzes did not reveal any signs of a recent bottleneck event. The allele frequency distribution showed a normal L-shaped distribution with no evidence of a loss of rare alleles. Additionally, the Wilcoxon test showed no evidence of heterozygosity excess under either the Two Phase Mutation Model (P = 0.348) or the Stepwise Mutation Model (P = 0.862).
Mitochondrial diversity was also high in this population. Six haplotypes were detected (Table 3), two of which were found in only one individual (one subadult male and one adult female; Table 4). Haplotype diversity was high at 0.741 ± 0.020, and nucleotide diversity was 0.011 ± 0.006. All six haplotypes differed from those of the same length reported previously for wild populations (Fernando et al. 2000; Vidya et al. 2005a, b). However, similarities were found between the shorter GenBank sequences (411 bp) from Thailand and either the Lao sequences or other published sequences: Thailand 2 matched Lao haplotype A, Thailand 3 matched Lao B, Thailand 13 matched Lao D, Thailand 6 matched Lao E, Thailand 7 matched Thailand 16, Thailand 5 differed from Thailand 19 at only two ambiguous bases, and Thailand 21 matched India BB. Thus, the 29 published haplotypes and six Lao haplotypes were reduced to a total of 28 unique Asian elephant haplotypes. Previous phylogeographic studies of Asian elephants using mitochondrial DNA have revealed two major clades that diverged approximately 1.6–2.1 mya (Fernando et al. 2000; Fleischer et al. 2001; Vidya et al. 2005a, b, 2009). Lao haplotypes A, B, C, and D were most similar to haplotypes in the alpha clade, and haplotypes E and F were most similar to those in the beta clade (Fig. 3), as defined by Fernando et al. (2000) and exemplified in Vidya et al. (2005a, b, 2009).
Mitochondrial haplotype network for Asian elephant haplotypes from Lao PDR (shaded) and the 22 other unique published haplotypes (India, Myanmar, Thailand, Vietnam). The alpha and beta clade designations follow Vidya et al. (2009)
Genetic diversity across the landscape
The spatial distribution of haplotypes revealed the highest diversity in blocks 9 and 10 and the lowest diversity in block 12 (Fig. 2). Four of the mitochondrial haplotypes (A, B, D, E) were found in blocks 5, 8, 9, and 10, and three of those four (all but haplotype E) were found in Block 12. The two unique haplotypes were only detected in blocks 9 and 10.
Individuals were concentrated in the center of the study area in sampling blocks 8, 9, and 10 (Table 5). This region encompassed all the haplotype diversity we detected in the study area, only 14 individuals were not detected in one of these central blocks. We did not detect spatial segregation of males and females; however, only one adult male was detected in block 10.
Genetic relationships
Females and juveniles detected near each other within a 48 h period had an average r-value that was significantly greater than zero (Table 6). Males detected near each other within a 48 h period had a negative average r-value that was not significantly different from zero. When relatedness values were examined for females and young of the same mitochondrial haplotype detected near each other within a 48 h period, the average relatedness values within all four haplotypes increased and were significantly greater than zero for the A, B, and E haplotypes (Table 6).
Across all individuals detected in the population, females (adults and subadults) and young have a significantly positive relatedness value (r = 0.012 ± 0.004, n = 77) while all adult and subadult males have a significantly negative relatedness value (r = -0.028 ± 0.013, n = 22). Furthermore, the older adult males (those with circumferences greater than 47 cm.) have a larger number of higher-order relationships (r ≥ 0.25) with other adult or subadult males (n = 13) in the population than with adult females (n = 8), while the younger adults (those with circumferences less than 47 cm) have a larger number of higher-order relationships with adult females (n = 33) than with subadult or adult males (n = 12).
Discussion
Genetic diversity is relatively high at all levels in this population. The mitochondrial haplotype diversity reported here (0.741 ± 0.020) is similar to that reported for Lao PDR and Vietnam previously (0.698 and 0.833, respectively; Fernando et al. 2000). However, previous studies have reported at most five haplotypes in the Laos/Vietnam region (Fernando et al. 2000; Vidya et al. 2007), while we detected six. Four of our Lao haplotypes matched shorter sequences from Thailand, which is not surprising given the geographic proximity of the two countries. Both major clades of mtDNA haplotypes were detected, and the four haplotypes of the alpha clade were found in over four times as many individuals (n = 82) as the two haplotypes of the beta clade (n = 17).
For microsatellite loci, the average number of alleles over ten loci (8.1) is higher than has been reported for India (3.4 over 6 loci; Vidya and Sukumar 2005b). However, the loci used in our study differ from those used by Vidya et al. (2005b) and may be more polymorphic overall. The average observed heterozygosity is higher than that reported for a small population in Vietnam (0.358; Vidya et al. 2007) and a much larger population in southern India (0.417; Vidya and Sukumar 2005b). It is not surprising that this population has higher genetic diversity than the population in Vietnam, which, unlike the Nakai elephants, appears to have gone through a bottleneck in the recent past (Vidya et al. 2007). The surprising finding is that the haplotype diversity is as high and the nuclear diversity is even higher than that of a much larger population in southern India.
There are several possible explanations for this high level of diversity. The Nakai Plateau’s elephants may be part of a larger regional metapopulation, and this region may be important for population connectivity. The occurrence of an adult female with a haplotype that does not match any other haplotype in the population supports this hypothesis. Alternatively, the area could harbor the remnants of populations that have moved into the region after losing habitat elsewhere. Further work will begin soon to distinguish between these hypotheses.
Recent work by Vidya et al. (2009) suggests that trade in Asian elephants may not have significantly altered the mtDNA population structure of the species. If this is found to be true for nuclear DNA population structure, it may be possible to construct a map of microsatellite allele frequencies for Asian elephants that can be used to infer the origin of seized ivory. This approach has been very successful in African elephants (Wasser et al. 2004), and recently allowed Wasser et al. (2007) to track the origin of a large shipment of ivory that was confiscated in Malawi to its probable origin in Zambia. The high level of polymorphism we found in this study, as well as in a second genetic survey in Cambodia (Pollard et al. 2008), suggests that the loci characterized by Kongrit et al. (2008), in combination with other loci described for this species, may be useful in constructing an Asian elephant microsatellite map. A tool such as this would be very useful to law enforcement efforts, as it would allow officials to focus their efforts on regions in which illegal killing is occurring (Wasser et al. 2007).
The relatively high relatedness values within sampling blocks suggest that female groups at this site represent traditional family groups. Females detected in the same sampling blocks at the same time had an average relatedness value significantly above zero (r = 0.061), and when broken down by haplotype, the relatedness values increased substantially (range 0.072–0.200) to values similar to those reported between adult females in savanna elephants (r = 0.1126; Archie et al. 2008). The relatedness values in this population are not quite as high as those reported within family groups (r = 0.365; Vidya and Sukumar 2005b) for a population in southern India, but they are well above those reported across family groups (r = 0.041). In addition, our estimates are likely biased low because we had no way to distinguish whether all the females detected together represented one family group or to detect individuals if they did not leave a dung pile. Therefore, more than one family group may be combined at some locations and some individuals may be missing from the group analyzes. However, despite the limitations of non-invasively collected dung samples, our relatedness values suggest familial relationships between females detected together.
For males, our results lend support to the locational dispersal model previously suggested by Vidya and Sukumar (2005a). The largest males in the population have a larger number of higher-order relationships (r ≥ 0.25) with other adult males than with adult females. However, higher-order relationships with females do exist, especially for the subadult males who may not have yet dispersed. If this is an area where populations mix, they may not have as great a need to disperse to mate with non-relatives. The average pairwise relatedness between all adult and subadult males in the population is significantly negative, suggesting that males may move into the area from more than one location. Finally, one subadult male detected in the population has a unique haplotype that is not shared by any females in the population, which could also be indicative of locational dispersal.
Our results confirm that the elephants of the Nakai Plateau represent a genetically diverse population. They suggest that the Nakai Plateau’s elephants may be part of a larger metapopulation, which has positive implications for population viability and supports the regional and international conservation significance of this population. The Nakai elephants appear to have social structure that is similar to that reported previously for larger Asian elephant populations (Vidya and Sukumar 2005b), which may mean that they will move as family groups as habitat is lost. The area along the Nam Theun River where it flows from the Nakai-Nam Theun NPA is an important area for the elephants because it has the highest genetic diversity and captures 86% of the population and all of its haplotype diversity. As development of the dam along the Nam Theun river and the relocation of people out of the inundated area continues, these data will be important in predicting and monitoring the effects of the project on an important Asian elephant population.
References
Amos W, Whitehead H, Ferrari MJ, Payne R, Gordon J (1992) Restrictable DNA from sloughed cetacean skin: its potential for use in population analyzes. Mar Mammal Sci 8:275–283
Archie EA, Maldonado JE, Hollister-Smith JA, Poole JH, Moss CJ, Fleischer RC, Alberts SC (2008) Fine-scale population genetic structure in a fission-fusion society. Mol Ecol 17:2666–2679
Clement M, Posada D, Crandall K (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1660
Cornuet J-M, Luikart G (1996) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144:2001–2014
Creel S, Spong G, Sands JL, Rotella J, Zeigle J, Joe L, Murphy KM, Smith D (2003) Population size estimation in Yellowstone wolves with error-prone noninvasive microsatellite genotypes. Mol Ecol 12:2003–2009
Duckworth JW, Hedges S (1998) Tracking Tigers: a review of the status of tiger, Asian elephant, gaur, and banteng in Vietnam, Lao, Cambodia, and Yunnan (China), with recommendations for future conservation action. WWF Indochina Programme, Hanoi, Vietnam
Duckworth JW, Salter RE, Khounboline K (1999) Wildlife in Lao PDR: the 1999 status report. IUCN-The World Conservation Union/Wildlife Conservation Society/Centre for Protected Areas and Watershed Management, Vientiane, Lao PDR
Eggert LS, Rasner CA, Woodruff DS (2002) The evolution and phylogeography of the African elephant (Loxodonta africana), inferred from mitochondrial DNA sequence and nuclear microsatellite markers. Proc R Soc Lond B 269:1993–2006
Eggert LS, Maldonado JE, Fleischer RC (2005) Nucleic Acid Isolation from Ecological Samples—Animal Scat and Other Associated Materials. In: Zimmer EA, Roalson EH (eds) Methods in Enzymology. Molecular Evolution: Producing the Biochemical Data, Part B. Academic Press, San Diego, pp 73–82
Eggert LS, Ahlering M, Manka S (2008) Lessons from genetic censuses of forest elephants. In: Olson, D (ed) Proceedings of the International Elephant Conservation and Research Symposium Nov, 2007. International Elephant Foundation and Ringling Brothers Barnum and Bailey Center for Elephant Conservation, pp 157–164
Fernando P, Lande R (2000) Molecular genetic and behavioral analysis of social organization in the Asian elephant (Elephas maximus). Behav Ecol Sociobiol 48:84–91
Fernando P, Pfrender ME, Encalada SE, Lande R (2000) Mitochondrial DNA variation, phylogeography and population structure of the Asian elephant. Heredity 84:362–372
Frantz AC, Pope LC, Carpenter PJ, Roper TJ, Wilson GJ, Delahay RJ, Burke T (2003) Reliable microsatellite genotyping of the Eurasian badger (Meles meles) using faecal DNA. Mol Ecol 12:1649–1661
Goodnight KF, Queller DC (1999) Computer software for performing likelihood tests of pedigree relationship using genetic markers. Mol Ecol 8:1231–1234
Hansen H, Ben-David M, McDonald DB (2008) Effects of genotyping protocols on success and errors in identifying individual river otters (Lontra canadensis) from their faeces. Mol Ecol Res 8:282–289
Hedges S (2006) Conservation. In: Fowler ME, Mikota SK (eds) Biology, Medicine and Surgery of Elephants. Blackwell, Ames, pp 475–490
Hedges S, Lawson D (2006) Monitoring the illegal killing of elephants: Dung survey standards for the MIKE Programme. CITES MIKE Programme Central Coordinating Unit (available at http://www.cites.org/common/prog/mike/survey/dung_standards.pdf)
Kongrit C, Siripunkaw C, Brockelman WY, Akkarapatumwong V, Wright TF, Eggert LS (2008) Isolation and characterization of dinucleotide microsatellite loci in the Asian elephant (Elephas maximus). Mol Ecol Res 8:175–177
Leimgruber P, Gagnon JB, Wemmer C, Kelly DS, Songer MA, Selig ER (2003) Fragmentation of Asia’s remaining wildlands: implications for Asian elephant conservation. Anim Cons 6:347–359
Luikart G, Alldendorf AW, Cornuet J-M, Sherwin WB (1998) Distortion of allele frequency distributions provides a test for recent population bottlenecks. J Hered 89:238–247
Munshi-South J, Tchignoumba L, Brown J, Abbondanza N, Maldonado JE, Henderson A, Alonso A (2008) Physiological indicators of stress in African forest elephants (Loxodonta africana cyclotis) in relation to petroleum operations in Gabon, central Africa. Divers Distrib 14:993–1001
Park SDE (2001) Trypanotolerance in west African cattle and the population genetic effects of selection. Ph.D. thesis, University of Dublin
Pollard E, Eggert L, Chanvibol C, Hedges S (2008) The status and conservation of Asian elephants in the Seima Biodiversity Conservation Area, Cambodia. Wildlife Conservation Society Cambodia Program, Phnom Penh
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Schneider S, Roessli D, Excoffier L (2000) Arlequin: a software for population genetics data analysis. Genetics and Biometry Laboratory, University of Switzerland, Geneva
Sukumar R (1989) The Asian elephant: ecology and management. Cambridge University Press, Cambridge
Tyson M, Hedges S, Sitompul AF (2002) WCS-Indonesia Sumatran Elephant Project: 6 month report January-June, 2002. Report to the Wildlife Conservation Society
van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) Micro-Checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Vidya TNC, Sukumar R (2005a) Social and reproductive behaviour in elephants. Curr Sci 89:1200–1207
Vidya TNC, Sukumar R (2005b) Social organization of the Asian elephant (Elephas maximus) in southern India inferred from microsatellite DNA. J Ethol 23:205–210
Vidya TNC, Fernando P, Melnick DJ, Sukumar R (2005a) Population differentiation within and among Asian elephant (Elephas maximus) populations in southern India. Heredity 94:71–80
Vidya TNC, Fernando P, Melnick DJ, Sukumar R (2005b) Population genetic structure and conservation of Asian elephants (Elephas maximus) across India. Anim Conserv 8:377–388
Vidya TNC, Varma S, Dang NX, Thanh TV, Sukumar R (2007) Minimum population size, genetic diversity, and social structure of the Asian elephant in Cat Tien National Park and its adjoining areas, Vietnam, based on molecular genetic analyzes. Conserv Genet 8:1471–1478
Vidya TNC, Sukumar R, Melnick DJ (2009) Range-wide mtDNA phylogeography yields insights into the origins of Asian elephants. Proc R Soc Lond B 276:893–902
Wasser SK, Shedlock A, Comstock K, Ostrander E, Mutayoba B, Stephens M (2004) Assigning African elephant DNA to geographic region of origin. Applications to the ivory trade. Proc Natl Acad Sci USA 101:14847–14852
Wasser S, Mailand C, Booth R, Mutayoba B, Kisamo E, Clark B, Stevens M (2007) Using DNA to track the origin of the largest ivory seizure since the 1989 trade ban. Proc Natl Acad Sci USA 104:4228–4233
Acknowledgments
We thank two anonymous reviewers for their insights and helpful comments. Funding for this project was provided by the Wildlife Conservation Society (WCS), the Nam Theun 2 Power Company Ltd., and the U.S. Fish and Wildlife Service (through Asian Elephant Conservation Fund grant number 98210-4-G800 to WCS); we are grateful to all these organizations. At the University of Missouri, Jessica Rabenold assisted with genetics laboratory work and data analyzes while the fieldwork would not have been possible without the willingness of the WCS Lao PDR Nakai Elephant Project’s numerous staff to work under what were sometimes trying circumstances; we are grateful to them all.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ahlering, M.A., Hedges, S., Johnson, A. et al. Genetic diversity, social structure, and conservation value of the elephants of the Nakai Plateau, Lao PDR, based on non-invasive sampling. Conserv Genet 12, 413–422 (2011). https://doi.org/10.1007/s10592-010-0148-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-010-0148-y